Abstract
L-type Ca2+ (CaV1) channels transduce channel activities into nuclear signals critical to neuritogenesis. Also, standalone peptides encoded by CaV1 DCT (distal carboxyl-terminus) act as nuclear transcription factors reportedly promoting neuritogenesis. Here, by focusing on exemplary CaV1.3 and cortical neurons under basal conditions, we discover that cytosolic DCT peptides downregulate neurite outgrowth by the interactions with CaV1’s apo-calmodulin binding motif. Distinct from nuclear DCT, various cytosolic peptides exert a gradient of inhibitory effects on Ca2+ influx via CaV1 channels and neurite extension and arborization, and also the intermediate events including CREB activation and c-Fos expression. The inhibition efficacies of DCT are quantitatively correlated with its binding affinities. Meanwhile, cytosolic inhibition tends to facilitate neuritogenesis indirectly by favoring Ca2+-sensitive nuclear retention of DCT. In summary, DCT peptides as a class of CaV1 inhibitors specifically regulate the channel activity-neuritogenesis coupling in a variant-, affinity-, and localization-dependent manner.
Similar content being viewed by others
Introduction
Voltage-gated Ca2+ channels (CaV) are closely involved in diverse pathophysiological processes, by generating Ca2+ signals in response to membrane potentials1,2. L-type Ca2+ channels (CaV1) from the CaV family, CaV1.2 and CaV1.3 in particular, are widely expressed in human tissues and organs including the nervous system3. Among its multiple roles, CaV1 channels mediate the signaling cascade known as excitation-transcription coupling, which transduces cellular stimuli into nuclear signals to regulate transcription of essential genes manifested into the growth conditions of neurites, which constitutes CaV1-dependent excitation-neuritogenesis coupling4,5,6,7. Notably, such coupling between CaV1 channels and neuritogenesis signaling is also functional even under basal conditions since Ca2+ channels are still active, e.g., to mediate the slow calcium oscillations8,9. That says, the essential factor to couple the downstreams is the activities of CaV1 channels, either at low (basal) or high (excited) levels, which explains the up- or downregulations of CaV1-mediated signaling and neuritogenesis upon high K+ stimulation and other perturbations10,11. Both CaV1 functions and neurite development are closely involved in learning and memory, and a broad spectrum of mental disorders12,13. The cascade linking CaV1 to neuritogenesis is coordinated by a number of key Ca2+-signaling proteins including calmodulin (CaM), Ca2+/CaM-dependent Kinase II (CaMKII), and calcineurin, to accomplish: CaV1 channel gating and Ca2+ influx, cytonuclear translocation of key molecules CaM and CaMKII, activation of nuclear transcription factors such as CREB (cAMP-response element-binding protein) and NFAT (nuclear factor of activated T-cells), expression of critical genes including c-Fos, and ultimately branching and elongation of neurites10,14,15,16,17,18. In parallel, the peptide fragments encoded by distal carboxyl-terminus (DCT) of CaV1 could also act as transcription factors in the nucleus that directly regulate gene transcription and expression, to promote neurite outgrowth19.
The effects of DCT as the key domain of CaV1 have been well characterized. By competing with apoCaM (Ca2+-free calmodulin), intramolecular DCT autonomously binds the canonical CaM-binding motif (the preIQ-IQ domain of CaV1), causing concurrent effects on channel gating, i.e., weaker Ca2+-dependent inactivation and reduced voltage-gated activation20,21,22,23. Chemical-induced dimerization of the proximal- and distal-DCT subdomains demonstrated that the reduction of Ca2+ influx is solely due to acute binding of DCT, ruling out other potential mechanisms of action21,24,25. Effects of DCT on CaV1 are in direct opposition to apoCaM, consistent with a mechanism of strict competition between DCT and apoCaM20,23. As summarized by the term CMI (C-terminus mediated inhibition)24, DCT is able to produce multifaceted and coherent effects, including competitive binding against apoCaM, reduction of Ca2+ influx, and concurrent attenuation of inactivation and activation.
Besides intramolecular DCT inhibition, standalone DCT peptides have also been reported to bind and inhibit CaV1 channels20,21,22,26,27,28,29,30,31. We postulated that DCT peptides through a mechanism of intermolecular CMI should attenuate Ca2+/CaV1 signaling to the nucleus. In support, some CaV1 inhibitors such as dihydropyridine (DHP) do exert inhibitory effects on CaV1-dependent signaling and neuritogenesis10,11,32. We pursued the above hypothetical inhibition on CaV1-dependent neuritogenesis by standalone DCT peptides, which, however, would be in direct contradiction to neuritogenic DCT effects as demonstrated by CCAT (Calcium Channel Associated Transcription regulator, encoded by CaV1.2 DCT)19, thus encountered with an immediate dilemma. In this work, we are motivated by such peptide CMI that supposedly downregulates the CaV1-neuritogenesis coupling through a unique mechanism of action: DCT peptides bind CaV1 at its CaM-binding motif.
DCT peptides have been found endogenously in native cells under physiological conditions33. These peptides could result from distinct production mechanisms in various cell types, and may have different lengths or compositions, but are all encoded by DCT fragments of high homology across the CaV1 family (Supplementary Fig. 1 and Supplementary Table 1). In skeletal muscle cells, CaV1.1-encoded peptides of ~30-40 kDa are produced by proteolysis34,35,36, named as CCTS (Cleaved Carboxyl-Terminal fragment from CaV1.1 pore-forming subunit α1S; and CCTC and CCTD from α1C and α1D, respectively). Similarly, CCTC is cleaved from CaV1.2 in neurons37 or cardiac myocytes38,39. Presumably by CaV1.3 cleavage, CCTD of ~40 kDa was found in cardiac myocytes40. Besides proteolytic cleavage, DCT peptides could also be generated by bicistronic mechanisms via exonic promoters for direct translation19,41,42,43, e.g., CCATC of ~60 kDa or more. In summary, two types of DCT peptides (~40 kDa and ~60 kDa) have been evidenced from diverse preparations33, alongside with a shorter peptide of ~15 kDa in neurons (encoded by the last ~100 a.a. at the C-terminal end of CaV1.2)41. Substantial discrepancies exist among various peptides in sequences, mechanisms of production, specificity to cell types, and effects on channels or cells. In this work, we undertook the task to clarify the actual roles of DCT-encoded peptides, focusing on the hypothesis that these peptides of diverse forms essentially share the same principle: affinity-dependent binding of CaV1 channels to downregulate CaV1 activities and channel activity-dependent neuritogenesis.
Results
Hints on DCT peptide inhibition of CaV1 channels and neuritogenesis
We first conducted sequence alignment for homologous DCT domains across CaV1.1-1.4, containing the major fragments of proximal C-terminal regulatory domain (PCRD), nuclear retention domain (NRD) and distal C-terminal regulatory domain (DCRD) (Supplementary Fig. 1 and Supplementary Table 1). The NRD motif is an indispensable region for Ca2+-dependent nuclear export as demonstrated in CCATC19 and CCTD40. The PCRD and DCRD cooperate to compete with apoCaM for binding the IQ motif of CaV1 as the molecular basis of CMI, where the DCRD plays a dominant role compared with the PCRD22,24. Two transcription activation domains are localized in the PCRD-NRD junction and the DCRD motif19,41, respectively. Based on these and earlier analyses, the representative peptides of three major categories have been focused on, including 1) DCT peptides (~60 kDa) via bicistronic transcription, such as CCATC that contains the entire DCT; 2) CCTC or CCTD (~40 kDa) from posttranslational cleavage lacking the PCRD domain but still incorporating the majority of DCT (from NRD to DCRD); 3) the short peptide of DCRD (~15 kDa) that is sufficient to modulate CaV1 gating, although its physiological relevance is relatively less established.
As proof of principle, the DCRDF was overexpressed in cortical neurons, considering that this short peptide encoded by the last ~100 a.a. of CaV1.4 DCT has been thoroughly characterized for its strong competition with apoCaM to bind onto the channel20,24,27,31. By the cocktail treatments of cultured cortical neurons, the relative contribution of CaV1.3 channels to CaV1 currents was evaluated (see Methods and Supplementary Fig. 2 for details). Based on the patch-clamp recordings by a voltage ramp and the representative step of −10 mV, CaV1.3 made a significant contribution to the total CaV1 currents in cortical neurons (~50%), in agreement with the previous reports suggesting that both CaV1.3 and CaV1.2 are critical to CaV1 signaling in cortical neurons and hippocampal neurons44,45,46,47,48. With the full cocktail recipe, Ca2+ currents mainly mediate by CaV1.3 were isolated and recorded to examine the effects of DCRDF peptides (Fig. 1a). The DCRDF potently attenuated cortical CaV1.3 currents at the peak; meanwhile, the steady-state amplitude (measured at 300 ms) was nearly unchanged. Such characteristic effects on native CaV1.3 channels are highly consistent with the CMI modulation of recombinant CaV1.3 channels24, where reduction of Ca2+ influx is ensured by concurrent attenuation of activation and inactivation evidenced from both acute and long-term effects. We then examined the hypothetical role of DCT inhibition on CaV1-dependent neuritogenesis signaling. As expected, the DCRDF peptide caused a significant reduction in neurite outgrowth and branching of cortical neurons under basal conditions, as measured by the total length and complexity (Sholl analysis) respectively (Fig. 1b). In contrast, the mutant peptide DCRDF_V/A (V/A denotes V41A, a loss-of-function mutation) produced no damage on neurite outgrowth of cortical neurons overexpressing DCRDF.
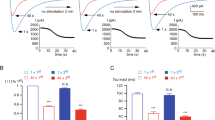
a Inhibition of endogenous CaV1.3 currents by DCRDF peptides. Cultured neurons were treated with the cocktail recipe to isolate CaV1.3 Ca2+ current (details in Supplementary Fig. 2). Exemplary Ca2+ current (upper, scale bar in red) was elicited by the standard protocol of voltage step (300 ms; -10 mV). Potential inhibitory effects on channel functions were evaluated by the indices of peak and steady-state current amplitudes (pA/pF, bottom). b Effects on neuronal morphology. Based on the confocal fluorescent images of peptide-expressing cortical neurons (upper row), neurite tracing for each neuron was performed (middle row), total neurite length and Sholl analyses summary (bottom row) were compared among the three groups: YFP, YFP-DCRDF and YFP-DCRDF_V/A. Sholl analyses are routinely accompanied by the quantitative index of AUC (area under the curve, lower right). c Effects of DCRDF peptides on pCREB signals. Cortical neurons were transfected with YFP, YFP-DCRDF or YFP-DCRDF_V/A (loss-of-function mutant), respectively. The pCREB signals were evaluated by immunofluorescence. Red and yellow fluorescence in confocal images represent pCREB signals and overexpressed YFP, respectively. pCREB signals were normalized over the YFP control group. d Effects on c-Fos signals. Cortical neurons expressing YFP, YFP-DCRDF or YFP-DCRDF_V/A were stained with c-Fos antibody (in red). Fluorescent intensities of c-Fos signals in the nuclei were summarized. Student’s t-test (a), one-way ANOVA followed by Bonferroni for post hoc tests (b, c) and Kruskal-Wallis and Dunn’s non-parametric test (d, non-normal distribution, checked by D’Agostino & Pearson omnibus normality test) were used (***p < 0.001; n.s., not significant, p > 0.05). Values are represented as mean ± SEM.
We further checked the major signals along the well-established cascade, including the phosphorylation of a key transcription factor CREB49,50. CaV1 would be the major path of Ca2+ entry preferred by downstream CaMKII/CREB signaling over CaV211, DCRDF peptides strongly attenuated pCREB signals (immunostaining of phosphorylated CREB or pCREB, Fig. 1c). In contrast, pCREB exhibited no difference between the control neurons and the mutant group DCRDF_V/A20,24. Furthermore, the expression level of c-Fos, one of the classical immediate early genes driven by pCREB15,16, was significantly reduced by DCRDF but not DCRDF_V/A (Fig. 1d). Additional stimulation to enhance membrane excitation or channel activation is expected to provide higher dynamic ranges, although the CaV1 channel activity-neuritogenesis coupling should function similarly in cortical neurons of either conditions (basal or excited). Indeed, when the neurons were stimulated by 40 mM K+, similar results from WT and mutant DCRDF were obtained when we reexamined the peptide effects on pCREB signals (Supplementary Fig. 3a). In addition, the enhanced signals by high K+ stimulation provided the opportunity to capture the potent inhibition of DCRDF on the translocation of CaM (from Cytosol to Nucleus, defined by N/C ratio, Supplementary Fig. 3b), another key event along the CaV1-triggered signaling pathway49,50. Here, the observed effects arise from DCRDF inhibition on CaV1 channels which can be well represented by CaV1.3, especially in cortical neurons under basal conditions.
