Abstract
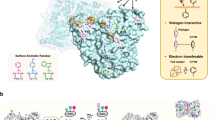
Ni–Fe carbon monoxide dehydrogenases (CODHs) are nearly diffusion-limited biocatalysts that oxidize CO. Their O2 sensitivity, however, is a major drawback for industrial applications. Here we compare the structures of a fast CODH with a high O2 sensitivity (ChCODH-II) and a slower CODH with a lower O2 sensitivity (ChCODH-IV) (Ch, Carboxydothermus hydrogenoformans). Some variants obtained by simple point mutations of the bottleneck residue (A559) in the gas tunnel showed 61–148-fold decreases in O2 sensitivity while maintaining high turnover rates. The variant structure A559W showed obstruction of one gas tunnel, and molecular dynamics supported the locked position of the mutated side chain in the tunnel. The variant was exposed to different gas mixtures, from simple synthetic gas to sophisticated real flue from a steel mill. Its catalytic properties remained unchanged, even at high O2 levels, and the efficiency was maintained for multiple cycles of CO detoxification/regeneration.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding authors upon reasonable request. MD trajectories of a total of 5 ns are available on Zenodo at https://doi.org/10.5281/zenodo.6865415. The atomic coordinates and structure factors for the ChCODH-II A559 variants have been deposited in the Protein Data Bank under accession codes PDB 7ERR, 7XDM, 7XDN and 7XDP.
References
Ragsdale, S. W. Life with carbon monoxide. Crit. Rev. Biochem. Mol. Biol. 39, 165–195 (2004).
Choi, E. S., Min, K., Kim, G. J., Kwon, I. & Kim, Y. H. Expression and characterization of Pantoea CO dehydrogenase to utilize CO-containing industrial waste gas for expanding the versatility of CO dehydrogenase. Sci. Rep. 7, 44323 (2017).
Hwang, H. W. et al. Two-stage bioconversion of carbon monoxide to biopolymers via formate as an intermediate. Chem. Eng. J. 389, 124394 (2020).
Kim, T. W. et al. A biological process effective for the conversion of CO-containing industrial waste gas to acetate. Bioresour. Technol. 211, 792–796 (2016).
Liew, F. et al. Gas fermentation—a flexible platform for commercial scale production of low-carbon-fuels and chemicals from waste and renewable feedstocks. Front. Microbiol. 7, 694 (2016).
Anderson, M. E., Derose, V. J., Hoffman, B. M. & Lindahl, P. A. Identification of a cyanide binding-site in CO dehydrogenase from Clostridium thermoaceticum using EPR and ENDOR spectroscopies. J. Am. Chem. Soc. 115, 12204–12205 (1993).
Shima, S. & Ataka, K. Isocyanides inhibit [Fe]-hydrogenase with very high affinity. FEBS Lett. 585, 353–356 (2011).
Ragsdale, S. W., Ljungdahl, L. G. & Dervartanian, D. V. Isolation of carbon monoxide dehydrogenase from Acetobacterium woodii and comparison of its properties with those of the Clostridium thermoaceticum enzyme. J. Bacteriol. 155, 1224–1237 (1983).
Domnik, L. et al. CODH-IV: a high-efficiency CO-scavenging CO dehydrogenase with resistance to O2. Angew. Chem. Int. Ed. 56, 15466–15469 (2017).
Jeoung, J. H., Martins, B. M. & Dobbek, H. X-ray crystallography of carbon monoxide dehydrogenases. Methods Mol. Biol. 1876, 167–178 (2019).
Rovaletti, A., Bruschi, M., Moro, G., Cosentino, U. & Greco, C. The challenging in silico description of carbon monoxide oxidation as catalyzed by molybdenum–copper CO dehydrogenase. Front. Chem. 6, 630 (2019).
Zhang, B., Hemann, C. F. & Hille, R. Kinetic and spectroscopic studies of the molybdenum–copper CO dehydrogenase from Oligotropha carboxidovorans. J. Biol. Chem. 285, 12571–12578 (2010).
Svetlitchnyi, V., Peschel, C., Acker, G. & Meyer, O. Two membrane-associated NiFeS–carbon monoxide dehydrogenases from the anaerobic carbon-monoxide-utilizing eubacterium Carboxydothermus hydrogenoformans. J. Bacteriol. 183, 5134–5144 (2001).
Merrouch, M. et al. O2 inhibition of Ni-containing CO dehydrogenase is partly reversible. Chemistry 21, 18934–18938 (2015).
Wu, M. et al. Life in hot carbon monoxide: the complete genome sequence of Carboxydothermus hydrogenoformans Z-2901. PLoS Genet. 1, 563–574 (2005).
Liebgott, P. P. et al. Relating diffusion along the substrate tunnel and oxygen sensitivity in hydrogenase. Nat. Chem. Biol. 6, 63–70 (2010).
Seravalli, J. & Ragsdale, S. W. 13C NMR characterization of an exchange reaction between CO and CO2 catalyzed by carbon monoxide dehydrogenase. Biochemistry 47, 6770–6781 (2008).
Zhu, G. H. & Spreitzer, R. J. Directed mutagenesis of chloroplast ribulose-1,5-bisphosphate carboxylase/oxygenase: loop 6 substitutions complement for structural stability but decrease catalytic efficiency. J. Biol. Chem. 271, 18494–18498 (1996).
Bonam, D., Murrell, S. A. & Ludden, P. W. Carbon monoxide dehydrogenase from Rhodospirillum rubrum. J. Bacteriol. 159, 693–699 (1984).
