Abstract
Runt-related transcription factor 2 (RUNX2) is critical for the development of the vertebrate bony skeleton. Unlike other RUNX family members, RUNX2 possesses a variable poly-glutamine, poly-alanine (QA) repeat domain. Natural variation within this repeat is able to alter the transactivation potential of RUNX2, acting as an evolutionary ‘tuning knob’ suggested to influence mammalian skull shape. However, the broader role of the RUNX2 QA repeat throughout vertebrate evolution is unknown. In this perspective, we examine the role of the RUNX2 QA repeat during skeletal development and discuss how its emergence and expansion may have facilitated the evolution of morphological novelty in vertebrates.
Similar content being viewed by others
Introduction
Runt-related (RUNX) proteins are a conserved family of DNA-binding transcription factors1 that play critical roles during development2. RUNX transcription factors are characterized by their conserved Runt domain3, first identified through sequence homology to Drosophila runt, a pair-rule segmentation gene with multiple developmental roles during embryogenesis4. Vertebrate RUNX transcription factors form heterodimeric complexes with core-binding factor-beta (CBFβ) to allosterically enhance DNA binding and promote transcription of their downstream targets5. RUNX proteins are fundamental in many developmental processes2 such as the regulation of cell cycle kinetics and proliferation6,7 and driving cell fate specification and cellular differentiation8. RUNX family members are necessary for development as loss-of-function mutations cause embryonic lethality2. In addition, the dysregulation of RUNX family members is associated with developmental disorders and cancer9.
Evolution of the RUNX family of transcription factors
RUNT/X genes are found broadly throughout Metazoans10. Vertebrates have evolved three paralogs, RUNX1, RUNX2, and RUNX3, suggested to have arisen through two independent duplications of the ancestral RUNT gene locus near the base of the vertebrate tree (Fig. 1)1,11. Modern RUNX paralogs have maintained strong structural homology and conserved protein domains, and are expressed in two main isoforms from a proximal (P2) or distal (P1) promoter (Fig. 1)1,2,12. While RUNX members play important and overlapping functions in many developmental processes (reviewed in ref. 2), they each have tissue-specific expression patterns indicating some exclusive roles2,8,13. For example, RUNX1 (AML1/CBFA2) is regarded as a master transcription factor regulating hematopoiesis and blood cell development14,15. RUNX3 plays an important role in inflammation and tumor suppression16,17, while RUNX2 (CBFA1) has evolved a unique role in bone development and is regarded as the master regulator of osteogenesis18,19. Unlike other RUNT/X family members, an additional poly-glutamine (Q) (polyQ), poly-alanine (A) (polyA) tandem (QA) repeat domain has evolved within RUNX2 (Fig. 1)20, which was found to play a role in protein transactivation21,22,23,24,25. Moreover, the RUNX2 QA repeat displays large length variation between species, which is often correlated with skull shape in mammals. RUNX2 QA repeat variation has been suggested to subtly impact its molecular function, leading to increased morphological variation within a population on which selection can act. Genes that play fundamental developmental roles and are prone to variation have been labeled as “evolutionary tuning knobs”26, where length variation within these can subtly alter their protein function. However, little is known about the precise role of the RUNX2 QA repeat during skeletal development, nor how and when it emerged during the evolution of vertebrates. In this study, we examine the roles of the RUNX2 QA repeat during osteogenesis and lend perspectives to how changes to its composition may have facilitated the evolution of the vertebrate skeleton. Furthermore, we perform a phylogenetic analysis of the RUNX2 QA repeat across the major vertebrate radiations and discuss how the emergence and expansion of the novel QA domain may have helped to shape vertebrate diversity.
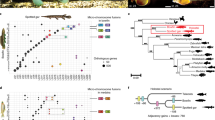
The ancestral metazoan RUNT gene locus underwent two independent rounds of duplication (stars, double lines) near the base of the vertebrate tree to derive the three modern RUNX paralogs in gnathostomes and cyclostomes. Each modern RUNX locus contains a conserved dual P1/P2 promoter, and overall structural homology sharing the RUNT DNA-binding domain, activation (AD) and inhibition (ID) domains, nuclear localization signal (NLS), and C-terminus VWRPY motif. Each member has diverged unique tissue specificity, with RUNX1 controlling hematopoiesis, RUNX2 regulating osteogenesis, and RUNX3 playing supporting roles in chondrogenesis and tumor suppression.
RUNX2 and osteogenic differentiation
RUNX2 plays an essential role during skeletal development, which we briefly summarize below (for detailed reviews see refs. 5,27,28). The two main RUNX2 isoforms (type I and II) are primarily expressed in developing bone and surrounding supporting tissue29,30,31,32,33,34. RUNX2 plays reciprocal roles during osteogenesis, regulating the commitment of undifferentiated mesenchyme towards an osteogenic fate through activation of the osteogenic gene expression network20,34, and in regulating cell cycle kinetics to determine the proliferation rate in osteogenic cell lineages7,35,36,37,38. RUNX2 initiates the osteogenic differentiation pathway through the activation of other transcriptional regulators, namely the zinc-finger transcription factor Sp7 (Osterix/OSX), to drive precursor cells towards a mature osteogenic cell fate39,40. Together, RUNX2 and OSX upregulate collagens (e.g., Col1a1, Col1a2) and osteogenic genes, including Osteocalcin (Bglap) and Osteopontin (Spp1) for osteoblast maturation, terminal differentiation, and formation of a mineralized, calcified matrix5,41. RUNX2 regulates bone formation through two distinct processes: endochondral ossification or intramembranous ossification. Endochondral ossification (e.g., development of the long bones of the limbs) occurs when mesenchymal stem cells differentiate into chondrocytes through the activation of the transcription factor SOX942. Chondrocytes grow and divide to form a cartilage anlage, which later matures and is converted to bone through the activation of RUNX243,44. Intramembranous ossification (e.g., development of the major facial bones and lateral end of the clavicles) occurs when RUNX2 directly regulates the differentiation of mesenchymal stem cells into osteoblasts40,45,46. Here, RUNX2 regulates the rate of mesenchymal cell proliferation and differentiation6,47, directly influencing the rate of growth and formation of bone.
The necessary role of RUNX2 during skeletal development is seen in Runx2-deficient mice, which fail to develop membranous and endochondral bone and have an atrophied cartilaginous skeleton18. Physiological development of the skeleton is RUNX2 dose-dependent, where alterations to RUNX2 expression levels impact the size and shape of bone48. For example, global RUNX2 haploinsufficiency produces the congenital disorder cleidocranial dysplasia (CCD)19. CCD patients display membranous bone abnormalities such as missing or reduced clavicles and craniofacial defects, including a prominent forehead, wide and short skull (brachycephaly), wide eyes (hypertelorism), flat nose, small upper jaw (maxillary hypoplasia), and incomplete closure of the cranial sutures19. Similarly, knockdown or overexpression of RUNX2 in targeted regions can specifically influence the rate of bone growth, without global effects49,50. As such, the temporospatial expression and activation of RUNX2 during skeletal development can ultimately determine the size and shape of individual membranous bones49,51. This demonstrates that not only is RUNX2 an essential regulator of osteogenesis and development of the cranial and postcranial skeleton20,52, but alterations to its expression, regulation, or activation during development may generate novel skeletal variation49.
