Abstract
Life has evolved in the ocean for 3.7 billion years, resulting in a rich ‘ocean genome’, the ensemble of genetic material present in all marine biodiversity, including both the physical genes and the information they encode. Rapid advances in sequencing technologies and bioinformatics have enabled exploration of the ocean genome and are informing innovative approaches to conservation and a growing number of commercial biotechnology applications. However, the capacity to undertake genomic research and to access and use sequence data is inequitably distributed among countries, highlighting an urgent need to build capacity, promote inclusive innovation and increase access to affordable technologies.
Similar content being viewed by others
Main
The ocean is a vast and diverse habitat that covers 70% of the Earth’s surface. Although estimates of extant species are based on indirect approaches and extrapolations can vary widely, some 2.2 million (range 0.3–10 million) eukaryotic marine species probably exist in the ocean, of which 230,000 are confirmed1,2. Comparisons with terrestrial life underscore the striking diversity of marine life: for instance, of the 34 major animal phyla, only 12 are found on land, while 33 have been recorded in the ocean3. The abundance and diversity of marine prokaryotes (bacteria and archaea) and viruses reaches monumental orders of magnitude, collectively accounting for the majority of living mass in the ocean, with estimates, extrapolated from mean values per unit volume of seawater, of 1.2 × 1029 prokaryote cells and 1.3 × 1030 virus particles found in ocean waters4,5. Some 24–98% of eukaryotic marine species, depending on the taxon group, remain undescribed, while even less is known about prokaryotic marine life, with estimates extrapolated using scaling laws, ranging from 1.0 × 106 to 3.0 × 1027 operational taxonomic units comparable, in taxonomic terms, to species2,6,7.
The diversity of marine life is closely associated with and dependent upon underlying genetic diversity, namely the total number of genetic characters in the genetic makeup of each species. Genetic diversity encodes the functional attributes of species, and their distribution and adaptability. Conserving genetic diversity provides more opportunities for evolution, and helps to foster the fitness of populations and their potential to recover from and adapt to threats ranging from disease to environmental changes8.
We define the ‘ocean genome’ as the ensemble of genetic material present in all marine biodiversity, including both the genes and the information they encode9. The explicit reference to the physical resources and informational component of genes reflects technological advances as well as the regulatory efforts striving to govern them. In recent decades, it has become possible not only to store the nucleotide sequences of DNA and RNA as digital information, but to then use this information to synthesize proteins, create molecular processes and innovation, and modify or even create organisms10,11. Genetic sequence data and innovations based on such digital information are now the subject of patent and ownership claims12. The complexity of regulating access to both informational and physical resources and equitably sharing benefits from the vast potential applications of these genetic resources across multiple industries remains unresolved and is the subject of negotiations in multiple international fora13.
In this Review, we address three questions. What are the benefits to the biosphere, to humanity and to other living organisms that arise from the ocean genome? What are the threats eroding genetic diversity in the ocean? How can the ocean genome be conserved and used in a more sustainable, fair and equitable manner?
Ecological benefits
Most attributes of organisms are encoded within their genomes, which determine much of their morphology, biology, behaviour and physiology. High levels of genetic variability and the presence of multiple genotypes within a species can result in functional redundancy that supports species resilience and adaptive capacity under environmental pressures and anomalous conditions14,15. Within the context of complex and dynamic systems, genetic diversity is therefore a crucial stabilizing factor. Such benefits have been of interest to fishery managers, as genetically diverse fish populations are better positioned to exploit a range of habitats, which adds flexibility in their responses to environmental change16. Alaska’s Bristol Bay salmon, for instance, is a highly heterogeneous population that includes over 100 discrete sub-populations, resulting in a portfolio effect, whereby the associated diversity has led not only to a more stable population, but also less frequent closures for fishing communities15. Elsewhere, the benefits of genetic variability have been recorded in restoration projects. For instance, in North America and Indonesia, plots of seagrass with higher levels of genetic diversity also exhibited higher rates of survival, plant density and growth17. The ecological benefits of genetic diversity extend beyond the resilience of individual populations. In both of the above cases, positive impacts were recorded—in the stability of populations feeding on salmon during spawning, and in increased levels of primary production and nutrient retention in restored seagrass beds in the Chesapeake Bay15,18.
Genetic variability also drives adaptive potential, which not only enhances resilience to anomalous conditions, but also enables persistence as environmental conditions change and evolve over time. Recent studies have demonstrated that this adaptive potential is of relevance even over short timeframes, for instance within the span of 200 to 600 generations (six months) of certain tropical diatoms19. Due to the prominence of coral reef ecosystems as hotspots of marine biodiversity and a crucial element of marine food webs, the bleaching and loss of corals is of special concern20. There is evidence suggesting that some corals may already have begun adapting to ocean warming caused by anthropogenic activity, rendering them more resistant in the context of mass-bleaching events21,22. This relatively rapid response is a function of genetic diversity and phenotypic plasticity at the holobiont level. Likewise, current changes in ocean conditions could alter the functional composition of marine phytoplankton communities, the foundation of virtually all marine food webs and the source of roughly half of the oxygen on the planet. In an experimental setting, cultures of marine phytoplankton with higher genetic diversity outperformed less diverse cultures with regard to their ability to withstand low salinities and maintain nitrogen uptake levels23.
