Abstract
Peroxygenases are attractive biocatalysts for the selective introduction of oxygen into organic molecules under mild conditions with hydrogen peroxide as the oxygen source. In addition to the identification of primary peroxygenases, different classes of enzymes were shown to display promiscuous peroxygenase activity. Even though enzymes with peroxygenase activity are promising industrial biocatalysts, further optimization of their properties is required for their effective use in industrial applications. Here we give a comprehensive overview of enzymes with peroxygenase activity and review diverse strategies, including directed evolution, rational approaches and the assistance of small functional molecules to improve the expression, catalytic activity, substrate scope or selectivity of these promising enzymes. Furthermore, we discuss the exploration of modified or unnatural cofactors to design artificial peroxygenases for desired reactions. The rapidly expanding field of hydrogen peroxide-utilizing enzymes bears a great potential to provide biocatalysts for selective oxyfunctionalization chemistry, contributing to the development of environmentally friendly and sustainable oxidation processes.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
White, M. C. Adding aliphatic C–H bond oxidations to synthesis. Science 335, 807–809 (2012).
Piera, J. & Bäckvall, J. E. Catalytic oxidation of organic substrates by molecular oxygen and hydrogen peroxide by multistep electron transfer — a biomimetic approach. Angew. Chem. Int. Ed. 47, 3506–3523 (2008).
Muzart, J. Chromium-catalyzed oxidations in organic synthesis. Chem. Rev. 92, 113–140 (1992).
Enthaler, S. & Company, A. Palladium-catalysed hydroxylation and alkoxylation. Chem. Soc. Rev. 40, 4912–4924 (2011).
Thiery, E., Chevrin, C., Le Bras, J., Harakat, D. & Muzart, J. Mechanistic insights into the palladiumII-catalyzed hydroxyalkoxylation of 2-allylphenols. J. Org. Chem. 72, 1859–1862 (2007).
Huybrechts, D. R. C., De Bruycker, L. & Jacobs, P. A. Oxyfunctionalization of alkanes with hydrogen peroxide on titanum silicalite. Nature 345, 240–242 (1990).
Hage, R. & Lienke, A. Applications of transition-metal catalysts to textile and wood-pulp bleaching. Angew. Chem. Int. Ed. 45, 206–222 (2006).
Hofrichter, M. & Ullrich, R. Oxidations catalyzed by fungal peroxygenases. Curr. Opin. Chem. Biol. 19, 116–125 (2014).
Ni, Y. et al. Peroxygenase-catalyzed oxyfunctionalization reactions promoted by the complete oxidation of methanol. Angew. Chem. Int. Ed. 55, 798–801 (2016).
Kiebist, J. et al. A peroxygenase from Chaetomium globosum catalyzes the selective oxygenation of testosterone. ChemBioChem 18, 563–569 (2017).
Ishimaru, A. & Yamazaki, I. The carbon monoxide-binding hemoprotein reducible by hydrogen peroxide in microsomal fractions of pea seeds. J. Biol. Chem. 252, 199–204 (1977).
Hanano, A. et al. Plant seed peroxygenase is an original haem-oxygenase with an EF-hand calcium binding motif. J. Biol. Chem. 281, 33140–33151 (2006).
Fuchs, C. & Schwab, W. Epoxidation, hydroxylation and aromatization is catalyzed by a peroxygenase from Solanum lycopersicum. J. Mol. Catal. B Enzym. 96, 52–60 (2013).
Tang, M. C., Fu, C. Y. & Tang, G. L. Characterization of SfmD as a haem peroxidase that catalyzes the regioselective hydroxylation of 3-methyltyrosine to 3-hydroxy-5-methyltyrosine in saframycin A biosynthesis. J. Biol. Chem. 287, 5112–5121 (2012).
Tuynman, A., Spelberg, J. L., Kooter, I. M., Schoemaker, H. E. & Wever, R. Enantioselective epoxidation and carbon-carbon bond cleavage catalyzed by Coprinus cinereus peroxidase and myeloperoxidase. J. Biol. Chem. 275, 3025–3030 (2000).
Ullrich, R., Nueske, J., Scheibner, K., Spantzel, J. & Hofrichter, M. Novel haloperoxidase from the agaric basidiomycete Agrocybe aegerita oxidizes aryl alcohols and aldehydes. Appl. Environ. Microbiol. 70, 4575–4581 (2004).
