Abstract
The dysregulation of gene dosage due to duplication or haploinsufficiency is a major cause of autosomal dominant diseases such as Alzheimer’s disease. However, there is currently no rapid and efficient method for manipulating gene dosage in a human model system such as human induced pluripotent stem cells (iPSCs). Here, we demonstrate a simple and precise method to simultaneously generate iPSC lines with different gene dosages using paired Cas9 nickases. We first generate a Cas9 nickase variant with broader protospacer-adjacent motif specificity to expand the targetability of double-nicking–mediated genome editing. As a proof-of-concept study, we examine the gene dosage effects on an Alzheimer’s disease patient-derived iPSC line that carries three copies of APP (amyloid precursor protein). This method enables the rapid and simultaneous generation of iPSC lines with monoallelic, biallelic, or triallelic knockout of APP. The cortical neurons generated from isogenically corrected iPSCs exhibit gene dosage-dependent correction of disease-associated phenotypes of amyloid-beta secretion and Tau hyperphosphorylation. Thus, the rapid generation of iPSCs with different gene dosages using our method described herein can be a useful model system for investigating disease mechanisms and therapeutic development.
Similar content being viewed by others
Introduction
The complexity of the human brain arises not only from the genes it expresses but also from the precise regulation of gene dosage and expression levels. Their dysregulation due to duplication or haploinsufficiency is a major cause of autosomal dominant diseases, such as Alzheimer’s disease (AD)1,2. As the most common type of neurodegenerative dementia, AD is characterized by two pathological hallmarks: extracellular amyloid plaques largely comprising amyloid-beta (Aβ) peptides and intracellular neurofibrillary tangles rich in hyperphosphorylated Tau protein3. In AD, the duplication of specific genes, such as APP (amyloid precursor protein) can cause rare early-onset forms of the disease termed “familial AD”4. Meanwhile, heterozygous premature termination codon and noncoding variants in a single allele of certain genes5,6 (e.g., ABCA7) confer a risk of late-onset sporadic AD. Given that numerous promising therapeutics showed efficacy in animal models of AD but failed in clinical trials7, there is a pressing need to establish relevant model systems, such as human neurons that better mimic the pathophysiology of AD in humans.
Technological developments have enabled the generation and subsequent differentiation of human-induced pluripotent stem cells (iPSCs) into various brain cell types and brain organoids8. Thus, iPSC-based models with AD-causing mutations can recapitulate certain phenotypes of familial AD, including increased Aβ production, Tau hyperphosphorylation, endosomal abnormalities, and oxidative stress9,10,11,12. However, when studying genes and variants associated with sporadic AD that might have fewer risk effects, interindividual genetic heterogeneity can hinder the accurate analysis of disease phenotypes in vitro13. Therefore, genome-editing tools, such as the bacterial CRISPR (clustered regularly interspaced short palindromic repeats)/Cas9 system, have great potential to generate isogenic iPSC lines that only differ with respect to a specific gene or mutation of interest but are otherwise genetically identical14,15. Although heterozygous knockout more accurately models the partial reduction in gene expression caused by noncoding variants, most sporadic AD genes have not been studied in heterozygous-knockout iPSC-based models in parallel to their homozygous-knockout counterparts (summarized in Supplementary Table 1)16, partly because there is currently no rapid and efficient method to manipulate gene dosage in iPSCs.
RNA-guided Cas9 nuclease enables sequence-specific, double-strand breaks by specifying a 20-nucleotide targeting sequence within its single-guide RNA (sgRNA) followed by a protospacer adjacent motif (PAM) sequence17. For the most commonly used Streptococcus pyogenes Cas9 (“Cas9” hereafter), the required PAM sequence is NGG17. When targeting the exonic regions of a gene, Cas9-induced double-strand breaks can result in frameshift mutations and gene knockout through DNA repair mediated by non-homologous end-joining18. As Cas9-mediated gene knockout appears to be highly efficient and insensitive to gene copy number19, Cas9 with a potent sgRNA tends to yield complete knockout of target genes, which can facilitate loss-of-function studies20,21,22,23 (summarized in Supplementary Table 2). However, given the considerable off-target effects of Cas9 nuclease24,25, the use of paired Cas9 nickases (Cas9n) can significantly reduce such off-target effects26. As both nickases must be functional to generate two single-strand breaks in close proximity (i.e., 0–20 nucleotides between two sgRNAs)27, the genome targetability of Cas9n is expected to be much lower than that of an individual Cas9 nuclease, thus necessitating further optimization. Furthermore, the ability of paired nickases to manipulate gene dosage in iPSCs has not been evaluated at the single-cell-derived clone level.
In this study, we expanded the genome-editing targetability of paired nickases by generating a Cas9nVQR variant (D1135V/R1335Q/T1337R) with an NGA PAM. In addition, by using an AD iPSC line carrying APP duplication (“APP cells” hereafter), we used paired Cas9nVQR for the one-step generation of isogenic iPSC lines with monoallelic, biallelic, or triallelic knockout of the target gene; among them was an isogenically corrected line (“Iso cells” hereafter) that had undergone monoallelic APP disruption (i.e., carrying two functional APP alleles) as an APP wild-type revertant. After the iPSCs differentiated into cortical neurons, compared to APP neurons, Iso neurons exhibited genotype-dependent rescue of Aβ production and Tau phosphorylation at Thr231. Furthermore, transcriptomic profiling and functional validation revealed a neuronal apoptotic pathway that is potentially implicated in APP dosage-dependent AD pathogenesis. Thus, our study demonstrates an efficient approach for rapidly manipulating gene dosage in iPSCs with paired Cas9n, which will facilitate the study of risk genes associated with human diseases.