Closely studied ecosystems and commercial fisheries are already providing some evidence of how genetic diversity contributes to ecosystem function and enhances adaptive potential. Yet the full value of marine genetic diversity for the ocean and the biosphere will become increasingly apparent as ocean systems continue to change and additive and synergistic impacts are better understood.
Commercial benefits
While the ocean genome provides the ecological foundation that sustains major commercial industries such as marine fisheries and tourism, commercial benefits are also derived directly from marine genes. Marine organisms, from microbes to large vertebrates, establish complex intra- and interspecific interactions mostly mediated by a variety of chemicals. These chemicals serve multiple purposes, including communication, chemical defence to predators, allelopathy, antifouling and many others24,25. These chemicals, also called secondary metabolites, are small molecules of a diverse nature (for example, peptides, sterols, phenols, terpenoids and alkaloids) often with a biological function yet to be resolved. The continuum of ‘omic’ approaches, extending from genomics and functional genomics to transcriptomics, proteomics and metabolomics provides a mechanistic pathway linking the ocean genome to the metabolites that play an important role as potential natural products for human applications, as well as a key role in modulating interactions among organisms. Much research has focused on finding useful biological activities for biomedical, cosmetic and other commercial purposes. With the recent advances in analytical techniques (mass spectrometry and nuclear magnetic resonance), new high-throughput metabolomics approaches are able to simultaneously unravel hundreds of novel compounds. When coupled to other omics technologies, such as next-generation sequencing, in a systems biology approach, insights of the complex picture of interaction among organisms can be obtained. Furthermore, the pathways for production of the metabolites can be obtained, which opens the way to their mass production using biotechnological methods26.
Perhaps the hallmark of human benefit from a marine gene is the discovery of green fluorescent protein, which produces bioluminescence in the jellyfish Aequorea victoria and has been used across a range of applications from protein tagging to identifying levels of environmental toxicity, contributions ultimately recognized with the 2008 Nobel Prize in Chemistry27. Further examples include bioprospecting for novel antifoulants and adhesives, and the search for novel antibiotics, which has increasingly focused on the bioactive compounds produced by marine invertebrates and microorganisms associated with sea sponges28,29,30. Other marine microorganisms produce a type of naturally occurring polymer (extracellular polymeric substance) that is of interest in bioremediation efforts due to its capacity to detoxify pollutants such as heavy metals31, while Pseudomonas spp. and Ideonella sakaiensis have the capacity to biodegrade certain plastics32. Additional categories of commercial activity focused on marine genetic resources are briefly introduced below. Controversies over the ownership and exclusive use of these genetic resources have persisted and present some unique challenges to existing international frameworks, as well as to potential pace of discovery.
Marine drug discovery
The marine environment has been an attractive source of bioactive compounds for the development of novel drugs. The approximately 34,000 marine natural products33 that have been reported have resulted in 8 clinically approved drugs, with a further 28 in clinical trials and 250 under preclinical investigation34. Compared with drug development from terrestrial natural products, this is a remarkable success rate35. A driving force behind the development of marine drugs has been extensive funding from the US National Cancer Institute and prospective efforts by private companies, as well as a focus on the collection of marine genetic resources globally, particularly from shallow tropical reefs and marine invertebrates. Consequently, five of the eight clinically approved drugs are treatments for cancer, with the remainder comprising treatments for neuropathic pain, the Herpes simplex virus and hypertriglyceridemia. Seven of the eight drugs were derived from sessile marine invertebrates, whose tendency to produce highly bioactive compounds—a virtual chemical arsenal—may be related to their lack of an adaptive immune system, predation pressure, and intense competition for space and resources, although the majority of these compounds have no apparent defensive function36.
Nutraceuticals and cosmeceuticals
With properties that provide medical or health benefits and also serve cosmetic or nutritional purposes, cosmeceuticals and nutraceuticals are a growing industry, with marine resources comprising an attractive source due to the wide range of exhibited metabolic pathways. The resulting diversity of bioactive compounds includes vitamins, carbohydrates, proteins and peptides, and—perhaps most prominently—omega-3 fatty acids37. While fish and crustaceans have long been exploited as sources of eicosapentaenoic acid and docosahexaenoic acid, overexploitation of fish stocks has led to research in alternative sources of omega-3 fatty acids, and the subsequent development of algal oils that can be produced in industrial quantities using phototrophic microalgae38,39. Cosmeceutical skin creams with purported anti-inflammatory and detoxifying agents have been developed from species as diverse as the Caribbean gorgonian (Pseudopterogorgia elisabethae) and bacteria isolated from deep-sea hydrothermal vents (Altermonas macleodi subsp. fijiensis biovar deepsane; Thermus thermophilus)40,41.