Protein coding repeats and the RUNX2 QA domain
Of the three RUNX family members, RUNX2 uniquely contains a polyQ, polyA tandem repeat domain. Tandem repeat domains can facilitate a diverse range of biological roles53,54,55, including gene transactivation21, intracellular protein translocation25,56,57, protein–protein, protein–DNA, and protein-RNA interactions55,58. Tandem repeat domains possess high length variability due to their tendency for mutations during DNA replication. High purity repeats with uninterrupted codon composition (e.g., CAG CAG CAG CAG : QQQQ) are more likely to stutter during replication causing homopolymeric expansions or contractions. On the other hand, low purity repeats with variable codon usage (e.g., CAG CAA CAG CAA : QQQQ) are generally more stable and less prone to mutation59. Repeat slippage mutations occur more frequently than point mutations60 and are able to rapidly generate multiallelic variation55 without deleterious consequences such as frameshifts or premature stop codons. As such, alleles with novel repeat lengths can facilitate variation within a population and those that have beneficial functions can become fixed by selection61. Therefore, protein coding repeats have been referred to as evolutionary “tuning knobs”26,60, acting to rapidly generate beneficial genetic variation, independent of longer-term evolutionary epistasis61,62,63,64. (Both nucleotide sequences code for the same homopolymeric amino acid sequence, consisting of 4 glutamines (QQQQ). Bold letters denote how changes to the codon nucelotide sequence do not affect the amino acid sequence, but do change the nucleotide composition that codons are alternated rather than being repetitive).
Tandem repeat domains are present in a wide range of developmental genes and transcription factors61,65,66 and possess substantial intra- and inter-species-specific variation67. However, the biological roles of repeat expansions and contractions are not entirely clear, and often only described in the context of disease. For example, polyQ repeats in coding genes can influence protein–protein interactions with transcriptional co-activators to regulate gene expression68,69 in a length-dependent manner56. However, long repeats can become unstable, forming toxic β-sheet aggregates. This is seen in Huntington’s disease, caused by large unstable polyQ expansions in the HTT (Huntingtin) gene70. The role of polyA repeats is less understood, although they are suggested to facilitate protein translocation between subcellular compartments25,71,72. For example, expanded polyA repeats in RUNX2 cause cytoplasmic aggregation, decreasing protein availability in the nucleus25. Humans and mice with homozygous polyA expansions in the homeobox gene Hoxd13 develop synpolydactyly; however, this phenotype is absent in heterozygous individuals73,74. Yet, while individual polyQ or polyA tracts are common in many proteins75, tandem polyQA repeat domains are exceedingly rare and their precise roles remain poorly understood. The best example is observed in RUNX2, which has been implicated in the fine-tuning of protein transactivation55.
The mammalian RUNX2 polyQ and polyA repeats sit within immediate proximity of each other, separated by a single glutamic acid (E) spacer. The polyQ and polyA domains each form individual α-helix coiled coils, which interact to form a super-coiled secondary structure25,57. In addition to variation in overall repeat length, the ratio of glutamine-to-alanine residues (Q:A ratio) in RUNX2 can also alter the stability of the coiled coil25, altering the affinity for protein–protein interactions and thus gene transactivation66. RUNX2 QA repeat length polymorphisms within a functional range influence its activity, providing a length-dependent mechanism to control gene transactivation. This has been empirically determined in vitro where RUNX2 QA alleles with increasing Q repeats promote transactivation and gene expression22,24. However, Q or A repeats outside the functional range become unstable, lose their transactivation potential, and form aggregates (Fig. 2a, b)21,23,25,76. Targeted deletion of the RUNX2 QA domain (Runx2ΔQA) results in a significant reduction in gene transactivation compared to the wild-type allele in vitro21. Also, conditional knock-in of wild-type Runx2 and the Runx2ΔQA allele in mouse hypertrophic chondrocytes (terminal cartilage) induced osteogenic differentiation and bone mineralization in vivo, although overall this level was reduced in ΔQA mice77,78. Together, these examples demonstrate that while the RUNX2 QA repeat influences osteogenic gene transactivation and the rate of osteogenesis, it is not essential for osteoblast differentiation and bone development. Here, the polyQ domain regulates transactivation levels, while the polyA may support translocation and protein stabilization, although these may be further controlled by variable Q:A ratios.
Schematic of RUNX2 QA repeat mode of action. a QA repeat lengths form a “goldilocks” range that determines function. Short repeats are less functional than medium length repeats, while expanded repeats form protein aggregates, reducing function. b Hypothetical mechanism of action where increasing repeat length promotes interactions with transcriptional co-factors, increasing gene expression (arrows) before hitting a critical threshold inhibiting activity. c QA repeat-driven changes to protein transactivation and downstream gene expression is suggested to fine-tune craniofacial length in several groups of mammals and can cause disease when in excess. OSE osteocalcin-specific element that occurs in osteogenic gene promotors.
Importantly, alterations to the QA ratio can influence the global development of the skeleton and impact morphology. Human RUNX2 variants have revealed that small QA length polymorphisms can subtly, but significantly alter bone mineral density23,24,79, while larger expansions cause subtle CCD-like craniofacial variation and brachydactyly76,80. These findings further demonstrate how variation of the QA domain can alter RUNX2 activity and osteogenic gene transactivation, before reaching a critical threshold and ultimately becoming unstable and causing disease (Fig. 2a, b).
The mammalian RUNX2 QA repeat and craniofacial evolution
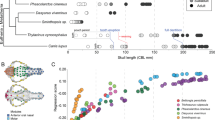
A potential role for the RUNX2 QA repeat in morphological evolution was first identified between breeds of domestic dogs (Canis lupus familiaris). Modern dog breeds display a wide range of skeletal and craniofacial variation as a result of their strong selective breeding (artificial selection). Fondon and Garner61 examined protein repeat length polymorphisms in 36 developmental genes across 92 dog breeds. Overall, dogs possessed elevated repeat purity (see also ref. 62) and many repeat length polymorphisms in key transcription factors61. Of these, large variability was detected in RUNX2 QA repeat length and ratio, which was found to be positively correlated with facial shape, namely the length (r2 = 0.63) and angle of facial bones (klinorhynchy; r2 = 0.51) relative to skull length, between breeds61. Lower QA ratios occurred in breeds with short-faced morphologies, and higher QA ratios in long-faced breeds. Experimental evidence has shown that RUNX2 QA repeat variation can alter its transactivation potential. This may impact the timing or rate of osteogenesis in the developing facial bones, with subtle QA variation influencing craniofacial shape and size (Fig. 2c). This supports the “tuning knob” hypothesis26,60 where RUNX2 QA repeat polymorphisms can impact its function, which in turn, may produce morphological variation that can be subsequently fixed through selection. However, it is important to note that dog breeds represent a single species under extreme artificial selection, and mechanisms that enable rapid variation (such as coding repeat domains) to promote trait evolvability will be favored under such extreme selective pressures. Thus, the question remained whether variation in the RUNX2 QA domain could explain craniofacial variation in naturally evolving species.