Results
Footprint-free editing of APP copy number by paired Cas9 nickases
To expand the genome-editing targetability of paired nickases, we generated a Cas9nVQR variant (D1135V/R1335Q/T1337R) with an NGA PAM28 (Supplementary Fig. 1a–d). Genome-wide coverage analysis demonstrated that Cas9n can target 58.1% of 3,209,286,105 sites in the human reference genome (GRCh38). Meanwhile, the Cas9nVQR variant developed in this study can increase the targetability to 79.8% when used alone and to 90.3% when used together with Cas9n; this expanded genome targetability translates into 695,373,139 and 1,032,629,268 genomic sites, respectively, that are not targetable by Cas9n (Supplementary Fig. 1e, Supplementary Software 1). To examine the ability and flexibility of the Cas9nVQR variant to manipulate gene copy number in iPSCs at the single-cell-derived clone level, we utilized an AD iPSC line carrying APP duplication (i.e., APP cells)9 and targeted APP exon 16, which is inaccessible to wild-type Cas9n (Fig. 1a). Accordingly, we designed a pair of sgRNAs—designated sgRNA1 and sgRNA2—that targeted this locus (Fig. 1a). To determine their genome-editing efficiency, we transfected constructs that express Cas9nVQR and corresponding sgRNAs into HEK 293T cells. We utilized the endogenous EcoRI site in the target sequence to rapidly screen the genome-editing events. An EcoRI-resistant band at 1.1 kb indicated the disruption of an endogenous EcoRI site and successful editing at the targeted locus (Fig. 1b). Successful genome editing occurred when both sgRNAs and Cas9nVQR were expressed in the HEK 293T cells, whereas no editing occurred when Cas9nVQR was expressed alone or with only one of the two sgRNAs (Fig. 1b).
a Design of a scarless, antibiotic marker-free, gene-editing strategy using a newly generated Cas9 nickase variant, Cas9nVQR (D1135V/R1335Q/T1337R), with paired single-guide RNAs (i.e., sgRNA1 and sgRNA2) that target APP exon 16 in neurons with APP duplication. Red arrowheads indicate the nickase cleavage site, blue boxes indicate the exon, and the red line within the blue boxes indicates Cas9-induced insertion/deletion (indel) mutations. b Genome-editing efficiency of Cas9nVQR with paired sgRNAs in HEK 293T cells. Cas9nVQR with or without corresponding sgRNAs was transfected into HEK 293T cells. The EcoRI-resistant band at 1.1 kb indicates successful editing. c Schematic diagram of genome editing in a human induced pluripotent stem cell (iPSC) line carrying duplication of the APP gene. d Screening of gene-edited iPSC clones by EcoRI digestion. e Specific deletion mutations detected in gene-edited iPSC clones by Sanger sequencing and next-generation deep sequencing. The percentage of deep-sequencing reads for each iPSC clone indicates the number of APP copies remaining intact or removed. One isogenic iPSC clone (Iso) had approximately one-third of 23-bp deletion reads and two-thirds of wild-type sequence reads, indicating that one copy of APP had been inactivated; therefore, it was used for subsequent experiments. PAM protospacer-adjacent motif.
To establish the APP copy number-corrected isogenic line (i.e., Iso cells), we first transfected APP iPSCs with Cas9nVQR-GFP constructs expressing sgRNA1 and sgRNA2 and isolated GFP-positive cells by cell sorting. Next-generation deep sequencing demonstrated an editing efficiency of 40 ± 3.5% in iPSCs after GFP sorting. We subsequently sorted cells for single-clone expansion and screened them by EcoRI digestion assay as described above (Fig. 1c). Out of the 12 screened iPSC clones, four exhibited potential genome editing at the target region (Fig. 1d). We did not detect large deletions in the 5.2-kb PCR amplicons surrounding the edited region in these clones; we subsequently genotyped them by Sanger sequencing and next-generation sequencing (Supplementary Fig. 2a). Accordingly, we generated an Iso line (i.e., a corrected line) with two copies of APP and lines with one or zero copies of APP (Fig. 1e, Supplementary Fig. 2b). Further array-based comparative genomic hybridization (CGH) assay showed that the edited iPSC lines with monoallelic disruption still carried a 722-kb duplication on chromosome 21 like the parent APP duplication iPSC line, whereas the iPSC lines with biallelic or triallelic disruption did not (Supplementary Table 3, Supplementary Data 1). These results collectively suggest that paired Cas9nVQR deletes the entire 722-kb duplicate region harboring the APP locus in iPSC lines with biallelic or triallelic disruption, which is more than 100-fold larger than that in the previous study (~6 kb)27. We used this approach to replicate the efficient generation of the three genotypes simultaneously, demonstrating its reproducibility (Supplementary Fig. 3). Therefore, the double-nicking method flexibly and efficiently manipulated gene copy numbers.
We subsequently examined whether the Iso iPSC lines maintained the pluripotent characteristics of the parental APP iPSC line, specifically pluripotency marker expression and a normal karyotype9. Indeed, an Iso iPSC clone stained positive for pluripotent stem cell markers including OCT4, SSEA4, and TRA-1-81 (Supplementary Fig. 4a) and had a normal karyotype upon CGH assay (Supplementary Fig. 4b).
Impact of APP copy correction on amyloid- and Tau-associated pathologies
We subsequently induced the formation of cortical neurons from corresponding iPSC lines according to the Ngn2 (neurogenin2)-mediated differentiation protocol15,29. After 28 days in vitro (DIV), APP, Iso, and nondemented control (NDC) iPSCs efficiently differentiated into forebrain cortical neurons that expressed the neuronal marker MAP2 (Fig. 2a). In addition, more than 90% of MAP2-positive neurons co-expressed the layer II–IV neuronal marker Cux1 (Fig. 2b), suggesting that a homogenous population of upper-layer cortical neurons had been generated. Moreover, RT-qPCR analysis indicated that the neurons induced from APP, Iso, and NDC iPSCs exhibited similar mRNA expression levels of NeuN (a mature neuronal marker), synaptophysin (a presynaptic marker), and PSD-95 (a postsynaptic marker) (Fig. 2c). These results indicate that these neurons had a similar differentiation status.