Aquaculture and new food products
While genetic modification has been used on a variety of commercial land crops, it remains in its infancy in the aquaculture industry. To date, only the Atlantic salmon (Salmo salar) has been commercialized using genetic engineering, namely through the insertion of growth hormone cDNA from Chinook salmon (Oncorhynchus tshawytscha) and regulated with antifreeze protein promoter sequences from the Ocean pout (Zoarces americanus) that enable it to survive in near-freezing temperatures42. The resulting transgenic salmon, which reached the market for the first time in 201743, can reach a marketable size within 16–18 months, as opposed to the three years it would otherwise require. But with just 40 fish species having fully sequenced genomes, and the recent advent of tools such as CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats) and associated enzymes (for example, Cas9) allowing selective gene editing, the number of transgenic aquaculture species is expected to increase, particularly due to a growing demand for seafood and increasing focus on lower trophic level species like seaweeds and bivalve molluscs44. However, questions of consumer acceptability, environmental risk and social desirability remain paramount and unresolved, alongside an uncertain regulatory framework45,46. Parallel research has also developed, focused on novel functional food ingredients with the potential to extend shelf-life and prevent spoilage (for example, chitosan and protein hydrolysates)47, and the use of enzymes from marine microorganisms as natural food processors48.
Bulk chemicals
Products and processes derived from marine genetic resources are of growing importance for the bulk chemical market, with applications ranging from novel laundry detergents to their use as emulsifiers and stabilizers in food production. Bioplastics derived from seaweed polymers are being used across a range of applications from straws and flip-flops to edible alternatives to plastic packaging48,49. The enzymes allowing species to flourish in extremely cold and hot marine environments have also attracted commercial interest. For example, a genetically modified version of a thermostable enzyme collected from a hydrothermal vent organism has been used for bioethanol production due to its capacity to function across wide pH and temperature ranges50. The addition of certain red seaweeds (Asparagopsis taxiformis and Asparagopsis armata) to ruminant feed has been shown to more than halve methane emissions, although concerns exist about the ozone-depleting properties of bromoform, a secondary metabolite produced by these seaweeds, if industrial-scale production for animal feed is pursued51,52.
Erosion of the ocean genome
All of these benefits—including products from marine genetic resources and ecosystem services delivered by diverse and fully functioning ocean systems—are predicated on the existing ocean genome. Yet this is threatened by the intensification of human activity around the world, which is contributing to a rapid loss of biodiversity in marine life and accelerating trends that are evident across multiple ocean-based industries53. Marine capture fisheries, coastal development and pollution have contributed to the loss of (sub-)populations and in extreme cases, species extinction, although these are rare in marine environments54. This leads to a decline in genetic diversity in the ocean, mostly concentrated, thus far, at the level of within species variability. The economic importance of the salmon industry has spurred close monitoring of the population dynamics of salmon, helping to understand the impact of human activities: within the Columbia River basin, for example, dam construction has resulted in the extinction of several sockeye salmon subpopulations, while chinook salmon have lost up to two-thirds of their genetic diversity55. The decline in genetic diversity has resulted in smaller and more variable salmon returns15. Even within subpopulations that persist, overfishing can result in the loss of genetic diversity over time, most likely reflected in loss of allelic diversity for specific genes, as the prominence of certain genotypes fluctuates and genetic drift reduces genetic diversity and lowers the capacity of species to persist and adapt to changing conditions.
Although documented species extinction has been rare in the ocean compared with recent rates of terrestrial species loss, climate change is expected to result in disproportionate levels of species loss in the ocean due to the narrow thermal range tolerated by marine ectotherms54,56. As the ocean warms and becomes more acidic and less oxygenated with climate change, the geographic distribution of species is also changing as they track their environmental niche57. At a genetic level, this implies an altered distribution of genetic variants in space and time, impacts on levels of phenotypic plasticity and changes to connectivity and population size58. Genetic variation is not uniform across species ranges, with populations in historic refuges often characterized by greater genetic diversity, and likewise threatened by shifts in distribution due to climate change59.
Efforts to optimize marine aquaculture have included the selective breeding of species and their introduction into non-native habitats. Careful monitoring and containment helps to maintain the integrity of local ecosystems, but escape events do occur, and have led to farmed species outcompeting native populations as well as the interbreeding of farmed and native species60. This results in genetic introgression and can lead to a rapid and irreversible loss of genetic diversity among the native fish populations, thereby lowering their adaptive capacity61. Such impacts could be accelerated by the accidental release of genetically modified strains in the ocean.
Much of the deep ocean and seabed remain unexplored, and scientific expeditions regularly result in the discovery of new species. A tendency for deep-sea life to exhibit slow growth rates and long lifespans renders deep-sea ecosystems particularly vulnerable to environmental disturbance62. The potential for large-scale commercial mining of the international seabed has therefore drawn particular concern within the scientific community due to uncertainty about the scale of physical and geochemical disturbance caused by mining operations and resulting sediment plumes in the deep sea, which are expected to lead to the loss of habitat and a potentially irreversible loss of biodiversity62. The scaly-foot snail (Chrysomallon squamiferum), for instance, is found exclusively on three hydrothermal vent systems in the Indian Ocean at depths of over 2,400 m, and was placed on the International Union for Conservation of Nature (IUCN) Red List in June 2019 due to two of the three systems falling within the boundaries of exploratory mining licenses granted by the International Seabed Authority63.
The acceleration of human activities on land and in the sea means that many marine species and communities are simultaneously facing multiple pressures with cumulative or synergistic effects on genetic structure and gene flow. Well-studied examples include the twin pressures of wild capture salmon fisheries and aquaculture escapement, which undermine the genetic variability of wild populations64. Looking beyond individual populations or species, multiple pressures can also result in a complete reordering of ecosystems into novel regimes. For instance, the combined and interlinked pressures from eutrophication, overfishing and the introduction of invasive species into the Black Sea led to abrupt transitions and the emergence of a new stable regime characterized by a low-energy food web dominated by jellyfish and the dinoflagellate Noctiluca scintillans65.