Correlations between RUNX2 QA repeat length/ratio and facial shape metrics have been previously investigated throughout several groups of naturally evolving mammals with diverse craniofacial morphologies. These relationships have been examined within marsupial (metatherian)81 and placental (eutherian) mammals82, including Primates83,84, Carnivora22, and Chiroptera (family Phyllostomidae)84. For each examined species, the RUNX2 polyQ and polyA repeats were determined, and the QA ratio was calculated. These data were then compared to specific facial shape metrics, namely the length or width of the membranous facial skeleton. Species QA ratios vs. facial “shape” ratios were individually plotted, and correlations were determined for each group.
Marsupials possess diverse facial shapes, but showed little to no variation in their RUNX2 QA repeat at any phylogenetic level, despite possessing high overall repeat purity81. This was suggested to be in response to the functional constraints placed on skeletal development in the highly altricial young of marsupials, which require accelerated bone development to support their unique mode of reproduction (see refs. 85,86,87). Therefore, the marsupial RUNX2 repeat was hypothesized to be under strong purifying selection81. However, in contrast, RUNX2 QA repeat ratios were found to be significantly correlated with the facial shape within all examined eutherian orders. Primates possess a small range of RUNX2 repeat ratios (Q:A = 1.1–1.7), which were positively correlated with facial length (relative to total skull length; r2 = 0.23, p < 0.005)83. Carnivorans possess a wide range of QA ratios (1.5–5.33), which were positively correlated with facial length (relative to total skull length, r2 = 0.24, p = 0.006)22 and chiropteran phyllostomids possess a small range of QA ratios (1.2–2.33), which were negatively correlated with palate length (r2 = −0.51, p = 0.003) and positively correlated with palate width (r2 = 0.55, p = 0.001), relative to the geometric mean of overall skull shape84. Although when RUNX2 repeat and facial length were compared more broadly across eutherian superorders (Laurasatheria, Xenartha, Eurachontoglires, and Afrotheria), correlations were absent (r2 = <0.1, n.s.)82, suggesting that this relationship exists in epistasis with other factors controlling bone development. However, across smaller evolutionary distances, QA repeat variation may facilitate adaptive morphological evolution, although the definitive role of RUNX2 QA repeat during craniofacial morphogenesis is yet to be determined.
Together, these data lend support to the hypothesis that RUNX2 QA repeat variation may function as an evolutionary “tuning knob”26,60 generating rapid, adaptable variation over short evolutionary distances. However, amino acid repeats are highly volatile sequences that can become unstable, causing detrimental cellular effects84. As such, over larger evolutionary distances, these inherent risks are likely compensated for by other osteogenic gene regulatory changes88,89, illustrated by the absence of RUNX2 repeat length vs. facial shape correlations between eutherian orders. Nevertheless, variability within the RUNX2 QA repeat appears to have played important roles in facilitating the evolution of the mammalian facial skeleton22,81,82,83,84. However, little is known about the QA repeat in other vertebrate clades, when it appeared during vertebrate evolution, nor the roles it may have played in the evolution of the vertebrate skeleton. For the remainder of this perspective, we examine when during vertebrate history the RUNX2 QA repeat emerged and how changes in its composition and structure correspond with the evolution of the major vertebrate radiations.
Evolution of the RUNX2 QA repeat across vertebrate history
The modern RUNX paralogs arose early during vertebrate evolution through whole-genome duplication events. Approximately 50 mya during the late Cambrian, the ancestral vertebrate genome underwent its first duplication, prior to the divergence of the agnathan and gnathostome lineages (Fig. 1)90,91,92. Around 50 mya later in the Ordovician, the agnathan and gnathostome lineages underwent additional, independent whole-genome duplications (Fig. 1)90,91,92. Signatures of these duplication events are observed through comparative sequence analysis of RUNX genes in chordates, where non-vertebrate cephalochordates (amphioxus) and tunicates possess a single RUNT/X gene1, while all extant lineages of vertebrates possess three RUNX paralogs93. However, the three RUNX genes in the basal cyclostome (agnathan) lamprey and hagfish (RunxA, RunxB, and RunxC) are not true one-to-one orthologs with the gnathostome RUNX genes (RUNX1, RUNX2, and RUNX3)11 supporting independent gene duplication events92 (Fig. 1). In addition, phylogenetic comparisons of modern RUNX paralogs reveal greater similarity between the RUNX2 and RUNX3 gene loci, suggesting that they diverged after the initial vertebrate whole-genome duplication1,92. Therefore, RUNX2 evolved in the last common ancestor of gnathostomes ~450 mya1,11,92.
It remained unclear when the QA repeat evolved after the divergence of the gnathostome RUNX2 paralog, and what role it may have played in the evolution of the vertebrate skeleton. As such, we examined the N terminus of RUNX2, containing the QA repeat and flanking sequences, from 409 vertebrate species covering all extant lineages (from publicly available sequences on GenBank, Ensembl, Short-read Archive (SRA), and published observations; Supplementary Methods). The RUNX2 QA repeat domain was defined as the intermittent region between a conserved, N-MSDVS and C-VPRLR motif, at the N terminus of the RUNX2 protein, immediately upstream of the RUNT domain. We defined sequences that lacked an obvious polyQ or polyA, but contained flanking motifs resembling MSDVS and VRPLR as the proto-QA domain. The presence of mostly conserved MSDVS and VPRLR motifs with a short or interrupted polyQ/polyA domain was defined as a primitive QA domain, while an uninterrupted and variable length RUNX2 QA repeat was defined as the variable polyQA domain. We then compared the evolution and expansion of the RUNX2 QA repeat domain across a consensus vertebrate phylogeny (Supplementary Data 1) with median divergence times94.
The divergence of the RUNX2 paralog occurred during the Ordovician and may have facilitated the evolution of complex vertebrates (Fig. 3). The duplication and divergence of RUNX2 (and RUNX3) occurred after the appearance of cartilage95, but before the evolution of the bony skeleton, suggesting it may have been co-opted to establish the primitive skeletal gene regulatory network96,97,98. We were unable to detect the presence of the proto-QA repeat domain in the non-vertebrate, cephalochordate or tunicate runt genes (Supplementary Fig. S1). In vertebrates, we failed to detect a proto-QA domain in the jawless cyclostome RunxA, B, or C, although we did identify some similar features such as sporadic polyQ/H tract in lamprey RunxA (Supplementary Figs. S1, 3). However, we detected the presence of at least the RUNX2 proto-QA domain in all available sequences from extant lineages of gnathostomes. The jawed, but cartilaginous, Chondrichthyes (sharks and rays) possessed a small domain with discernable flanking motifs and a short polyA, but lacked a polyQ (Fig. 3, Supplementary Fig. S1, and Supplementary Data 2).
