Abstract
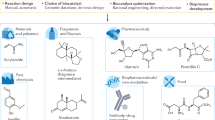
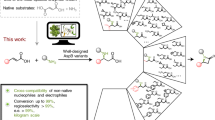
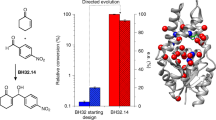
Enzymes are renowned for their catalytic efficiency and selectivity. Despite the wealth of carbon–carbon bond-forming transformations in traditional organic chemistry and nature, relatively few C–C bond-forming enzymes have found their way into the biocatalysis toolbox. Here we show that the enzyme UstD performs a highly selective decarboxylative aldol addition with diverse aldehyde substrates to make non-standard γ-hydroxy amino acids. We increased the activity of UstD through three rounds of classic directed evolution and an additional round of computationally guided engineering. The enzyme that emerged, UstDv2.0, is efficient in a whole-cell biocatalysis format. The products are highly desirable, functionally rich bioactive γ-hydroxy amino acids that we demonstrate can be prepared stereoselectively on the gram scale. The X-ray crystal structure of UstDv2.0 at 2.25 Å reveals the active site and provides a foundation for probing the UstD mechanism.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Code availability
The linear regression modelling code used during the final round of protein engineering is available through GitHub42 under the MIT License.
References
Nestl, B. M., Hammer, S. C., Nebel, B. A. & Hauer, B. New generation of biocatalysts for organic synthesis. Angew. Chem. Int. Ed. 53, 3070–3095 (2014).
Zhang, X. et al. Divergent synthesis of complex diterpenes through a hybrid oxidative approach. Science 369, 799–806 (2020).
Brown, D. G. & Boström, J. Analysis of past and present synthetic methodologies on medicinal chemistry: where have all the new reactions gone? J. Med. Chem. 59, 4443–4458 (2016).
Fesko, K. & Gruber-Khadjawi, M. Biocatalytic methods for C–C bond formation. ChemCatChem 5, 1248–1272 (2013).
Schmidt, N. G., Eger, E. & Kroutil, W. Building bridges: biocatalytic C–C-bond formation toward multifunctional products. ACS Catal. 6, 4286–4311 (2016).
Fujii, I. Heterologous expression systems for polyketide synthases. Nat. Prod. Rep. 26, 155–169 (2009).
Heine, A. et al. Observation of covalent intermediates in an enzyme mechanism at atomic resolution. Science 294, 369–374 (2001).
Wang, Z. J. et al. Improved cyclopropanation activity of histidine-ligated cytochrome P450 enables the enantioselective formal synthesis of levomilnacipran. Angew. Chem. Int. Ed. 53, 6810–6813 (2014).
Berkeš, D., Kolarovič, A., Manduch, R., Baran, P. & Považanec, F. Crystallization-induced asymmetric transformations (CIAT): stereoconvergent acid-catalyzed lactonization of substituted 2-amino-4-aryl-4-hydroxybutanoic acids. Tetrahedron Asymm. 16, 1927–1934 (2005).
Goldberg, S. L. et al. Preparation of β-hydroxy-α-amino acid using recombinant d-threonine aldolase. Org. Process Res. Dev. 19, 1308–1316 (2015).
Steinreiber, J. et al. Overcoming thermodynamic and kinetic limitations of aldolase-catalyzed reactions by applying multienzymatic dynamic kinetic asymmetric transformations. Angew. Chem. Int. Ed. 46, 1624–1626 (2007).
Zetzsche, L. E. & Narayan, A. R. H. Broadening the scope of biocatalytic C–C bond formation. Nat. Rev. Chem. 4, 334–346 (2020).
Ye, Y. et al. Unveiling the biosynthetic pathway of the ribosomally synthesized and post-translationally modified peptide ustiloxin B in filamentous fungi. Angew. Chem. Int. Ed. 55, 8072–8075 (2016).
Prier, C. K. & Arnold, F. H. Chemomimetic biocatalysis: exploiting the synthetic potential of cofactor-dependent enzymes to create new catalysts. J. Am. Chem. Soc. 137, 13992–14006 (2015).
Di Salvo, M. L. et al. On the catalytic mechanism and stereospecificity of Escherichia coli l-threonine aldolase. FEBS J. 281, 129–145 (2014).