a, b Generation of a homogenous population of cortical neurons from nondemented control (NDC), APP duplication parent (APP), and APP copy number-corrected isogenic (Iso) cell lines. a Neurons at 28 days in vitro (DIV) were stained for MAP2 (neuronal marker) and Cux1 (cortical layer II–IV marker). b Percentages of Cux1- and MAP2-positive neurons. c Neuronal differentiation status as assessed by qRT-PCR for MAP2, NeuN (mature neuron marker), Synaptophysin (presynaptic marker), and PSD-95 (postsynaptic marker). d Levels of secreted amyloid-beta (Aβ)38, Aβ40, and Aβ42 as well as Aβ42/Aβ40 ratios in conditioned media from NDC, APP, and Iso neurons at 28 DIV. e Western blot analysis of full-length APP (FL-APP), phosphorylated Tau at Thr231 (pThr231 Tau), and total Tau in NDC, APP, and Iso neurons at 28 DIV. βIII-tubulin (βIII-tub) and GAPDH served as loading controls for neurons and total protein, respectively. f Immunocytochemical analysis of pThr231 Tau and doublecortin (DCX) in APP and Iso neurons at 28 DIV. g Quantification of APP/βIII-tub, pThr231 Tau/βIII-tub, total Tau/βIII-tub, and pThr231/total Tau ratios relative to those in NDC neurons. Values are mean ± SEM (n = 4 independent biological replicates per cell line; *p < 0.05, **p < 0.01, or ***p < 0.001 vs. APP neurons, Student’s t-test). Scale bars: 100 μm.
Neurons derived from APP iPSCs exhibit abnormal Aβ peptide secretion and Tau hyperphosphorylation9. Indeed, compared to NDC neurons, APP neurons secreted significantly more Aβ38, Aβ40, and Aβ42 peptides into the medium. Meanwhile, subsequent correction of APP gene dosage in Iso iPSCs restored the normal secretion levels of these Aβ peptides (Fig. 2d). Furthermore, in Iso iPSC-derived neurons, the APP protein level decreased close to that in neurons derived from the iPSCs of NDCs, confirming the correction of APP copy number in Iso iPSCs (Fig. 2e, g).
We subsequently examined whether APP gene copy number affected Tau phosphorylation in Ngn2-induced neurons from APP, Iso, and NDC iPSCs. Tau phosphorylated at Thr231, a component of paired helical filaments Tau, is an early marker of AD pathology correlated with cognitive decline30. Compared to NDC neurons, APP neurons exhibited significantly higher Tau phosphorylation at Thr231, total Tau level, and pTau231/total Tau ratio, which is consistent with the previous studies9,11. Meanwhile, APP copy number correction in Iso neurons reduced the levels of phosphorylated and total Tau (Fig. 2e–g), suggesting that deleting the extra copy of APP restored Tau phosphorylation at Thr231, total Tau level, and the pTau231/total Tau ratio. These results are concordant with a recent study showing that the pTau231/total Tau ratio is regulated through mechanisms involving cholesterol metabolism, cholesteryl esters, and proteasome degradation in iPSC-derived neurons with APP duplication31.
Screening of pathways associated with APP gene dosage
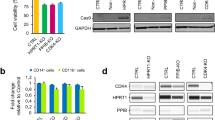
To screen for the molecular pathways associated with APP gene dosage, we profiled the transcriptomes of neurons from APP and Iso iPSC lines. Differential gene expression analysis revealed 376 differentially expressed genes (DEGs) between the APP and Iso lines termed “isogenic DEGs.” Hierarchical clustering analysis revealed that three batches of APP and Iso neurons clustered together within each respective group (Fig. 3a). Among them, 208 and 168 genes were up and downregulated, respectively, in APP neurons compared to Iso neurons. This suggests that APP gene dosage broadly affects gene expression in a human diploid cell system. In parallel, differential gene expression analysis between the APP and non-isogenic NDC lines yielded 2,158 DEGs termed “non-isogenic DEGs” (including 1050 and 1108 up and downregulated genes, respectively). Venn diagram analysis of these two groups of DEGs revealed 207 common genes (Fig. 3b), which accounted for 55% of the isogenic DEGs (i.e., APP vs. Iso) and less than 10% of non-isogenic DEGs (i.e., APP vs. NDC). Therefore, we analyzed these 207 common genes, which exhibited both APP-dependent effects and dysregulation in APP neurons.
a Hierarchical clustering analysis of the 376 differentially expressed genes (DEGs) in APP duplication parent (APP), APP copy number-corrected isogenic (Iso), and nondemented control (NDC) neurons. Among them, 208 and 168 genes were up and downregulated, respectively, in APP neurons compared to Iso neurons (false discovery rate [FDR] < 0.05, fold change ≥1.5). b Venn diagram illustrating overlap between isogenic DEGs (APP vs. Iso neurons) and non-isogenic DEGs (APP vs. NDC neurons) (FDR < 0.05, fold change ≥1.5). c Enriched Gene Ontology (GO) terms for the 207 common DEGs. Volcano plots showing the DEGs between the APP and NDC neurons (d) and between the Iso and APP neurons (e). Four selected genes upregulated in APP neurons compared to NDC neurons that were the top genes downregulated in copy number-corrected Iso neurons compared to APP neurons. f RT-qPCR validation of top DEGs common to both NDC and Iso neurons compared to APP neurons. Values are mean ± SEM (n = 4 independent biological replicates per condition; *p < 0.05, **p < 0.01, or ***p < 0.001 vs. APP neurons, Student’s t-test).
Functional annotation of these 207 common DEGs revealed several significantly enriched gene ontology categories including “regulation of transcription” (false discovery rate [FDR] = 4.82E−4), “regulation of response to stimulus” (FDR = 2.88E−2), “regulation of the Wnt signaling pathway” (FDR = 3.84E−2), and “homophilic cell adhesion via plasma membrane adhesion molecules” (FDR = 4.06E−2) (Fig. 3c). Subsequent analysis of the top genes using FDR filters and visualization using volcano plots yielded candidates that might have functional implications in AD pathogenesis (Fig. 3d, e; Table 1). Therefore, we performed RT-qPCR to validate the top four DEGs between the APP and NDC lines as well as between the APP and Iso lines (Fig. 3f).