Simplified phylogeny showing the evolution of the RUNX2 QA repeat during vertebrate history. Evolutionary relationships between major vertebrate radiations are shown with median divergence times. Representative species of each group are shown by silhouettes, and repeat structures with Q, spacer, and A residues are shown for each group on the right. The QA repeat emerged in the last common ancestor of gnathostomes and can be observed in all subsequent extant lineages. The repeat continued to expand throughout early tetrapods and amniotes, before reaching its long and variable modern condition in eutherian mammals <100 million years ago. Taxa silhouettes were created from images under a CC BY 4.0 open license.
We also detected a RUNX2 primitive QA domain throughout all sampled lineages of Osteichthyes bony fish, namely the actinopterygian (ray-finned) Holostei and Teleostei radiations, and sarcopterygian (lobe-finned) Coelacanth (subclass Actinistia) and air-breathing lungfish (subclass Dipnoi)94 (Fig. 3, Supplementary Fig. S1, and Supplementary Data 2). The osteichthyan primitive QA repeat was observed as a small domain with conserved flanking motifs and short polyQ and polyA tracts (Fig. 3). Interestingly, Teleost fish underwent an additional lineage-specific whole-genome duplication99, producing two Runx2 copies (Runx2a and Runx2b) expressed in skeletal tissues100. However, limited sequence data suggest that both orthologs possess the primitive RUNX2 QA repeat, so the influence of this repeat duplication remains unclear. Together, the lack of a QA repeat in cephalochordate/tunicate runt, cyclostome RunxB, and gnathostome RUNX1/3, combined with the presence of a proto-QA domain in extant Chondrichthyes and primitive QA domain in extant Osteichthyes RUNX2, strongly suggests that the RUNX2 QA repeat first evolved after the duplication and divergence of RUNX2, but in the last common ancestor of extant gnathostomes <450 mya.
Curiously, prior to the duplication and divergence of the RUNX2 paralog and evolution of the gnathostomes, several lineages of jawless fish populated the ancient oceans, which possessed “bony” exoskeletal headshields (i.e., the Galeaspida, Pituriaspida, and Osteostraci). However, it is unclear as to whether these early lineages possessed or utilized RUNX2 to develop their ossified headshields due to a lack of extant representatives. Similarly, among the first jawed vertebrates were the extinct Placoderms, known for their large “bony” exoskeletal armor plating97. Although it remains unclear whether they possessed RUNX2 and a proto- or primitive QA repeat. The evolution of the Osteichthyan fish marks a major transition in vertebrate evolution, signified by the first emergence of a true bony endoskeleton and skull96,97,101. The evolution of the RUNX2 QA domain, coinciding with this major vertebrate transition, may have established a novel molecular function during early ossification and skeletal development, particularly given the known transcriptional activation roles of the polyQ56,68,69 and protein translocation roles of the polyA domains71,72. Although small, the proto- and primitive QA domains may have established novel protein–protein interactions or affinities for transcriptional co-activators56,68,69,71,72 enhancing ossification capacity in the early skeleton. The early emergence of the proto-QA provided a template for the evolutionary expansion of the modern QA domain, which may have acted as a primer for the evolution of morphological complexity in tetrapods.
The emergence of a variable length RUNX2 polyQA repeat occurred ~350 mya during the tetrapod radiation, first observed in the amphibia94. Amphibians represent the most basal tetrapod lineage and possess a primitive/variable QA repeat with conserved flanking residues, a distinct spacer residue, and polyQ and polyA domains with interrupting residues, which vary in length between groups. Caecillians (order Gymnophiona) possess a short QA domain (2 Q:3–5 A), while frogs and toads (order Anura) have evolved an expanded polyQ domain (6–10 Q:2 A) with several proline interruptions (Supplementary Data 3 and Fig. 3). The initial expansion of the repeat domain may have required compensatory changes for novel protein functions. For example, proline interruptions in both polyQ and polyA repeats are found to decrease coiled-coil stabilization, reducing expansion-related protein aggregation25,57.
Expansion of the variable RUNX2 QA repeat occurred with the emergence of amniotes ~312 mya, observed in all extant lineages of Reptilia, Aves, and Mammalia94. Reptilian clades, such as the Rhynchocephalia (7 Q:6 A), Testudinata (turtles and tortoises; 6–9 Q:3–7 A), and Crocodilia (10 Q:4 A) have a short repeat with minor interspecific length variation (Supplementary Data 4). However, groups such as Squamata (snakes, lizards, and geckos) and Aves (birds) have a large repeat with intraspecific variability. For example, snakes (Serpentes, Squamata) have evolved a long polyQ domain (12–21 Q; QA ratio 2.0–5.25; Supplementary Data 4 and Fig. 3). Similar length expansions and variation were observed in taxa from the Iguania (Squamata), largely in Anolis lizards. Anoles are well known for their extraordinary adaptive radiation102 and convergent ecomorphs in cranial shape arising over the past 50 million years103,104,105,106. Members of the Anolis genus showed large variation in repeat length and composition (Supplementary Data 4), having a short polyA repeat but expanded and variable polyQ domain (10–27 Q:4–6 A) with several histidine, alanine, proline, and serine interruptions. As mentioned previously, these may have evolved to compensate for the rapid and hyper-expansion of their repeat structure to reduce protein aggregation and stabilize function25,57. Interestingly, the adaptive diversification of the anole skull may have been facilitated by RUNX2 QA variation to enable the rapid evolution of disparate and convergent ecomorphs. As such, the disparity in Anolis RUNX2 QA repeat length and ratio provides a unique opportunity to examine whether QA repeat variation facilitates the adaptive evolution of facial shape in a naturally evolving family, acting as a natural counterpoint to observations in domestic dogs61.
Birds (Aves) represent a relatively recent vertebrate evolutionary radiation, diverging from their therapod dinosaurian ancestors ~160 mya, with the modern avian crown group appearing ~111 mya. Modern birds have since diversified into over 10,000 different species94,102, displaying a broad spectrum of sizes and body forms in response to their unique ecologies. This is reflected by many specific skeletal adaptations in response to their locomotor demands (i.e., powered flight, swimming, gliding, or terrestrial bipedalism) and remarkable diversity in beak shape reflective of their specialized diets107. Interestingly, birds possess minor RUNX2 QA repeat length variation within sampled Neognathae and Palaeognathae orders, although some groups display large inter-order variation. For example, sampled Tinamiformes possess low Q:A ratios (n = 3; Q:A ≤ 1.80), while sampled Galliformes possess conserved, high QA repeat ratios (n = 8; Q:A 6.0; Supplementary Data 5). This high intra-order conservation of QA repeat length is in contrast to mammals, which show large intra-order variation suggested to facilitate facial shape evolution22,82,83,84. Instead, the ancestors of modern bird groups may have utilized repeat length variation to promote lineage-specific radiations and adaptive beak evolution107,108,109, subsequently preserved within extant species. An unpublished study examining correlations between RUNX2 QA repeat composition and beak length in shorebirds (Charadriiformes) found that, in contrast to our hypothesis, that this order possesses highly variable QA ratios (1.86–4.25) that are weakly correlated with beak length (R2 = 0.13)110. However, RUNX2 QA repeat sequences for these taxa are not publicly available, so this result could not be verified. Additional correlative examinations of RUNX2 QA repeat vs. beak shape will help define the role of RUNX2 in avian morphological diversification.
