Abstract
Nitrogenases uniquely reduce atmospheric N2 to bioavailable ammonium. They group into three isoforms that primarily differ in the architecture of their active-site cofactors. A molybdenum or vanadium ion is introduced into a common precursor cluster to form Mo- and V-dependent nitrogenases, respectively. In contrast, the third class of the enzyme only utilizes abundant iron to reduce N2 under ambient conditions and is consequently of high interest for mechanistic studies and catalyst design. Here we report the three-dimensional structure of Fe-nitrogenase from Azotobacter vinelandii and its FeFe cofactor, a [8Fe:9S:C] cluster with an interstitial carbide and an organic homocitrate ligand at the apical iron that substitutes for Mo or V in the other isoforms. The structure reveals lability of sulfide S2B, the proposed binding site for substrate in other nitrogenases, further supporting a general mechanism of proton and electron transfer for all nitrogenases and all their substrates.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The atomic coordinates and structure factors for A. vinelandii Fe-only nitrogenase FeFe protein have been deposited with the Protein Data Bank at http://www.pdb.org with accession code 8BOQ. All other data are available from the authors upon reasonable request. Source data are provided with this paper.
References
Canfield, D. E., Glazer, A. N. & Falkowski, P. G. The evolution and future of Earth’s nitrogen cycle. Science 330, 192–196 (2010).
Rees, D. C. Dinitrogen reduction by nitrogenase—if N2 isn’t broken, it can’t be fixed. Curr. Opin. Struct. Biol. 3, 921–928 (1993).
Wilson, P. W. & Burris, R. H. The mechanism of biological nitrogen fixation. Bacteriol. Rev. 11, 41–73 (1947).
Smil, V. The world’s greatest fix: a history of nitrogen in agriculture. Nature 431, 909–910 (2004).
Norskov, J. K. & Chen, J. Sustainable Ammonia Synthesis, Round Table Report (US DoE, 2016).
Chen, J. G. et al. Beyond fossil fuel-driven nitrogen transformations. Science 360, eaar6611 (2018).
Ribbe, M. W., Hu, Y. L., Hodgson, K. O. & Hedman, B. Biosynthesis of nitrogenase metalloclusters. Chem. Rev. 114, 4063–4080 (2014).
Jimenez-Vicente, E. et al. Sequential and differential interaction of assembly factors during nitrogenase MoFe protein maturation. J. Biol. Chem. 293, 9812–9823 (2018).
Einsle, O. & Rees, D. C. Structural enzymology of nitrogenase enzymes. Chem. Rev. 120, 4969–5004 (2020).
Bulen, W. A. & LeComte, J. R. Nitrogenase system from Azotobacter: 2-enzyme requirement for N2 reduction, ATP-dependent H2 evolution and ATP hydrolysis. Proc. Natl Acad. Sci. USA 56, 979–986 (1966).
Einsle, O. Nitrogenase FeMo cofactor: an atomic structure in three simple steps. J. Biol. Inorg. Chem. 19, 737–745 (2014).
Einsle, O. et al. Nitrogenase MoFe-protein at 1.16 Å resolution: a central ligand in the FeMo-cofactor. Science 297, 1696–1700 (2002).
Spatzal, T. et al. Evidence for interstitial carbon in nitrogenase FeMo cofactor. Science 334, 940 (2011).
Lancaster, K. M. et al. X-ray emission spectroscopy evidences a central carbon in the nitrogenase iron-molybdenum cofactor. Science 334, 974–977 (2011).
Sippel, D. & Einsle, O. The structure of vanadium nitrogenase reveals an unusual bridging ligand. Nat. Chem. Biol. 13, 956–960 (2017).
Yang, J. G., Xie, X. Q., Wang, X., Dixon, R. & Wang, Y. P. Reconstruction and minimal gene requirements for the alternative iron-only nitrogenase in Escherichia coli. Proc. Natl Acad. Sci. USA 111, E3718–E3725 (2014).
