Abstract
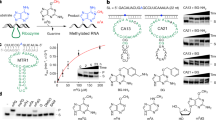
RNA methylations contribute to a wide range of cellular functions. Cellular RNAs are usually methylated by protein methyltransferases using S-adenosyl-l-methionine (SAM) as a cofactor. Here we report the in vitro selection of a 33-nucleotide SAM-dependent methyltransferase ribozyme RNA from a randomized sequence. Detection and mapping of the methyl group on the RNA demonstrates site-specific methylation of the N7 position of guanine by SAM. The ribozyme is active over a wide range of pH and temperatures and is robust compared to protein enzymes. The ribozyme structures in the presence and absence of SAM show a dramatic local conformational change associated with cofactor binding. The ribozyme motif was found to be distributed in nature, and candidate sequences were shown to be active in vitro. The discovery of this ribozyme that uses the cofactor SAM to specifically methylate RNA opens up the possibility that methyltransferase ribozymes may contribute to cellular RNA methylation.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data supporting the findings of this study are available within the paper (and its supplementary information files and source data files). Structural factors and coordinates of the SMRZ-1 RNA–U1A protein complex have been deposited in the Protein Data Bank under accession codes 7DLZ and 7DWH for the SMRZ-1 RNA–U1A complex in the absence and presence of SAM, respectively. Source data are provided with this paper.
Code availability
The script for searching the Refseq genomic database is available at: https://github.com/threadtag/SMRZ-1.
References
Roundtree, I. A., Evans, M. E., Pan, T. & He, C. Dynamic RNA modifications in gene expression regulation. Cell 169, 1187–1200 (2017).
Benne, R. & Grosjean, H. Modification and Editing of RNA (American Society for Microbiology, 1998).
Helm, M. & Alfonzo, J. D. Posttranscriptional RNA modifications: playing metabolic games in a cell’s chemical Legoland. Chem. Biol. 21, 174–185 (2014).
Zhang, L.-S. et al. Transcriptome-wide mapping of internal N7-methylguanosine methylome in mammalian mRNA. Mol. Cell 74, 1304–1316.e8 (2019).
Pandolfini, L. et al. METTL1 promotes let-7 microRNA processing via m7G methylation. Mol. Cell 74, 1278–1290.e9 (2019).
Cantoni, G. L. Biological methylation: selected aspects. Annu. Rev. Biochem. 44, 435–451 (1975).
Loenen, Wa. M. S-Adenosylmethionine: jack of all trades and master of everything? Biochem. Soc. Trans. 34, 330–333 (2006).
Schubert, H. L., Blumenthal, R. M. & Cheng, X. Many paths to methyltransfer: a chronicle of convergence. Trends Biochem. Sci. 28, 329–335 (2003).
Traube, F. R. & Carell, T. The chemistries and consequences of DNA and RNA methylation and demethylation. RNA Biol. 14, 1099–1107 (2017).
Rydberg, B. & Lindahl, T. Nonenzymatic methylation of DNA by the intracellular methyl group donor S-adenosyl-l-methionine is a potentially mutagenic reaction. EMBO J. 1, 211–216 (1982).
Benner, S. A., Ellington, A. D. & Tauer, A. Modern metabolism as a palimpsest of the RNA world. Proc. Natl Acad. Sci. USA 86, 7054–7058 (1989).
Harms, J. et al. High resolution structure of the large ribosomal subunit from a mesophilic eubacterium. Cell 107, 679–688 (2001).
Korostelev, A., Trakhanov, S., Laurberg, M. & Noller, H. F. Crystal structure of a 70S ribosome–tRNA complex reveals functional interactions and rearrangements. Cell 126, 1065–1077 (2006).
Nissen, P., Hansen, J., Ban, N., Moore, P. B. & Steitz, T. A. The structural basis of ribosome activity in peptide bond synthesis. Science 289, 920–930 (2000).
Schuwirth, B. S. et al. Structures of the bacterial ribosome at 3.5 Å resolution. Science 310, 827–834 (2005).
Selmer, M. et al. Structure of the 70S ribosome complexed with mRNA and tRNA. Science 313, 1935–1942 (2006).