Mammals diverged from other amniotes ~177 million years94,111 and have since evolved a remarkable array of adaptations in response to terrestrial, arboreal, aerial, subterranean, and aquatic environments. Mammals are characterized by several unique craniofacial traits, such as the secondary palate and temporo-mandibular joint, which have established novel feeding ecologies through jaw articulation101,112,113,114,115. These novel morphological characteristics coincide with the longest RUNX2 QA repeat of all vertebrate lineages examined. Extant monotremes (n = 2, 16–20 Q:6–19 A) and marsupials (n = 26, 16–24 Q:19–22 A) possess some minor RUNX2 repeat variation, while eutherian mammals (n = 162, 7–31 Q:4–19 A) display extreme repeat length variation across extant lineages (Supplementary Data 6 and Fig. 3). For example, the naked mole-rat (Heterocephalus glaber) has the largest ratio (31 Q:5 A, QA ratio = 6.2), while the Baiji, or Yangtze River dolphin (Lipotes vexillifer) has the shortest ratio (7 Q:16 A; QA ratio = 0.44; Supplementary Data 6). Eutherian mammals diverged from the marsupials and monotremes, ~160 and ~177 mya, respectively, and radiated into four superorders ~105 mya94,111. As such, the extreme RUNX2 QA repeat variation observed between eutherian mammals is a recent phenomenon occurring within the past ~100 mya.
Future directions
We highlight several independent studies with consistently recovered correlations between RUNX2 QA repeat variability and facial length morphology at multiple taxonomic levels22,82,83,84. However, additional examinations in other groups of naturally evolving vertebrates will further support the role of the RUNX2 QA repeat variation underlying morphological evolution. For example, the previously mentioned Anoles represent a natural case study by which RUNX2 repeat evolution may influence adaptive convergence of cranial ecomorphs107 and studies in birds will elucidate whether RUNX2 QA repeat variation promotes adaptive changes in beak shape diversity107. In addition, leporid rabbits and hares (order Lagomorpha) are a relatively recent (~20 mya) evolutionary radiation that display substantial craniofacial length and angle variation in response to different ecologies116,117. However, how RUNX2 QA repeat variation corresponds with the natural craniofacial variation in this group remains unknown.
The reported within-group correlations, combined with several lines of empirical evidence for variable polyQ and polyA repeats altering RUNX2 transactivation, suggest that the RUNX2 QA repeat is a putative mechanism that can produce morphological variation that selection can act upon. However, it is important to note that while these anecdotal correlations and empirical evidence exist, the direct influence of RUNX2 QA repeat variation on skeletal morphogenesis is still yet to be determined. Experimental approaches utilizing genome editing in a range of model vertebrates118 will unequivocally reveal the role of RUNX2 QA repeat variation on the formation of the vertebrate skeleton. Particularly, replacement of the endogenous RUNX2 QA domain with varying length QA polymorphisms will allow precise quantification of its contribution to bone development and morphological evolution.
Concluding remarks
In this perspective, we examined the origin and emergence of the RUNX2 QA repeat across the evolution of vertebrates and drew comparisons with empirical studies to determine how this may have shaped vertebrate diversity. This is the first study to assess broad taxonomic sampling with deep evolutionary coverage to analyze sequence variation in the RUNX2 repeat domain. Through this investigation, we have uncovered several fascinating concordances of the emergence and expansion of the RUNX2 QA repeat domain with the major vertebrate transitions. The duplication and divergence of the RUNX2 paralog from the ancestral RUNT gene ~450 mya may have primed the development of a bone-specific program, establishing the evolution of the vertebrate bony skeleton. The skeleton supported the emergence of morphological novelty across vertebrates, acting as a scaffold for unique adaptations. The internal RUNX2 protein QA repeat, absent in the structurally homologous RUNX1 and RUNX3, has provided a novel functional mechanism to fine-tune osteogenesis through enhanced protein–protein interactions and gene transactivation. Therefore, the evolution of the RUNX2 QA repeat has likely played a critical role in shaping the wide range of diversity seen across vertebrates.
Since first appearing in the ancestor of gnathostomes, the QA repeat has sequentially emerged, evolved, and expanded through the divergence of vertebrates, reaching its highly variable configuration with the evolution of amniotes ~312 mya. However, the broad repeat variability observed in eutherian mammals is a recent evolutionary event occurring within the past ~100 million years. The gradual evolution of the internal QA repeat throughout vertebrates may have promoted an increasing ability to subtly alter the development of the craniofacial skeleton through its direct role in intramembranous ossification. The progressive expansion and stabilization of the QA repeat throughout vertebrates demonstrate that it has been fixed during evolution, emphasizing its important roles. While additional studies are required to define the precise role of the RUNX2 QA repeat during skeletal development, the evolution and emergence of the RUNX2 QA repeat provide an intriguing putative mechanism underlying vertebrate evolution.
Data availability
All sequence data that support the findings of this study are deposited in the GenBank, SRA, Whole-Genome Shotgun, and TSA repositories with accession codes listed in the Supplementary files.
Change history
16 January 2021
A Correction to this paper has been published: https://doi.org/10.1038/s42003-021-01687-0
25 January 2021
A Correction to this paper has been published: https://doi.org/10.1038/s42003-021-01687-0
References
Rennert, J., Coffman, J. A., Mushegian, A. R. & Robertson, A. J. The evolution of Runx genes I. A comparative study of sequences from phylogenetically diverse model organisms. BMC Evol. Biol. 3, 1–11 (2003).
Mevel, R., Draper, J. E., Lie-a-Ling, M., Kouskoff, V. & Lacaud, G. RUNX transcription factors: orchestrators of development. Development 146, dev148296 (2019).
Bruno, L. et al. Selective deployment of transcription factor paralogs with submaximal strength facilitates gene regulation in the immune system. Nat. Immunol. 20, 1372–1380 (2019).
Kagoshima, H. et al. The runt domain identifies a new family of heterometric transcriptional regulators. Trends Genet. 9, 338–341 (1993).
Komori, T. Requisite roles of Runx2 and Cbfb in skeletal development. J. Bone Miner. Metab. 21, 193–197 (2003).
Teplyuk, N. M. et al. Runx2 regulates G protein-coupled signaling pathways to control growth of osteoblast progenitors. J. Biol. Chem. 283, 27585–27597 (2008).
Shen, R. et al. Cyclin D1-Cdk4 induce Runx2 ubiquitination and degradation. J. Biol. Chem. 281, 16347–16353 (2006).
Liu, P., Neil, J. C. & Speck, N. A. RUNX Proteins in Development and Cancer, Vol. 962 (Springer Singapore, 2017).