Marsden, S. R., Gjonaj, L., Eustace, S. J. & Hanefeld, U. Separating thermodynamics from kinetics—a new understanding of the transketolase reaction. ChemCatChem 9, 1808–1814 (2017).
Ariza, J., Font, J. & Ortuño, R. M. An efficient and concise entry to (–)-4,5-dihydroxy-d-threo-l-norvaline. Formal synthesis of clavalanine. Tetrahedron Lett. 32, 1979–1982 (1991).
Blaskovich, M. A. T. Unusual amino acids in medicinal chemistry. J. Med. Chem. 59, 10807–10836 (2016).
Moreno, C. J. et al. Synthesis of γ-hydroxy-α-amino acid derivatives by enzymatic tandem aldol addition–transamination reactions. ACS Catal. 11, 4660–4669 (2021).
Hernandez, K. et al. Combining aldolases and transaminases for the synthesis of 2-amino-4-hydroxybutanoic acid. ACS Catal. 7, 1707–1711 (2017).
Vargas-Rodriguez, O., Sevostyanova, A., Söll, D. & Crnković, A. Upgrading aminoacyl-tRNA synthetases for genetic code expansion. Curr. Opin. Chem. Biol. 46, 115–122 (2018).
Marchand, J. A. et al. Discovery of a pathway for terminal-alkyne amino acid biosynthesis. Nature 567, 420–424 (2019).
Yang, J. et al. The I-TASSER suite: protein structure and function prediction. Nat. Methods 12, 7–8 (2014).
Ho, T. H. et al. Catalytic intermediate crystal structures of cysteine desulfurase from the Archaeon Thermococcus onnurineus NA1. Archaea 2017, 1–11 (2017).
Kumar, P. et al. l-Threonine transaldolase activity is enabled by a persistent catalytic intermediate. ACS Chem. Biol. 16, 95 (2021).
Reetz, M. T., Prasad, S., Carballeira, J. D., Gumulya, Y. & Bocola, M. Iterative saturation mutagenesis accelerates laboratory evolution of enzyme stereoselectivity: rigorous comparison with traditional methods. J. Am. Chem. Soc. 132, 9144–9152 (2010).
Romero, P. A. & Arnold, F. H. Exploring protein fitness landscapes by directed evolution. Nat. Rev. Mol. Cell Biol. 10, 866–876 (2009).
Reetz, M. T., Bocola, M., Carballeira, J. D., Zha, D. & Vogel, A. Expanding the range of substrate acceptance of enzymes: combinatorial active-site saturation test. Angew. Chem. Int. Ed. 44, 4192–4196 (2005).
Romney, D. K., Sarai, N. S. & Arnold, F. H. Nitroalkanes as versatile nucleophiles for enzymatic synthesis of noncanonical amino acids. ACS Catal. 9, 8726–8730 (2019).
Marfey, P. Determination of d-amino acids. II. Use of a bifunctional reagent, 1,5-difluoro-2,4-dinitrobenzene. Carlsberg Res. Commun. 49, 591–596 (1984).
Wu, G. et al. Proline and hydroxyproline metabolism: implications for animal and human nutrition. Amino Acids 40, 1053–1063 (2011).
Müller, J.-C., Toome, V., Pruess, D. L., Blount, J. F. & Weigele, M. Ro 22-5417, a new clavam antibiotic from Streptomyces clavuligerus. III Absolute stereochemistry. J. Antibiot. 36, 217–225 (1983).
Wahab, R. A., Elias, N., Abdullah, F. & Ghoshal, S. K. On the taught new tricks of enzymes immobilization: an all-inclusive overview. React. Func. Pol. 152, 104613 (2020).
Wachtmeister, J. & Rother, D. Recent advances in whole cell biocatalysis techniques bridging from investigative to industrial scale. Curr. Opin. Biotechnol. 42, 169–177 (2016).
Al-Ayyoubi, M., Gettins, P. G. W. & Volz, K. Crystal structure of human maspin, a serpin with antitumor properties: reactive center loop of maspin is exposed but constrained. J. Biol. Chem. 279, 55540–55544 (2004).
Fox, R. Directed molecular evolution by machine learning and the influence of nonlinear interactions. J. Theor. Biol. 234, 187–199 (2005).