Lee, D. S. et al. Substrate recognition and molecular mechanism of fatty acid hydroxylation by cytochrome P450 from Bacillus subtilis: crystallographic, spectroscopic, and mutational studies. J. Biol. Chem. 278, 9761–9767 (2003).
Burek, B. O., Bormann, S., Hollmann, F., Bloh, J. Z. & Holtmann, D. Hydrogen peroxide driven biocatalysis. Green. Chem. 21, 3232–3249 (2019).
Seelbach, K., Van Deurzen, M. P. J., Van Rantwijk, F., Sheldon, R. A. & Kragl, U. Improvement of the total turnover number and space-time yield for chloroperoxidase catalyzed oxidation. Biotechnol. Bioeng. 55, 283–288 (1997).
Hofrichter, M. & Ullrich, R. Haem-thiolate haloperoxidases: versatile biocatalysts with biotechnological and environmental significance. Appl. Microbiol. Biotechnol. 71, 276–288 (2006).
Harrison, J. E. & Schultz, J. Studies on the chlorinating activity of myeloperoxidase. J. Biol. Chem. 251, 1371–1374 (1976).
Joo, H., Lin, Z. & Arnold, F. H. Laboratory evolution of cytochrome P450 hydroxylation. Nature 399, 670–673 (1999).
Bissaro, B. et al. Oxidative cleavage of polysaccharides by monocopper enzymes depends on H2O2. Nat. Chem. Biol. 13, 1123–1128 (2017). In this work, the ability of lytic polysaccharide monooxygenases to use H2O2 instead of O2 as an oxygen source for oxygenation reactions was demonstrated, which might be beneficial for their application in the enzymatic conversion of biomass.
Picard, M. et al. Metal-free bacterial haloperoxidases as unusual hydrolases: activation of H2O2 by the formation of peracetic acid. Angew. Chem. Int. Ed. 36, 1196–1199 (1997).
Björkling, F., Godtfredsen, S. E. & Kirk, O. Lipase-mediated formation of peroxycarboxylic acids used in catalytic epoxidation of alkenes. J. Chem. Soc. Chem. Commun. 19, 1301–1303 (1990).
Xu, G., Crotti, M., Saravanan, T., Kataja, K. M. & Poelarends, G. J. Enantiocomplementary epoxidation reactions catalyzed by an engineered cofactor-independent non-natural peroxygenase. Angew. Chem. Int. Ed. 59, 1–6 (2020).
Cirino, P. C. & Arnold, F. H. A self-sufficient peroxide-driven hydroxylation biocatalyst. Angew. Chem. Int. Ed. 42, 3299–3301 (2003).
Wang, Y., Lan, D., Durrani, R. & Hollmann, F. Peroxygenases en route to becoming dream catalysts. What are the opportunities and challenges? Curr. Opin. Chem. Biol. 37, 1–9 (2017).
Bormann, S., Gomez Baraibar, A., Ni, Y., Holtmann, D. & Hollmann, F. Specific oxyfunctionalisations catalysed by peroxygenases: opportunities, challenges and solutions. Catal. Sci. Technol. 5, 2038–2052 (2015).
Morris, D. R. & Hager, L. P. Chloroperoxidase, I. Isolation and properties of the crystalline glycoprotein. J. Biol. Chem. 241, 1763–1768 (1966).
Morgan, J. A., Lu, Z. & Clark, D. S. Toward the development of a biocatalytic system for oxidation of p-xylene to terephthalic acid: oxidation of 1,4-benzenedimethanol. J. Mol. Catal. B Enzym. 18, 147–154 (2002).
Morozov, A. N., Pardillo, A. D. & Chatfield, D. C. Chloroperoxidase-catalyzed epoxidation of cis-β-methylstyrene: NH–S hydrogen bonds and proximal helix dipole change the catalytic mechanism and significantly lower the reaction barrier. J. Phys. Chem. B 119, 14350–14363 (2015).
Rai, G. P., Sakai, S., Florez, A. M., Mogollon, L. & Hager, L. P. Directed evolution of chloroperoxidase for improved epoxidation and chlorination catalysis. Adv. Synth. Catal. 343, 638–645 (2001).
Allain, E. J., Hager, L. P., Deng, L. & Jacobsen, E. N. Highly enantioselective epoxidation of disubstituted alkenes with hydrogen peroxide catalyzed by chloroperoxidase. J. Am. Chem. Soc. 115, 4415–4416 (1993).