Ito, Y., Bae, S. C. & Chuang, L. S. H. The RUNX family: Developmental regulators in cancer. Nat. Rev. Cancer 15, 81–95 (2015).
Duncan, E. J., Wilson, M. J., Smith, J. M. & Dearden, P. K. Evolutionary origin and genomic organisation of runt-domain containing genes in arthropods. BMC Genomics 9, 558 (2008).
Nah, G. S. S., Tay, B. H., Brenner, S., Osato, M. & Venkatesh, B. Characterization of the runx gene family in a jawless vertebrate, the Japanese lamprey (Lethenteron japonicum). PLoS ONE 9, e113445 (2014).
Stock, M. & Otto, F. Control of RUNX2 isoform expression: The role of promoters and enhancers. J. Cell. Biochem. 95, 506–517 (2005).
Levanon, D. & Groner, Y. Structure and regulated expression of mammalian RUNX genes. Oncogene 23, 4211–4219 (2004).
Okuda, T., Nishimura, M., Nakao, M. & Fujita, Y. RUNX1/AML1: a central player in hematopoiesis. Int. J. Hematol. 74, 252–257 (2001).
De Bruijn, M. & Dzierzak, E. Runx transcription factors in the development and function of the definitive hematopoietic system. Blood 129, 2061–2069 (2017).
Guo, W. H. et al. Inhibition of growth of mouse gastric cancer cells by Runx3, a novel tumor suppressor. Oncogene 21, 8351–8355 (2002).
Cohen, M. M. Perspectives on RUNX genes: an update. Am. J. Med. Genet. A 149, 2629–2646 (2009).
Komori, T. et al. Targeted disruption of Cbfa1 results in a complete lack of bone formation owing to maturational arrest of osteoblasts. Cell 89, 755–764 (1997).
Otto, F. et al. Cbfa1, a candidate gene for cleidocranial dysplasia syndrome, is essential for osteoblast differentiation and bone development. Cell 89, 765–771 (1997).
Ducy, P., Zhang, R., Geoffroy, V., Ridall, A. L. & Karsenty, G. Osf2/Cbfa1: a transcriptional activator of osteoblast differentiation. Cell 89, 747–754 (1997).
Thirunavukkarasu, K., Mahajan, M., McLarren, K. W., Stifani, S. & Karsenty, G. Two domains unique to osteoblast-specific transcription factor Osf2/Cbfa1 contribute to its transactivation function and its inability to heterodimerize with Cbfβ. Mol. Cell. Biol. 18, 4197–4208 (1998).
Sears, K. E., Goswami, A., Flynn, J. J. & Niswander, L. A. The correlated evolution of Runx2 tandem repeats, transcriptional activity, and facial length in Carnivora. Evol. Dev. 9, 555–565 (2007).
Morrison, N. A. et al. Glutamine repeat variants in human RUNX2 associated with decreased femoral neck BMD, broadband ultrasound attenuation and target gene transactivation. PLoS ONE 7, e42617 (2012).
Morrison, N. A. et al. Polyalanine repeat polymorphism in RUNX2 is associated with site-specific fracture in post-menopausal females. PLoS ONE 8, e72740 (2013).
Pelassa, I. et al. Association of polyalanine and polyglutamine coiled coils mediates expansion disease-related protein aggregation and dysfunction. Hum. Mol. Genet. 23, 3402–3420 (2014).
King, D. G., Soller, M. & Kashi, Y. Evolutionary tuning knobs. Endeavour 21, 36–40 (1997).
Bruderer, M., Richards, R. G., Alini, M. & Stoddart, M. J. Role and regulation of runx2 in osteogenesis. Eur. Cells Mater. 28, 269–286 (2014).
Vimalraj, S., Arumugam, B., Miranda, P. J. & Selvamurugan, N. Runx2: Structure, function, and phosphorylation in osteoblast differentiation. Int. J. Biol. Macromol. 78, 202–208 (2015).
Choi, K.-Y. Y. et al. Spatio-temporal expression patterns of Runx2 isoforms in early skeletogenesis. Exp. Mol. Med. 34, 426–433 (2002).
Xiao, Z. S., Liu, S. G., Hinson, T. K. & Quarles, L. D. Characterization of the upstream mouse Cbfa1/Runx2 promoter. J. Cell. Biochem. 82, 647–659 (2001).
Xiao, Z. S., Thomas, R., Hinson, T. K. & Quarles, L. D. Genomic structure and isoform expression of the mouse, rat and human Cbfa1/Osf2 transcription factor. Gene 214, 187–197 (1998).
Li, Y. i., Xiao, Z.-s. & Sheng, W. Advances in Runx2 regulation and its isoforms. Med. Hypotheses 68, 169–175 (2006).
Park, M. H. et al. Differential expression patterns of Runx2 isoforms in cranial suture morphogenesis. J. Bone Miner. Res. 16, 885–892 (2001).
Tai, P. W. L. et al. Epigenetic landscape during osteoblastogenesis defines a differentiation-dependent Runx2 promoter region. Gene 550, 1–9 (2014).
San Martin, I. A. et al. Impaired cell cycle regulation of the osteoblast-related heterodimeric transcription factor Runx2-Cbfβ in osteosarcoma cells. J. Cell. Physiol. 221, 560–571 (2009).
Pratap, J. et al. Cell growth regulatory role of Runx2 during proliferative expansion of preosteoblasts. Cancer Res. 63, 5357–5362 (2003).
Galindo, M. et al. The bone-specific expression of Runx2 oscillates during the cell cycle to support a G1-related antiproliferative function in osteoblasts. J. Biol. Chem. 280, 20274–20285 (2005).
Qiao, M. et al. Cell cycle-dependent phosphorylation of the RUNX2 transcription factor by cdc2 regulates endothelial cell proliferation. J. Biol. Chem. 281, 7118–7128 (2006).
Hojo, H., Ohba, S., He, X., Lai, L. P. & McMahon, A. P. Sp7/Osterix is restricted to bone-forming vertebrates where it acts as a Dlx co-factor in osteoblast specification. Dev. Cell 37, 238–253 (2016).
Takarada, T. et al. Genetic analysis of Runx2 function during intramembranous ossification. Development 143, 211–218 (2016).
Komori, T. Regulation of bone development and extracellular matrix protein genes by RUNX2. Cell Tissue Res. 339, 189–195 (2010).
Akiyama, H., Chaboissier, M. C., Martin, J. F., Schedl, A. & De Crombrugghe, B. The transcription factor Sox9 has essential roles in successive steps of the chondrocyte differentiation pathway and is required for expression of Sox5 and Sox6. Genes Dev. 16, 2813–2828 (2002).
Zhou, N. et al. BMP2 induces chondrogenic differentiation, osteogenic differentiation and endochondral ossification in stem cells. Cell Tissue Res. 366, 101–111 (2016).
Eames, B. F., Sharpe, P. T. & Helms, J. A. Hierarchy revealed in the specification of three skeletal fates by Sox9 and Runx2. Dev. Biol. 274, 188–200 (2004).