Good, A. Toward nitrogen-fixing plants. Science 359, 869–870 (2018).
Curatti, L. & Rubio, L. M. Challenges to develop nitrogen-fixing cereals by direct nif-gene transfer. Plant Sci. 225, 130–137 (2014).
Lee, C. C., Hu, Y. L. & Ribbe, M. W. Vanadium nitrogenase reduces CO. Science 329, 642 (2010).
Zheng, Y. N. et al. A pathway for biological methane production using bacterial iron-only nitrogenase. Nat. Microbiol. 3, 281–286 (2018).
Diederichs, K. et al. Crystal structure of MalK, the ATPase subunit of the trehalose/maltose ABC transporter of the archaeon Thermococcus litoralis. EMBO J. 19, 5951–5961 (2000).
Oehlmann, N. N. & Rebelein, J. G. The conversion of carbon monoxide and carbon dioxide by nitrogenases. ChemBioChem 23, e202100453 (2022).
Sippel, D. et al. Production and isolation of vanadium nitrogenase from Azotobacter vinelandii by molybdenum depletion. J. Biol. Inorg. Chem. 22, 161–168 (2017).
Trncik, C., Müller, T., Franke, P. & Einsle, O. Structural analysis of the reductase component AnfH of iron-only nitrogenase from Azotobacter vinelandii. J. Inorg. Biochem. 227, 111690 (2022).
Dos Santos, P. C. Molecular biology and genetic engineering in nitrogen fixation. Methods Mol. Biol. 766, 81–92 (2011).
Chatterjee, R., Ludden, P. W. & Shah, V. K. Characterization of VnfG, the delta subunit of the vnf-encoded apodinitrogenase from Azotobacter vinelandii—implications for its role in the formation of functional dinitrogenase 2. J. Biol. Chem. 272, 3758–3765 (1997).
Danyal, K., Dean, D. R., Hoffman, B. M. & Seefeldt, L. C. Electron transfer within nitrogenase: evidence for a deficit-spending mechanism. Biochemistry 50, 9255–9263 (2011).
Duval, S. et al. Electron transfer precedes ATP hydrolysis during nitrogenase catalysis. Proc. Natl Acad. Sci. USA 110, 16414–16419 (2013).
Davydov, R. et al. Exploring electron/proton transfer and conformational changes in the nitrogenase MoFe protein and FeMo-cofactor through cryoreduction/EPR measurements. Isr. J. Chem. 56, 841–851 (2016).
Decamps, L., Rice, D. B. & DeBeer, S. An Fe6C Core in all nitrogenase cofactors. Angew. Chem. Int. Ed. 61, e202209190 (2022).
Lukoyanov, D. A. et al. 13C-ENDOR characterization of the central carbon within the nitrogenase catalytic cofactor indicates that the CFe6 core is a stabilizing ‘Heart of Steel’. J. Am. Chem. Soc. 144, 18315–18328 (2022).
Rohde, M., Sippel, D., Trncik, C., Andrade, S. L. A. & Einsle, O. The critical E4 state of nitrogenase catalysis. Biochemistry 57, 5497–5504 (2018).
Sippel, D. et al. A bound reaction intermediate sheds light on the mechanism of nitrogenase. Science 359, 1484–1489 (2018).
Harris, D. F. et al. Mo-, V- and Fe-nitrogenases use a universal eight-electron reductive-elimination mechanism to achieve N2 reduction. Biochemistry 58, 3293–3301 (2019).
Peters, J. W., Fisher, K., Newton, W. E. & Dean, D. R. Involvement of the P-cluster in intramolecular electron-transfer within the nitrogenase MoFe protein. J. Biol. Chem. 270, 27007–27013 (1995).
Danyal, K. et al. Uncoupling nitrogenase: catalytic reduction of hydrazine to ammonia by a MoFe protein in the absence of Fe protein-ATP. J. Am. Chem. Soc. 132, 13197–13199 (2010).