CMI effects on recombinant CaV1.3 channels by various DCT peptides
Encouraged by the results that DCT peptides downregulated cortical CaV1 channel-dependent transcription and neuritogenesis, we proceeded further with the recombinant CaV1.3 channels for the details on DCT effects. We chose five variants encoded by DCT of CaV1.2 or CaV1.3 which represent the native forms of DCT peptides: one long-form variant CCATC (~60 kDa), two medium-form variants CCTC and CCTD (~40 kDa), and two short-form variants DCRDC and DCRDD (~15 kDa) (Fig. 2a). To quantify their effects on the gating of full-length CaV1.3 (α1DL) channels, the two major indices were routinely examined: inactivation (the strength of Ca2+-dependent inactivation, SCa) and activation (the peak amplitude of Ca2+ current, ICa)24 (Fig. 2b, left column). Firstly, consistent with the previous report27, the DCRDF peptides generated characteristic CMI effects: concurrent attenuation of both inactivation and activation, as illustrated by the altered profiles (the green shades to illustrate the actual attenuation) (Supplementary Fig. 4a). In direct contrast, DCRDF_V/A did not cause any appreciable change in gating indices (SCa or ICa). CMI effects on activation were also evidenced from Ba2+ currents which were significantly inhibited by DCRDF but not by DCRDF_V/A (Supplementary Fig. 4b). Notably, demonstrated by voltage-dependent steady-state (at 300 ms) currents, DCRDF and DCRDF_V/A peptides are essentially indistinguishable from the α1DL control, supporting the notion that intermolecular CMI (by standalone DCT peptides) shares similar mechanisms with intramolecular CMI (by the DCT motif covalently-linked to the channel), supposedly in an acute manner as previously proved24.
a Illustration of serial representative peptides encoded from DCT domains of CaV1.2 (blue) and CaV1.3 (red). The long-form peptides (~60 kDa) represented by CCATC contain the complete set of motifs including PCRD, NRD and DCRD. The medium-form (~40 kDa) representatives CCTC and CCTD start from the very end of the PCRD till the end of the DCRD thus containing both the NRD and the DCRD. The short-form peptides (~15 kDa) are represented here by the peptides DCRDC and DCRDD. b CMI effects of representative variants on CaV1.3 (α1DL) in HEK293 cells. As illustrated by the control group (left column), exemplary Ca2+ current (trace with scale bar, red) and Ba2+ current (rescaled to Ca2+ current at the peaks, gray) were elicited by voltage step at −10 mV (top traces). The next two rows show the profiles of inactivation and activation respectively, with r50 (ration between current amplitudes at 50 ms and the peak) for inactivation and Ipeak (Ca2+ current) for activation across the full range of membrane potentials (V). Based on Ca2+ currents at −10 mV, SCa (in fraction, 1-r50) and ICa (in pA/pF, Ipeak) serve as the major indices for inactivation and activation, respectively. Five representative peptides of CCATC, CCTC, CCTD, DCRDC and DCRDD (from left to right) as in (a) are compared with α1DL control (the leftmost column) for their inactivation and activation profiles (lines in orange indicate the control). Green areas are to highlight peptide effects. c Statistical summary of the extent of Ca2+-dependent inactivation (SCa) and peptide’s CMI potency (CMI in percentage). To evaluate the attenuation of DCT peptide on inactivation (SCa), CMI potency is defined as the change in Ca2+-dependent inactivation (SCa): (SCa,Control − SCa,Peptide)/SCa,Control, which is equivalent to fractional change in apoCaM-bound channels (ΔFCaM) before (FCaM) and after DCT peptide’s competition. Thus, CMI essentially indicates what percentage of apoCaM-bound channels are converted to peptide-bound channels (fPeptide). See Eqs. E2 and E3 in Methods for more details. One-way ANOVA followed by Dunnett for post hoc tests were used for (c): *p < 0.05; **p < 0.01; ***p < 0.001. Values are represented as mean ± SEM.
Following the initial evidence from the CaV1.4 DCRDF, we performed a systematic comparison for the five representative peptides (Fig. 2b), each of which was co-expressed with CaV1.3 channels. To better quantify DCT effects, the inhibition potency (CMI, in percentage) is defined as the normalized fraction of channels that switch from apoCaM-bound to DCT-bound, which can be directly calculated from the inactivation parameter SCa before and after peptide inhibition (Eqs. E2 and E3 in Methods). CMI is inversely proportional to SCa, therefore strong DCT such as DCRDF should have higher CMI than weaker DCT such as DCRDF_V/A.
By YFP fluorescence intensities, individual cells expressing YFP-tagged DCT were scrutinized to ensure that the expression levels of these peptides were at the comparable (high) levels. DCT peptides of DCRDC, CCTC, DCRDD and CCTD clearly produced CMI effects of substantial potency, except that CCATC only slightly attenuated CaV1.3 gating (weak CMI) (Fig. 2c). Electrophysiological profiling of peptide CMI lays the foundation for our subsequent investigations into the effects of DCT peptides on CaV1-dependent neuritogenesis. CCATC exhibited rather weak (insignificant) effects on channel gating, which appears to agree with the earlier report where the long DCTC was found to have no effect on Ca2+-dependent inactivation of CaV1.222. However, CCTC, the shorter motif encoded by a portion of CCATC, is capable of strong CMI. Also, the key segment DCRDC has the capability to attenuate Ca2+-dependent inactivation. In this work, one of our aims is to clarify the above discrepancy regarding various DCTC peptides. In fact, all the four DCT domains across the CaV1 family are homologous (including the DCRD domains), suggesting high similarities in their functional roles (Supplementary Fig. 1). Taking the DCRDF as the exemplar, its core segment was further narrowed down to DCDRF_17-66 (the residues between S17 and L66) well conserved among CaV1.2-1.4 (but not CaV1.1), which may account for the potent inhibition observed from DCRDC, DCRDD or DCRDF peptides (Supplementary Fig. 5).
CaV1 channels and neuritogenesis are inhibited by cytosolic DCT peptides
In the context of CaV1-dependent neuritogenesis, modulation of CaV1 channels would make changes in the growth of neurites, provided that the channel activity-transcription coupling is coherently regulated. Based on the potent inhibition by DCRDF (Fig. 1), we pursued the hypothesis further that the DCT peptides of native forms may induce inhibitory effects in accordance with CMI potency (Fig. 2c). Since the signals or events at all the checkpoints (CaV1 gating, Ca2+ influx, CaM translocation, pCREB, c-Fos and neuritogenesis) were consistently attenuated by DCRDF in cortical neurons (Fig. 1), CaV1 gating and neurite growth, the major input and output respectively, were selected as the two major checkpoints to represent the full cascade of channel activity-neuritogenesis coupling. By overexpressing DCT peptides in cortical neurons with careful scrutinization of cellular fluorescence as in electrophysiology (Fig. 2), the potential CMI on neuritogenesis was examined for the representative peptides of CCATC, CCTC, and CCTD (long and medium forms), along with the peptides DCRDC and DCRDD (short form). To our surprise, statistically none of CCATC, CCTC, and CCTD exhibited any significant effect on neurite length and branching even for ensured overexpression (Fig. 3a–c). Meanwhile, resembling DCRDF inhibition, DCRDC and DCRDD peptides induced significant neurite retractions. Since functional CaV1 channels are located at the plasma membrane, CMI effects by DCRD peptides should take place in the cytosol. We then examined the cytosolic-nuclear distribution for each peptide variant, indexed by its N/C ratio (Fig. 3d). On average, the N/C ratio values for DCRDC and DCRDD fell below the control level (YFP, with N/C ratio ~1.5)51, showing a pattern of cytosolic distribution; in contrast, the peptides CCATC, CCTC, and CCTD were more distributed into the nucleus (N/C ratio>1.5). For each variant, by applying N/C ratio criteria (cut-off value of 1.5) the neurons could be divided into two distinct (cytosolic versus nuclear) subgroups. For the cytosolic subgroup, similar to DCRDC and DCRDD, cytosolic CCTC and CCTD significantly attenuated neurite outgrowth (Fig. 3e, f). Notably, although cytosolic CCATC appeared to have a tendency of attenuation, its actual effects on neurites turned out to be rather mild with no statistical significance, consistent with its weak CMI on CaV1.3 gating. We then postulated that cytosolic DCT peptides would downregulate the CaV1-dependent neuritogenesis. In support, for the five representative peptides we tested in cortical neurons, an inverse correlation appeared to exist between CMI potency of cytosolic peptides and neurite length (Fig. 3g). Hence, it is likely that the DCT peptides present in the cytosol share the same mechanisms with DCRDF to inhibit CaV1 gating and signaling (Fig. 1).
a Fluorescence images of cortical neurons (DIV-7) expressing CFP and DCT variants tagged with YFP. Representative confocal images (upper row, merged CFP in cyan with YFP in yellow) and corresponding neurite tracing (bottom row) are shown for YFP control, CCATC, CCTC, CCTD, DCRDC and DCRDD, respectively. b, c Statistical summary of total neurite length (b) and Sholl analyses (c). d Statistical summary of N/C ratio for all the five groups of DCT peptides. Horizontal line indicates N/C ratio of YFP as the criteria (for DCT distribution) to assign each neuron to either the nuclear subgroup (N/C ratio>1.5) or the cytosolic subgroup (N/C ratio<1.5). e Representative confocal images, neurite tracings and detailed cytonuclear distribution are shown for neurons with cytosolic CCATC, CCTC and CCTD, respectively. The envelope of the nucleus is highlighted by live-cell Hoechst 33342 staining (dotted lines). f Total length for the neurons from the cytosolic groups: peptides of CCATC, CCTC and CCTD, and also the groups of DCRDC and DCRDD peptides. g A potential correlation between CMI values (adopted from Fig. 2c) and total neurite length (R2 = 0.93). One-way ANOVA followed by Bonferroni or Dunnett for post hoc tests were used for (b, c) and (f), respectively (*p < 0.05; **p < 0.01; ***p < 0.001). Values are represented as mean ± SEM.
Both PCRD and DCRD tune CMI potency of DCT variants
No structural information of DCT is available thus far52,53,54. To gain further insights into the mechanisms underlying DCT effects, we firstly focused on CCATC, unexpectedly exerting rather mild inhibition on CaV1.3 channels and cortical neurons (Fig. 3g). In contrast to CCATC, the shorter peptides of DCRDC and CCTC both encoded by CaV1.2 DCT have strong CMI, suggesting a self-limiting mechanism within the longer CCATC. Moreover, CaV1.2 has been considered to have the same level of Ca2+-dependent inactivation with or without its DCT domain22,55, inconsistent with strong CMI of DCRDC in our experiments (Fig. 2). To resolve these discrepancies, we performed systematic analysis with the representative DCT peptide variants. By utilizing 2-hybrid 3-cube FRET (Förster resonance energy transfer), a quantitative imaging assay for protein-protein interactions in live cells24,56, the capabilities of DCRD peptides to bind the channel were quantified by dose-dependent binding (FR-Dfree) curves (Fig. 4a). Following the convention, we employed the (effective) dissociation equilibrium constant (Kd, units in fluorescence intensities through the donor cube) as the index of binding affinities. Utilizing CFP-tagged DCRDX peptides (X = S, D, C, and F representing CaV1.1-1.4) and YFP-tagged preIQ3-IQD-PCRDD (CaM-binding motif of CaV1.3) as the FRET pairs, a series of binding curves were achieved by iterative fitting processes (Fig. 4a). Among a gradient of Kd values from the four pairs of binding, DCRDF encoded by CaV1.4 resulted in the strongest affinity (Kd = 1.8 × 103), followed by the peptides DCRDD (Kd = 4.3 × 103), DCRDC (Kd = 16.5 × 103) and DCRDS (Kd = 29.0 × 103).
a Binding curves for the interactions between key channel motifs and DCRD peptides were quantified by 2-hybrid 3-cube FRET, for each pair between CFP-DCRDX from CaV1 (X = S, D, C and F) and YFP-preIQ3-IQD-PCRDD. Dfree and FR represent free donor concentration and FRET ratio, respectively. The binding affinity Kd for each pair was achieved by iterative fitting. In particular, the pair of PCDRD and DCRDF serves as the major reference for subsequent analyses. b Comparison of CMI potency among DCRDX peptides. As in the cartoon illustration of peptide CMI (top), the peptide DCRDX could coordinate with the IQD and PCRDD motifs on α1DΔ-PCRDD to compete apoCaM off the channel. Ca2+ trace exemplars and inactivation profiles are shown for α1DΔ-PCRDD alone or with different DCRDX isoforms. c Relationship between Kd and CMI for DCRDX peptides. Four peptides directly from CaV1.1-CaV1.4, plus one additional mutant peptide DCRDF_V/A (Supplementary Fig. 6). The relationship between Kd and CMI for DCRDX peptides was fit by Eq. E4 (see Methods). d Similar to DCRD measurements (a), PCRDX peptides across CaV1 family (X = S, D, C, and F) were also quantified by FRET for the interactions between YFP-preIQ3-IQD-PCRDX and CFP-DCRDF. The two sets of measurements share the common pair PCRDD and DCRDF as the major index. See Kd values of PCRDX-DCRDX in Supplementary Fig. 7. e Comparison of CMI potency among PCRDX-DCRDF peptides. As illustrated by the cartoon (top), PCRDX-DCRDF could form the complex with IQD of α1DΔ to compete with apoCaM. In similar fashion, Ca2+ trace exemplars and inactivation profiles are shown for α1DΔ alone or with long-form peptides PCRDX-DCRDF. f Further support for the proposed CMI-Kd correlation. Additional data points (CMI and Kd) of PCRDX-PCRDF (e) and PCRDC-DCRDC (Supplementary Fig. 8) are superimposed onto the ligand-binding curve from (c). g The tuning curve of Kd-dependent CMI with the summary of peptides. For compound effects of long-form DCT peptides on α1DΔ-PCRDD, three more data points (black squares) were added onto the tuning curve, based on CMI measurements and Kd estimations for PD+PC-DC (or PD+CCATC), PD+PC-DF and PD+PF-DF (Supplementary Fig. 9). Values are represented as mean ± SEM.