A more sustainable and equitable future
How can a growing understanding of the ecological and commercial benefits associated with the ocean genome be leveraged to promote conservation efforts and mitigate the drivers of genetic diversity loss in the ocean? Below, we detail not only what can be done to conserve the ocean genome, but also avenues through improved regulatory frameworks and models of inclusive innovation that can render use of the ocean genome more sustainable, equitable and fair.
Towards conservation of the ocean genome
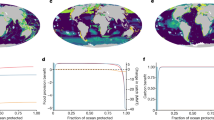
In marine systems, there are opportunities for the conservation of genetic diversity via key tools, among them ecosystem-based approaches to fisheries management, spatial planning, effective quotas, marine protected areas (MPAs), protecting and managing key marine biodiversity areas, reducing run-off pollution into oceans, and working closely with producers and consumers of ocean products66. Among these, the imperative of conserving the ocean genome would appear to be on firm footing in existing international frameworks: the importance of genetic diversity was already emphasized in the Convention on Biological Diversity (CBD) (1992), while the target of protecting at least 10% of the ocean is found in both Sustainable Development Goal 14 and Aichi Target 11. Yet genetic diversity has been largely overlooked in conservation policies and action plans67, and only 8% of the ocean is set aside for biodiversity conservation, while just 2.5% is considered fully or highly protected68.
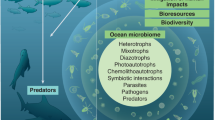
MPAs are considered one of the most effective tools for achieving the conservation of genetic diversity on an ecosystem scale69. Fully or highly protected large-scale MPAs and networks of MPAs can encompass multiple sites of importance for the life cycle of marine species. Well-managed MPAs with adequate protection levels function as storehouses of genetic diversity that simultaneously serve as important reference points for understanding changes to the ocean70. MPA networks can be designed with a specific focus on areas where genetic diversity is exceptionally high, or where particular adaptation potential lies. Such MPA adaptation networks are relevant for instance in coral reef systems71, which have been the focus of empirical work to map their adaptation potential72. The capacity for a single coral species to inhabit a range of environments characterized by high genetic diversity and on scales of less than 100 m underscores the need for protected areas to be designed with a consideration not only for potential shifts in species distribution across latitudes, but also different water depths14,73 (Fig. 1).
Effective conservation hinges on using multiple tools, including area-based conservation measures such as fully and highly protected MPAs, that provide the greatest protection from the impacts of extractive and destructive activities. Coupling these with effective management of sustainable use can ensure wide-ranging benefits that are ecological, sustaining, provisional and commercial.
Recognizing the importance of MPAs and other effective area-based conservation measures (OECMs) as tools for conserving the ocean genome, there is a particular need to optimize design to also conserve the genetic component of marine biodiversity. This remains a substantial challenge due to the rarity of temporal genetic diversity datasets or baselines, although a number of novel genetic technologies are becoming available with the potential to overcome this barrier. These include Sanger sequencing, with a history of applications ranging from wildlife conservation and management to the identification of mislabelled seafood74, as well as next-generation sequencing (NGS), a high-throughput DNA-sampling tool that can provide large-scale spatial and temporal syntheses for both individual species and community assemblages75. Several community initiatives using NGS are starting to change this landscape, such as the Earth BioGenome Project which aims to sequence, catalogue and characterize the genomes of all of Earth’s eukaryotic biodiversity over a period of 10 years76; the sponge microbiome project, a comprehensive resource of sponge-associated microbial communities based on 16S ribosomal RNA (rRNA) gene sequences that can be used to address overarching hypotheses regarding host-associated prokaryotes77; and the Earth Microbiome Project to characterize microbial life on the planet using DNA sequencing and mass spectrometry78. Emerging data on diversity within the microbiome of marine holobionts are revealing a vast reservoir of hitherto largely ignored microbial biodiversity79.
When coupled with another novel molecular approach, environmental DNA (eDNA) analysis, NGS has also been used for the detection and monitoring of marine invasive species80,81. The passive sampling techniques employed for eDNA analysis provide multiple benefits of interest to marine conservation. Due to the constant shedding of DNA by species as they interact with their environment, analysis of eDNA samples can indicate whether certain species are present in a given geography82. As a result of the rapid decomposition of eDNA over the span of days or even hours in seawater, eDNA analysis provides an almost real-time picture of species presence, including rare or elusive species, and invasive species83,84. The flexibility of the approach allows for simultaneous identification of hundreds of species in a single sample, providing insight into areas of particular species richness and potential priority for area-based protection. Recent advances in interpreting eDNA are also enabling quantification of population genetic structure and insights into trophic connectivity85. Sampling and analysis of eDNA is most powerful in well-studied marine ecosystems with substantial barcode reference collections, but may become an increasingly useful tool for the design of marine conservation interventions.
Some have touted the potential of new genome-editing techniques such as CRISPR for conservation, but their application remains theoretical. Extensive work done on corals, for example, reveals limited knowledge about potential candidate genes to target, whether this would result in phenotypic changes, whether the modified genome would be stable, and what unintended consequences gene editing could generate86,87. Moreover, a scarcity of information about the environmental, social and ethical risks of existing and new genetic engineering tools, especially in marine environments, have raised important questions about the governance and regulation of such technologies, necessitating a precautionary approach to the introduction of such technologies for conservation and fisheries management.