Andersson, M., Willetts, A. & Allenmark, S. Asymmetric sulfoxidation catalyzed by a vanadium-containing bromoperoxidase. J. Org. Chem. 62, 8455–8458 (1997).
Yi, X., Mroczko, M., Manoj, K. M., Wang, X. & Hager, L. P. Replacement of the proximal haem thiolate ligand in chloroperoxidase with a histidine residue. Proc. Natl Acad. Sci. USA 96, 12412–12417 (1999).
Sundaramoorthy, M., Terner, J. & Poulos, T. L. Stereochemistry of the chloroperoxidase active site: crystallographic and molecular-modeling studies. Chem. Biol. 5, 461–473 (1998).
Huang, X. & Groves, J. T. Oxygen activation and radical transformations in haem proteins and metalloporphyrins. Chem. Rev. 118, 2491–2553 (2018). Extensive overview of haem protein-mediated O2 activation processes and the reactivity of important iron−oxygen intermediates, elucidating fundamental mechanistic features of these metalloenzymes.
Matsunaga, I. & Shiro, Y. Peroxide-utilizing biocatalysts: structural and functional diversity of haem-containing enzymes. Curr. Opin. Chem. Biol. 8, 127–132 (2004).
Murphy, C. D. New frontiers in biological halogenation. J. Appl. Microbiol. 94, 539–548 (2003).
Green, M. T., Dawson, J. H. & Gray, H. B. Oxoiron(IV) in chloroperoxidase compound II is basic: implications for P450 chemistry. Science 304, 1653–1656 (2004).
Groves, J. T. Enzymatic C–H bond activation: using push to get pull. Nat. Chem. 6, 89–91 (2014).
Wang, X., Peter, S., Ullrich, R., Hofrichter, M. & Groves, J. T. Driving force for oxygen-atom transfer by haem-thiolate enzymes. Angew. Chem. Int. Ed. 52, 9238–9241 (2013).
Yosca, T. H. et al. Iron(IV)hydroxide pKa and the role of thiolate ligation in C–H bond activation by cytochrome P450. Science 342, 825–829 (2013).
Wang, X., Ullrich, R., Hofrichter, M. & Groves, J. T. Haem-thiolate ferryl of aromatic peroxygenase is basic and reactive. Proc. Natl Acad. Sci. USA 112, 3686–3691 (2015).
Conesa, A. et al. Expression of the Caldariomyces fumago chloroperoxidase in Aspergillus niger and characterization of the recombinant enzyme. J. Biol. Chem. 276, 17635–17640 (2001).
Hrycay, E. G. & Bandiera, S. M. Monooxygenase, peroxidase and peroxygenase properties of cytochrome P450 enzymes. Adv. Exp. Med. Biol. 522, 71–89 (2012).
Rydberg, P., Ryde, U. & Olsen, L. Sulfoxide, sulfur, and nitrogen oxidation and dealkylation by cytochrome P450. J. Chem. Theory Comput. 4, 1369–1377 (2008).
McKay, C. P. & Hartman, H. Hydrogen peroxide and the evolution of oxygenic photosynthesis. Orig. Life Evol. Biosph. 21, 157–163 (1991).
Meunier, B., de Visser, S. P. & Shaik, S. Mechanism of oxidation reactions catalyzed by cytochrome P450 enzymes. Chem. Rev. 104, 3947–3980 (2004).
Imai, M. et al. Uncoupling of the cytochrome P450cam monooxygenase reaction by a single mutation, threonine-252 to alanine or valine: a possible role of the hydroxy amino acid in oxygen activation. Proc. Natl Acad. Sci. USA 86, 7823–7827 (1989).
Yeom, H., Sligar, S. G., Li, H., Poulos, T. L. & Fulco, A. J. The role of Thr268 in oxygen activation of cytochrome P450 BM-3. Biochemistry 34, 14733–14740 (1995).
Munro, A. W., McLean, K. J., Grant, J. L. & Makris, T. M. Structure and function of the cytochrome P450 peroxygenase enzymes. Biochem. Soc. Trans. 46, 183–196 (2018).
Guengerich, F. P. & Munro, A. W. Unusual cytochrome P450 enzymes and reactions. J. Biol. Chem. 288, 17065–17073 (2013).