Zhang, L. M., Morrison, C. N., Kaiser, J. T. & Rees, D. C. Nitrogenase MoFe protein from Clostridium pasteurianum at 1.08 Å resolution: comparison with the Azotobacter vinelandii MoFe protein. Acta Crystallogr. D 71, 274–282 (2015).
Mayer, S. M., Lawson, D. M., Gormal, C. A., Roe, S. M. & Smith, B. E. New insights into structure-function relationships in nitrogenase: a 1.6 Å resolution X-ray crystallographic study of Klebsiella pneumoniae MoFe-protein. J. Mol. Biol. 292, 871–891 (1999).
Owens, C. P., Katz, F. E. H., Carter, C. H., Oswald, V. F. & Tezcan, F. A. Tyrosine-coordinated P-cluster in G. diazotrophicus nitrogenase: evidence for the importance of O-based ligands in conformationally gated electron transfer. J. Am. Chem. Soc. 138, 10124–10127 (2016).
Zheng, L. M., White, R. H. & Dean, D. R. Purification of the Azotobacter vinelandii nifV-encoded homocitrate synthase. J. Bacteriol. 179, 5963–5966 (1997).
Kennedy, C. & Dean, D. The NifU, NifS and NifV gene-products are required for activity of all 3 nitrogenases of Azotobacter vinelandii. Mol. Gen. Genet. 231, 494–498 (1992).
Hwang, J. C., Chen, C. H. & Burris, R. H. Inhibition of nitrogenase-catalyzed reductions. Biochim. Biophys. Acta 292, 256–270 (1973).
Spatzal, T., Perez, K. A., Einsle, O., Howard, J. B. & Rees, D. C. Ligand binding to the FeMo-cofactor: structures of CO-bound and reactivated nitrogenase. Science 345, 1620–1623 (2014).
Spatzal, T., Perez, K. A., Howard, J. B. & Rees, D. C. Catalysis-dependent selenium incorporation and migration in the nitrogenase active site iron-molybdenum cofactor. eLife 4, e11620 (2015).
Thorneley, R. N. F. & Lowe, D. J. in Molybdenum Enzymes 1 (ed. Spiro, T. G.) 221–284 (Wiley, 1985).
Lukoyanov, D. et al. Reductive elimination of H2 activates nitrogenase to reduce the N-N triple bond: characterization of the E4(4H) Janus intermediate in wild-type enzyme. J. Am. Chem. Soc. 138, 10674–10683 (2016).
Lukoyanov, D. et al. Unification of reaction pathway and kinetic scheme for N2 reduction catalyzed by nitrogenase. Proc. Natl Acad. Sci. USA 109, 5583–5587 (2012).
Lukoyanov, D. et al. Identification of a key catalytic intermediate demonstrates that nitrogenase is activated by the reversible exchange of N2 for H2. J. Am. Chem. Soc. 137, 3610–3615 (2015).
Rohde, M., Grunau, K. & Einsle, O. CO binding to the FeV cofactor of CO-reducing vanadium nitrogenase at atomic resolution. Angew. Chem. Int. Ed. 59, 23626–23630 (2020).
Rohde, M., Laun, K., Zebger, I., Stripp, S. T. & Einsle, O. Two ligand-binding sites in CO-reducing V nitrogenase reveal a general mechanistic principle. Sci. Adv. 7, eabg4474 (2021).
Buscagan, T. M., Perez, K. A., Maggiolo, A. O., Rees, D. C. & Spatzal, T. Structural characterization of two CO molecules bound to the nitrogenase active site. Angew. Chem. Int. Ed. 60, 5704–5707 (2021).
Kang, W., Lee, C. C., Jasniewski, A. J., Ribbe, M. W. & Hu, Y. Structural evidence for a dynamic metallocofactor during N2 reduction by Mo-nitrogenase. Science 368, 1381–1385 (2020).