In parallel with FRET binding analyses, the whole-cell electrophysiology was performed for functional characterizations. DCRDX peptides were overexpressed with α1DΔ-PCRDD, a channel variant producing ultra-strong Ca2+-dependent inactivation (due to lacking the critical DCRD domain) thus providing an ample dynamic range to evaluate CMI effects. All of the four DCRDX peptides caused inhibitory effects of different potency on α1DΔ-PCRDD channels, illustrated by their inactivation (SCa) profiles (Fig. 4b). The classic ligand binding (Langmuir isotherm) equation (see Methods: Eq. E4) between inhibition potency CMI and binding affinity Kd was utilized to describe the differential effects among the peptide variants (Fig. 4c). For the mutant DCRDF_V/A, both peptide binding and channel inhibition were severely perturbed by the critical mutation (Supplementary Fig. 6), also agreeing well with the tuning curve of Kd-dependent CMI (Fig. 4c).
In parallel, the potencies of PCRDX (X = S, D, C and F) were examined with FRET pairs YFP-preIQ3-IQD-PCRDX and CFP-DCRDF. Similar to DCRDX, Kd values were obtained for the PCRDX peptides, unveiling the relative order of strength in binding (starting from the strongest): PCRDF, PCRDD, PCRDC, and PCRDS (Fig. 4d). The difference in Kd between PCRDC (Kd = 11.3 × 103) and PCRDD (Kd = 1.8 × 103) is even more pronounced than that between DCRDC and DCRDD (6.3-fold versus 3.8-fold), suggesting that the rather weak inhibition by DCTC (either as the intramolecular motif or the intermolecular peptide) is mainly attributed to its proximal domain PCRDC. Such result is unexpected, since the PCRD motif has been considered to play a much lesser role (than the DCRD motif) in DCT effects. For instance, it has been reported that PCRD is not required for channels inhibition under the low CaM conditions whereas DCRD still remains indispensable to CMI24. For fair comparison, the pair of DCRDF and preIQ3-IQD-PCRDD is taken as the principle reference (noted as PCRDD/DCRDF or its abbreviation PD/DF) (Supplementary Table 1). All combinations of PCRDX/DCRDX (abbreviated as PX/DX) are summarized to compare their Kd values (Supplementary Fig. 7). Besides experimental values from FRET, Kd for other PX/DX combinations can also be roughly estimated according to the values assigned to PX and DX. For validation purposes, FRET experiments were conducted for PC/DC (CaV1.2) and PS/DS (CaV1.1) (Supplementary Fig. 8), resulted in rather weak binding affinities (Kd), consistent with the predictions from PX and DX (Supplementary Fig. 7).
Similar to DCRD, the functional role of PCRD was also examined, but by co-expressing PCRDX-DCRDF (i.e., PX-DF) with α1DΔ channels. PS-DF and PC-DF peptides exhibited much weaker inhibition than PD-DF and PF-DF, indicated by less changes in Ca2+-dependent inactivation of α1DΔ channels (Fig. 4e). The four peptides of PX-DF comply with the same tuning curve of CMI-Kd in Fig. 4c, but here with PX as the factor subject to variations (Fig. 4f). In addition, the relatively weak Kd and CMI for PC-DC (Supplementary Fig. 8) agree well with the tuning curve, as one additional validation for its applicability to PX-DX peptides.
In summary, both PCRD and DCRD may underlie the distinct effects of DCT peptides across the CaV1 family20,22,57,58. Here, the importance of PCRD is unmasked. The ultra-weak CMI potency of DCTS or DCTC is mainly attributed to its PCRD domain, in that DCRDS and DCRDC are fully capable of strong CMI effects (Fig. 4b). In this context, for CCTD, CCTC and CCATC (here equivalent to DCRDD, DCRDC, and DCTC), the potencies of CMI effects (from strong to weak inhibition) on CaV1 (represented by CaV1.3, Fig. 2c) are expected to be in the same order as their binding affinities: PD/DD, PD/DC, and PC/DC (from strong to weak binding).
Compound effects on α1DL channels by long DCT peptides
Although the weak inhibition by the long peptides DCTC or CCATC could be attributed to its PCRDC, it is still unclear why CaV1.3 (the full-length channel containing intramolecular PCRDD) is barely regulated by CCATC peptides. To fully elucidate the mechanism underlying the weak CCATC, we decomposed its effects on α1DΔ-PCRDD channels into two scenarios (Fig. 4g). The first component (I) represents the combination of PD from the channel and DC from the peptide, which produces strong inhibitory effects. The second component (II) represents the combination of PC and DC both from the long peptide DCTC (equivalent to CCATC), which has rather weak CMI (Supplementary Fig. 9a, b). Overall, CMI potency of CCATC on α1DΔ-PCRDD is expected to fall into the range defined by both components (I and II) corresponding to the upper- and lower-limit respectively. The compound effects of CCATC resulted in weak CMI potency toward its lower limit, suggesting a dominant role of PCRDC in this particular scenario. For validation purposes, PC-DF (PCRDC fused with DCRDF) was constructed and applied as an artificial type of DCT peptides (Supplementary Fig. 9c, d). Similar to PC-DC, the effects of engineered PC-DF on α1DΔ-PD could also be decomposed into two combinations, where PCRDC compromised the ultrastrong CMI of DCRDF and thus the overall CMI only reached an intermediate level.
Collecting the data from PCRD or DCRD variants (Fig. 4c, f), a tuning curve between CMI potency and binding affinity (CMI-Kd) has been established, applicable to a broad scope of channel and peptide variants (Fig. 4g). In principle, for any DCT peptide variants, of either native or engineered and either WT or mutant forms, when applied to CaV1.3 channels (supposedly to CaV1 channels in general), the potency of CMI quantitatively would correlate with the affinity between peptides and channels, which is also a measure of the competition (against apoCaM) introduced by DCT peptides. Based on the tuning curve, Kd values for particular peptides can be estimated from their CMI values measured in electrophysiology, which has been demonstrated by PC-DC (or CCATC) and PC-DF, and also by another long peptide PF-DF (Supplementary Fig. 9e, f). Furthermore, CCTS36, hypothetical CCATD or CCTF, and more other variants, can also be evaluated or predicated for the effects on CaV1 channels according to such unified tuning curve of CMI-Kd.
Cytosol/nucleus-dependent effects of DCT peptides reconcile the discrepancy in neurons
Our data thus far demonstrate that cytosolic DCT peptides negatively regulate neurite outgrowth, intrinsically tuned by CMI (channel inhibition) or Kd (peptide binding) in a variant-dependent manner. The apparent contradictions regarding DCT effects (inhibitory versus facilitatory) may simply reflect the differential roles of peptides in the cytosol versus in the nucleus. In order to test this hypothesis, we first revisited the overexpressing CCTD which was widely distributed across the whole cell, featuring a broad range of N/C ratio values (Fig. 3d). Dual directional effects are evident: for the nuclear group CCTD (N) with N/C ratio>1.5, neurite outgrowth was promoted; in contrast, for neurons from the cytosolic group (N/C ratio<1.5) or CCTD (C), neurite outgrowth was significantly reduced similar to DCRDD (Fig. 5a–c). To confirm this result, the short tags of nuclear export signal (NES) and nuclear localization signal (NLS) were fused to the N-terminus of CCATC or CCTC. In doing so, NES-tagged CCATC and CCTC were predominantly expressed in the cytosol (Fig. 5d, e). In comparison with the minor effects of NES-CCATC, neurite retractions were evidenced from NES-CCTC as indicated by shorter neurites (Fig. 5f) and reduced complexity (Fig. 5g), consistent with CMI-dependent inhibition of neuritogenesis we observed earlier (Fig. 3g). In contrast, NLS-tagged CCATC and CCTC were constrained in the nucleus, presumably acting as neuritogenic transcription factors19 (Fig. 5d–g). Indeed, both peptides led to the significant promotion of neurite outgrowth, whereas the longer peptide NLS-CCATC was slightly more potent than the shorter NLS-CCTC (Fig. 5f), suggesting that the second TA (transcription activation) region (roughly overlapped with DCRD) may play a major role and the first TA region (largely overlapped with PCRD) would be relatively less significant (Supplementary Fig. 1). By revisiting Fig. 3, the actual effects on neurite outgrowth have been clarified to be highly dependent on subcellular localization of DCT peptides, which could be inhibitory when present in the cytosol by attenuating CaV1 activities and signals, or could be neuritogenic when localized in the nucleus as transcription factors (Fig. 5h). Furthermore, such opposing effects have been confirmed with more mature neurons (>DIV 15) of which neurite outgrowth was suppressed or facilitated by CCTD explicitly tagged with NES or NLS, respectively (Supplementary Fig. 10).
a Cortical neurons with CCTD fragments are grouped into two categories: nuclear versus cytosolic, i.e., CCTD (N) and CCTD (C) by the same criteria of N/C ratio as in Fig. 3 (CCTD fluorescence with N/C ratio >1.5 for the nuclear group; and N/C ratio <1.5 for the cytosolic group). b, c Total neurite length (b) and Sholl analyses (c) for neuron groups of Control, total CCTD, CCTD (C) and CCTD (N). Original data for Control and total CCTD groups are adopted from Fig. 3b-e NES and NLS were fused to N-terminus of YFP-CCATC (d) or YFP-CCTC (e) to constrain the distribution of DCT peptides within the cytosol or the nucleus, respectively. f, g Total neurite length (f) and Sholl analyses (g) for the groups of Control, NES-YFP-CCATC versus NLS-YFP-CCATC, and NES-YFP-CCTC versus NLS-YFP-CCTC. h Total neurite lengths for neurons overexpressing CCATC, CCTC and CCTD peptides are summarized and compared among nuclear versus cytosolic subgroups, together with DCRDC and DCRDD groups (adopted from Fig. 3f). i Cyto-nuclear distribution (indexed by N/C ratio) of DCT peptides in correlation with the total neurite length. Horizontal and vertical lines in green represent the control group (YFP expressed in cortical neurons). The range of N/C ratio is indicated on top of the scatter plot for each peptide group of CCATC, CCTC, CCTD, or the mutant CCTD_V/A. The grey shades for peptide variants are to illustrate the potential correlation between neurite length and N/C ratio. j Correlation between CMI strength and cyto-nuclear distribution. For four peptide variants of CCATC, CCTC, CCTD and CCTD_V/A, standard deviations of N/C ratio values are represented by σ (N/C ratio) to quantify spatial dynamics for each peptide variant. CMI and σ are highly correlated (linear fit, R2 = 0.90). One-way ANOVA followed by Dunnett for post hoc tests were used for (b, c), and (f–h) (*p < 0.05; **p < 0.01; ***p < 0.001). Values are represented as mean ± SEM.