Matsunaga, I., Ueda, A., Fujiwara, N., Sumimoto, T. & Ichihara, K. Characterization of the ybdT gene product of Bacillus subtilis: novel fatty acid β-hydroxylating cytochrome P450. Lipids 34, 841–846 (1999).
Faponle, A. S., Quesne, M. G. & De Visser, S. P. Origin of the regioselective fatty-acid hydroxylation versus decarboxylation by a cytochrome P450 peroxygenase: what drives the reaction to biofuel production? Chem. Eur. J. 22, 5478–5483 (2016).
Matthews, S. et al. Catalytic determinants of alkene production by the cytochrome P450 peroxygenase OleTJE. J. Biol. Chem. 292, 5128–5143 (2017).
Ullrich, R. & Hofrichter, M. The haloperoxidase of the agaric fungus Agrocybe aegerita hydroxylates toluene and naphthalene. FEBS Lett. 579, 6247–6250 (2005).
Peter, S. et al. Selective hydroxylation of alkanes by an extracellular fungal peroxygenase. FEBS J. 278, 3667–3675 (2011).
Peter, S., Kinne, M., Ullrich, R., Kayser, G. & Hofrichter, M. Epoxidation of linear, branched and cyclic alkenes catalyzed by unspecific peroxygenase. Enzym. Microb. Technol. 52, 370–376 (2013).
Bassanini, I. et al. Peroxygenase-catalyzed enantioselective sulfoxidations. Eur. J. Org. Chem. 2017, 7186–7189 (2017).
Ullrich, R., Dolge, C., Kluge, M. & Hofrichter, M. Pyridine as novel substrate for regioselective oxygenation with aromatic peroxygenase from Agrocybe aegerita. FEBS Lett. 582, 4100–4106 (2008).
Martínez, A. T. et al. Search, engineering, and applications of new oxidative biocatalysts. Biofuels, Bioprod. Bioref. 8, 819–835 (2014).
Wang, X., Peter, S., Kinne, M., Hofrichter, M. & Groves, J. T. Detection and kinetic characterization of a highly reactive haem-thiolate peroxygenase compound I. J. Am. Chem. Soc. 134, 12897–12900 (2012).
Olmedo, A. et al. Fatty acid chain shortening by a fungal peroxygenase. Chem. Eur. J. 23, 16985–16989 (2017).
Faiza, M., Huang, S., Lan, D. & Wang, Y. New insights on unspecific peroxygenases: superfamily reclassification and evolution. BMC Evol. Biol. 19, 76–95 (2019).
Vaaje-Kolstad, G. et al. An oxidative enzyme boosting the enzymatic conversion of recalcitrant polysaccharides. Science 330, 219–222 (2010).
Vaaje-Kolstad, G., Horn, S. J., Van Aalten, D. M. F., Synstad, B. & Eijsink, V. G. H. The non-catalytic chitin-binding protein CBP21 from Serratia marcescens is essential for chitin degradation. J. Biol. Chem. 280, 28492–28497 (2005).
Horn, S. J., Vaaje-Kolstad, G., Westereng, B. & Eijsink, V. G. Novel enzymes for the degradation of cellulose. Biotechnol. Biofuels 5, 45–57 (2012).
Busk, P. K. & Lange, L. Classification of fungal and bacterial lytic polysaccharide monooxygenases. BMC Genomics 16, 368–381 (2015).
Ciano, L., Davies, G. J., Tolman, W. B. & Walton, P. H. Bracing copper for the catalytic oxidation of C–H bonds. Nat. Catal. 1, 571–577 (2018).
Quinlan, R. J. et al. Insights into the oxidative degradation of cellulose by a copper metalloenzyme that exploits biomass components. Proc. Natl Acad. Sci. USA 108, 15079–15084 (2011).
Hangasky, J. A., Iavarone, A. T. & Marletta, M. A. Reactivity of O2 versus H2O2 with polysaccharide monooxygenases. Proc. Natl Acad. Sci. USA 115, 4915–4920 (2018).
Wang, B. et al. QM/MM studies into the H2O2-dependent activity of lytic polysaccharide monooxygenases: evidence for the formation of a caged hydroxyl radical intermediate. ACS Catal. 8, 1346–1351 (2018).
Martínez, A. T. et al. Oxidoreductases on their way to industrial biotransformations. Biotechnol. Adv. 35, 815–831 (2017).