Inhibition of PEG3–Bax rescues neuronal cell death with APP duplication
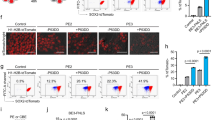
To investigate the possible biological relevance of the identified DEGs, we focused on PEG3 (paternally expressed gene 3), a gene implicated in p53-mediated cell death32,33. We first performed western blot analysis to validate that the changes in PEG3 mRNA levels observed in iPSC-derived neurons paralleled the changes in protein levels. PEG3 protein level was significantly higher in APP neurons than NDC neurons but reverted to the basal level upon APP copy number correction (Fig. 4a, b). Given that PEG3 is implicated in the DNA damage-induced neuronal apoptotic pathway34, we investigated neuronal death in both APP and Iso iPSC-derived neurons. Genetic correction reduced apoptosis as indicated by the number of cleaved caspase 3-positive apoptotic cells in APP and Iso iPSC-derived neurons at 28 DIV (Fig. 4c, d).
a Western blot analysis of PEG3 protein in nondemented control (NDC), APP duplication parent (APP), and APP copy number-corrected isogenic (Iso) neurons. b Quantification of PEG3 expression levels normalized to that of βIII-tubulin (βIII-tub) and relative to those in the NDC neurons. c Immunofluorescence images showing CASP3 (cleaved caspase 3)-positive apoptotic cells with pyknotic condensed nuclei in APP and Iso neurons at 28 days in vitro. Scale bars: 50 and 10 μm. d Percentage of CASP3-positive apoptotic cells in all neurons. The proportion of apoptotic cells was lower in Iso neurons than that in APP neurons. b, d Values are mean ± SEM (n = 3 independent biological replicates per line; **P < 0.01 vs. APP line; Student’s t-test). e Immunofluorescence images showing CASP3-positive cells in APP neurons at 28 days in vitro treated with vehicle control (Con), 100 μM BIP-V5 (Bax inhibitor peptide V5), or 5 μM CRI (cytochrome C release inhibitor) for 7 days. Scale bars: 50 and 10 μm. f Percentage of CASP3-positive apoptotic cells among all neurons. The proportion of apoptotic cells was lower in BIP-V5- or CRI-treated neurons than that in Con neurons. Values are mean ± SEM (n = 3 independent biological replicates per line; **p < 0.01 vs. Con neurons; Student’s t-test).
Next, we elucidated which downstream effector(s) mediates the action of PEG3 during neuronal death in APP iPSC-derived neurons. Given that PEG3 promotes apoptosis by inducing Bax translocation from the cytosol to the mitochondria32,34, we examined if inhibition of Bax translocation alleviates neuronal death in APP iPSC-derived neurons. Bax, a pro-apoptotic member of the Bcl-2 protein family, exerts its function in mitochondria-dependent apoptosis35, while its mitochondrial translocation can be inhibited by BIP-V5 (Bax inhibitor peptide V5), a cell-permeable synthetic peptide inhibitor of Bax36. Bax translocation induces the release of cytochrome C in apoptosis, which can be inhibited by CRI (cytochrome C release inhibitor; a Bax channel blocker)37. BIP-V5 or CRI administration significantly reduced the percentage of cleaved caspase 3-positive or propidium iodide-positive neurons compared to the vehicle control-treated APP iPSC-derived neurons (Fig. 4e, f; Supplementary Fig. 5), suggesting that inhibiting either Bax translocation or cytochrome C release can rescue apoptosis in these neurons. These results collectively reveal that APP duplication induced PEG3 upregulation and BAX-mediated neuronal apoptosis in a human iPSC model of familial AD carrying APP duplication.
Discussion
This study demonstrates that the copy number of a given gene can be efficiently manipulated by paired Cas9 nickases. Correcting the copy number of APP rescues several pathological features of AD, including the production of Aβ and hyperphosphorylated Tau. Our transcriptomic analysis reveals key dysregulated pathways in neuronal death and cell stress response among others. Specifically, we show that APP duplication induces PEG3 upregulation and BAX-mediated neuronal apoptosis. Furthermore, inhibition of Bax rescues the neuronal loss in APP-overexpressing, iPSC-derived neurons, suggesting that inhibiting Bax can be explored as a potential therapeutic approach for AD associated with increased APP expression.
The CRISPR/Cas9-mediated manipulation of gene copy number holds great promise for modeling genetic disorders caused by gene duplication or haploinsufficiency, such as AD4,38. Genome-wide association studies have identified thousands of risk variants associated with AD; most of these variants reside in noncoding regions surrounding ~30 risk genes39,40,41 and have been suggested to modify disease risk by affecting the expression of their respective target genes42,43. Although it is impossible to generate an iPSC line for each variant, it is both feasible and of great interest to manipulate the gene dosage by inducing heterozygous and homozygous knockout of these risk genes in iPSCs in order to mimic variant effects, elucidate disease mechanisms, or create relevant models for therapeutic development. Despite the enhanced specificity of Cas9n, the rapid, flexible, and efficient manipulation of gene dosage using Cas9n in human iPSCs have not been demonstrated. One potential obstacle is the restricted targetability of Cas9n. Therefore, we generated a Cas9nVQR variant with significantly expanded targetability, which can potentially edit 90.3% of the 3,209,286,105 sites in the human reference genome (GRCh38); this represents 1,032,629,268 additional sites that are not targetable by Cas9n. Furthermore, while Cas9n exhibits enhanced specificity, it is assumed to be less efficient for genome editing than Cas9 nuclease. In this study, we readily generated integration-free isogenic clones with different gene dosages of APP from the parent iPSCs by screening fewer than 20 clones within 1 month. Considering that there are ~30 uncharacterized sporadic AD risk genes (Supplementary Table 1), it would be possible to simultaneously generate heterozygous- and homozygous-knockout iPSC lines within a reasonable timeframe (e.g., ten genes in 1 month). Hence, our protocol could enable the generation of a large panel of isogenic iPSC lines that are haploinsufficient in the ever-growing number of known disease risk genes, which could be used to uncover shared and distinct pathophysiological mechanisms.
There are several differentiation protocols for generating cortical excitatory neurons in the literature. In this study, we utilized the Ngn2-mediated differentiation protocol to generate cortical neurons with homogenous differentiation status and neuronal identity (i.e., Cux1+ superficial layer neurons)29. In contrast, other protocols utilize a chemical-based, dual-SMAD inhibition method44, which recapitulates the temporal development of the cerebral cortex and produces mixed populations of deep- and upper-layer neurons as well as astrocytes45. Such a complex cell type composition might affect results regarding specific AD phenotypes. Thus, our study demonstrates the utility of using an iPSC-derived, single-neuron population to examine APP duplication-specific effects on AD pathogenesis.