Huffman, M. A. et al. Design of an in vitro biocatalytic cascade for the manufacture of islatravir. Science 366, 1255–1259 (2019).
Eliot, A. C. & Kirsch, J. F. Pyridoxal phosphate enzymes: mechanistic, structural, and evolutionary considerations. Annu. Rev. Biochem. 73, 383–415 (2004).
Romney, D. K., Murciano-Calles, J., Wehrmüller, J. E. & Arnold, F. H. Unlocking reactivity of TrpB: a general biocatalytic platform for synthesis of tryptophan analogues. J. Am. Chem. Soc. 139, 10769–10776 (2017).
Boville, C. E. et al. Engineered biosynthesis of β-alkyl tryptophan analogues. Angew. Chem. Int. Ed. 57, 14764–14768 (2018).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Ellis, J. M. Linear regression analysis of UstD-TLM. Zenodo https://doi.org/10.5281/zenodo.5719389 (2021).
Acknowledgements
We thank I. Guzei for small-molecule X-ray structure determinations and S.H. Gellman and members of the Buller group for critical reading of the manuscript. The crystal mounting and data collection were mediated by the Collaborative Crystallography Core, Department of Biochemistry, UW–Madison, and data were collected at the Life Sciences Collaborative Access Team beamline 21ID-D at the Advanced Photon Source, Argonne National Laboratory, and we thank Z. Wawrzak for technical assistance during data collection. Use of LS-CAT Sector 21 was supported by the Michigan Economic Development Corporation and the Michigan Technology Tri-Corridor (grant 085P1000817). This work was supported by the Office of the Vice Chancellor for Research and Graduate Education at the University of Wisconsin-Madison, Wisconsin Alumni Research Foundation, National Institute of Health (grant DP2-GM137417, A.R.B.), Morgridge Institute for Research—Metabolism Theme Fellowship (P.K.) and the NIH Biotechnology Training Grant (T32-GM008349, J.M.E.). The Bruker AVANCE III-500 NMR spectrometers were supported by the Bender Fund. The Advanced Photon Source was supported by the US Department of Energy, Office of Science, Office of Basic Energy Sciences, under contract no. W-31-109-Eng-38. The Bruker D8 VENTURE Photon III X-ray diffractometer was partially funded by a NSF Award (no. CHE-1919350) to the UW–Madison Department of Chemistry.
Author information
Authors and Affiliations
Contributions
A.R.B. and J.M.E. conceptualized the goals and aims of the project. J.M.E., M.E.C., P.K., E.P.G., C.A.B. and A.R.B. carried out the development of the chemistry and enzymes. J.M.E. developed the code for data analysis and developed the linear regression model. J.M.E. and M.E.C. verified the results. J.M.E., M.E.C., P.K. and A.R.B. prepared the figures and data visualizations. A.R.B. secured funding for the project that led to this publication. A.R.B. coordinated team members for the development of the chemistry and enzyme evolution. C.A.B. supervised the data acquisition of protein crystals that led to the resolved crystal structure. A.R.B. supervised the research activity planning and execution. J.M.E., M.E.C. and A.R.B. prepared the initial manuscript. J.M.E., M.E.C., P.K. and A.R.B. reviewed and edited the initial manuscript and provided critical commentary and revisions.
Corresponding author
Ethics declarations
Competing interests
A.R.B., J.M.E. and P.K. have a patent pending on the use of engineered UstD for the synthesis of nsAAs, US Patent application no. 20210115480A1. All other authors declare no competing interests.
Peer review
Peer review information
Nature Catalysis thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–28, Tables 1–4 and Methods.
Rights and permissions
About this article
Cite this article
Ellis, J.M., Campbell, M.E., Kumar, P. et al. Biocatalytic synthesis of non-standard amino acids by a decarboxylative aldol reaction. Nat Catal 5, 136–143 (2022). https://doi.org/10.1038/s41929-022-00743-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41929-022-00743-0
This article is cited by
-
A pyridoxal 5′-phosphate-dependent Mannich cyclase
Nature Catalysis (2023)
-
Recent advances in the biosynthesis of ribosomally synthesized and posttranslationally modified peptides of fungal origin
The Journal of Antibiotics (2023)
-
Substrate multiplexed protein engineering facilitates promiscuous biocatalytic synthesis
Nature Communications (2022)