Ranganathan, S., Zeitlhofer, S. & Sieber, V. Development of a lipase-mediated epoxidation process for monoterpenes in choline chloride-based deep eutectic solvents. Green. Chem. 19, 2576–2586 (2017).
van de Velde, F., Könemann, L., van Rantwijk, F. & Sheldon, R. A. Enantioselective sulfoxidation mediated by vanadium-incorporated phytase: a hydrolase acting as a peroxidase. Chem. Commun. 29, 1891–1892 (1998).
Aharoni, A. et al. The ‘evolvability’ of promiscuous protein functions. Nat. Genet. 37, 73–76 (2005).
Miura, Y. & Fulco, A. J. ω-1, ω-2 and ω-3 Hydroxylation of long-chain fatty acids, amides and alcohols by a soluble enzyme system from Bacillus megaterium. Biochim. Biophys. Acta 388, 305–317 (1975).
Capdevila, J. H. et al. The highly stereoselective oxidation of polyunsaturated fatty acids by cytochrome P450 BM-3. J. Biol. Chem. 271, 22663–22671 (1996).
Ost, T. W. B. et al. Rational re-design of the substrate binding site of flavocytochrome P450 BM3. FEBS Lett. 486, 173–177 (2000).
Cirino, P. C. & Arnold, F. H. Regioselectivity and activity of cytochrome P450 BM-3 and mutant F87A in reactions driven by hydrogen peroxide. Adv. Synth. Catal. 344, 932–937 (2002).
Li, Q. S., Ogawa, J. & Shimizu, S. Critical role of the residue size at position 87 in H2O2-dependent substrate hydroxylation activity and H2O2 inactivation of cytochrome P450 BM-3. Biochem. Biophys. Res. Commun. 280, 1258–1261 (2001).
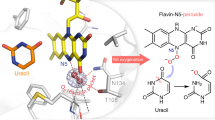
Ma, N. et al. Dual-functional small molecules for generating an efficient cytochrome P450 BM3 peroxygenase. Angew. Chem. Int. Ed. 57, 7628–7633 (2018). In this study, the P450 monooxygenase BM-3 was converted into a peroxygenase by the exogenous addition of a small molecule comprising a substrate mimicking part to activate the enzyme towards low-molecular-weight substrates and to anchor the molecule to the active site and an acid–base catalyst to enable H2O2-usage.
Shoji, O. et al. A substrate-binding-state mimic of H2O2-dependent cytochrome P450 produced by one-point mutagenesis and peroxygenation of non-native substrates. Catal. Sci. Technol. 6, 5806–5811 (2016).
Behera, R. K., Goyal, S. & Mazumdar, S. Modification of the haem active site to increase the peroxidase activity of thermophilic cytochrome P450: a rational approach. J. Inorg. Biochem. 104, 1185–1194 (2010).
Ozaki, S., Matsui, T. & Watanabe, Y. Conversion of myoglobin into a peroxygenase: a catalytic intermediate of sulfoxidation and epoxidation by the F43H/H64L mutant. J. Am. Chem. Soc. 119, 6666–6667 (1997).
Kawakami, N., Shoji, O. & Watanabe, Y. Direct hydroxylation of primary carbons in small alkanes by wild-type cytochrome P450 BM3 containing perfluorocarboxylic acids as decoy molecules. Chem. Sci. 4, 2344–2348 (2013).
Shoji, O. et al. Direct hydroxylation of benzene to phenol by cytochrome P450 BM3 triggered by amino acid derivatives. Angew. Chem. Int. Ed. 129, 10460–10465 (2017).
Shoji, O., Kunimatsu, T., Kawakami, N. & Watanabe, Y. Highly selective hydroxylation of benzene to phenol by wild-type cytochrome P450 BM3 assisted by decoy molecules. Angew. Chem. Int. Ed. 52, 6606–6610 (2013).
Kawakami, N., Shoji, O. & Watanabe, Y. Use of perfluorocarboxylic acids to trick cytochrome P450 BM3 into initiating the hydroxylation of gaseous alkanes. Angew. Chem. Int. Ed. 50, 5315–5318 (2011).
Cong, Z. et al. Activation of wild-type cytochrome P450 BM3 by the next generation of decoy molecules: enhanced hydroxylation of gaseous alkanes and crystallographic evidence. ACS Catal. 5, 150–156 (2015).