Cytosolic and nuclear DCT peptides are in direct opposition to each other (inhibition versus facilitation) in regulating neuritogenesis, as the plausible reason to account for the less-pronounced overall effects observed from overexpressing CCATC, CCTC, and CCTD (Fig. 3a–c). On the one hand, nuclear DCT peptides are able to promote neuritogenesis; but on the other hand, cytosolic peptides have distinct CMI potency to induce differential levels of inhibition on neurite outgrowth. Is there any mechanism to regulate/maintain the potential balance between the opposing (cytosolic versus nuclear) effects? First, the spatial distribution was examined for DCT peptides in accordance to neurite outgrowth. As depicted by the scatter plots to correlate N/C ratio with total neurite length, the peptides CCATC, CCTC and CCTD (compared with CCTD_V/A) spread across the cytosol/nucleus of each neuron (Fig. 5i). In contrast, the DCRDC and DCRDD peptides were exclusively constrained within the cytosol thus solely functioning as inhibitors of neurite outgrowth (Supplementary Fig. 11). Roughly at the same expression levels for whole cells, these peptides exhibited different patterns of subcellular distribution, as illustrated by the dynamic range between the minimum and maximum N/C ratios for each peptide. For CCATC, the rather narrow range of N/C ratio is consistent with its least localization in the nucleus. In comparison, CCTD appears to spread into the nucleus with a broader range of N/C ratio. We quantified the dynamic range of peptide distribution by the standard deviation (σ) of N/C ratio, which is closely correlated with CMI potency in the order of CCATC, CCTD_V/A, CCTC, and CCTD from weak to strong (Fig. 5j). Regarding the mutant peptide CCTD_V/A, the single-residue V/A mutation in the DCRD domain significantly attenuated its inhibitory effects on channel gating (Supplementary Fig. 12a–c), and neurite outgrowth (Supplementary Fig. 12d–f). Notably, nuclear CCTD_V/A still promoted neurite outgrowth, similar to WT CCTD in the nucleus. In agreement with the proposed σ-CMI correlation, CCTD_V/A is less distributed in the nucleus compared with WT CCTD (Fig. 5j); also, the clear differences between CCTD_V/A and WT CCTD resemble CCATC versus CCTC. One potential explanation could be that the distributions and effects of DCT peptides are subject to certain autonomous regulations in neurons, presumably by way of CaV1/Ca2+ influx and Ca2+-sensitive NRD (the nucleus retention domain contained in the long or medium peptides but not in the short peptides, Supplementary Fig. 1).
Downregulations of neuritogenesis signaling by CaV1.3-encoded peptides
The localization-dependent regulation of neuritogenesis reconciles the opposing roles of exogenous DCT peptides in neurons. Before we proceeded further with its potential Ca2+/CaV1 dependence, we examined the key signals that are involved. For CaV1.3-encoded peptides, CCTD should serve as one representative form, considering that all the key domains such as NRD are included and there is no issue of weak PCRD as in CCATC (Supplementary Fig. 1). As expected, the exogenous peptides of NES-CCTD demonstrated that the cytosolic peptides suppressed pCREB in direct contrast to the nuclear peptides of NLS-CCTD (Fig. 6a). c-Fos, a hallmark gene of DCT effects (Fig. 1d), was also examined here for its expression driven by CaV1/pCREB. Resembling pCREB, c-Fos expression was significantly reduced by NES-tagged but not by NLS-tagged peptides (Fig. 6b), consistent with that nuclear DCT peptides are able to directly serve as transcription factors to promote expression of neuritogenic genes. Next, we attempted to explore whether CaV1.3-encoded peptides would play an endogenous role in cortical neurons. By anti-CTD immunostaining, the subcellular distribution of CTD peptides exhibited a potential correlation with neurite outgrowth (Supplementary Fig. 13a, b). In agreement with exogeneous peptide effects, endogenous CTD in the cytosol appeared to suppress pCREB signaling (Supplementary Fig. 13c). The antibodies of anti-CTD and anti-CTC resulted into western-blot bands of differential sizes (Supplementary Fig. 14a, b and Supplementary Fig. 17a, b), supporting antibody specificity with no cross-reactivity (Supplementary Figs. 14c, 17c). Meanwhile, a bicistronic mechanism19,43 may underlie the peptide production from the full-length α1CL or α1DL transfected into HEK cells (Supplementary Figs. 14d–f, Fig. 17d–f).
a, b Inhibitory effects on pCREB and c-Fos by cytosolic peptides of exogenous CCTD. Confocal fluorescence images of cortical neurons expressing YFP, NLS-YFP-CCTD or NES-YFP-CCTD. YFP indicates CCTD distribution, and CFP illustrates cell bodies, respectively. Normalized pCREB (a) and c-Fos fluorescence (b) in the nuclei are summarized and compared. One-way ANOVA followed by Dunnett for post hoc test (a), and Kruskal-Wallis and Dunn’s non-parametric test (b, non-normal distribution, checked by D’Agostino & Pearson omnibus normality test) (*p < 0.05; ***p < 0.001; n.s. denotes not significant, p > 0.05). Values are represented as mean ± SEM.
In summary, CaV1.3-encoded DCT peptides, if exogenously expressed in the cytosol, suppressed CaV1/CREB-mediated neuritogenesis, which may represent the role of endogenous DCT peptides in cortical neurons awaiting future investigations.
CaV1/Ca2+ influx and CMI are critical to peptide distributions in neurons
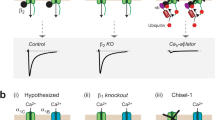
We discovered that subcellular distributions of DCT peptides are subject to autoregulation in accordance with peptide CMI, potentially through DCT inhibition of CaV1/Ca2+ influx (Fig. 5i, j). In support, nuclear export of DCT peptides is enhanced by intracellular Ca2+ rise as previously reported19,40. Ca2+ influx, via CaV1 channels in particular, is critical to subcellular distributions (or cytosol-nucleus translocation) of DCT peptides, which was confirmed in this work (Supplementary Fig. 15). When CaV1 activities under basal conditions (5 mM [K+]o) were blocked by 50 μM nifedipine (DHP derivative), the tendency of CCTD to translocate into the nucleus was significantly enhanced. In addition, N/C ratio of CCTD returned back to the control level when DHP-insensitive channels (α1DL DHP-) were employed instead (Fig. 7a, b), supporting the unique importance of CaV1 in cytosol-nucleus translocation of CCTD. In cortical neurons (supplied with 50 μM nifedipine) where endogenous CaV1 channels were replaced with DHP-insensitive channels, we further investigated whether different DCT motifs contained within the channels (through differential overall CMI) could produce any effect on peptide translocation. Four DHP-resistant CaV1.3 variants, in the order of α1DΔ, α1DL_V/A, α1DL and α1DΔ-DCTF, exhibited increasingly stronger DCT effects, i.e., leading to weaker activation/inactivation, and less fraction of apoCaM-bound channels (Fig. 7c). In this context, covalently-linked (intramolecular) DCT motifs comply with the same principle as standalone (intermolecular) DCT peptides to induce inhibitory effects (CMI), both able to elevate DCT-bound fraction (and reduce apoCaM-bound fraction) of channels, thus inhibiting Ca2+ influx and Ca2+/CaV1 signaling. Since α1DΔ-DCTF (DHP-) channels would have the largest DCT-bound fraction (fDCT) when expressing in neurons, more CCTD peptides translocated into the nucleus and exhibited the largest variation in peptide’s localization (σ of N/C ratio) (Fig. 7d). In contrast, for neurons expressing the α1DΔ DHP- channels (the weakest CMI due to completely lacking DCT), the least nuclear retention of CCTD (the smallest σ) was observed. This is presumably due to the fact that Ca2+ influx via CaV1 channels and the related Ca2+-sensitive nuclear export of CCTD would be the most pronounced for α1DΔ DHP- among the four variants.
a, b Effects of dihydropyridine (DHP). As shown in the cartoon (a, left), endogenous CaV1 channels in cortical neurons mediate Ca2+ influx (upper); DHP (50 μM nifedipine) specifically blocked cortical CaV1 channels thus reducing Ca2+ influx (middle); and overexpression of α1DL (DHP-) channels (mutant CaV1.3 insensitive to DHP) rescued Ca2+ influx in the presence of 50 μM nifedipine (bottom). Exemplary fluorescence confocal images through three channels: YFP (CCTD distribution), CFP (cell contour) and Hoechst (nuclear envelop) (a, right). N/C ratios of individual neurons from the three groups: control, DHP, and α1DL (DHP-) with DHP (b). c CaV1 variants containing DCT of different potency regulated cytosolic-nuclear distribution of CCTD. In addition to CCTD, cortical neurons were overexpressed with one of the four DHP-resistant CaV1.3 variants: α1DΔ (lacking DCT), α1DL_V/A (with the key mutation of Valine to Alanine at DCRD), α1DL (control), and α1DΔ-DCTF (chimera with ultra-strong DCT). These CaV1.3 variants were characterized by exemplar current traces from HEK293 cells expressing the variants illustrated in the cartoons. Cortical neurons were treated with 50 μM nifedipine to block endogenous CaV1 channels while sparing the exogenous DHP- CaV1.3 channels. Confocal images depict cytosolic-nuclear localizations of CCTD peptides in a similar fashion to (a). d Correlation between DCT inhibition of each CaV1.3 variant and spatial dynamics for CCTD peptides in cortical neurons. DCT/CMI potency (DCT-bound fraction fDCT) here is directly linked to inactivation (fraction of apoCaM-bound channels, FCaM): fDCT = 1 − FCaM (Eq. E5 in Methods). DCT/CMI potency and standard deviation of CCTD distribution (σ) exhibited a tight correlation (linear fit, R2 = 0.95). One-way ANOVA followed by Bonferroni for post hoc tests were used for (b) (**p < 0.01; ***p < 0.001). Data are represented as mean ± SEM.
Effective CMI (incorporating both intra- and inter-molecular CMI) could be tuned by a series of standalone DCT peptides (acting on the same channel CaV1.3, Fig. 5j), or by replacing the full-length CaV1.3 with various DCT motifs (modulated by the same peptide CCTD, Fig. 7d). Taken together, a tight correlation has been unveiled between DCT-peptide localization and CaV1-channel activities, where the central factor is overall DCT inhibition of CaV1 (effective CMI). To this point, CaV1-encoded DCT peptides comply with a type of self-regulatory scheme that DCT distributions are adjusted between the cytosol versus the nucleus through DCT inhibition of CaV1 channels.
Discussion
In this study, we systematically examined a series of DCT-encoded peptides across the CaV1 family; and by focusing on the representative CaV1.3 in cortical neurons, we unveiled that DCT peptides through CMI inhibit the signaling cascade from CaV1 to neuritogenesis. One determining factor of the overall CMI effects is the DCT affinity with CaV1, contributed by both PCRD and DCRD segments of the peptide-channel complex. In parallel, the reduction of Ca2+ influx by cytosolic DCT is in favor of nuclear localization of DCT acting as neurotrophic transcription factors. In all, CaV1 channel activities, CaV1 channel-mediated signaling to the nucleus, gene transcription related to neuritogenesis, and CaV1/Ca2+-sensitive nuclear export of DCT are all downregulated by cytosolic DCT peptides that bind and inhibit CaV1 channels (Fig. 8).
Regardless of the distinctions in mechanisms of production, molecular compositions, or modulatory effects, diverse peptides have been unified into a tuning curve of the central principle: DCT/CaV1 affinity-dependent inhibition of the channel activity-neuritogenesis coupling, which has been demonstrated by representative CaV1.3 channels in cortical neurons. A series of DCT peptide variants (color-coded to illustrate the difference in origin), by interacting with CaV1 channels, inhibit Ca2+ influx (color-coded to illustrate the difference in potency) and regulate CaV1 signaling to the nucleus and gene expression, all quantitatively in accordance with peptide/channel affinity (CMI-Kd relationship). For CaV1 channel bound with CaM at the preIQ-IQ domain, cytosolic DCT peptides compete against CaM to form peptide-channel complex. The capability of DCT competition is quantified as CMI potency to represent the fraction of channels being switched from CaM-bound to DCT-bound. DCT-channel affinities or CMI potencies are drastically different among serial peptide variants including those endogenous to native cells, varying in size (long, medium or short) and/or origin (CaV1.1-1.4). In close correlation with CMI potency, DCT peptides inhibit CaV1 gating and Ca2+ influx, reduce nuclear translocation of key signaling molecules CaM (and CaMKII), and attenuate CaV1-mediated transcription (e.g., pCREB) and expression of essential genes (e.g., c-Fos), eventually leading to inhibition of neuritogenesis (in the blue shade). As collateral effects (in the pink shade), CMI also downregulates nuclear export of DCT peptides, and thus facilitates gene transcription directly mediated by nuclear DCT peptides, leading to promotion of neurite arborization and extension. In summary, we discover cytosolic DCT inhibition of the CaV1 activity-neuritogenesis coupling, which is in direct opposition to neurotrophic signaling of nuclear DCT.
In this study, our major strategy was to utilize a series of representative DCT peptides covering the major variants across the CaV1 family. The central hypothesis has been that the DCT peptides with apparent distinctions share the same principles: the capabilities to downregulate CaV1 activity-dependent neuritogenesis via the interactions between DCT peptides and CaV1 channels. On the other hand, it would be unrealistic to exclusively examine all the signals or events along the whole pathway. Instead of multiple checkpoints with less rigor, our resolution was to focus on the key signals or indices, e.g., CaV1 gating or neurite morphology, but taking advantage of multiple peptides with a gradient of binding affinities and neuronal effects for quantitative consolidations.