Bergmann, J., Oksanen, E. & Ryde, U. Critical evaluation of a crystal structure of nitrogenase with bound N2 ligands. J. Biol. Inorg. Chem. 26, 341–353 (2021).
Peters, J. W. et al. Comment on ‘Structural evidence for a dynamic metallocofactor during N2 reduction by Mo-nitrogenase’. Science 371, eabe5481 (2021).
Lee, C. C. et al. Evidence of substrate binding and product release via belt-sulfur mobilization of the nitrogenase cofactor. Nat. Catal. 5, 443–454 (2022).
Eady, R. R. Structure-function relationships of alternative nitrogenases. Chem. Rev. 96, 3013–3030 (1996).
Garcia, A. K., Kolaczkowski, B. & Kaçar, B. Reconstruction of nitrogenase predecessors suggests origin from maturase-like proteins. Genome Biol. Evol. 14, evac031 (2022).
Perez-Gonzalez, A. et al. Specificity of NifEN and VnfEN for the assembly of nitrogenase active site cofactors in Azotobacter vinelandii. mBio 12, e0156821 (2021).
Liang, J. H., Madden, M., Shah, V. K. & Burris, R. H. Citrate substitutes for homocitrate in nitrogenase of a nifV mutant of Klebsiella pneumoniae. Biochemistry 29, 8577–8581 (1990).
Mayer, S. M., Gormal, C. A., Smith, B. E. & Lawson, D. M. Crystallographic analysis of the MoFe protein of nitrogenase from a nifV mutant of Klebsiella pneumoniae identifies citrate as a ligand to the molybdenum of iron molybdenum cofactor (FeMoco). J. Biol. Chem. 277, 35263–35266 (2002).
Cao, L. L., Caldararu, O. & Ryde, U. Protonation states of homocitrate and nearby residues in nitrogenase studied by computational methods and quantum refinement. J. Phys. Chem. B 121, 8242–8262 (2017).
Nicolet, Y. et al. Crystallographic and FTIR spectroscopic evidence of changes in Fe coordination upon reduction of the active site of the Fe-only hydrogenase from Desulfovibrio desulfuricans. J. Am. Chem. Soc. 123, 1596–1601 (2001).
Fontecilla-Camps, J. C., Volbeda, A., Cavazza, C. & Nicolet, Y. Structure/function relationships of [NiFe]- and [FeFe]-hydrogenases. Chem. Rev. 107, 4273–4303 (2007).
Van Stappen, C. et al. Spectroscopic description of the E1 state of Mo nitrogenase based on Mo and Fe X-ray absorption and Mössbauer studies. Inorg. Chem. 58, 12365–12376 (2019).
Van Stappen, C., Thorhallsson, A. T., Decamps, L., Björnsson, R. & DeBeer, S. Resolving the structure of the E1 state of Mo nitrogenase through Mo and Fe K-edge EXAFS and QM/MM calculations. Chem. Sci. 10, 9807–9821 (2019).
Lukoyanov, D. A. et al. Electron redistribution within the nitrogenase active site FeMo-cofactor during reductive elimination of H2 to achieve N≡N triple-bond activation. J. Am. Chem. Soc. 142, 21679–21690 (2020).
Yu, Y. et al. The reactivity patterns of low-coordinate iron-hydride complexes. J. Am. Chem. Soc. 130, 6624–6638 (2008).
Coric, I. & Holland, P. L. Insight into the iron molybdenum cofactor of nitrogenase from synthetic iron complexes with sulfur, carbon and hydride ligands. J. Am. Chem. Soc. 138, 7200–7211 (2016).
Rittle, J., McCrory, C. C. L. & Peters, J. C. A 106-fold enhancement in N2-binding affinity of an Fe2(µ-H)2 core upon reduction to a mixed-valence FeIIFeI state. J. Am. Chem. Soc. 136, 13853–13862 (2014).