While brain atrophy caused by neuronal loss is a prominent pathological feature of AD46, whether and how familial AD-associated APP duplication leads to neuronal loss remains poorly understood. Transcriptomic analysis revealed dysregulation in apoptosis-associated genes in APP neurons compared to Iso neurons. Among them, PEG3 is a tumor-suppressor protein involved in the DNA damage-induced apoptotic pathway via Bax translocation32,34. Accordingly, APP neurons exhibited dysregulated PEG3 expression and elevated apoptosis, which were rescued by correcting the extra copy of APP. Furthermore, specific inhibition of Bax translocation or cytochrome C release rescued the elevated cell death of APP neurons. These results suggest that APP duplication contributes to the increased susceptibility neurons to death via a PEG3–Bax cytochrome C pathway. Together with reports on the neuroprotective role of the inhibition of Bax translocation or cytochrome C release in ischemia47 and Huntington’s disease48, our findings suggest that inhibiting Bax can be exploited as a potential therapeutic approach to alleviating neuronal loss in AD.
In summary, by using APP as a proof-of-concept, we developed a Cas9nVQR-based genome-editing method that can efficiently manipulate the copy number of specific risk genes with expanded targetability and no footprint. Our study also demonstrates that APP regulates Tau expression and hyperphosphorylation and contributes to PEG3 upregulation and BAX-mediated neuronal apoptosis in an isogenic background. Thus, our method can rapidly and simultaneously generate heterozygous and homozygous gene knockout in iPSCs and can be utilized as a model system to mimic gene dosage effects, elucidate disease mechanisms, and facilitate therapeutic development.
Methods
Plasmids and antibodies
The sgRNA sequences for APP exon 16 were as follows: sgRNA1: 5′-TCTGCATCCAGATTCACTTC-3′, sgRNA2: 5′-TTCCGACATGACTCAGGATA-3′. We generated the Cas9nVQR (D1135V/R1335Q/T1337R) variant with altered PAM specificity to the NGA sequence by site-directed mutagenesis (QuikChange II XL Site-Directed Mutagenesis Kit, Stratagene) based on the pSpCas9n(BB)-2A-GFP (PX461) plasmid (Addgene #48140). The newly generated plasmid has been deposited in Addgene (accession code: 165033). For the double-nicking method, we inserted sgRNA1 and sgRNA2 into pSpCas9nVQR(BB)-2A-GFP via the BbsI site (New England Biolabs). Then, we transformed the ligation product into NEB Stable Competent E. coli (New England Biolabs) for molecular cloning. We prepared plasmid DNA using the E.Z.N.A. Plasmid DNA Mini Kit I (Omega Bio-tek) according to the manufacturer’s instructions. We prepared large-scale plasmids in endotoxin-free conditions using the Qiagen Plasmid Maxi Kit (Qiagen).
We used the following primary antibodies for immunofluorescence and western blot analysis: mouse anti-Oct-3/4 (C-10; 1:1,000, #SC-5279, Santa Cruz), anti-SSEA4 (1:1,000, #414000, Life Technologies), anti-TRA-1-81 (1:1,000, #MAB4381, Millipore), anti-Cux1 (1:500, #ab54583, Abcam), anti-APP (6E10; 1:2,000, #803016, Biolegend), anti-human phospho-Tau at Thr231 (AT180; 1:1,000, #MN1040, Thermo Fisher Scientific), anti-total Tau (TAU-5; 1:20,000, #AHB0042, Life Technologies), anti–βIII-tubulin (TUJ1; 1:5,000, #MAB1637, Millipore), anti-GAPDH (1:10,000, GAPDH, Thermo Fisher Scientific), rabbit anti-cleaved Caspase 3 (Asp175; 1:1,000, #9661, Cell Signaling), guinea pig anti-DCX (1:2,000, #AB2253, Millipore), and chicken anti-MAP2 (1:5,000, #ab5392, Abcam).
Cell lines and cell culture
Dr. Lawrence Goldstein (University of California, San Diego) generously provided one NDC-derived iPSC line (NDC1.1) and one AD patient-derived iPSC line (APP1.1) carrying APP gene duplication, both from male donors9. The culture of iPSCs was approved by the Committee on Research Practices of the Hong Kong University of Science and Technology. We cultured iPSCs in feeder-free conditions on Geltrex (Thermo Fisher Scientific) in Essential 8 medium (E8, Thermo Fisher Scientific). We monitored cell quality daily and mechanically removed differentiated cells under a light microscope in a biosafety hood. We cultured iPSCs to 80% confluence and passaged them using Versene EDTA solution (Thermo Fisher Scientific).
We incubated HEK 293T cells in Dulbecco’s modified Eagle medium (DMEM, Invitrogen) with 10% heat-inactivated fetal bovine serum (v/v, GIBCO) plus 1% penicillin/streptomycin. We transfected HEK 293T cells using Lipofectamine 3000 Reagent (Thermo Fisher Scientific) according to the manufacturer’s instructions. We harvested cells 24–48 h post transfection for genotyping.
Genome editing and karyotyping of iPSCs
To manipulate the copy number of APP in the APP1.1 line, we pretreated iPSCs with 10 μM Rock inhibitor (Y-27632 dihydrochloride, Tocris) 2 h before nucleofection. We then dissociated iPSCs with Accutase (Thermo Fisher Scientific) into single-cell suspensions and filtered them through a 100-μm filter. We nucleofected 2.0 × 106 iPSCs with pSpCas9nVQR(BB)-2A-GFP constructs expressing sgRNA1 and sgRNA2 by using a Lonza Nucleofector 2b Device (Program B-016) and Human Stem Cell Nucleofector Kit 1 (Lonza).