Although some CaV1 inhibitors such as dihydropyridine (DHP) do exert inhibitory effects on CaV1-dependent signaling and neuritogenesis10,11, it is still a difficult task to identify effective CaV1 inhibitors for CaV1-dependent neuritogenesis. First, the linkage from Ca2+ influx to downstream signaling is not guaranteed, i.e., the potential decoupling between CaV1 channels and gene expressions, known as flux independence. In fact, the channel pore blocker Cd2+ and the gating blocker nimodipine may behave very differently in their effects on pCREB signals59. Also, instead of neuritogenic effects, CaV1 agonist Bay-K-8644 causes neural toxicity60,61. In Timothy syndrome, gain-of-function CaV1.2 mutations that promote Ca2+ influx cause neural damages due to ectopic activation of retractive signals17. Therefore, careful experiments and analyses are required to link modulation of CaV1 to neuritogenesis. Second, multiple factors besides Ca2+ influx should work synergistically to ensure the complete signaling cascade. Less noticeable factors, e.g., voltage-dependent conformational changes of α or β channel subunits, may also play important roles to ensure proper signaling from CaV1 to the nucleus59,62. Channel inhibitors which only reduce the Ca2+ influx may not attenuate neuritogenesis as effectively as expected. In this context, we have unveiled a class of CaV1-encoded peptide inhibitors endogenously present in neurons that effectively and consistently downregulate CaV1-dependent neuritogenesis, presumably by stripping apoCaM from the IQ domain of the channel.
The effects of DCT on channel gating appear to be divergent among CaV1 family members before this work. Whether CCTs could affect CaV1.1’s functions has been debated, perhaps due to different cellular environments and/or different truncation sites in these studies26,57. Moreover, for CaV1.2 channels, it has been reported that DCTC attenuates channel activation but does not affect Ca2+-dependent inactivation22, inconsistent with attenuation of inactivation evidenced from other reports24,27. In contrast, DCTD causes strong attenuation concurrently on both activation and inactivation, as the resolution of the contradictory effects on Ca2+ influx; and DCTF and DCTD resemble each other except that DCTF inhibition is of even higher potency20,21,24. Here, we have provided a tuning scheme of CMI unified across CaV1.1-1.4 DCT (Fig. 8), demonstrated by representative CaV1.3 channels, which is expandable onto CaV1.2 (Supplementary Fig. 16) and other CaV1 channels. In particular, DCRDS and DCRDC are actually able to exert inhibition (CMI) of substantial potency, as opposed to previous observations or estimations although relatively less potent than DCRDD and DCRDF. Importantly, we have clarified that the existing discrepancy in CMI potency among the DCT variants is critically dependent on the differences in PCRD isoforms. The weak effects of DCTS and DCTC are mainly attributed to PCRDS or PCRDC. Future structure-function analyses are needed to identify key PCRD residues and related mechanisms in detail. Two arginine residues reported earlier (R1696 and R1697 of CaV1.2) serve as the potential candidates on PCRD22. A few residues away from the above sites, i.e., S1575/T1579 or S1700/T1704 on CaV1.1 and CaV1.2 respectively, may provide some additional clues63,64. Our tuning curves could make the predictions for CMI and/or Kd of diverse DCT peptides in principle (Fig. 4g and Supplementary Fig. 7). For example, regarding CCTS peptides generated by cleavage of CaV1.1 in skeletal muscle36, its CMI potency by estimations would be moderate if acting on CaV1.3 (PCRDD), and weak on CaV1.1 (PCRDS) according to CMI-Kd relationship. Hence, it is unlikely that (cytosolic) CCTS could cause any strong inhibition of Ca2+ influx via CaV1.1 in smooth muscle cells, which may help elucidate the existing arguments36,57. Despite important progress in CaV1 structures, none of these structures has acquired atomic details on DCT52,53,54, which are foundational to understand DCT functions20,24. Our data here provide both the properties in common and the critical differences among DCT variants. FRET binding and electrophysiology data suggest that DCT is subject to the tight competition with CaM before the channel permits Ca2+, based on which we postulate that DCT peptides may acquire the (apo)CaM-like structures. In this context, DCRD and PCRD may mimic the C- and N-lobe of CaM respectively. First, apoCaM usually binds the target (such as the IQ domain of neuromodulin or neurogranin) with its C-lobe65; similarly, DCRD as one of the two helical subdomains plays a dominant role in the DCT/apoCaM competition (for binding the IQ domain of CaV1). In comparison, PCRD appears to be assistive, e.g., to properly anchor DCRD in the close vicinity (of the channel). Second, the interactions between the CaM-binding motif and EF-hand containing CaM-like proteins are mainly mediated by charged and aromatic/hydrophobic residues65,66, which are also similarly enriched in DCT. The functional and structural details of CMI/DCT would advance our understanding about how apoCaM binds CaV1 and promotes its functions23,67.
In this work, a neurotrophic role has been confirmed for DCT peptides localized in the nucleus; meanwhile, cytosolic DCT peptides inhibit neurite outgrowth. Potentially, the overall effects may constitute a homeostatic balance sustained by two signaling opponents (cytosolic versus nuclear) in neurite morphogenesis (Fig. 3b, Fig. 5h and Supplementary Fig. 13b). DCT/CMI inhibits the CaV1 activity-neuritogenesis coupling represented by the following crucial signals or events: translocation of CaM/CaMKII from cytosol to nucleus, phosphorylation of CREB, and transcription and expression of hallmark genes (e.g., c-Fos)11,14,49,68,69. Higher CMI potency leads to less Ca2+ influx via CaV1, eventually causing more pronounced retraction of neurites. On the other hand, less Ca2+ influx resulted from potent DCT peptides tends to cause more nuclear retention due to Ca2+-dependent nuclear export of DCT peptides. Nuclear DCT peptides as transcription factors drive the expression of a spectrum of neurotrophic genes19. Notably, DCT in the nucleus is autonomously regulated by DCT in the cytosol through its inhibition on CaV1. For example, under our experimental conditions of basal (channel/neuron) activities, CCATC exhibits ultraweak inhibition of CaV1 but with a larger fraction of cytosolic distribution, opposed to nuclear DCT peptides of a relatively smaller fraction. CCTD is much more potent in CaV1 inhibition but a relatively small cytosolic fraction (in opposition to nuclear CCTD of a larger fraction), hence the tendencies of retraction/maintenance versus outgrowth could be substantially balanced out. For CCATC or CCTD, similar autoregulatory mechanisms may account for the rather mild CMI effects on neuritogenesis (Fig. 3b). We postulate that CMI regulation of endogenous DCT distribution would be employed to maintain a delicate balance for neuritogenesis, the setting points of which may vary with developmental stages, external stimuli or cues, and actual peptide variants, in addition to expression levels. Such tuning scheme of homeostasis is expected to generally apply to other types of neurons. Besides CaV1.3 and CaV1.3-encoded DCT peptides (e.g., CCTD and CTD) as the focus of this work, CaV1.2 and CaV1.2 DCT peptides (e.g., CCATC and CCTC) are widely expressed in the brain3,43,44. Compared with CaV1.3, CaV1.2 channels should have less potent CMI effects due to the weak PCRDC motif, as another reason for this work to focus on CaV1.3. In cerebellar granule cells, CCATC peptides serve as nuclear transcription factors that promote neurite ourgrowth19. CCATC has also been evidenced in cell nuclei of the cerebellum and thalamus in embryonic brain, exporting to the cytosol along with aging and development41. CCTC by proteolysis has been found in hippocampus neurons37, and hippocampal CaV1 channels are required for normal neurogenesis45.
In addition to CaV1, CaV2.1 and CaV3.2 could encode peptides targeting the nucleus to regulate gene transcription by the bicistronic mechanism, which might be conserved across the superfamily of voltage-gated Ca2+ channels41,42,43,70. C-terminal fragments of α1A act as transcription factors to promote neuronal development42,43, resembling the effects of CaV1 DCT in the nucleus. Unlike the dual roles of CaV1 DCT peptides in this study, CaV2 CT has not been found to have any effect on channel gating20,71. Besides the autoregulatory scheme proposed here (Fig. 8), some other forms of feedbacks may also exist, e.g., CaV1.2-encoded peptides could reduce the transcription of CaV1.2 gene when located in the nucleus of cardiac myocytes38,72. Although no such downregulation has been evidenced from CaV1.3 in either recombinant systems or cortical neurons, more direct examinations are necessary to confirm the actual expression of functional CaV1.3 channels in different scenarios. CaV1-encoded polypeptides exhibit Ca2+-dependent nuclear export, with the aid of its nucleus retention domain NRD19, also supported by the distinctions between the short and long/medium peptides demonstrated in this study (Fig. 5h, i and Supplementary Fig. 11). Ca2+-dependent DCT translocation is particularly sensitive to Ca2+ via CaV1 (Fig. 7), while the exact mechanisms of DCT translocation are still awaiting future investigations19.
The specificity of the antibodies is critical given that non-specific activities of Cav antibodies are not uncommon, in part due to the low expression levels of these membrane channels compared to other proteins. In the future, rigorous validations, e.g., with CaV1.3 or CaV1.2 knock-out (ideally conditional knock-out) neurons and/or the control of blocking peptides, are expected to confirm the western-blot and immunostaining data. A set of consistent data would strengthen the conclusions, if multiple approaches including electrophysiology, biochemistry and imaging could be combined together. For future work, additional methods/tools are expected, such as CaV1.3 antibodies with knock-out validations. Meanwhile, due to the compensatory effects on Ca2+ channels in CaV1.3-/- and CaV1.2-/- mice as reported73,74, cautions also need to be taken in interpreting the data from these knock-out mice. Alternatively, knock-in mice such as CaV1.2 DHP-/- and/or CaV1.3 DHP-/- may be advantageous for the purpose of identifying and isolating CaV1.3 and CaV1.2 channels.
Our data demonstrate that inhibition of CaV1/Ca2+ influx is highly correlated with attenuation of CaV1 signaling and neuritogenesis. As mentioned earlier, for particular modulation or perturbation of CaV1 channels, it may not be as effective as expected for downstream signals and neuritogenesis. In this work, based on the fact that apoCaM/CaV1 binding is the critical linkage from channel gating to nuclear signaling20,23,75, we propose DCT/CMI as an outstanding modality with high specificity and effectiveness compared to other CaV1 inhibitors known thus far. Particularly targeting CaV1, DCT consistently generates inhibitory effects across the signaling cascade. We expect that small molecules or biologics mimicking DCT/CMI would provide new interventions for potential therapeutics of diseases related to the CaV1 activity-neuritogenesis coupling. Both CaV1 channels and neurite outgrowth are involved in a variety of neuropsychiatric and neurodegenerative diseases, such as autism, bipolar disorder, schizophrenia, Parkinson’s disease, and Alzheimer’s disease76,77,78. Ca2+ dysregulations associated with CaV have gained increasing support for its close relevance to neurodegenerative diseases, known as the ‘Ca2+ hypothesis’79,80. However, DCT peptides in these diseases are largely unexplored despite the observations indicating that the amount and distribution of DCT peptides are age-dependent19,41. In this regard, there are still unresolved questions pertaining to CaV1 channels, DCT peptides, and CaV1/DCT-dependent neuritogenesis. Exemplars of such questions include: whether and why CaV1 genes (compared with other CaV) play uniquely important roles in certain pathological processes; whether and how DCT and CMI (compared with other modulations) would play unique roles; whether DCT is prone to disease-associated mutations and how exactly mutant DCT would affect healthy neurons; and eventually what need to do to rescue the defective DCT and neurons. Notably, the expression levels of CaM are downregulated in Parkinson’s disease and Alzheimer’s disease77,81, for which the overall inhibitory effects of DCT peptides on CaV1 and CaV1-mediated downstream signals would be even more profound due to less apoCaM competition (Fig. 8).
Methods
Molecular biology
The plasmids of channels and peptides were constructed from α1S (rabbit CaV1.1, NM_001101720.1, GenbankTM accession number), α1C (human CaV1.2, AF465484.1), α1DL (human CaV1.3 α1DL_human, EU363339.1; or rat CaV1.3 chimera α1DL_rat: backbone a.a. 1-1625 from rat AF370009.1 and DCTD 1626-2155 from a.a. 1674-2203 of rat NM_017298.1), and α1F (human CaV1.4 NP005174). In particular, α1DΔ was generated by truncation of α1DL (rat AF370009.1) with a unique XbaI site following the IQ domain (ending with G1625). For chimeric α1DΔ-PCRDD and α1DΔ-DCTF, desired segments (PCRDD from 1626-1780 in α1DL_rat and DCTF from 1596-1966 in α1F) were PCR-amplified with SpeI and XbaI sites and cloned into aforementioned α1DΔ. Rat CaV1.3 DHP– was generated by single point mutation T1033Y46 on α1DΔ, α1DL_rat, α1DL_V/A (V2075A in α1DL_rat) or α1DΔ-DCTF, respectively. α1DL-Flag was generated by fusing PCR-amplified 3xFlag (DYKDHDGDYKDHDIDYKDDDDK) to the C-terminus of α1DL_rat by KpnI and SacII. α1CL-Flag was generated by replacing the EGFP in a customized pEGFP-N1 vector (modified by inserting a 3xHA tag before MCS) with PCR-amplified α1CL_human fused with 3xFlag via XhoI and NotI sites.