Although genetic techniques are rapidly evolving, policymakers do not have the luxury of waiting to make interventions until comprehensive inventories of marine genetic diversity are available, or until the extent of varied threats are fully understood. A delayed response risks resulting in the loss of rapidly deteriorating storehouses of genetic information due to over-harvesting of species and habitat degradation. Scientific recommendations to protect 30% of the ocean from all but the most minimal extraction focus on encompassing sufficient biodiversity, species biomass and representative habitats88. This requires that ongoing management outside protected areas complements these efforts by ensuring sustainable use, minimizing habitat destruction and avoiding overexploitation of resources. Such strategies include sustainable management of fisheries with a focus on ecosystem-based fisheries management, affording special protections for rare, vulnerable, threatened or endangered genotypes, populations and species, and using precautionary approaches when initiating exploitation of previously unexploited species or places.
Towards equitable benefit sharing
Investments in marine biodiscovery are typically extremely costly and risky due in part to the high costs of sampling in areas like the deep sea, the low chances of success, the technical, financial and scientific investments required, and the considerable regulatory hurdles for product approval (Fig. 2)89,90. The nature of the research enterprise is also changing, as research shifts towards bioinformatics and the mining and exploration of these vast and growing datasets of genetic information (Fig. 3), which requires advanced computational resources that are not broadly available91. As a result, most exploration has been undertaken by high-income countries, especially with regard to deep-sea research36,92,93. Disparities in research capacity, technology, finances and intellectual property rights represent major constraints that prevent the inclusion of low- and middle-income countries in marine biotechnology efforts. Biodiversity and molecular expertise is unevenly spread94, and research vessels or submersibles are typically owned only by a few high-income nations, and require substantial operational costs95, representing a major barrier to sampling of the deep ocean or in areas beyond national jurisdiction (see Box 1). While there are growing numbers of collaborations between high-income and lower-income countries96, the model of international collaboration is still characterized by a pharmaceutical or biotechnology company working with established centres of excellence located in high-income countries.
A number of international governance instruments and bodies provide an important platform where new models of equitable benefit sharing and research partnerships can evolve, and conservation of the ocean genome can be strengthened. These include the CBD, its Nagoya and Cartagena (Biosafety) Protocols, the International Treaty on Plant Genetic Resources for Food and Agriculture, the United Nations Convention on the Law of the Sea (UNCLOS) and the World Intellectual Property Organization (WIPO). Among other provisions, these agreements place the responsibility on states to conserve their biological diversity or to enter into meaningful management discussions with other countries with which they share resources (for example, transboundary fish stocks). In the context of genetic resource use, the CBD’s Nagoya Protocol sets forth the requirement for provider countries (where genetic resources are located) and user countries (those accessing and developing the genetic resources) to enter into mutually agreed terms based on prior informed consent before access to genetic resources is granted. In areas beyond national jurisdiction (ABNJ)—roughly 64% of the ocean—no restrictions currently exist on access to genetic resources, or regulations for the sharing of benefits based on their potential commercialization. However, this is one of four main elements of ongoing UN treaty negotiations of cross-cutting importance for the ocean genome (Box 1; see also Box 2).
A further complication is the so-called ‘definitional mistake’ of the CBD and Nagoya Protocol, whereby focus is placed on genetic resources in terms of their physical form rather than explicitly including the intangible informational aspects97. The development of novel genomic techniques has contributed to a 4,000-fold drop in sequencing costs over the past decade98, and has resulted in vast and exponentially growing databases of genetic sequence data and hundreds of millions of predicted genes (Fig. 3)99,100,101. While many of these databases are in the public domain and freely accessible, the growing tendency towards ‘big data’ applications means that leveraging novel genomic techniques for conservation or other uses is becoming increasingly dependent on computational and bioinformatics capacity, including access to technologies protected by intellectual property rights. Additionally, the private appropriation of genetic resources through intellectual property rights such as patents, because they result in exclusivity of use, can exacerbate existing gaps in the ability to benefit from their exploitation102. Because countries of the Global North and South do not have equal capacities or technologies to exploit these resources, there is a risk of inequitable outcomes out of sync with the 2030 Agenda for Sustainable Development and the CBD and its Nagoya Protocol103.
One opportunity rests in the development of research partnerships that connect countries that have high molecular research capacity and biotechnology infrastructure with those that do not, with such partnerships guided by norms of inclusive innovation and those of responsible research and innovation90,104. Taking an explicit focus on those excluded from the development mainstream, inclusive innovation is a conceptual approach for ensuring that innovation both addresses the problems faced by the poorest and most marginalized communities105, and also involves these communities in crafting a range of legal, technical and governance-based solutions101,102. The related concept of responsible research and innovation (RRI) envisages a transparent, interactive process by which societal actors and innovators become mutually responsive to each other with a view to the (1) ethical acceptability, (2) sustainability and (3) societal desirability of the innovation process and its marketable products106.