James, M. J., Järvinen, E., Wang, X. P. & Thesleff, I. Different roles of Runx2 during early neural crest-derived bone and tooth development. J. Bone Miner. Res. 21, 1034–1044 (2006).
Abzhanov, A., Rodda, S. J., McMahon, A. P. & Tabin, C. J. Regulation of skeletogenic differentiation in cranial dermal bone. Development 134, 3133–3144 (2007).
Kawane, T. et al. Runx2 is required for the proliferation of osteoblast progenitors and induces proliferation by regulating Fgfr2 and Fgfr3. Sci. Rep. 8, 1–17 (2018).
Zhang, S. et al. Dose-dependent effects of Runx2 on bone development. J. Bone Miner. Res. 24, 1889–1904 (2009).
Hall, J. et al. Evolution of a developmental mechanism: Species-specific regulation of the cell cycle and the timing of events during craniofacial osteogenesis. Dev. Biol. 385, 380–395 (2014).
Takarada, T. et al. An analysis of skeletal development in osteoblast-specific and chondrocyte-specific runt-related transcription factor-2 (Runx2) knockout mice. J. Bone Miner. Res. 28, 2064–2069 (2013).
Shirai, Y. et al. Runx2 function in cells of neural crest origin during intramembranous ossification. Biochem. Biophys. Res. Commun. 509, 1028–1033 (2019).
Ducy, P. & Karsenty, G. Two distinct osteoblast-specific cis-acting elements control expression of a mouse osteocalcin gene. Mol. Cell. Biol. 15, 1858–1869 (2015).
Hannan, A. J. Tandem repeat polymorphisms: modulators of disease susceptibility and candidates for ‘missing heritability’. Trends Genet. 26, 59–65 (2010).
Hannan, A. J. Tandem repeats mediating genetic plasticity in health and disease. Nat. Rev. Genet. 19, 286–298 (2018).
Lynch, V. J. & Wagner, G. P. Resurrecting the role of transcription factor change in developmental evolution. Evolution 62, 2131–2154 (2008).
Gemayel, R. et al. Variable glutamine-rich repeats modulate transcription factor activity. Mol. Cell 59, 615–627 (2015).
Fiumara, F., Fioriti, L., Kandel, E. R. & Hendrickson, W. A. Essential role of coiled coils for aggregation and activity of Q/N-rich prions and polyq proteins. Cell 143, 1121–1135 (2010).
Emili, A., Greenblatt, J. & Ingles, C. J. Species-specific interaction of the glutamine-rich activation domains of Sp1 with the TATA box-binding protein. Mol. Cell. Biol. 14, 1582–1593 (1994).
Tautz, D. & Schlötterer, C. Simple sequences. Curr. Opin. Genet. Dev. 4, 832–837 (1994).
Kashi, Y. & King, D. G. Simple sequence repeats as advantageous mutators in evolution. Trends Genet. 22, 253–259 (2006).
Fondon, J. W. & Garner, H. R. Molecular origins of rapid and continuous morphological evolution. Proc. Natl Acad. Sci. USA 101, 18058–18063 (2004).
Southwestern, U. T. et al. Elevated basal slippage mutation rates among the Canidae. J. Hered. 98, 452–460 (2007).
Caburet, S., Cocquet, J., Vaiman, D. & Veitia, R. A. Coding repeats and evolutionary ‘agility’. BioEssays 27, 581–587 (2005).
Gemayel, R., Vinces, M. D., Legendre, M. & Verstrepen, K. J. Variable tandem repeats accelerate evolution of coding and regulatory sequences. Annu. Rev. Genet. 44, 445–477 (2010).
Wren, J. D. et al. Repeat polymorphisms within gene regions: phenotypic and evolutionary implications. Am. J. Hum. Genet. 67, 345–356 (2000).
Rose, A. & Meier, I. Scaffolds, levers, rods and springs: diverse cellular functions of long coiled-coil proteins. Cell. Mol. Life Sci. 61, 1996–2009 (2004).
Mortlock, D. P., Sateesh, P. & Innis, J. W. Evolution of N-terminal sequences of the vertebrate HOXA13 protein. Mamm. Genome 11, 151–158 (2000).
Nucifora, J. et al. Interference by huntingtin and atrophin-1 with CBP-mediated transcription leading to cellular toxicity. Science 291, 2423–2428 (2001).
Steffan, J. S. et al. Histone deacetylase inhibitors arrest polyglutamine-dependent neurodegeneration in Drosophila. Nature 413, 739–743 (2001).
Bates, G. P. et al. Huntington disease. Nat. Rev. Dis. Prim. 1, 15005 (2015).
Li, L., Ng, N. K. L., Koon, A. C. & Chan, H. Y. E. Expanded polyalanine tracts function as nuclear export signals and promote protein mislocalization via eEF1A1 factor. J. Biol. Chem. 292, 5784–5800 (2017).
Banerjee, A. et al. Proteomic analysis reveals that wildtype and alanine-expanded nuclear poly(A)-binding protein exhibit differential interactions in skeletal muscle. J. Biol. Chem. 294, 7360–7376 (2019).
Brown, L. Y. & Brown, S. A. Alanine tracts: the expanding story of human illness and trinucleotide repeats. Trends Genet. 20, 51–58 (2004).
Johnson, K. R. et al. A new spontaneous mouse mutation of Hoxd13 with a polyalanine expansion and phenotype similar to human synpolydactyly. Hum. Mol. Genet. 7, 1033–1038 (1998).
Almeida, B., Fernandes, S., Abreu, I. A. & Macedo-Ribeiro, S. Trinucleotide repeats: a structural perspective. Front. Neurol. 4, 1–24 (2013).
Mastushita, M. et al. A glutamine repeat variant of the RUNX2 gene causes cleidocranial dysplasia. Mol. Syndromol. 6, 50–53 (2015).
Sato, S. et al. The distinct role of the runx proteins in chondrocyte differentiation and intervertebral disc degeneration: findings in murine models and in human disease. Arthritis Rheum. 58, 2764–2775 (2008).
Takeda, S., Bonnamy, J. P., Owen, M. J., Ducy, P. & Karsenty, G. Continuous expression of Cbfa1 in nonhypertrophic chondrocytes uncovers its ability to induce hypertrophic chondrocyte differentiation and partially rescues Cbfa1-deficient mice. Genes Dev. 15, 467–481 (2001).
Vaughan, T., Pasco, J. A., Kotowicz, M. A., Nicholson, G. C. & Morrison, N. A. Alleles of RUNX2/CBFA1 gene are associated with differences in bone mineral density and risk of fracture. J. Bone Miner. Res. 17, 1527–1534 (2002).
Mundlos, S. et al. Mutations involving the transcription factor CBFA1 cause cleidocranial dysplasia. Cell 89, 773–779 (1997).
Newton, A. H., Feigin, C. Y. & Pask, A. J. RUNX2 repeat variation does not drive craniofacial diversity in marsupials. BMC Evol. Biol. 17, 1–9 (2017).