CFP/YFP-DCRDF in pcDNA3 were constructed as the templates for peptide plasmids. In brief, CFP or YFP was inserted into pcDNA3 vector with the unique KpnI and NotI sites, then DCRDF was fused to the C-terminus of CFP/YFP by unique NotI and XbaI sites. Other CFP/YFP-tagged constructs were generated by replacing DCRDF with appropriate PCR amplified segments, via unique NotI and XbaI sites. The constructs we have made include: YFP-DCRDF truncations, CFP-DCRDS/C/D (DCRDD from α1DL human EU363339.1), YFP-preIQ3-IQD-PCRDS/C/D/F (preIQ3-IQD from 1576-1625 and PCRDD from 1626-1780 in α1DL_rat), CFP/YFP-PCRDS/C/D/F-DCRDF (PCRDD from 1626-1780 in α1DL_rat), YFP-PCRDC-DCRDC and YFP-CCATC. DCRDS/C/D-YFP (DCRDD from α1DL_human), CCTC-YFP and CCTD-YFP (CCTD from α1DL_human) were based on another pcDNA3/YFP vector with the cloning sites of KpnI and NotI on 5’. The pcDNA3/YFP vector was made by inserting YFP into pcDNA3 vector with the unique NotI and XbaI sites. Single point mutants such as CFP/YFP-DCRDF_V/A and CCTD_V/A-YFP were made by overlap PCR. For DCT peptides to target nucleus or cytosol, nuclear localization signal (NLS) (PKKKRKV) or nuclear export signal (NES) (LALKLAGLDIGS) was fused to N-terminus of YFP-DCT peptides by overlap PCR, to achieve NLS-YFP-CCATC, NES-YFP-CCATC, NLS-YFP-CCTC, NES-YFP-CCTC, NLS-YFP-CCTD and NES-YFP-CCTD. For 3xFlag-DCTC/D, 3xFlag tag was firstly inserted into pcDNA3 vector with the unique KpnI and NotI sites, then DCTC/D (from α1CL_human or α1DL_human) peptides were PCR-amplified with NotI and XbaI sites and fused to the C-terminus of 3xFlag.
Dissection and culturing of cortical neurons
Cortical neurons were dissected from postnatal day 0 (P0, either sex) newborn ICR mice. Isolated cortex tissues were digested with 0.25% trypsin for 15 min at 37 °C, followed by terminating the enzymatic reaction by DMEM supplemented with 10% FBS. The suspension of cells was sieved through a filter then centrifuged at 1000 rpm for 5 minutes. The cell pellet was resuspended in DMEM supplemented with 10% FBS and then plated on poly-D-lysine-coated 35-mm No. 0 confocal dishes (In Vitro Scientific) or poly-D-lysine-coated coverslips. After 4 hours, neurons were maintained in Neurobasal medium supplemented with 2% B27, 1% glutaMAX-I (growth medium). Temperature of 37 °C with 5% CO2 was controlled in the incubator. All animals were obtained from the laboratory animal research centers at Tsinghua University and Peking University. Procedures involving animals have been approved by local institutional ethical committees of Tsinghua University and Beihang University.
Transfection of cDNA constructs in cell lines and cultured neurons
For electrophysiological recording, HEK293 cells (ATCC), checked by PCR with primers 5′-GGCGAATGGGTGAGTAACACG-3′ and 5′-CGGATAACGCTTGCGACCTATG-3′ to ensure free of mycoplasma contamination, were cultured in 60-mm dishes, and recombinant channels were transiently transfected according to established calcium phosphate protocol20,24. 5 μg of cDNA encoding channel α1 subunit, along with 4 μg of rat brain β2a (M80545) and 4 μg of rat brain α2δ (NM012919.2) subunits were applied to HEK293 cells. To enhance expression, cDNA for simian virus 40 T antigen (1 μg) was also co-transfected. For each additional construct, 2 μg cDNA was added. All of the above cDNA constructs were driven by a cytomegalovirus (CMV) promoter. Cells were washed with PBS 6–8 h after transfection and maintained in supplemented DMEM, then incubated for at least 48 h in a water-saturated 5% CO2 incubator at 37 °C before usage.
For transfection in neurons, 2 μg of cDNA encoding the desired peptides were transiently transfected by Lipofectamine 2000 (Invitrogen) for each confocal dish with a typical protocol according to the manual. The mixture of plasmids and Lipofectamine 2000 in opti-MEM was added to the Neurobasal medium for transfection. After 2 hours, neurons were maintained in Neurobasal medium supplemented with 2% B27, 1% glutaMAX-I for 48 hours.
For 2-hybrid 3-cube FRET experiments, HEK293 cells were cultured on confocal dishes. FRET cDNA constructs of 2 μg each were transfected by Lipofectamine 2000 for 6 hours. Cells were used after 24 hours.
For western blot experiments, CHO (Cell Resource Center, IBMS, CAMS/PUMC) or HEK293 cells were cultured on 60 mm dishes. cDNA constructs were transfected by Lipofectamine for at least 6 hours. Cells were collected after 2 days.
Whole-cell electrophysiology
Whole-cell recordings of transfected HEK293 cells were performed at room temperature (25 °C) using an Axopatch 200B amplifier (Molecular Devices). Electrodes were pulled with borosilicate glass capillaries by a programmable puller (P-1000, Sutter Instrument) and heat-polished by a microforge (MF-830, Narishige), resulting in 2–5 MΩ resistances before 70% of compensation. The internal/pipette solution contained (in mM): CsMeSO3, 135; CsCl, 5; MgCl2, 1; MgATP, 4; HEPES, 5; and EGTA, 5; with ~290 mOsm adjusted with glucose and pH 7.3 adjusted with CsOH. The extracellular/bath solution contained (in mM): TEA-MeSO3, 135; HEPES, 10; CaCl2 or BaCl2, 10; with ~300 mOsm, adjusted with glucose and pH 7.3 adjusted with TEA-OH, similar to the previous protocols24. Whole-cell currents were generated from a family of step depolarizations (−70 to +50 mV from a holding potential of −70 mV and step increase of 10 mV). Current traces were recorded at 2 kHz low-pass filtering in response to voltage steps with minimum interval of 30 s. P/8 leak subtraction was used throughout. Ca2+ current was normalized over different cells by cell capacitance (Cm, in pF), and the current amplitude (peak, 50 ms or 300 ms, in pA/pF) was measured at −10mV.
Neuronal patch-clamp recording was performed according to our previous protocol51. In brief, isolated cortical neurons were cultured in coverslips. To record neuronal CaV1.3 current, neurons were pre-incubated in Tyrode’s solution containing 1 μM nimodipine (Sigma-Aldrich), 1 μM ω-conotoxin GVIA (Sigma-Aldrich, or alomone labs) and 1 μM ω-conotoxin MVIIC (Sigma-Aldrich, or alomone labs) for 30 min to block endogenous CaV1.2, N- and P/Q-type Ca2+ current, according to the cocktail recipes75,82,83. Under the conditions of our evaluations (−10 mV, full cocktail recipes), CaV1.3 current appeared to be the dominant component (~80%) after the treatment, and CaV2.3 (16%) and CaV1.2 (4%) contributed to the rest (details see Supplementary Fig. 2). The voltage ramp protocol (holding at −60 mV, ramping from −60 to +50 mV at 0.2 mV/ms) was applied to cortical neurons in the bath solution containing 10 mM Ba2+. The resulted I-V curves were fitted by Boltzmann-based equations (OriginPro) to obtain the half-activation voltage (Vhalf) of voltage-dependent channel activation. Isradipine (Sigma-Aldrich) at 100 nM was used to further isolate/confirm CaV1.3 currents, and 20 μM isradipine would eliminate all the CaV1 (CaV1.2 and CaV1.3) currents84. Treated neurons were recorded in various bath solutions containing appropriate blockers within one hour.
2-hybrid 3-cube FRET
2-hybrid 3-cube FRET experiments were carried out with standard protocols similarly shared by several groups20,24,56. Briefly, experiments were performed on an inverted epi-fluorescence microscope (Ti-U, Nikon), with computer-controlled filter wheels (Sutter Instrument) to coordinate with diachronic mirrors for appropriate imaging at excitation, emission, and FRET channels. The filters used in the experiments were excitation: 438/24 (FF01-438/24-25, Semrock) and 480/30 (FITC, Nikon); emission: 483/32 (FF01-483/32-25, Semrock) and 535/40 (FITC, Nikon); dichroic mirrors: 458 nm (FF458-Di02-25 × 36, Semrock) and 505 nm (FITC, Nikon). Fluorescence images were acquired by Neo sCMOS camera (Andor Technology) and analyzed with 33-FRET algorithms coded in Matlab (Mathworks), mainly based on the following formula:
FRmax represents the maximum FRET ratio, and Dfree denotes the equivalent free donor (CFP-tagged) concentration. Kd (effective dissociation equilibrium constant) is calculated from an iterative procedure to evaluate the binding affinity for each pair of binding partners. FRET imaging experiments were performed with HEK293 cells in Tyrode’s buffer containing 2 mM Ca2+.
Confocal fluorescence imaging and analysis
Cultured neurons were transfected with CFP (to label the soma area and neurites) and DCT peptides tagged with YFP on 5th day (DIV-5) and used on DIV-7, or transfected on DIV 12-15 and used on DIV 15–18 (Supplementary Fig. 10). Neurons were loaded with Hoechst 33342 for 5 min to label the nuclei and then imaged by Zeiss LSM710 confocal Scanning Microscope. Fluorescent intensity was quantified and analyzed with ImageJ (NIH). Calculation of nuclear intensity was based on the nuclear contour indicated by Hoechst 33342. Cytosolic intensity was calculated by intermediate region between nucleus and plasma membrane. N/C ratio of DCT peptides was calculated by the ratio of fluorescence intensity (nuclear/cytosolic). Measurements of the total length and Sholl analysis for neurites were performed with Imaris 7.7.2 (Bitplane) through CFP channel. Only non-overlapping neurons were selected for analysis of morphogenesis. Neurite tracings were depicted with Imaris 7.7.2 and further processed with Photoshop 7.0 (Adobe).
To observe the cytosolic-nuclear translocation of DCT peptides, neurons were pre-incubated in 5 mM [K+]o solution (130 mM NaCl, 5 mM KCl, 1 mM MgCl2, 15 mM HEPES, 2 mM CaCl2, at 300 mOsm adjusted with glucose) and perfused with 40 mM [K+]o solution (95 mM NaCl, 40 mM KCl, 1 mM MgCl2, 15 mM HEPES, 2 mM CaCl2, at 300 mOsm adjusted with glucose) or 5 mM [K+]o with 50 μM Nifedipine for 0.5-1 hour, then washed out by 5 mM [K+]o when needed. For the experiments with DHP-insensitive variants of CaV1, neurons were incubated with 50 μM Nifedipine for at least 1 hour, and neurons without clear damages were selected to calculate N/C ratio for the peptides.
Analyses on neurite morphology and cytosolic-nuclear translocation were performed over cultured neurons from at least two culture preparations and two independent experiments, adding up to the total number for each data group (20 cells or more).
Immunocytochemistry
DIV-5 cultured cortical neurons were transfected with DCRDF and used on DIV-7. Firstly, to stimulate neurons, 1 μM TTX (sodium channel blocker) was applied to neurons for 6 hours to suppress action potential. 40 mM [K+]o solution containing 1 μM TTX was applied for 5 min for CaM staining, or 30 min for pCREB staining before fixation. To measure desired signals in physiological condition, neurons were maintained in growth medium until fixation. Secondly, neurons were rapidly rinsed with ice-cold PBS and fixed with ice-cold 4% paraformaldehyde in PBS (pH 7.4) for 15–20 min. Fixed neurons were washed by ice-cold PBS for 3 times and permeabilized with 0.3% Trition X-100 for 5 minutes. Then neurons were blocked by 10% normal goat serum in PBS for 1 hour and incubated with the primary antibodies overnight at 4 oC. The following antibodies were used: CaM (Rabbit mAb #5197-1, Epitomics, Species Cross-Reactivity: Human, Mouse, Rat, Dilutions: 1:500 in PBS)51, pCREB (Rabbit mAb #9198, Cell Signaling Technology, Species Cross-Reactivity: Human, Mouse, Rat, Dilutions: 1:500 in PBS)49,51, c-Fos (Rabbit mAb [EPR21930-238] #ab222699, Abcam, Species Cross-Reactivity: Mouse, Human, Dilutions: 1:1000 in PBS, https://www.abcam.com/nav/primary-antibodies/rabbit-monoclonal-antibodies/c-fos-antibody-epr21930-238-ab222699.html), and CaV1.3 CT (a.a. 2025-2161) (Mouse mAb [N38/8] #ab84811, Abcam, Species Cross-Reactivity: Mouse, Rat, Rabbit, Human, Dilution: 1:500 in PBS)85. Finally, the next day, neurons were washed with PBS for 3 times and incubated with the secondary antibodies (Goat anti-Rabbit Alexa Fluor 647, Invitrogen, Dilutions: 1:800 in PBS; Goat anti-Mouse Alexa Fluor 568, Invitrogen, Dilutions: 1:800 in PBS; Goat anti-Mouse IgG(H+L)-DyLight 488, Gene-Protein Link, Dilutions: 1:500) for 2 hours. Then neurons were washed with PBS for 3 times and treated with Hoechst 33342 (Invitrogen) for 5 min for nuclear counterstain. Mounted neurons on confocal dishes were imaged with a confocal microscope (LSM710, Carl Zeiss) and ZEN software. Nuclear and cytosolic fluorescence intensities of endogenous CaM in cortical neurons were analyzed by ImageJ (NIH). Neuronal culture preparations of each round supplied one or two independent experiments. Neurons (>15) were assayed or evaluated from at least two culture preparations and two independent experiments unless otherwise noted.