Historically, the majority of exploration of the ocean genome has been funded by high-income countries such as the United States, Japan, Russia and European Union (EU) states, and commercial activities and benefits continue to be concentrated within a handful of highly industrialized countries12,50. However, sustained commitments to research partnerships and inclusive and responsible research and innovation, including through capacity building and the transfer of marine technology, could result over time in a growing number of dynamic knowledge hubs and diffuse scientific collaborations outside the Global North107. Indeed, it may well be that enabling virtual access to data and the ability to use it might prove an easier task than equalizing physical access to marine genetic resources.
While the concept of inclusive and responsible research and innovation is appealing and in line with existing governance and regulatory frameworks, there are substantial and unresolved legal and ethical issues related to the use and sharing of genomic information. The science bodies of the EU, United States and Australia, for instance, require genomic data collected over the course of funded projects to be deposited in open-access databases. While regulations on disclosure of origin and other measures aimed at increasing transparency in the use of genetic sequence data are being negotiated within multiple international fora108, the informational component of the ocean genome is increasingly entering the public domain and becoming a de jure universal resource, that is de facto only accessible to those with corresponding capacity. The capacity for industry actors to access public databases, while remaining unbound by benefit-sharing requirements, has compounded concerns among some countries of both a loss of control over national patrimony and the management of global resources and a loss of opportunity to reap benefits109.
Scientists and policymakers have noted the importance of respecting such concerns and avoiding inequitable exploitation—commercial or otherwise—while also ensuring that scientific progress can continue with as few impediments as possible. Some have criticized the Nagoya Protocol for inadvertently hampering taxonomic research and international collaborations110, while being unable to enforce meaningful benefits sharing. Others have underscored that UNCLOS ensures freedom to undertake scientific research, including in ABNJ (Articles 256–257)111 while still others have rejected the legitimacy of legal claims that arise from marine scientific research (Article 241), including in the form of intellectual property rights. The outcome of the biodiversity in areas beyond national jurisdiction (BBNJ) negotiations (see Box 1) has the potential to not only strike this balance for ABNJ, but also to illustrate alternative pathways for regulating the use and circulation of genetic resources internationally.
Conclusion
The future state of ocean ecosystems will depend in large part on recognizing that human activity has already substantially eroded the ocean genome and that this has been to the detriment of the biosphere, humanity and other life forms. Encouraging signals are emerging, including recent calls to move beyond current international targets by ensuring that in the future at least 30% of the ocean is fully or highly protected88, the declaration of a UN Decade of Ocean Science for Sustainable Development (2021–2030), and the consensus decision among UN member states to move forward with negotiating a legally binding instrument for conservation and sustainable use of BBNJ. Yet human impacts on the ocean are growing alongside accelerating commercial use of its resources and space53,112, while the potential for new industries such as mining of the international seabed and methane hydrates poses vast risks for the ocean genome62. Ensuring that the ocean genome is conserved will require effective regulation and governance based on inclusive and iterative dialogue processes that connect diverse stakeholders, are based on principles of fairness, equity and inclusivity, and are informed by the latest scientific techniques and knowledge of the ocean genome.
References
Strother, P. K., Battison, L., Brasier, M. D. & Wellman, C. H. Earth’s earliest non-marine eukaryotes. Nature 473, 505–509 (2011).
Louca, S., Mazel, F., Doebeli, M. & Parfrey, L. A census-based estimate of Earth’s bacterial and archaeal diversity. PLoS Biol. 17, e3000106 (2019).
Jaume, D. & Duarte, C. M. in The Exploration of Marine Biodiversity–Scientific and Technological Challenges (ed. Duarte, C. M.) 17–30 (Fundación BBVA, 2006).
Bar-On, Y. M., Phillips, R. & Milo, R. The biomass distribution on Earth. Proc. Natl Acad. Sci. USA 115, 6506–6511 (2018).
Cobián Güemes, A. G. et al. Viruses as winners in the game of life. Annu. Rev. Virol. 3, 197–214 (2016).
Locey, K. J. & Lennon, J. T. Scaling laws predict global microbial diversity. Proc. Natl Acad. Sci. USA 113, 5970–5975 (2016).
Mora, C., Tittensor, D. P., Adl, S., Simpson, A. G. B. & Worm, B. How many species are there on Earth and in the ocean? PLoS Biol. 9, e1001127 (2011).
Reed, D. H. & Frankham, R. Correlation between Fitness and Genetic Diversity. Conserv. Biol. 17, 230–237 (2003).
Blasiak, R. et al. The Ocean Genome: Conservation and the Fair, Equitable and Sustainable Use of Marine Genetic Resources (High Level Panel for a Sustainable Ocean Economy, 2020).
Gibson, D. G. et al. Creation of a bacterial cell controlled by a chemically synthesized genome. Science 329, 52–56 (2010).
Hutchison, C. A. III et al. Design and synthesis of a minimal bacterial genome. Science 351, aad6253 (2016).
Blasiak, R., Jouffray, J.-B., Wabnitz, C. C., Sundström, E. & Österblom, H. Corporate control and global governance of marine genetic resources. Sci. Adv. 4, eaar5237 (2018).
Blasiak, R. International regulatory changes poised to reshape access to marine genes. Nat. Biotechnol. 37, 357–358 (2019).
Webster, M. S. et al. Who should pick the winners of climate change? Trends Ecol. Evol. 32, 167–173 (2017).
Schindler, D. E. et al. Population diversity and the portfolio effect in an exploited species. Nature 465, 609–612 (2010).