Cao, L. L. & Ryde, U. Putative reaction mechanism of nitrogenase after dissociation of a sulfide ligand. J. Catal. 391, 247–259 (2020).
Thorhallsson, A. T. & Björnsson, R. The E2 state of FeMoco: hydride formation versus Fe reduction and a mechanism for H2 evolution. Chem. Eur. J. 27, 16788–16800 (2021).
Rees, J. A. et al. Comparative electronic structures of nitrogenase FeMoco and FeVco. Dalton Trans. 46, 2445–2455 (2017).
Spatzal, T. et al. Nitrogenase FeMoco investigated by spatially resolved anomalous dispersion refinement. Nat. Commun. 7, 10902 (2016).
Cummins, C. C. Three-coordinate complexes of ‘Hard’ ligands: advances in synthesis, structure and reactivity. Prog. Inorg. Chem. 47, 685–836 (1998).
Smith, J. M., Lachicotte, R. J. & Holland, P. L. N=N bond cleavage by a low-coordinate iron(II) hydride complex. J. Am. Chem. Soc. 125, 15752–15753 (2003).
Lipman, J. G. Experiments on the transformation and fixation of nitrogen by bacteria. Rep. N. Jersey Agric. Exp. Stat. 24, 217–285 (1903).
Subba Rao, N. S. Soil Microorganisms and Plant Growth 3rd edn (Science, 1995).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Vagin, A. & Teplyakov, A. Molecular replacement with MOLREP. Acta Crystallogr. D 66, 22–25 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Murshudov, G. N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D 67, 355–367 (2011).
Weiss, M. & Hilgenfeld, R. On the use of the merging R factor as a quality indicator for X-ray data. J. Appl. Crystallogr. 30, 203–205 (1997).
Karplus, P. A. & Diederichs, K. Linking crystallographic model and data quality. Science 336, 1030–1033 (2012).
Okada, T., Tomita, T., Wulandari, A. P., Kuzuyama, T. & Nishiyama, M. Mechanism of substrate recognition and insight into feedback inhibition of homocitrate synthase from Thermus thermophilus. J. Biol. Chem. 285, 4195–4205 (2010).
Acknowledgements
This work was supported by the European Research Council (grant no. 310656) and Deutsche Forschungsgemeinschaft (PP 1927, project ID 311061829, and RTG 1976, project ID 235777276). We thank P. dos Santos, M. Rohde, K. Parison, J. Gies-Elterlein, F. Schneider, S.L.A. Andrade and P. Franke for helpful discussions and the beamline staff at the Swiss Light Source, Villigen, Switzerland, for excellent assistance with data collection.
Author information
Authors and Affiliations
Contributions
C.T. and O.E. designed the experiments. C.T. and F.D. produced protein and generated crystals. C.T. and O.E. built and refined the crystal structure. C.T., F.D. and O.E. analysed data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Catalysis thanks Per Siegbahn, Tristan Wagner and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
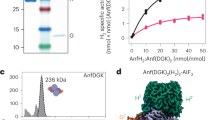
Extended Data Fig. 1 Isolation and characterization of Fe nitrogenase from Azotobacter vinelandii.
a) Analytical size exclusion chromatography of FeFe protein on Superdex S200 (Cytiva), using a triple detector array with UV absorbance (purple), right-angle light scattering (green) and refractive index (red). The derived absolute molecular mass is shown in black. b) SDS-PAGE of preparations of the three Fe proteins of A. vinelandii (left) and the three dinitrogenases (right). c) N2 reduction activities of the three A. vinelandii nitrogenase isoenzymes with their respective Fe proteins. Data from three technical replicates from the protein batch used for structure determination, presented as mean values + /– standard deviation.
Extended Data Fig. 2 Structural comparison of the dinitrogenase core subunits of A. vinelandii.