Rodnina, M. V. The ribosome as a versatile catalyst: reactions at the peptidyl transferase center. Curr. Opin. Struct. Biol. 23, 595–602 (2013).
Belousoff, M. J. et al. Ancient machinery embedded in the contemporary ribosome. Biochem. Soc. Trans. 38, 422–427 (2010).
Guerrier-Takada, C., Gardiner, K., Marsh, T., Pace, N. & Altman, S. The RNA moiety of ribonuclease P is the catalytic subunit of the enzyme. Cell 35, 849–857 (1983).
Cech, T. R., Zaug, A. J. & Grabowski, P. J. In vitro splicing of the ribosomal RNA precursor of Tetrahymena: involvement of a guanosine nucleotide in the excision of the intervening sequence. Cell 27, 487–496 (1981).
Peebles, C. L. et al. A self-splicing RNA excises an intron lariat. Cell 44, 213–223 (1986).
van der Veen, R. et al. Excised group II introns in yeast mitochondria are lariats and can be formed by self-splicing in vitro. Cell 44, 225–234 (1986).
Prody, G. A., Bakos, J. T., Buzayan, J. M., Schneider, I. R. & Bruening, G. Autolytic processing of dimeric plant virus satellite RNA. Science 231, 1577–1580 (1986).
Buzayan, J. M., Gerlach, W. L. & Bruening, G. Satellite tobacco ringspot virus RNA: a subset of the RNA sequence is sufficient for autolytic processing. Proc. Natl Acad. Sci. USA 83, 8859–8862 (1986).
Winkler, W. C., Nahvi, A., Roth, A., Collins, J. A. & Breaker, R. R. Control of gene expression by a natural metabolite-responsive ribozyme. Nature 428, 281–286 (2004).
Tarasow, T. M., Tarasow, S. L. & Eaton, B. E. RNA-catalysed carbon–carbon bond formation. Nature 389, 54–57 (1997).
Kraut, S. et al. Three critical hydrogen bonds determine the catalytic activity of the Diels–Alderase ribozyme. Nucleic Acids Res. 40, 1318–1330 (2012).
Seelig, B. & Jäschke, A. A small catalytic RNA motif with Diels–Alderase activity. Chem. Biol. 6, 167–176 (1999).
Wilson, C. & Szostak, J. W. In vitro evolution of a self-alkylating ribozyme. Nature 374, 777–782 (1995).
Lohse, P. A. & Szostak, J. W. Ribozyme-catalysed amino-acid transfer reactions. Nature 381, 442–444 (1996).
Beaudry, A. A. & Joyce, G. F. Directed evolution of an RNA enzyme. Science 257, 635–641 (1992).
Tjhung, K. F., Shokhirev, M. N., Horning, D. P. & Joyce, G. F. An RNA polymerase ribozyme that synthesizes its own ancestor. Proc. Natl Acad. Sci. USA 117, 2906–2913 (2020).
Attwater, J., Wochner, A. & Holliger, P. In-ice evolution of RNA polymerase ribozyme activity. Nat. Chem. 5, 1011–1018 (2013).
Attwater, J., Raguram, A., Morgunov, A. S., Gianni, E. & Holliger, P. Ribozyme-catalysed RNA synthesis using triplet building blocks. eLife 7, e35255 (2018).
Scheitl, C. P. M., Ghaem Maghami, M., Lenz, A.-K. & Höbartner, C. Site-specific RNA methylation by a methyltransferase ribozyme. Nature 587, 663–667 (2020).
Dominissini, D., Moshitch-Moshkovitz, S., Salmon-Divon, M., Amariglio, N. & Rechavi, G. Transcriptome-wide mapping of N6-methyladenosine by m6A-seq based on immunocapturing and massively parallel sequencing. Nat. Protoc. 8, 176–189 (2013).
Burke, D. H. & Gold, L. RNA aptamers to the adenosine moiety of S-adenosyl methionine: structural inferences from variations on a theme and the reproducibility of SELEX. Nucleic Acids Res. 25, 2020–2024 (1997).
Su, D. et al. Quantitative analysis of ribonucleoside modifications in tRNA by HPLC-coupled mass spectrometry. Nat. Protoc. 9, 828–841 (2014).