Western blot
Cortical neurons for western blot were isolated from newborn ICR mice or from cultured cortical neurons around DIV-14. Liver tissues were from newborn ICR mice. HEK293 Cells and CHO (Chinese Hamster Ovary) cells served as the blank control or the recombinant system to express proteins. Cells lysates were prepared by lysis buffer RIPA (with protease inhibitor cocktail, cat#P1265, Applygen Tech) and centrifuged at 15,000 × g. Proteins were separated using 8% sodium dodecyl sulphate polyacrylamide gel electrophoresis and transferred to a nitrocellulose membrane for 100 min. Then nitrocellulose membrane was blocked in 5% non-fat dry milk and incubated with primary antibodies: anti-CaV1.3 CT, identical to the antibody used in immunocytochemistry, Dilutions: 1:1000; anti-CaV1.2 CT (a.a. 1835-2135) (Rabbit pAb #21774-1-AP, proteintech, Species Cross-Reactivity: human, mouse, rat, Dilutions: 1:1000, https://www.ptgcn.com/products/L-VOCC-Antibody-21774-1-AP.htm); anti-CaV1.2 II-III loop (#ACC-003, allomone, Source: Rabbit, Species Cross-Reactivity: Human, Mouse, Rat, Dilutions: 1:1000, https://www.alomone.com/p/anti-cav1-2-antibody/ACC-003), anti-Flag (#20543-1-AP, proteintech, Source: Rabbit, Species Cross-Reactivity: recombinant protein with Flag tag, Dilutions: 1:3000); and anti-GAPDH (#P01L081, Gene-Protein Link, Source: Rabbit, Species Cross-Reactivity: Human, Mouse, Rat, Bovine, Pig, Chicken, Zebrafish, Green Monkey, Dilutions: 1:5000) overnight at 4 °C. Next, the nitrocellulose membrane was washed three times with TBST and incubated with secondary antibody (Goat anti-Mouse, # SA00001-1, Proteintech, Dilution: 1:3000; Goat anti-Rabbit, #ZRA03, YTHXbio, Dilution: 1:5000) for 1–2 h and then washed with TBST for three times again. The membrane was coved with ECL chemiluminescent liquid (#P1010, Applygen Tech) before detection with an enhanced chemiluminescence system. Three or more independent replicates were performed for each experiment.
Definition of CMI potency and related curve fitting
For α1DL channels with covalently-linked DCT (Figs. 2, 7), there are two subgroups: DCT-bound and apoCaM-bound. For these channels subject to modulation by standalone DCT peptides, CMI potency (CMI, in percentage) is quantified as the normalized fractional change of apoCaM-bound channels switching to peptide-bound channels (fPeptide):
Since apo-CaM bound fraction (FCaM) is proportional to Ca2+-dependent inactivation (SCa)20, we have:
Hence, CMI potency is inversely and linearly correlated with actual inactivation of channels under peptide modulation. Meanwhile, interactions between peptides and channels should follow the relationship defined by the classical ligand-binding equation:
Here, [Peptide] is the concentration of DCT peptide, and Kd is the dissociation constant of peptide interactions with channels. Fluorescence intensities (a.u., arbitrary units) are used to calculate [Peptide] and Kd. In our electrophysiology experiments, patch-clamped cells are supposed to have comparable (over)expression levels of peptides, thus Eq. E4 represents the relationship between CMI and Kd across a series of peptide variants (Fig. 4c). The concentration of DCT peptides ([Peptide]) on average was estimated to be ~23,000 (a.u.), by directly applying Eq. E4 to fit Fig. 4c. To estimate the binding affinities between YFP-preIQ3-IQD and CFP-PCRDX-DCRDF, we proceeded to perform FRET experiments between YFP-preIQ3-IQD-PCRDX and CFP-DCRDF. The reason for taking such indirect approach is in part due to severe aggregation (puncta) for YFP-preIQ3-IQD (but not for YFP-preIQ3-IQD-PX) when overexpressed in cells. Correspondingly, CMI values of PCRDX-DCRDF peptides to α1DΔ were from Fig. 4e. Kd for IQD and PD-DF was estimated to be 25-fold of the Kd value listed for IQD-PD and DF in Fig. 4c. Similar calculations (25-fold) can be conducted to estimate Kd values for other peptides in Fig. 4f from Supplementary Fig. 7.
In Fig. 4g, results from Fig. 4c, f were combined and plotted together with the tuning curve (Eq. E4, ([Peptide] = 23,000). In addition, compound CMI effects for PD+PF-DF, PD+PC-DF, and PD+PC-DC were also illustrated, based on CMI values from electrophysiology and estimated Kd values by interpolation.
Regarding full-length channels with covalently-linked DCT of its own (such as α1DL and DCT mutants), DCT potency (of competing with apoCaM to bind the channel) can be derived from Eqs. E2 and E3 into similar formulas for the intramolecular CMI by the covalently-linked DCT (Figs. 5j, 7d). Considering α1DΔ as the control (100% apoCaM-bound), (CMI) potency of linked DCT domains represents the fraction of channels bound with DCT (fDCT) for particular channel/DCT variant. We have
In principle, CMI refers to fractional changes of channels switched from apoCaM-bound to apoCaM-unbound (i.e., DCT-bound), which could be achieved by standalone DCT peptides and/or covalently-linked DCT domains in a similar fashion.
Statistics and reproducibility
Data were analyzed in Matlab, OriginPro and GraphPad Prism software. Data were shown as mean ± SEM (Standard Error of the Mean). Unpaired or paired Student’s t-test (two-tailed with criteria of significance) was performed to compare two groups. One-way ANOVA followed by Dunnett or Bonferroni for post hoc tests were performed to compare more than two groups with or without a restrictive control group, respectively, provided the normal distribution of the data. Kruskal-Wallis and Dunn’s non-parametric test was performed if the data did not follow the normal distribution. D’Agostino & Pearson omnibus normality test was used before column analyses. Significance *p < 0.05; **p < 0.01; ***p < 0.001 and n.s. denotes ‘not significant’.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The plasmids of pcDNA3-NLS-YFP-CCTD (#184325), pcDNA3-NES-YFP-CCTD (#184326), pcDNA3-NLS-YFP-CCTC (#184327) and pcDNA3-NES-YFP-CCTC (#184328) are available on Addgene. Source data underlying the figures are organized as Supplementary Data 1. Uncropped blots are presented in Supplementary Fig. 17. The data in details associated with the main figures have been deposited to Dryad (https://doi.org/10.5061/dryad.cvdncjt63)86. Other data and information are available from the corresponding author upon reasonable request.
References
Catterall, W. A. Voltage-gated calcium channels. Cold Spring Harb. Perspect. Biol. 3, a003947 (2011).
Berridge, M. J., Bootman, M. D. & Roderick, H. L. Calcium signalling: dynamics, homeostasis and remodelling. Nat. Rev. Mol. cell Biol. 4, 517–529 (2003).
Simms, B. A. & Zamponi, G. W. Neuronal voltage-gated calcium channels: structure, function, and dysfunction. Neuron 82, 24–45 (2014).
Ma, H., Cohen, S., Li, B. & Tsien, R. W. Exploring the dominant role of Cav1 channels in signalling to the nucleus. Biosci. Rep. 33, 97–101 (2012).
Wong, R. O. L. & Ghosh, A. Activity-dependent regulation of dendritic growth and patterning. Nat. Rev. 3, 803–812 (2002).
Dolmetsch, R. Excitation-transcription coupling: signaling by ion channels to the nucleus. Sci. STKE 2003, PE4 (2003).
Deisseroth, K. et al. Excitation-neurogenesis coupling in adult neural stem/progenitor cells. Neuron 42, 535–552 (2004).
Kamijo, S. et al. A Critical Neurodevelopmental Role for L-Type Voltage-Gated Calcium Channels in Neurite Extension and Radial Migration. J. Neurosci. 38, 5551–5566 (2018).
Gomez, T. M. & Zheng, J. Q. The molecular basis for calcium-dependent axon pathfinding. Nat. Rev. Neurosci. 7, 115–125 (2006).
Redmond, L., Kashani, A. H. & Ghosh, A. Calcium regulation of dendritic growth via CaM kinase IV and CREB-mediated transcription. Neuron 34, 999–1010 (2002).
Wheeler, D. G. et al. Ca(V)1 and Ca(V)2 channels engage distinct modes of Ca(2+) signaling to control CREB-dependent gene expression. Cell 149, 1112–1124 (2012).
Andrade A., et al. Genetic Associations between Voltage-Gated Calcium Channels and Psychiatric Disorders. Int. J. Mol. Sci. 20, 3537 (2019).
Forrest, M. P., Parnell, E. & Penzes, P. Dendritic structural plasticity and neuropsychiatric disease. Nat. Rev. Neurosci. 19, 215–234 (2018).
Wheeler, D. G., Barrett, C. F., Groth, R. D., Safa, P. & Tsien, R. W. CaMKII locally encodes L-type channel activity to signal to nuclear CREB in excitation-transcription coupling. J. Cell Biol. 183, 849–863 (2008).
Sheng, M. & Greenberg, M. E. The regulation and function of c-fos and other immediate early genes in the nervous system. Neuron 4, 477–485 (1990).
Gil, G. A. et al. c-Fos activated phospholipid synthesis is required for neurite elongation in differentiating PC12 cells. Mol. Biol. Cell 15, 1881–1894 (2004).
Krey, J. F. et al. Timothy syndrome is associated with activity-dependent dendritic retraction in rodent and human neurons. Nat. Neurosci. 16, 201–209 (2013).
Wild, A. R. et al. Synapse-to-Nucleus Communication through NFAT Is Mediated by L-type Ca2+ Channel Ca2+ Spike Propagation to the Soma. Cell Rep. 26, 3537–3550.e3534 (2019).
Gomez-Ospina, N., Tsuruta, F., Barreto-Chang, O., Hu, L. & Dolmetsch, R. The C terminus of the L-type voltage-gated calcium channel Ca(V)1.2 encodes a transcription factor. Cell 127, 591–606 (2006).
Liu, X., Yang, P. S., Yang, W. & Yue, D. T. Enzyme-inhibitor-like tuning of Ca(2+) channel connectivity with calmodulin. Nature 463, 968–972 (2010).
Singh, A. et al. C-terminal modulator controls Ca2+-dependent gating of Ca(v)1.4 L-type Ca2+ channels. Nat. Neurosci. 9, 1108–1116 (2006).
Hulme, J. T., Yarov-Yarovoy, V., Lin, T. W., Scheuer, T. & Catterall, W. A. Autoinhibitory control of the CaV1.2 channel by its proteolytically processed distal C-terminal domain. J. Physiol. 576, 87–102 (2006).
Adams, P. J., Ben-Johny, M., Dick, I. E., Inoue, T. & Yue, D. T. Apocalmodulin itself promotes ion channel opening and Ca(2+) regulation. Cell 159, 608–622 (2014).
Liu N., et al. Cooperative and acute inhibition by multiple C-terminal motifs of L-type Ca2+ channels. Elife 6, e21989 (2017).
Kuzmenkina, E., Novikova, E., Jangsangthong, W., Matthes, J. & Herzig, S. Single-Channel Resolution of the Interaction between C-Terminal CaV1.3 Isoforms and Calmodulin. Biophys. J. 116, 836–846 (2019).
Morrill, J. A. & Cannon, S. C. COOH-terminal truncated alpha(1S) subunits conduct current better than full-length dihydropyridine receptors. J. Gen. Physiol. 116, 341–348 (2000).
Sang, L., Dick, I. E. & Yue, D. T. Protein kinase A modulation of CaV1.4 calcium channels. Nat. Commun. 7, 12239 (2016).
Bock, G. et al. Functional properties of a newly identified C-terminal splice variant of Cav1.3 L-type Ca2+ channels. J. Biol. Chem. 286, 42736–42748 (2011).
Wei, X. et al. Modification of Ca2+ channel activity by deletions at the carboxyl terminus of the cardiac alpha 1 subunit. J. Biol. Chem. 269, 1635–1640 (1994).