Ruzzante, D. E. et al. Biocomplexity in a highly migratory pelagic marine fish, Atlantic herring. Proc. Biol. Sci. 273, 1459–1464 (2006).
Reynolds, P. L., Richardson, J. P. & Duffy, J. E. Field experimental evidence that grazers mediate transition between microalgal and seagrass dominance. Limnol. Oceanogr. 59, 1053–1064 (2014).
Reynolds, L. K., McGlathery, K. J. & Waycott, M. Genetic diversity enhances restoration success by augmenting ecosystem services. PLoS ONE 7, e38397 (2012).
Jin, P. & Agustí, S. Fast adaptation of tropical diatoms to increased warming with trade-offs. Sci. Rep. 8, 17771 (2018).
Norström, A. V. et al. Guiding coral reef futures in the Anthropocene. Front. Ecol. Environ. 14, 490–498 (2016).
Morikawa, M. K. & Palumbi, S. R. Using naturally occurring climate resilient corals to construct bleaching-resistant nurseries. Proc. Natl Acad. Sci. USA 116, 10586–10591 (2019).
Ziegler, M., Seneca, F. O., Yum, L. K., Palumbi, S. R. & Voolstra, C. R. Bacterial community dynamics are linked to patterns of coral heat tolerance. Nat. Commun. 8, 14213 (2017).
Sjöqvist, C. O. & Kremp, A. Genetic diversity affects ecological performance and stress response of marine diatom populations. ISME J. 10, 2755–2766 (2016).
Brown, E. R., Cepeda, M. R., Mascuch, S. J., Poulson-Ellestad, K. L. & Kubanek, J. Chemical ecology of the marine plankton. Nat. Prod. Rep. 36, 1093–1116 (2019).
Puglisi, M. P., Sneed, J. M., Sharp, K. H., Ritson-Williams, R. & Paul, V. J. Marine chemical ecology in benthic environments. Nat. Prod. Rep. 31, 1510–1553 (2014).
Kuhlisch, C. & Pohnert, G. Metabolomics in chemical ecology. Nat. Prod. Rep. 32, 937–955 (2015).
Rogers, A. D. The Deep: The Hidden Wonders of Our Oceans and How We Can Protect Them (Wildfire, 2019).
El Samak, M., Solyman, S. M. & Hanora, A. Antimicrobial activity of bacteria isolated from Red Sea marine invertebrates. Biotechnol. Rep. (Amst.) 19, e00275 (2018).
Tortorella, E. et al. Antibiotics from deep-sea microorganisms: current discoveries and perspectives. Mar. Drugs 16, 355 (2018).
Tincu, J. A. & Taylor, S. W. Antimicrobial peptides from marine invertebrates. Antimicrob. Agents Chemother. 48, 3645–3654 (2004).
Pal, A. & Paul, A. K. Microbial extracellular polymeric substances: central elements in heavy metal bioremediation. Indian J. Microbiol. 48, 49–64 (2008).
Wilkes, R. A. & Aristilde, L. Degradation and metabolism of synthetic plastics and associated products by Pseudomonas sp.: capabilities and challenges. J. Appl. Microbiol. 123, 582–593 (2017).
A Database of Marine Natural Products Literature (MarinLit, 2019).
Clinical Pipeline (Midwestern University, accessed 9 November 2019); https://go.nature.com/3c1snHp
Gerwick, W. H. & Moore, B. S. Lessons from the past and charting the future of marine natural products drug discovery and chemical biology. Chem. Biol. 19, 85–98 (2012).
Leal, M. C., Puga, J., Serôdio, J., Gomes, N. C. M. & Calado, R. Trends in the discovery of new marine natural products from invertebrates over the last two decades – where and what are we bioprospecting? PLoS ONE 7, e30580 (2012).
Suleria, H. A. R., Osborne, S., Masci, P. & Gobe, G. Marine-based nutraceuticals: an innovative trend in the food and supplement industries. Mar. Drugs 13, 6336–6351 (2015).
Rodolfi, L. et al. Microalgae for oil: strain selection, induction of lipid synthesis and outdoor mass cultivation in a low-cost photobioreactor. Biotechnol. Bioeng. 102, 100–112 (2009).
Chauton, M. S., Reitan, K. I., Norsker, N. H., Tveterås, R. & Kleivdal, H. T. A techno-economic analysis of industrial production of marine microalgae as a source of EPA and DHA-rich raw material for aquafeed: research challenges and possibilities. Aquaculture 436, 95–103 (2015).
Jaspars, M. et al. The marine biodiscovery pipeline and ocean medicines of tomorrow. J. Mar. Biol. Assoc. UK 96, 151–158 (2016).
Cambon-Bonavita, M. A., Raguenes, G., Vincent, P. & Guezennec, J. A novel polymer produced by a bacterium isolated from a deep-sea hydrothermal vent polychaete annelid. J. Appl. Microbiol. 93, 310–315 (2002).
Smith, M. D., Asche, F., Guttormsen, A. G. & Wiener, J. B. Genetically modified salmon and full impact assessment. Science 330, 1052–1053 (2010).
Waltz, E. First genetically engineered salmon sold in Canada. Nat. News 548, 148 (2017).
Zhu, B. & Ge, W. Genome editing in fishes and their applications. Gen. Comp. Endocrinol. 257, 3–12 (2018).