Stereo images of the D- and K-subunits of Fe-, V-, and Mo-nitrogenases (top to bottom) in identical orientation, colored from blue at the N-terminus to red on the C-terminus. The respective cofactors are shown in the D-subunits, P-clusters are shown in both subunits to emphasize their position in the interface. Note the extended C-terminus of AnfD and N-terminus of NifK that in part occupy the same position within the quaternary structure of the respective dinitrogenases (Fig. 1a, Extended Data Fig. 3).
Extended Data Fig. 3 Structural Differences and Similarities between Mo- and Fe-nitrogenase.
a) Schematic representation of Fe-nitrogenase (left) and Mo-nitrogenase (right) from A. vinelandii. Although the surfaces of both enzymes appear distinct, the extended C-terminus of AnfD (red) occupies a very similar position on the protein surface as the C-terminus of NifK (blue). b) Detail views of A) with surface representations for AnfD with its C-terminus (left) and NifK with its N-terminus.
Extended Data Fig. 4 The P-cluster in A. vinelandii Fe nitrogenase.
The [8Fe:7S] cluster in the structure is observed in the all-ferrous PN state. The labelled Fe6 that moves towards the conserved serine S143K upon oxidation to the P+1 state does not show the dual conformation that is frequently observed in crystal structures. The stereo image shows the 2Fo–Fc electron density map contoured at the 2σ level (grey) and the 6σ level (blue), as well as an anomalous difference electron density map collected at the Fe K-edge contoured at the 5σ level (orange).
Extended Data Fig. 5 Homocitrate synthase from A. vinelandii.
a) Structure of the NifV gene product as predicted by Alphafold2. Like other known enzymes of this type, NifV forms a homodimer with a TIM-barrel domain that coordinates a Zn2+ ion in its centre as the active site. The metal ion and a homocitrate ligand were modelled based on the structure of R-homocitrate synthase from Thermus thermophilus (PDB 2ZTK)84. Homocitrate is formed by the condensation of 2-oxoglutarate with acetyl-CoA. b) The modelled metal site of NifV. Zn2+ is coordinated by two histidine and one aspartate residue. During catalysis, 2-oxoglutarate binds as a bidentate ligand to the metal and is then reacted with the acetyl group of acetyl-CoA.
Extended Data Fig. 6 Kinetic Scheme for N2 reduction by nitrogenases according to Lowe and Thorneley.
Catalysing an eight-electron process, the enzyme cycles through eight states E0–E7, in which each electron transfer (blue arrows) is accompanied by a protonation event for charge compensation. In an initial charging phase (grey), the enzyme must be reduced from its resting state E0 to at least E3, more likely E4, to gain the ability to bind and activate N2. From E2 on, unproductive H2 release is observed, which is commonly interpreted as the (unwanted) protonation of a surface hydride on the cofactor, turning the enzyme back two states in the cycle. In contrast, the H2 released in E4 is the result of the reductive elimination of H2 from two hydrides, leaving the cofactor in a 2-electron-reduced state that is uniquely capable of N2 binding and activation. The remaining steps of substrate reduction (E5–E7) are then facile. Lowe and Thorneley did not observe H2 formation from these states, indicating that no further surface hydrides are formed, and reduction occurs directly on the bound intermediates.
Supplementary information
Source data
Source Data Extended Data Fig. 1
Unprocessed SDS–PAGE for Extended Data Fig. 1B.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Trncik, C., Detemple, F. & Einsle, O. Iron-only Fe-nitrogenase underscores common catalytic principles in biological nitrogen fixation. Nat Catal 6, 415–424 (2023). https://doi.org/10.1038/s41929-023-00952-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41929-023-00952-1
This article is cited by
-
Structural insights into the iron nitrogenase complex
Nature Structural & Molecular Biology (2024)
-
Unveiling the final nitrogenase
Nature Reviews Chemistry (2023)
-
The structural components of the Azotobacter vinelandii iron-only nitrogenase, AnfDKG, form a protein complex within the plant mitochondrial matrix
Plant Molecular Biology (2023)