Pointer, M. A. et al. RUNX2 tandem repeats and the evolution of facial length in placental mammals. BMC Evol. Biol. 12, 103 (2012).
Ritzman, T. B. et al. Facing the facts: the Runx2 gene is associated with variation in facial morphology in primates. J. Hum. Evol. 111, 139–151 (2017).
Ferraz, T. et al. Contrasting patterns of RUNX2 repeat variations are associated with palate shape in phyllostomid bats and New World primates. Sci. Rep. 8, 1–10 (2018).
Tyndale-Biscoe, C. H. & Janssens, P. A. The Developing Marsupial. The British Journal of Psychiatr, Vol. 111 (Springer Berlin Heidelberg, 1988).
Smith, K. K. Craniofacial development in marsupial mammals: developmental origins of evolutionary change. Dev. Dyn. 235, 1181–1193 (2006).
Smith, K. K. Comparative patterns of craniofacial development in Eutherian and Metatherian mammals. Evolution 51, 1663 (1997).
Carroll, S. B. Endless forms: the evolution of gene regulation and morphological diversity. Cell 101, 577–580 (2000).
Carroll, S. B. Evo-devo and an expanding evolutionary synthesis: a genetic theory of morphological evolution. Cell 134, 25–36 (2008).
Sacerdot, C., Louis, A., Bon, C., Berthelot, C. & Roest Crollius, H. Chromosome evolution at the origin of the ancestral vertebrate genome. Genome Biol. 19, 166 (2018).
Kuraku, S., Meyer, A. & Kuratani, S. Timing of genome duplications relative to the origin of the vertebrates: did cyclostomes diverge before or after? Mol. Biol. Evol. 26, 47–59 (2008).
Simakov, O. et al. Deeply conserved synteny resolves early events in vertebrate evolution. Nat. Ecol. Evol. 4, 820–830 (2020).
Hecht, J. et al. Evolution of a core gene network for skeletogenesis in chordates. PLoS Genet. 4, https://doi.org/10.1371/journal.pgen.1000025 (2008).
Kumar, S., Stecher, G., Suleski, M. & Hedges, S. B. TimeTree: a resource for Timelines, Timetrees, and divergence times. Mol. Biol. Evol. 34, 1812–1819 (2017).
Jandzik, D. et al. Evolution of the new vertebrate head by co-option of an ancient chordate skeletal tissue. Nature 518, 534–537 (2015).
Fisher, S. & Franz-Odendaal, T. Evolution of the bone gene regulatory network. Curr. Opin. Genet. Dev. 22, 390–397 (2012).
Kaucka, M. & Adameyko, I. Evolution and development of the cartilaginous skull: From a lancelet towards a human face. Semin. Cell Dev. Biol. 91, 2–12 (2019).
Kuratani, S., Kusakabe, R. & Hirasawa, T. The neural crest and evolution of the head/trunk interface in vertebrates. Dev. Biol. 444, S60–S66 (2018).
Glasauer, S. M. K. & Neuhauss, S. C. F. Whole-genome duplication in teleost fishes and its evolutionary consequences. Mol. Genet. Genomics 289, 1045–1060 (2014).
Vega, M. et al. Duplicate zebrafish runx2 orthologues are expressed in developing skeletal elements. Gene Expr. Patterns 4, 573–581 (2004).
Fish, J. L. Evolvability of the vertebrate craniofacial skeleton. Semin. Cell Dev. Biol. 91, 13–22 (2019).
Brusatte, S. L., O’Connor, J. K. & Jarvis, E. D. The origin and diversification of birds. Curr. Biol. 25, R888–R898 (2015).
Pinto, G., Mahler, D. L., Harmon, L. J. & Losos, J. B. Testing the island effect in adaptive radiation: rates and patterns of morphological diversification in Caribbean and mainland Anolis lizards. Proc. R. Soc. Ser. B 275, 2749–2757 (2008).
Mahler, D. L., Revell, L. J., Glor, R. E. & Losos, J. B. Ecological opportunity and the rate of morphological evolution in the diversification of greater Antillean anoles. Evolution 64, 2731–2745 (2010).
Mahler, D. L., Ingram, T., Revell, L. J. & Losos, J. B. Exceptional convergence on the macroevolutionary landscape in island lizard radiations. Science 341, 292–295 (2013).
Stayton, C. T. The definition, recognition, and interpretation of convergent evolution, and two new measures for quantifying and assessing the significance of convergence. Evolution 69, 2140–2153 (2015).
Cooney, C. R. et al. Mega-evolutionary dynamics of the adaptive radiation of birds. Nature 542, 344–347 (2017).
Bright, J. A., Marugán-Lobón, J., Cobb, S. N. & Rayfield, E. J. The shapes of bird beaks are highly controlled by nondietary factors. Proc. Natl Acad. Sci. USA 113, 5352–5357 (2016).
Tokita, M., Yano, W., James, H. F. & Abzhanov, A. Cranial shape evolution in adaptive radiations of birds: comparative morphometrics of Darwin’s finches and Hawaiian honeycreepers. Philos. Trans. R. Soc. Ser. B 372, 20150481 (2017).
Green, R. M. & Kimball, R. T. Analysis of RUNX2 Gene’s Influence on Bill Morphology Within Shore Birds. Thesis 3737, The University of Florida (2012).
Bininda-Emonds, O. R. P. et al. The delayed rise of present-day mammals. Nature 446, 507–512 (2007).
Hinton, R. J., Jing, J. & Feng, J. Q. Genetic influences on temporomandibular joint development and growth. Curr. Top. Dev. Biol. 115, 85–109 (2015).
Allin, E. F. Evolution of the mammalian middle ear. J. Morphol. 147, 403–437 (1975).
Anthwal, N., Joshi, L. & Tucker, A. S. Evolution of the mammalian middle ear and jaw: adaptations and novel structures. J. Anat. 222, 147–160 (2013).
Weisbecker, V. Monotreme ossification sequences and the riddle of mammalian skeletal development. Evolution 65, 1323–1335 (2011).
Kraatz, B. & Sherratt, E. Evolutionary morphology of the rabbit skull. PeerJ 2016, 1–23 (2016).
Kraatz, B. P., Sherratt, E., Bumacod, N. & Wedel, M. J. Ecological correlates to cranial morphology in Leporids (Mammalia, Lagomorpha). PeerJ 2015, 1–20 (2015).
Yang, L., Mali, P., Kim-Kiselak, C. & Church, G. CRISPR-Cas-mediated targeted genome editing in human cells. Methods Mol. Biol. 1114, 245–267 (2014).
Author information
Authors and Affiliations
Contributions
A.H.N. and A.J.P. conceived the study and wrote and edited the manuscript. A.H.N. performed data analysis and generated figures.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Newton, A.H., Pask, A.J. Evolution and expansion of the RUNX2 QA repeat corresponds with the emergence of vertebrate complexity. Commun Biol 3, 771 (2020). https://doi.org/10.1038/s42003-020-01501-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-020-01501-3
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.