Peattie, D. A. Direct chemical method for sequencing RNA. Proc. Natl Acad. Sci. USA 76, 1760–1764 (1979).
Kirpekar, F., Douthwaite, S. & Roepstorff, P. Mapping posttranscriptional modifications in 5S ribosomal RNA by MALDI mass spectrometry. RNA 6, 296–306 (2000).
De Bie, L. G. S. et al. The yggH gene of Escherichia coli encodes a tRNA (m7G46) methyltransferase. J. Bacteriol. 185, 3238–3243 (2003).
Barbés, C., Sánchez, J., Yebra, M. J., Robert-Geró, M. & Hardisson, C. Effects of sinefungin and S-adenosylhomocysteine on DNA and protein methyltransferases from Streptomyces and other bacteria. FEMS Microbiol. Lett. 57, 239–243 (1990).
Schluckebier, G., Zhong, P., Stewart, K. D., Kavanaugh, T. J. & Abad-Zapatero, C. The 2.2 Å structure of the rRNA methyltransferase ErmC′ and its complexes with cofactor and cofactor analogs: implications for the reaction mechanism. J. Mol. Biol. 289, 277–291 (1999).
Ferré-D’Amaré, A. R. & Doudna, J. A. Crystallization and structure determination of a hepatitis delta virus ribozyme: use of the RNA-binding protein U1A as a crystallization module. J. Mol. Biol. 295, 541–556 (2000).
Price, I. R., Grigg, J. C. & Ke, A. Common themes and differences in SAM recognition among SAM riboswitches. Biochim. Biophys. Acta 1839, 931–938 (2014).
Gruber, A. R., Lorenz, R., Bernhart, S. H., Neuböck, R. & Hofacker, I. L. The Vienna RNA websuite. Nucleic Acids Res. 36, W70–W74 (2008).
Wolk, S. K. et al. Modified nucleotides may have enhanced early RNA catalysis. Proc. Natl Acad. Sci. USA 117, 8236–8242 (2020).
White, H. B. Coenzymes as fossils of an earlier metabolic state. J. Mol. Evol. 7, 101–104 (1976).
Laurino, P. & Tawfik, D. S. Spontaneous emergence of S-adenosylmethionine and the evolution of methylation. Angew. Chem. Int. Ed. 56, 343–345 (2017).
Minor, W., Cymborowski, M., Otwinowski, Z. & Chruszcz, M. HKL-3000: the integration of data reduction and structure solution—from diffraction images to an initial model in minutes. Acta Crystallogr. D Biol. Crystallogr. 62, 859–866 (2006).
McCoy, A. J. Solving structures of protein complexes by molecular replacement with Phaser. Acta Crystallogr. D Biol. Crystallogr. 63, 32–41 (2007).
Potterton, E., Briggs, P., Turkenburg, M. & Dodson, E. A graphical user interface to the CCP4 program suite. Acta Crystallogr. D Biol. Crystallogr. 59, 1131–1137 (2003).
Murshudov, G. N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D Biol. Crystallogr. 67, 355–367 (2011).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D Biol. Crystallogr. 68, 352–367 (2012).
Soto, A. M., Gmeiner, W. H. & Marky, L. A. Energetic and conformational contributions to the stability of Okazaki fragments. Biochemistry 41, 6842–6849 (2002).
Acknowledgements
We thank members of the Murchie and Gan laboratories for discussion, C. Zhang (IBS, Fudan University) for assistance with LC–MS/MS analysis, X. Zhou (IBS, Fudan University) for assistance with MALDI–TOF mass spectrometry, W. Zhang (Shanghai Jiao Tong University) for assistance with differential scanning calorimetry analysis and the BL19U1 beamline of the National Facility for Protein Science in Shanghai for X-ray diffraction facilities. We also thank the late K. Nagai (MRC Cambridge) for the gift of the U1A expression plasmid. This work was supported by National Key R&D Program of China grant 2016YFA0500604 to A.M., National Natural Science Foundation grants 31420103907, 31770873 and 31330022 to A.M., National Natural Science Foundation grant 31370107 to D.C, Fudan University Original Research Grant 31470777 to H.J. and National Natural Science Foundation of China grant 31870721 to J.G.