Haines, D. C. et al. Crystal structure of inhibitor-bound P450 BM-3 reveals open conformation of substrate access channel. Biochemistry 47, 3662–3670 (2008).
Shoji, O. et al. Hydrogen peroxide dependent monooxygenations by tricking the substrate recognition of cytochrome P450BSβ. Angew. Chem. Int. Ed. 46, 3656–3659 (2007). In this study, the usage of short-alkyl-chain carboxylic acids was explored to mimic the acid–base catalyst of the natural substrate of a fatty acid peroxygenase, broadening its substrate scope towards small non-natural substrates.
Kluge, M., Ullrich, R., Scheibner, K. & Hofrichter, M. Stereoselective benzylic hydroxylation of alkylbenzenes and epoxidation of styrene derivatives catalyzed by the peroxygenase of Agrocybe aegerita. Green. Chem. 14, 440–446 (2012).
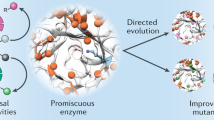
Molina-Espeja, P., De Santos, P. G. & Alcalde, M. Directed evolution of unspecific peroxygenase from Agrocybe aegerita. Appl. Environ. Microbiol. 80, 3496–3507 (2014).
Ramirez-Escudero, M. et al. Structural insights into the substrate promiscuity of a laboratory-evolved peroxygenase. ACS Chem. Biol. 13, 3259–3268 (2018).
Molina-Espeja, P., Ma, S., Mate, D. M., Ludwig, R. & Alcalde, M. Tandem-yeast expression system for engineering and producing unspecific peroxygenase. Enzym. Microb. Technol. 73–74, 29–33 (2015).
Molina-Espeja, P. et al. Synthesis of 1-naphthol by a natural peroxygenase engineered by directed evolution. ChemBioChem 17, 341–349 (2016).
Gomez De Santos, P. et al. Selective synthesis of the human drug metabolite 5′-hydroxypropranolol by an evolved self-sufficient peroxygenase. ACS Catal. 8, 4789–4799 (2018). Structure-guided evolution of the recently discovered unspecific peroxygenase AaeUPO for the synthesis of 5′-hydroxypropranolol with high enantioselectivity and reduced oxidative side reactions, making the use of radical scavengers redundant.
Gomez de Santos, P. et al. Benchmarking of laboratory evolved unspecific peroxygenases for the synthesis of human drug metabolites. Tetrahedron 75, 1827–1831 (2019).
Poraj-Kobielska, M. et al. Immobilization of unspecific peroxygenases (EC 1.11.2.1) in PVA/PEG gel and hollow fiber modules. Biochem. Eng. J. 98, 144–150 (2015).
Peng, L. et al. Peroxygenase based sensor for aromatic compounds. Biosens. Bioelectron. 26, 1432–1436 (2010).
Peng, L. et al. Bioelectrocatalytic properties of Agrocybe aegerita peroxygenase. Electrochim. Acta 55, 7809–7813 (2010).
Karich, A., Scheibner, K., Ullrich, R. & Hofrichter, M. Exploring the catalase activity of unspecific peroxygenases and the mechanism of peroxide-dependent haem destruction. J. Mol. Catal. B Enzym. 134, 238–246 (2016).
Valderrama, B., Ayala, M. & Vazquez-Duhalt, R. Suicide inactivation of peroxidases and the challenge of engineering more robust enzymes. Chem. Biol. 9, 555–565 (2002).
Ayala, M., Batista, C. V. & Vazquez-Duhalt, R. Haem destruction, the main molecular event during the peroxide-mediated inactivation of chloroperoxidase from Caldariomyces fumago. J. Biol. Inorg. Chem. 16, 63–68 (2011).
Hernández-Ruiz, J., Arnao, M. B., Hiner, A. N. P., García-Cánovas, F. & Acosta, M. Catalase-like activity of horseradish peroxidase: relationship to enzyme inactivation by H2O2. Biochem. J. 354, 107–114 (2001).
Vidal-Limón, A., Águila, S., Ayala, M., Batista, C. V. & Vazquez-Duhalt, R. Peroxidase activity stabilization of cytochrome P450 BM3 by rational analysis of intramolecular electron transfer. J. Inorg. Biochem. 122, 18–26 (2013).
Albertolle, M. E. & Peter Guengerich, F. The relationships between cytochromes P450 and H2O2: production, reaction, and inhibition. J. Inorg. Biochem. 186, 228–234 (2018).