After recovery in culture for 2 days, we isolated GFP-positive iPSCs by Accutase dissociation followed by FACS (FACS Aria IIa, BD Biosciences), plated them at 1000–2000 cells per 32-mm Geltrex-coated plate, and cultured them in E8 media containing 10 μM Rock inhibitor for 2 days. After 10–14 days, we manually selected individual iPSC clones. We cultured half of each clone in individual wells of 48-well plates and submitted the other half to genomic DNA extraction using QuickExtract DNA Extraction Solution (Epicentre) and screening for EcoRI disruption. We excluded the possibility of large deletions by generating ~5.2-kb PCR amplicons surrounding the APP exon 16 using primers (5′-GCACGTTGCCAGATCTTAATAACAG-3′ and 5′-CCTGCAGCAGTGAACATCAAGG-3′). We validated potentially edited clones by Sanger sequencing. Briefly, we generated ~1.2-kb PCR amplicons surrounding the target genomic region using primers (5′-AACAAAGCCCCAAAGTAGCAG-3′ and 5′- GCATGTGTTTGATCCTTCAAATAAACC-3′), ligating them to a cloning vector, and subjecting them to Sanger sequencing. We quantitatively evaluated iPSC clones containing frameshifting mutations by next-generation deep sequencing of ~150-bp PCR amplicons using primers (5′-CAGGTTCTGGGTTGACAAATATCAAG-3′ and 5′-GCAAGACAAACAGTAGTGGAAAGAGG-3′) and analyzed them using CRISPResso249. The percentage of deep-sequencing reads for each iPSC clone indicated the number of APP copies remaining intact or removed.
We determined the karyotypes and genomic amplification and deletion regions of the parent and isogenic iPSCs by array-based CGH assay using Agilent Human Catalog CGH 8x60K Microarrays.
Availability of iPSC lines
The gene-edited iPSC lines with monoallelic, biallelic, or triallelic knockout of APP are available upon request from the Ip Laboratory.
Differentiation of iPSCs into cortical neurons
We generated cortical neurons from iPSCs as previously described15,29 with minor modifications. On day −2, we plated iPSCs on six-well Geltrex-coated plates 1 day before infection. On day −1, we suspended lentivirus with rtTA and Ngn2-GFP-puro vectors in E8 media containing 10 μM Rock inhibitor and infected the iPSCs for 24 h. On day 0, to induce the expression of NEUROG2 (the Ngn2 gene), we replaced the culture medium with N2 medium (DMEM/F12, 1× N2, and 1× NEAA; Thermo Fisher Scientific) containing 2 μg mL−1 doxycycline (Sigma). On day 1, we replaced the culture medium with N2 medium containing 2 μg mL−1 doxycycline and 1 μg mL−1 puromycin (Thermo Fisher Scientific). On days 2–3, we replaced the medium with B27 medium (Neurobasal medium, 1× B27, and 1× GlutaMAX; Thermo Fisher Scientific) containing 2 μg mL−1 doxycycline, 1 μg mL−1 puromycin, and 0.2 μg mL−1 laminin (Sigma). On day 4, we dissociated the induced cells with Accutase and plated them onto six-well PDL/laminin-coated plates at 1.0 × 106 cells per well or 24-well plates with coverslips at 50,000 cells per coverslip. For further experiments, until day 28, we removed half of the medium every 4 days and replaced it with B27 medium containing 0.2 μg mL−1 laminin, 0.5 μg mL−1 doxycycline, 10 ng mL−1 BDNF, and 10 ng mL−1 NT-3.
Immunocytochemistry
We fixed cells grown on coverslips or chamber slides with 4% paraformaldehyde (Electron Microscopy Services) for 15 min at room temperature. We then permeabilized the cells using 0.1% Triton X-100 in Dulbecco’s phosphate-buffered saline (DPBS) for 15 min at room temperature, blocked them in 2% normal goat serum in DPBS for 1 h at room temperature, and incubated them with primary antibodies overnight at 4 °C. We then washed the cells three times in DPBS, incubated them with the appropriate secondary antibodies for 1 h at room temperature, washed them twice with DPBS, and incubated them with DAPI for 10 min. We acquired images with a Leica SP8 inverted confocal microscope.
Protein extraction and western blot analysis
We lysed cells with RIPA buffer containing protease inhibitor cocktail (Roche). We measured protein concentration by Bradford Assay (Bio-Rad). Briefly, we denatured samples at 95 °C for 5 min in sample buffer. We separated 20 μg protein per sample on 4–12% Bis-Tris or 10–20% Tricine gradient gels (Thermo Fisher Scientific) at 100 V for 1 h and subsequently transferred samples to nitrocellulose membranes (iBlot Transfer Stack, Thermo Fisher Scientific). We then used HRP-conjugated secondary antibodies and ECL substrate for detection.
RNA extraction and RT-PCR
We purified total RNA from iPSCs and iPSC-derived neurons by using the NucleoSpin RNA Kit (Macherey-Nagel) according to the manufacturer’s instructions. We assessed RNA concentration and quality by using BioDrop μLite (BioDrop). We performed reverse-transcription by using a PrimeScript RT Reagent Kit (TaKaRa). We performed real-time qPCR with TaqMan gene expression assays and the Premix Ex Taq qPCR assay (TaKaRa). Finally, we used the 2exp (i.e., −ΔΔCt) method to determine the relative expression of each gene with HPRT as a reference.
RNA sequencing analysis of iPSC-derived neurons
We performed the transcript-level expression analysis as previously described50. Briefly, quality control was first accomplished using FastQC and RSeQC. The FASTQ files were aligned to the human GRCh38.91 reference genome using the HISAT2 2.0 alignment program. The aligned files were assembled into potential transcripts using the StringTie assembler program. Next, the potential transcripts were passed to the prepDE program to generate read counts. Minimal pre-filtering with row counts greater than ten reads was performed. We performed differential gene expression analysis using DESeq2 from Bioconductor version 3.1051. We used an FDR cutoff of 0.05 with a fold change ratio cutoff of 1.5 and a mean FPKM cutoff of 1 for differential gene expression analysis. We created the pipeline to run the analysis using the Snakemake workflow management system52.
Quantification of secreted Aβ
We replaced the media 2 days before quantifying secreted Aβ. We quantified secreted Aβ by using MSD Human (6E10) Abeta3-Plex Kits (Meso Scale Discovery) according to the manufacturer’s instructions. We normalized secreted Aβ levels to total protein levels determined by BCA assay (Thermo Fisher Scientific).