Author information
Authors and Affiliations
Contributions
A.I.H.M., D.C. and H.J. were responsible for the initial experimental design. H.J. performed the SELEX and methylation analysis. H.J. and L.Z. performed the mass spectrometry. H.J., Y.G. and J.G. undertook the crystallization and crystallography. Y.G. and J.G. were responsible for structure solution and modelling. H.J. performed the bioinformatic analysis. A.I.H.M., D.C. and J.G. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Catalysis thanks Wen Zhang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 In vitro selection strategy.
a, Schematic of in vitro selection procedure for the methyltransferase ribozyme. b, Level of radioactivity incorporation for RNAs from round 3 and round 6 after reaction with [methyl-3H] SAM. Error bars are the standard deviation from the mean of 3 independent experiments, data points are shown (as dots).
Extended Data Fig. 2 Identification of a minimal methyltransferase sequence.
a, Schematic for the identification of a minimal sequence, truncated RNA from the 59nt RNA. b, Level of radioactivity incorporation for truncated RNAs after reaction with [methyl-3H] SAM, RNAs were incubated at 45 °C for 2 hours and assayed by centrifugal dialysis. Error bars are the standard deviation from the mean of 3 independent experiments, data points are shown.
Extended Data Fig. 3 Differential Scanning Calorimetry of SMRZ-1.
a, Differential scanning calorimetry curve for SMRZ-1 in 20 mM Tris–HCl, 20 mM KCl, 1 mM MgCl2, 0.1 mM CuSO4 buffer, pH 7.4 at total strand concentrations with 88.7 μM SMRZ-1 RNA. The calculated Tm and ΔH are indicated in the box. b, The structures of the methyl donor SAM, SAH, and the protein methyltransferase inhibitor Sinefungin SFG.
Extended Data Fig. 4 The crystal structure of SMRZ-1 RNA/U1A protein complex.
a, Primary sequence of the SMRZ-1-U1Abd RNA. The U1A linker region of the RNA is coloured in cyan. b, Cartoon presentation showing the U1A proteins and the SMRZ-1-U1Abd RNA molecules packed in the asymmetric unit of the crystal. U1A proteins are coloured in green, cyan, magenta, and yellow. RNA molecules are coloured in wheat and grey. c-d, Structural superposition of the two SMRZ-1-U1Abd RNA molecules.
Extended Data Fig. 5 The activity of the ribozyme with the U1A binding domain embedded.
Embedding the U1A binding domain did not affect ribozyme activity, nor did the inclusion of U1A protein in the reaction. Error bars are the standard deviation from the mean of 3 independent experiments, data points are shown. Error bars are the standard deviations of 3 independent experiments (shown as contrasting dots).
Extended Data Fig. 6 Structure of SMRZ-1-U1A RNA.
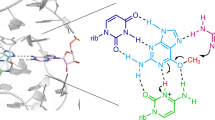
a, The 2Fo-Fc electron density maps of the SMRZ-1-U1Abd RNA. The maps are contoured at 1.2 level. The 5'- and 3'-ends of SMRZ-1 are shown as sticks in magenta and yellow, respectively. The U1A linker is coloured in green. b, The detailed sequence and secondary structure of SMRZ-1-U1Abd RNA. c, The detailed conformations of the nucleotides near the active site of SMRZ-1. Inset, the U6:G40 wobble base pair. For clarity, the stacking interactions between the nucleobases of U9, A35, A10, and C34 are omitted.
Extended Data Fig. 7 Mutational analysis of the methyltransferase ribozyme.
a, Secondary structure of the minimal SMRZ-1 (WT) sequence. The stacked planar pseudo triples are indicated by the grey shading. b, Normalised activity (relative to SMRZ-1) for point mutated RNAs after reaction with [methyl-3H] SAM. c, Further mutational analysis of SMRZ-1 to the A7 and the pseudo-triple nucleotides A10 and A24 (equivalent to the A35 position in the crystal structure RNA), normalised to SMRZ-1. For panels b and c error bars are the standard deviation from the mean of 3 independent experiments, data points are shown as dots.