Gerhardstein, B. L. et al. Proteolytic processing of the C terminus of the alpha(1C) subunit of L-type calcium channels and the role of a proline-rich domain in membrane tethering of proteolytic fragments. J. Biol. Chem. 275, 8556–8563 (2000).
Sang. L., Vieira D. C. O., Yue D. T., Ben-Johny M. & Dick I. E. The molecular basis of the inhibition of CaV1 calcium dependent inactivation by the distal carboxy tail. J. Biol. Chem. 296, 100502 (2021).
Tang, F., Dent, E. W. & Kalil, K. Spontaneous calcium transients in developing cortical neurons regulate axon outgrowth. J. Neurosci. 23, 927–936 (2003).
Abele, K. & Yang, J. Regulation of voltage-gated calcium channels by proteolysis. Sheng li xue bao: [Acta physiologica Sin.] 64, 504–514 (2012).
De Jongh, K. S., Warner, C., Colvin, A. A. & Catterall, W. A. Characterization of the two size forms of the alpha 1 subunit of skeletal muscle L-type calcium channels. Proc. Natl Acad. Sci. USA 88, 10778–10782 (1991).
De Jongh, K. S., Colvin, A. A., Wang, K. K. & Catterall, W. A. Differential proteolysis of the full-length form of the L-type calcium channel alpha 1 subunit by calpain. J. Neurochem 63, 1558–1564 (1994).
Hulme, J. T. et al. Sites of proteolytic processing and noncovalent association of the distal C-terminal domain of CaV1.1 channels in skeletal muscle. Proc. Natl Acad. Sci. USA 102, 5274–5279 (2005).
Hell, J. W. et al. N-methyl-D-aspartate receptor-induced proteolytic conversion of postsynaptic class C L-type calcium channels in hippocampal neurons. Proc. Natl Acad. Sci. USA 93, 3362–3367 (1996).
Schroder, E., Byse, M. & Satin, J. L-type calcium channel C terminus autoregulates transcription. Circ. Res. 104, 1373–1381 (2009).
Yang, L. et al. beta-adrenergic regulation of the L-type Ca2+ channel does not require phosphorylation of alpha1C Ser1700. Circ. Res. 113, 871–880 (2013).
Lu, L. et al. Regulation of gene transcription by voltage-gated L-type calcium channel, Cav1.3. J. Biol. Chem. 290, 4663–4676 (2015).
Gomez-Ospina, N. et al. A promoter in the coding region of the calcium channel gene CACNA1C generates the transcription factor CCAT. PLoS One 8, e60526 (2013).
Du, X. et al. Second cistron in CACNA1A gene encodes a transcription factor mediating cerebellar development and SCA6. Cell 154, 118–133 (2013).
Du, X. et al. alpha1ACT Is Essential for Survival and Early Cerebellar Programming in a Critical Neonatal Window. Neuron 102, 770–785 e777 (2019).
Hell, J. W. et al. Identification and differential subcellular localization of the neuronal class C and class D L-type calcium channel alpha 1 subunits. J. Cell Biol. 123, 949–962 (1993).
Marschallinger, J. et al. The L-type calcium channel Cav1.3 is required for proper hippocampal neurogenesis and cognitive functions. Cell Calcium 58, 606–616 (2015).
Zhang, H. et al. Association of Ca(V)1.3 L-type calcium channels with shank. J. Neurosci. 25, 1037–1049 (2005).
Zhang, H. et al. Ca1.2 and CaV1.3 neuronal L-type calcium channels: differential targeting and signaling to pCREB. Eur. J. Neurosci. 23, 2297–2310 (2006).
Kim, S. et al. Functional interaction of neuronal Cav1.3 L-type calcium channel with ryanodine receptor type 2 in the rat hippocampus. J. Biol. Chem. 282, 32877–32889 (2007).
Ma, H. et al. gammaCaMKII shuttles Ca(2)(+)/CaM to the nucleus to trigger CREB phosphorylation and gene expression. Cell 159, 281–294 (2014).
Deisseroth, K., Heist, E. K. & Tsien, R. W. Translocation of calmodulin to the nucleus supports CREB phosphorylation in hippocampal neurons. Nature 392, 198–202 (1998).
Yang, Y. X. et al. Improved calcium sensor GCaMP-X overcomes the calcium channel perturbations induced by the calmodulin in GCaMP. Nat. Commun. 9, 18 (2018).
Wu, J. et al. Structure of the voltage-gated calcium channel Cav1.1 complex. Science 350, aad2395 (2015).
Wu, J. et al. Structure of the voltage-gated calcium channel Ca(v)1.1 at 3.6 A resolution. Nature 537, 191–196 (2016).
Zhao, Y. et al. Molecular Basis for Ligand Modulation of a Mammalian Voltage-Gated Ca(2+) Channel. Cell 177, 1495–1506 e1412 (2019).
Peterson, B. Z., DeMaria, C. D., Adelman, J. P. & Yue, D. T. Calmodulin is the Ca2+ sensor for Ca2+ -dependent inactivation of L-type calcium channels. Neuron 22, 549–558 (1999).
Butz, E. S. et al. Quantifying macromolecular interactions in living cells using FRET two-hybrid assays. Nat. Protoc. 11, 2470–2498 (2016).
Ohrtman, J. D. et al. Apparent lack of physical or functional interaction between CaV1.1 and its distal C terminus. J. Gen. Physiol. 145, 303–314 (2015).
Stroffekova, K. Ca2+/CaM-dependent inactivation of the skeletal muscle L-type Ca2+ channel (Cav1.1). Pflug. Arch. 455, 873–884 (2008).
Li, B., Tadross, M. R. & Tsien, R. W. Sequential ionic and conformational signaling by calcium channels drives neuronal gene expression. Science 351, 863–867 (2016).
Barger, S. W. Complex influence of the L-type calcium-channel agonist BayK8644(+/-) on N-methyl-D-aspartate responses and neuronal survival. Neuroscience 89, 101–108 (1999).
Jinnah, H. A. et al. Calcium channel agonists and dystonia in the mouse. Mov. Disord. 15, 542–551 (2000).
Servili, E., Trus, M., Maayan, D. & Atlas, D. beta-Subunit of the voltage-gated Ca(2+) channel Cav1.2 drives signaling to the nucleus via H-Ras. Proc. Natl Acad. Sci. USA 115, E8624–E8633 (2018).
Emrick, M. A., Sadilek, M., Konoki, K. & Catterall, W. A. Beta-adrenergic-regulated phosphorylation of the skeletal muscle Ca(V)1.1 channel in the fight-or-flight response. Proc. Natl Acad. Sci. USA 107, 18712–18717 (2010).
Fuller, M. D., Emrick, M. A., Sadilek, M., Scheuer, T. & Catterall, W. A. Molecular mechanism of calcium channel regulation in the fight-or-flight response. Sci. Signal 3, ra70 (2010).
Kumar, V. et al. Structural basis for the interaction of unstructured neuron specific substrates neuromodulin and neurogranin with Calmodulin. Sci. Rep. 3, 1392 (2013).
Van Petegem, F., Chatelain, F. C. & Minor, D. L. Jr. Insights into voltage-gated calcium channel regulation from the structure of the CaV1.2 IQ domain-Ca2+/calmodulin complex. Nat. Struct. Mol. Biol. 12, 1108–1115 (2005).
Ben-Johny, M. & Yue, D. T. Calmodulin regulation (calmodulation) of voltage-gated calcium channels. J. Gen. Physiol. 143, 679–692 (2014).
Impey, S. et al. Defining the CREB regulon: a genome-wide analysis of transcription factor regulatory regions. Cell 119, 1041–1054 (2004).
Cohen, S. M. et al. Calmodulin shuttling mediates cytonuclear signaling to trigger experience-dependent transcription and memory. Nat. Commun. 9, 2451 (2018).
Kordasiewicz, H. B., Thompson, R. M., Clark, H. B. & Gomez, C. M. C-termini of P/Q-type Ca2+ channel alpha1A subunits translocate to nuclei and promote polyglutamine-mediated toxicity. Hum. Mol. Genet 15, 1587–1599 (2006).
Lubbert, M. et al. A novel region in the CaV2.1 alpha1 subunit C-terminus regulates fast synaptic vesicle fusion and vesicle docking at the mammalian presynaptic active zone. Elife 6, e28412 (2017).
Satin, J., Schroder, E. A. & Crump, S. M. L-type calcium channel auto-regulation of transcription. Cell Calcium 49, 306–313 (2011).
Namkung, Y. et al. Requirement for the L-type Ca(2+) channel alpha(1D) subunit in postnatal pancreatic beta cell generation. J. Clin. Investig. 108, 1015–1022 (2001).
Xu, M., Welling, A., Paparisto, S., Hofmann, F. & Klugbauer, N. Enhanced expression of L-type Cav1.3 calcium channels in murine embryonic hearts from Cav1.2-deficient mice. J. Biol. Chem. 278, 40837–40841 (2003).
Dolmetsch, R. E., Pajvani, U., Fife, K., Spotts, J. M. & Greenberg, M. E. Signaling to the nucleus by an L-type calcium channel-calmodulin complex through the MAP kinase pathway. Science 294, 333–339 (2001).
Smoller, J. W. et al. Identification of risk loci with shared effects on five major psychiatric disorders: a genome-wide analysis. Lancet 381, 1371–1379 (2013).
Hurley, M. J., Brandon, B., Gentleman, S. M. & Dexter, D. T. Parkinson’s disease is associated with altered expression of CaV1 channels and calcium-binding proteins. Brain 136, 2077–2097 (2013).
Anekonda, T. S. et al. L-type voltage-gated calcium channel blockade with isradipine as a therapeutic strategy for Alzheimer’s disease. Neurobiol. Dis. 41, 62–70 (2011).
Chan, C. S. et al. ‘Rejuvenation’ protects neurons in mouse models of Parkinson’s disease. Nature 447, 1081–1086 (2007).
Chakroborty, S. & Stutzmann, G. E. Early calcium dysregulation in Alzheimer’s disease: setting the stage for synaptic dysfunction. Sci. China Life Sci. 54, 752–762 (2011).
McLachlan, D. R., Wong, L., Bergeron, C. & Baimbridge, K. G. Calmodulin and calbindin D28K in Alzheimer disease. Alzheimer Dis. associated Disord. 1, 171–179 (1987).
Gao, L., Blair, L. A., Salinas, G. D., Needleman, L. A. & Marshall, J. Insulin-like growth factor-1 modulation of CaV1.3 calcium channels depends on Ca2+ release from IP3-sensitive stores and calcium/calmodulin kinase II phosphorylation of the alpha1 subunit EF hand. J. Neurosci. 26, 6259–6268 (2006).
Hall, D. D. et al. Competition between alpha-actinin and Ca(2)(+)-calmodulin controls surface retention of the L-type Ca(2)(+) channel Ca(V)1.2. Neuron 78, 483–497 (2013).
Koschak, A. et al. alpha 1D (Cav1.3) subunits can form l-type Ca2+ channels activating at negative voltages. J. Biol. Chem. 276, 22100–22106 (2001).
Chovancova, B. et al. Calcium signaling affects migration and proliferation differently in individual cancer cells due to nifedipine treatment. Biochem Pharm. 171, 113695 (2020).
Yang, Y., Yu Z., Geng, J., Liu, X. Cytosolic peptides encoding CaV1 C-termini downregulate the calcium channel activity-neuritogenesis coupling. Dryad, https://doi.org/10.5061/dryad.cvdncjt5063 (2022).
Acknowledgements
We thank all X-Lab members for discussions and help. This work is supported by grants from Natural Science Foundation of China (81971728 and 21778034 for XDL, 11902021 for YXY, U20A20390 and 11827803 for YBF) and Natural Science Foundation of Beijing Municipality (7191006 for XDL, 5204037 for YXY) and China Postdoctoral Science Foundation (BX20180027 and 2018M641146 for YXY), and by an open fund from Laboratory for Biomedical Engineering of Ministry of Education, Zhejiang University.
Author information
Authors and Affiliations
Contributions
X.D.L. and Y.X.Y. conceived the project; X.D.L., Y.B.F., X.M.L., S.S., and P.W. provided general help; M.L. and N.L. made important contributions by conducting the pilot experiments and analyses; Y.X.Y., Z.Y., and J.L.G. performed the experiments; Y.X.Y. and X.D.L analyzed the data; P.L., W.L.H., S.H.Y., H.J., H.Y.G., F.Q., and W.X. provided technical assistance; X.D.L. and Y.X.Y. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Boxing Li, Amy Lee and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: George Inglis. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yang, Y., Yu, Z., Geng, J. et al. Cytosolic peptides encoding CaV1 C-termini downregulate the calcium channel activity-neuritogenesis coupling. Commun Biol 5, 484 (2022). https://doi.org/10.1038/s42003-022-03438-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-022-03438-1
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.