Gonzalez-Perez, D., Garcia-Ruiz, E., Ruiz-Dueñas, F. J., Martinez, A. T. & Alcalde, M. Structural determinants of oxidative stabilization in an evolved versatile peroxidase. ACS Catal. 4, 3891–3901 (2014).
Ogola, H. J. O. et al. Enhancement of hydrogen peroxide stability of a novel Anabaena sp. DyP-type peroxidase by site-directed mutagenesis of methionine residues. Appl. Microbiol. Biotechnol. 87, 1727–1736 (2010).
Opperman, D. J. & Reetz, M. T. Towards practical Baeyer–Villiger-monooxygenases: design of cyclohexanone monooxygenase mutants with enhanced oxidative stability. ChemBioChem 11, 2589–2596 (2010).
Ziegler, D. Recent studies on the structure and function of multisubstrate flavin-containing monooxygenases. Annu. Rev. Pharmacol. Toxicol. 33, 179–199 (1993).
de Gonzalo, G. & Fraaije, M. W. Recent developments in flavin-based catalysis. ChemCatChem 5, 403–415 (2013).
De Gonzalo, G., Smit, C., Jin, J., Minnaard, A. J. & Fraaije, M. W. Turning a riboflavin-binding protein into a self-sufficient monooxygenase by cofactor redesign. Chem. Commun. 47, 11050–11052 (2011). The combination of N-alkylated flavins able to form hydroperoxyflavins and oxygenate organic substrates, with a flavin-binding protein introducing enantioselectivity, yielded an artificial peroxygenase, which can help with elucidating requirements for the cofactor and protein scaffold.
Massey, V. Activation of molecular oxygen by flavins and flavoproteins. J. Biol. Chem. 269, 22459–22462 (1994).
Kemal, C. & Bruice, T. C. Simple synthesis of a 4a-hydroperoxy adduct of a 1,5-dihydroflavine: preliminary studies of a model for bacterial luciferase. Proc. Natl Acad. Sci. USA 73, 995–999 (1976).
Kemal, C., Chan, T. W. & Bruice, T. C. Chemiluminescent reactions and electrophilic oxygen donating ability of 4a-hydroperoxyflavins: general synthetic method for the preparation of N5-alkyl-1,5-dihydroflavins. Proc. Natl Acad. Sci. USA 74, 405–409 (1977).
Smit, C., Fraaije, M. W. & Minnaard, A. J. Reduction of carbon–carbon double bonds using organocatalytically generated diimide. J. Org. Chem. 73, 9482–9485 (2008).
Hayashi, T. et al. Blue myoglobin reconstituted with an iron porphycene shows extremely high oxygen affinity. J. Am. Chem. Soc. 124, 11226–11227 (2002).
Oohora, K., Kihira, Y., Mizohata, E., Inoue, T. & Hayashi, T. C(sp3)–H bond hydroxylation catalyzed by myoglobin reconstituted with manganese porphycene. J. Am. Chem. Soc. 135, 17282–17285 (2013).
Oohora, K. et al. Manganese(v) porphycene complex responsible for inert C–H bond hydroxylation in a myoglobin matrix. J. Am. Chem. Soc. 139, 18460–18463 (2017).
Leone, L. et al. Mn–Mimochrome VI*a: an artificial metalloenzyme with peroxygenase activity. Front. Chem. 6, (2018).
Caserta, G. et al. Enhancement of peroxidase activity in artificial mimochrome vi catalysts through rational design. ChemBioChem 19, 1823–1826 (2018).
Nastri, F. et al. Haemoprotein models based on a covalent helix–haem–helix sandwich: design, synthesis, and characterization. Angew. Chem. Int. Ed. Engl. 3, 340–349 (1997).
Lombardi, A. et al. Design of a new mimochrome with unique topology. Chem. Eur. J. 9, 5643–5654 (2003).
Nastri, F. et al. A haem-peptide metalloenzyme mimetic with natural peroxidase-like activity. Chem. Eur. J. 17, 4444–4453 (2011).
Van De Velde, F., Arends, I. W. C. E. & Sheldon, R. A. Biocatalytic and biomimetic oxidations with vanadium. J. Inorg. Biochem. 80, 81–89 (2000).
Fernández-Gacio, A., Codina, A., Fastrez, J., Riant, O. & Soumillion, P. Transforming carbonic anhydrase into epoxide synthase by metal exchange. ChemBioChem 7, 1013–1016 (2006).