Drake, H. L. Occurrence of nickel in carbon monoxide dehydrogenase from Clostridium pasteurianum and Clostridium thermoaceticum. J. Bacteriol. 149, 561–566 (1982).
Chovancova, E. et al. CAVER 3.0: a tool for the analysis of transport pathways in dynamic protein structures. PLoS Comput. Biol. 8, e1002708 (2012).
DeMirci, H. et al. Structural adaptation of oxygen tolerance in 4-hydroxybutyrl-CoA dehydratase, a key enzyme of archaeal carbon fixation. Preprint at bioRxiv https://doi.org/10.1101/2020.02.05.935528 (2020).
Fritsch, J. et al. The crystal structure of an oxygen-tolerant hydrogenase uncovers a novel iron-sulphur centre. Nature 479, 249–252 (2011).
Montet, Y. et al. Gas access to the active site of Ni–Fe hydrogenases probed by X-ray crystallography and molecular dynamics. Nat. Struct. Biol. 4, 523–526 (1997).
Kalms, J. et al. Krypton derivatization of an O2-tolerant membrane-bound [NiFe] hydrogenase reveals a hydrophobic tunnel network for gas transport. Angew. Chem. Int. Ed. 55, 5586–5590 (2016).
Feng, J. & Lindahl, P. A. Effect of sodium sulfide on Ni-containing carbon monoxide dehydrogenases. J. Am. Chem. Soc. 126, 9094–9100 (2004).
Nakamura, M., Saeki, K. & Takahashi, Y. Hyperproduction of recombinant ferredoxins in Escherichia coli by coexpression of the ORF1-ORF2-iscS-iscU-iscA-hscB-hscA-fdx-ORF3 gene cluster. J. Biochem. 126, 10–18 (1999).
Miller, E., Wohlfarth, G. & Diekert, G. Comparative studies on tetrachloroethene reductive dechlorination mediated by Desulfitobacterium sp. strain PCE-S. Arch. Microbiol. 168, 513–519 (1997).
Munasinghe, P. C. & Khanal, S. K. Syngas fermentation to biofuel: evaluation of carbon monoxide mass transfer coefficient (kla) in different reactor configurations. Biotechnol. Progr. 26, 1616–1621 (2010).
Yung-Chi, C. & Prusoff, W. H. Relationship between the inhibition constant (KI) and the concentration of inhibitor which causes 50 percent inhibition (I50) of an enzymatic reaction. Biochem. Pharmacol. 22, 3099–3108 (1973).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Dobbek, H., Svetlitchnyi, V., Liss, J. & Meyer, O. Carbon monoxide induced decomposition of the active site [Ni–4Fe–5S] cluster of CO dehydrogenase. J. Am. Chem. Soc. 126, 5382–5387 (2004).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D 75, 861–877 (2019).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011).
Brooks, B. R. et al. CHARMM: a program for macromolecular energy, minimization, and dynamics calculations. J. Comput. Chem. 4, 187–217 (1983).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Darden, T., York, D. & Pedersen, L. Particle mesh Ewald: an Nlog(N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089–10092 (1993).
Ryckaert, J.-P., Ciccotti, G. & Berendsen, H. J. C. Numerical integration of the Cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes. J. Comput. Phys. 23, 327–341 (1977).
DeVane, R. et al. A molecular dynamics method for calculating molecular volume changes appropriate for biomolecular simulation. Biophys. J. 85, 2801–2807 (2003).
Zhang, Y. & Skolnick, J. TM-align: a protein structure alignment algorithm based on the TM-score. Nucleic Acids Res. 33, 2302–2309 (2005).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 35, 1547–1549 (2018).
Huerta-Cepas, J. et al. eggNOG 5.0: a hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Res. 47, D309–D314 (2019).
Acknowledgements
This research was supported by the C1 Gas Refinery Program and the Engineering Research Center Program through the National Research Foundation of Korea (NRF), funded by the Ministry of Science, ICT & Future Planning (2015M3D3A1A01064919 and 2020R1A5A1019631, respectively).
Author information
Authors and Affiliations
Contributions
Y.H.K. and S.M.K. conceived and planned all the experiments. S.M.K. performed the bioinformatic analysis and gene cloning. S.M.K. and J.L. performed the biochemical characterization, kinetic analysis and feasibility evaluation, all under the supervision of Y.H.K. S.M.K. and S.H.K. engineered the gas tunnels and performed their structural analysis. Y.H.K., S.M.K. and J.-S.H. wrote the manuscript. H.H.L., H.-J.Y. and Y.H. determined the crystal structure. Y.H.K., H.H.L. and J.-S.H. reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Catalysis thanks Tristan Wagner, Anna Rovaletti, Stephen Ragsdale and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Tables 1–9 and Figs. 1–17.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kim, S.M., Lee, J., Kang, S.H. et al. O2-tolerant CO dehydrogenase via tunnel redesign for the removal of CO from industrial flue gas. Nat Catal 5, 807–817 (2022). https://doi.org/10.1038/s41929-022-00834-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41929-022-00834-y
This article is cited by
-
Identifying a key spot for electron mediator-interaction to tailor CO dehydrogenase’s affinity
Nature Communications (2024)
-
Improved Functional Expression of Carbon Monoxide Dehydrogenase from Carboxydothermus hydrogenoformans Using Genetically Engineered Escherichia coli Under Aerobic Conditions
Korean Journal of Chemical Engineering (2024)