Extended Data Fig. 8 Structure of SAM-complexed SMRZ-1-U1A RNA.
a, Overall structure of SMRZ-1-U1A RNA/U1A in the presence of SAM and Cu2+. The 5'- and 3'-ends of SMRZ-1 are coloured in magenta and yellow, respectively. The U1A linker is coloured in green. b-c, The 2Fo-Fc electron density maps of the SMRZ-1-U1A RNA and SAM and Cu2+, respectively. The maps are contoured at 1.0 level. d, Structural superposition of the SMRZ-1-U1A RNA molecules, which is coloured in white in the absence of SAM. Upon binding of SAM, the RNA is coloured as in a. Cu2+ is shown as black sphere in c. The SAM molecules are shown as red spheres in a and d, but as sticks in atomic colours (C, green; N, blue; O, red; S, orange) in c.
Extended Data Fig. 9 Distribution of candidate methyltransferase ribozyme sequences.
a, Table of natural SMRZ-1 ribozyme candidate sequences 16-40. The sequences are listed by classification and accession numbers. The alignment and activity relative to SMRZ-1 of each sequence and their relative position in the secondary structure in the RNA is shown. b, Methylation activity of sequences 16-40. The histograms show (Mauve, left axis) uncorrected incorporation of radioactivity (cpm) for each sample compared to the background measurement (Bkg) and the internal SMRZ-1 control for each experiment (separated by vertical dashed lines). The axis break from 300-1000 cpm shows the relatively low background signal (mean of 187 cpm ± 34 (n=15)), error bars are the standard deviations from the mean of 3 independent experiments (shown as contrasting dots). The right-hand (light blue) axis is the normalised (%) activity for each sequence relative to the SMRZ-1 control, for comparison between experiments, error bars are the standard deviations from the mean of 3 independent experiments (shown as contrasting dots).
Extended Data Fig. 10 The distribution of potential ribozyme sequences.
a, The numbers of candidate sequences classified by Kingdom, the number of unique sequences and the number of species harbouring candidate sequences. b, The numbers of candidate sequences identified in Bacteria and Archaea. Potential ribozyme sequences are classified by relative genome position, as sense or antisense strand, coding sequence (CDS), location within CDS (5’ or 3’), intergenic (between CDS) and sequences that have not been annotated. c, The numbers of candidate sequences present in Eukaryotes. Sequences are classified by relative genome position, as sense or antisense strand, exon or intron location, mix (exon and intron) or by annotation.
Supplementary Information
Supplementary Information
Supplementary Fig. 1 and Tables 1, 3 and 4.
Supplementary Table
Supplementary Tables 2, 5 and 6. Microsoft Excel file annotating sequences tabulated in Fig. 4c and Extended Data Fig. 9A.
Supplementary Data
Source Data Supplementary Figure 1
Source data
Source Data Fig. 1
Uncropped sequence gel.
Source Data Fig. 2
Source data Fig. 2
Source Data Fig. 4
Source data Fig. 4
Source Data Extended Data Fig. 1
Source data Extended data Fig 1
Source Data Extended Data Fig. 2
Source data Extended data Fig 2
Source Data Extended Data Fig. 5
Source data Extended data Fig 5
Source Data Extended Data Fig. 7
Source data Extended data Fig7
Source Data Extended Data Fig. 9
Source data Extended data Fig 9
Rights and permissions
About this article
Cite this article
Jiang, H., Gao, Y., Zhang, L. et al. The identification and characterization of a selected SAM-dependent methyltransferase ribozyme that is present in natural sequences. Nat Catal 4, 872–881 (2021). https://doi.org/10.1038/s41929-021-00685-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41929-021-00685-z
This article is cited by
-
Ribozyme for stabilized SAM analogue modifies RNA in cells
Nature Chemistry (2023)
-
A SAM analogue-utilizing ribozyme for site-specific RNA alkylation in living cells
Nature Chemistry (2023)
-
Blocker-dUThiophene poly tailing-based method for assessing methyl transferase activity
Analytical and Bioanalytical Chemistry (2023)
-
Structure-based insights into recognition and regulation of SAM-sensing riboswitches
Science China Life Sciences (2023)
-
Structure and mechanism of a methyltransferase ribozyme
Nature Chemical Biology (2022)