Statistics and reproducibility
All data are presented as mean ± SEM unless otherwise specified. We examined the significance of differences by unpaired Student’s t-tests across three or four independent biological replicates using GraphPad prism version 6 (GraphPad Software). The level of statistical significance was set at *p < 0.05, **p < 0.01, or ***p < 0.001.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The deep-sequencing data are deposited in the NCBI Sequence Read Archive (SRA; accession code: PRJNA681926). The RNA sequencing data are deposited in the NCBI Gene Expression Omnibus (GEO; accession code: GSE160224). The CGH data are available in Supplementary Data 1. The source data for the graphs and charts presented in the main figures are available in Supplementary Data 2. Full blots are included in Supplementary Fig. 6.
Code availability
Custom code for genome-wide coverage analysis of various Cas9 nickase configurations in the GRCh38 human reference genome is available in Supplementary Software 1.
References
Huang, N., Lee, I., Marcotte, E. M. & Hurles, M. E. Characterising and predicting haploinsufficiency in the human genome. PLoS Genet. 6, e1001154 (2010).
Rice, A. M. & McLysaght, A. Dosage-sensitive genes in evolution and disease. BMC Biol. 15, 78 (2017).
Scheltens, P. et al. Alzheimer’s disease. Lancet 388, 505–517 (2016).
Rovelet-Lecrux, A. et al. APP locus duplication causes autosomal dominant early-onset Alzheimer disease with cerebral amyloid angiopathy. Nat. Genet. 38, 24–26 (2006).
Steinberg, S. et al. Loss-of-function variants in ABCA7 confer risk of Alzheimer’s disease. Nat. Genet. 47, 445–447 (2015).
De Roeck, A., Van Broeckhoven, C. & Sleegers, K. The role of ABCA7 in Alzheimer’s disease: evidence from genomics, transcriptomics and methylomics. Acta Neuropathol. 138, 201–220 (2019).
Huang, Y. & Mucke, L. Alzheimer mechanisms and therapeutic strategies. Cell 148, 1204–1222 (2012).
Hong, Y. J. & Do, J. T. Neural lineage differentiation from pluripotent stem cells to mimic human brain tissues. Front. Bioeng. Biotechnol. 7, 400 (2019).
Israel, M. A. et al. Probing sporadic and familial Alzheimer’s disease using induced pluripotent stem cells. Nature 482, 216–220 (2012).
Kondo, T. et al. Modeling Alzheimer’s disease with iPSCs reveals stress phenotypes associated with intracellular Abeta and differential drug responsiveness. Cell Stem Cell 12, 487–496 (2013).
Moore, S. et al. APP metabolism regulates Tau proteostasis in human cerebral cortex neurons. Cell Rep. 11, 689–696 (2015).
Gonzalez, C. et al. Modeling amyloid beta and Tau pathology in human cerebral organoids. Mol. Psychiatry 23, 2363–2374 (2018).
Carcamo-Orive, I. et al. Analysis of transcriptional variability in a large human iPSC library reveals genetic and non-genetic determinants of heterogeneity. Cell Stem Cell 20, 518–532 (2017). e519.
Hockemeyer, D. & Jaenisch, R. Induced pluripotent stem cells meet genome editing. Cell Stem Cell 18, 573–586 (2016).
Lin, Y. T. et al. APOE4 causes widespread molecular and cellular alterations associated with alzheimer’s disease phenotypes in human iPSC-derived brain cell types. Neuron 98, 1141–1154 (2018). e1147.
Penney, J., Ralvenius, W. T. & Tsai, L. H. Modeling Alzheimer’s disease with iPSC-derived brain cells. Mol. Psychiatry 25, 148–167 (2020).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Hsu, P. D., Lander, E. S. & Zhang, F. Development and applications of CRISPR-Cas9 for genome engineering. Cell 157, 1262–1278 (2014).
Yuen, G. et al. CRISPR/Cas9-mediated gene knockout is insensitive to target copy number but is dependent on guide RNA potency and Cas9/sgRNA threshold expression level. Nucleic Acids Res. 45, 12039–12053 (2017).
Gonzalez, F. et al. An iCRISPR platform for rapid, multiplexable, and inducible genome editing in human pluripotent stem cells. Cell Stem Cell 15, 215–226 (2014).
Fong, L. K. et al. Full-length amyloid precursor protein regulates lipoprotein metabolism and amyloid-beta clearance in human astrocytes. J. Biol. Chem. 293, 11341–11357 (2018).
Robbins, J. P. et al. Clusterin is required for beta-amyloid toxicity in human iPSC-derived neurons. Front. Neurosci. 12, 504 (2018).
Sullivan, S. E. et al. Candidate-based screening via gene modulation in human neurons and astrocytes implicates FERMT2 in Abeta and TAU proteostasis. Hum. Mol. Genet. 28, 718–735 (2019).
Fu, Y. et al. High-frequency off-target mutagenesis induced by CRISPR-Cas nucleases in human cells. Nat. Biotechnol. 31, 822–826 (2013).
Pattanayak, V. et al. High-throughput profiling of off-target DNA cleavage reveals RNA-programmed Cas9 nuclease specificity. Nat. Biotechnol. 31, 839–843 (2013).
Cho, S. W. et al. Analysis of off-target effects of CRISPR/Cas-derived RNA-guided endonucleases and nickases. Genome Res. 24, 132–141 (2014).
Ran, F. A. et al. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 154, 1380–1389 (2013).
Kleinstiver, B. P. et al. Engineered CRISPR-Cas9 nucleases with altered PAM specificities. Nature 523, 481–485 (2015).
Zhang, Y. et al. Rapid single-step induction of functional neurons from human pluripotent stem cells. Neuron 78, 785–798 (2013).
Luna-Munoz, J., Chavez-Macias, L., Garcia-Sierra, F. & Mena, R. Earliest stages of Tau conformational changes are related to the appearance of a sequence of specific phospho-dependent Tau epitopes in Alzheimer’s disease. J. Alzheimer’s Dis. 12, 365–375 (2007).
van der Kant, R. et al. Cholesterol metabolism is a druggable axis that independently regulates Tau and amyloid-beta in iPSC-derived Alzheimer’s disease neurons. Cell Stem Cell 24, 363–375 (2019). e369.