Fujieda, N. et al. A well-defined osmium-cupin complex: hyperstable artificial osmium peroxygenase. J. Am. Chem. Soc. 139, 5149–5155 (2017). This work shows an example of the application of a robust protein scaffold providing regioselectivity together with osmium complexed by well-exposed histidine residues to generate a thermostable artificial peroxygenase.
Carey, J. R. et al. A site-selective dual anchoring strategy for artificial metalloprotein design. J. Am. Chem. Soc. 126, 10812–10813 (2004).
Garner, D. K., Liang, L., Barrios, D. A., Zhang, J. L. & Lu, Y. The important role of covalent anchor positions in tuning catalytic properties of a rationally designed Mnsalen-containing metalloenzyme. ACS Catal. 1, 1083–1089 (2011).
Linde, D. et al. Two new unspecific peroxygenases from heterologous expression of fungal genes in Escherichia coli. Appl. Environ. Microbiol. 86, 1–16 (2020).
Carro, J. et al. Modulating fatty acid epoxidation vs hydroxylation in a fungal peroxygenase. ACS Catal. 9, 6234–6242 (2019).
Gumulya, Y. et al. Engineering highly functional thermostable proteins using ancestral sequence reconstruction. Nat. Catal. 1, 878–888 (2018). An inspiring example how ancestral sequence reconstruction can provide a tool to obtain enzymes with improved properties like enhanced thermostability.
Ander, P. & Marzullo, L. Sugar oxidoreductases and veratryl alcohol oxidase as related to lignin degradation. J. Biotechnol. 53, 115–131 (1997).
Kirk, T. K. & Farrell, R. L. Enzymatic ‘combustion’: the microbial degradation of lignin. Ann. Rev. Microbiol. 41, 465–505 (1987).
Hollmann, F. et al. Formate oxidase (FOx) from Aspergillus oryzae: one catalyst enables diverse H2O2-dependent biocatalytic oxidation reactions. Angew. Chem. Int. Ed. 58, 7873–7877 (2019).
Willot, S. J. P. et al. Expanding the spectrum of light-driven peroxygenase reactions. ACS Catal. 9, 890–894 (2019).
Horst, A. E. W. et al. Electro-enzymatic hydroxylation of ethylbenzene by the evolved unspecific peroxygenase of Agrocybe aegerita. J. Mol. Catal. B Enzym. 133, S137–S142 (2016).
Wu, S. et al. Highly regio- and enantioselective multiple oxy- and amino-functionalizations of alkenes by modular cascade biocatalysis. Nat. Commun. 7, 11917 (2016). Elegant design of four enzyme modules comprising two to three enzymes each and stable starting and end products, which can be combined in various ways to yield enzyme cascades converting alkenes to hydroxyacids, aminoalcohols or aminoacids.
Acknowledgements
We acknowledge financial support from the Netherlands Organization of Scientific Research (VICI grant 724.016.002), the European Research Council under the European Community’s Seventh Framework Programme (FP7/2007-2013)/ERC Grant agreement no. 242293, and the European Union’s Horizon 2020 research and innovation programme under the Marie Skłodowska–Curie grant agreement no. 722390.
Author information
Authors and Affiliations
Contributions
M.-C.S. examined data for the article, wrote the manuscript and prepared the figures. Both M.-C.S. and G.J.P. contributed to the discussion, reviewing and editing of the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Sigmund, MC., Poelarends, G.J. Current state and future perspectives of engineered and artificial peroxygenases for the oxyfunctionalization of organic molecules. Nat Catal 3, 690–702 (2020). https://doi.org/10.1038/s41929-020-00507-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41929-020-00507-8
This article is cited by
-
Identification, heterologous expression and characterization of a new unspecific peroxygenase from Marasmius fiardii PR-910
Bioresources and Bioprocessing (2024)
-
Reaction engineering blocks ether cleavage for synthesizing chiral cyclic hemiacetals catalyzed by unspecific peroxygenase
Nature Communications (2024)
-
Recent Progress on Peroxidase Modification and Application
Applied Biochemistry and Biotechnology (2024)
-
Heat-fueled enzymatic cascade for selective oxyfunctionalization of hydrocarbons
Nature Communications (2022)
-
A modular two yeast species secretion system for the production and preparative application of unspecific peroxygenases
Communications Biology (2021)