Deng, Y. & Wu, X. Peg3/Pw1 promotes p53-mediated apoptosis by inducing Bax translocation from cytosol to mitochondria. Proc. Natl Acad. Sci. USA 97, 12050–12055 (2000).
Relaix, F. et al. Pw1/Peg3 is a potential cell death mediator and cooperates with Siah1a in p53-mediated apoptosis. Proc. Natl Acad. Sci. USA 97, 2105–2110 (2000).
Johnson, M. D., Wu, X., Aithmitti, N. & Morrison, R. S. Peg3/Pw1 is a mediator between p53 and Bax in DNA damage-induced neuronal death. J. Biol. Chem. 277, 23000–23007 (2002).
Kuwana, T. & Newmeyer, D. D. Bcl-2-family proteins and the role of mitochondria in apoptosis. Curr. Opin. Cell Biol. 15, 691–699 (2003).
Gomez, J. A. et al. Bax-inhibiting peptides derived from Ku70 and cell-penetrating pentapeptides. Biochemical Soc. Trans. 35, 797–801 (2007).
Pawlowski, J. & Kraft, A. S. Bax-induced apoptotic cell death. Proc. Natl Acad. Sci. USA 97, 529–531 (2000).
Sleegers, K. et al. APP duplication is sufficient to cause early onset Alzheimer’s dementia with cerebral amyloid angiopathy. Brain 129, 2977–2983 (2006).
Lambert, J. C. et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat. Genet. 45, 1452–1458 (2013).
Jansen, I. E. et al. Genome-wide meta-analysis identifies new loci and functional pathways influencing Alzheimer’s disease risk. Nat. Genet. 51, 404–413 (2019).
Kunkle, B. W. et al. Genetic meta-analysis of diagnosed Alzheimer’s disease identifies new risk loci and implicates Abeta, Tau, immunity and lipid processing. Nat. Genet. 51, 414–430 (2019).
Zhang, F. & Lupski, J. R. Non-coding genetic variants in human disease. Hum. Mol. Genet. 24, R102–R110 (2015).
Rosenthal, S. L. & Kamboh, M. I. Late-onset Alzheimer’s disease genes and the potentially implicated pathways. Curr. Genet Med. Rep. 2, 85–101 (2014).
Ovchinnikov, D. A., Korn, O., Virshup, I., Wells, C. A. & Wolvetang, E. J. The impact of APP on Alzheimer-like pathogenesis and gene expression in down syndrome iPSC-derived neurons. Stem Cell Rep. 11, 32–42 (2018).
Shi, Y., Kirwan, P., Smith, J., Robinson, H. P. & Livesey, F. J. Human cerebral cortex development from pluripotent stem cells to functional excitatory synapses. Nat. Neurosci. 15, 477–486 (2012). S471.
Serrano-Pozo, A., Frosch, M. P., Masliah, E. & Hyman, B. T. Neuropathological alterations in Alzheimer disease. Cold Spring Harb. Perspect. Med. 1, a006189 (2011).
Hetz, C. et al. Bax channel inhibitors prevent mitochondrion-mediated apoptosis and protect neurons in a model of global brain ischemia. J. Biol. Chem. 280, 42960–42970 (2005).
Wang, X. et al. Inhibitors of cytochrome c release with therapeutic potential for Huntington’s disease. J. Neurosci. 28, 9473–9485 (2008).
Clement, K. et al. CRISPResso2 provides accurate and rapid genome editing sequence analysis. Nat. Biotechnol. 37, 224–226 (2019).
Pertea, M., Kim, D., Pertea, G. M., Leek, J. T. & Salzberg, S. L. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat. Protoc. 11, 1650–1667 (2016).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Koster, J. & Rahmann, S. Snakemake–a scalable bioinformatics workflow engine. Bioinformatics 28, 2520–2522 (2012).
Acknowledgements
We thank Dr. Tom H. Cheung, Dr. Wing-Yu Fu, Wayne Chi Wai Ng, Jingyi Zhang, Wei Wu, Dr. Carole Shum, Dr. Kwok-Wang Hung, Dr. Xiaopu Zhou, Dr. Nandita Mullapudi, Shun Fat Lau, Cara Kwong, Ka Chun Lok, Jingwen Lin, Hongruo Zhang, and Paul Chow for their excellent advice and technical assistance. We thank Dr. Lawrence Goldstein at the University of California, San Diego, for providing induced pluripotent stem cells. We are grateful to all members of the Ip Laboratory for their helpful discussions. This study was supported in part by the National Key R&D Program of China (2017YFE0190000 and 2018YFE0203600); the Hong Kong Research Grants Council Theme-based Research Scheme (T13-605/18-W); the Area of Excellence Scheme of the University Grants Committee (AoE/M-604/16); the Innovation and Technology Commission (ITCPD/17-9); the Lee Hysan Foundation (LHF17SC01); the Guangdong Provincial Key S&T Program (2018B030336001); the Shenzhen Knowledge Innovation Program (JCYJ20180507183642005, JCYJ20170413165053031, JCYJ20200109115631248, and JCYJ20170413173717055); and the Shenzhen–Hong Kong Institute of Brain Science-Shenzhen Fundamental Research Institutions (project number: 2019SHIBS0001).
Author information
Authors and Affiliations
Contributions
T.Y., A.K.Y.F., and N.Y.I. designed the research. T.Y., Y.D., H.W.S.T., H.X., Yuewen C., and Yu C. conducted the experiments. H.C. performed the bioinformatics analysis. T.Y., A.K.Y.F., and N.Y.I. analyzed the data. T.Y., A.K.Y.F., and N.Y.I. wrote the paper with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ye, T., Duan, Y., Tsang, H.W.S. et al. Efficient manipulation of gene dosage in human iPSCs using CRISPR/Cas9 nickases. Commun Biol 4, 195 (2021). https://doi.org/10.1038/s42003-021-01722-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-021-01722-0
This article is cited by
-
A tensor decomposition-based integrated analysis applicable to multiple gene expression profiles without sample matching
Scientific Reports (2022)
-
CRISPR and iPSCs: Recent Developments and Future Perspectives in Neurodegenerative Disease Modelling, Research, and Therapeutics
Neurotoxicity Research (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.