Otts, S. S. US regulatory framework for genetic biocontrol of invasive fish. Biol. Invasions 16, 1289–1298 (2014).
Van Eenennaam, A. L., Wells, K. D. & Murray, J. D. Proposed US regulation of gene-edited food animals is not fit for purpose. NPJ Sci. Food 3, 3 (2019).
Shahidi, F. & Ambigaipalan, P. Novel functional food ingredients from marine sources. Curr. Opin. Food Sci. 2, 123–129 (2015).
Beygmoradi, A. & Homaei, A. Marine microbes as a valuable resource for brand new industrial biocatalysts. Biocatal. Agric. Biotechnol. 11, 131–152 (2017).
Guedes, A. C., Amaro, H. M., Sousa-Pinto, I. & Malcata, F. X. in Biofuels from Algae 2nd edn (eds Pandey, A. et al.) 397–433 (Elsevier, 2019).
Leary, D., Vierros, M., Hamon, G., Arico, S. & Monagle, C. Marine genetic resources: a review of scientific and commercial interest. Mar. Policy 33, 183–194 (2009).
Roque, B. M., Salwen, J. K., Kinley, R. & Kebreab, E. Inclusion of Asparagopsis armata in lactating dairy cows’ diet reduces enteric methane emission by over 50 percent. J. Clean. Prod. 234, 132–138 (2019).
Carpenter, L. J. & Liss, P. S. On temperate sources of bromoform and other reactive bromine gases. J. Geophys. Res.: Atmos. 105, 20539–20547 (2000).
Jouffray, J.-B., Blasiak, R., Norström, A. V., Österblom, H. & Nyström, M. The blue acceleration – the trajectory of human expansion into the ocean. One Earth 2, 43–54 (2020).
McCauley, D. J. et al. Marine defaunation: animal loss in the global ocean. Science 347, 1255641 (2015).
Johnson, B. M., Kemp, B. M. & Thorgaard, G. H. Increased mitochondrial DNA diversity in ancient Columbia River basin Chinook salmon Oncorhynchus tshawytscha. PLoS ONE 13, e0190059 (2018).
Pinsky, M. L., Eikeset, A. M., McCauley, D. J., Payne, J. L. & Sunday, J. M. Greater vulnerability to warming of marine versus terrestrial ectotherms. Nature 569, 108–111 (2019).
Poloczanska, E. S. et al. Responses of marine organisms to climate change across oceans. Front. Mar. Sci. 3, 62 (2016).
Hoffmann, A. A. & Sgrò, C. M. Climate change and evolutionary adaptation. Nature 470, 479–485 (2011).
Provan, J. & Maggs, C. A. Unique genetic variation at a species’ rear edge is under threat from global climate change. Proc. Biol. Sci. 279, 39–47 (2012).
Fleming, I. A. et al. Lifetime success and interactions of farm salmon invading a native population. Proc. R. Soc. Lond. B: Biol. Sci. 267, 1517–1523 (2000).
Glover, K. A. et al. Half a century of genetic interaction between farmed and wild Atlantic salmon: status of knowledge and unanswered questions. Fish Fish. 18, 890–927 (2017).
Van Dover, C. L. et al. Biodiversity loss from deep-sea mining. Nat. Geosci. 10, 464–465 (2017).
Sigwart, J. D. et al. Red Listing can protect deep-sea biodiversity. Nat. Ecol. Evol. 3, 1134 (2019).
Waples, R. S., Hindar, K. & Hard, J. J. Genetic Risks Associated with Marine Aquaculture NOAA Technical Memo NMFD-NWFSC-119 (US Department of Commerce, NOAA, 2012).
Oguz, T. & Velikova, V. Abrupt transition of the northwestern Black Sea shelf ecosystem from a eutrophic to an alternative pristine state. Mar. Ecol. Prog. Ser. 405, 231–242 (2010).
Summary for Policymakers of The Global Assessment Report on Biodiversity and Ecosystem Services – Unedited Advance Version (IPBES 2019).
Laikre, L. Genetic diversity is overlooked in international conservation policy implementation. Conserv. Genet. 11, 349–354 (2010).
Sala, E. et al. Assessing real progress towards effective ocean protection. Mar. Policy 91, 11–13 (2018).
Sala, E. & Giakoumi, S. No-take marine reserves are the most effective protected areas in the ocean. ICES J. Mar. Sci. 75, 1166–1168 (2018).
Grorud-Colvert, K. et al. Marine protected area networks: assessing whether the whole is greater than the sum of its parts. PLoS ONE 9, e102298 (2014).
Costello, M. J. Long live Marine Reserves: a review of experiences and benefits. Biol. Conserv. 176, 289–296 (2014).
Hughes, T. P. et al. Global warming transforms coral reef assemblages. Nature 556, 492–496 (2018).
Barshis, D. J. et al. Genomic basis for coral resilience to climate change. Proc. Natl Acad. Sci. USA 110, 1387–1392 (2013).
Tinacci, L. et al. DNA barcoding for the verification of supplier’s compliance in the seafood chain: how the lab can support companies in ensuring traceability. Ital. J. Food Saf. 7, 83–88 (2018).
Djurhuus, A. et al. Evaluation of marine zooplankton community structure through environmental DNA metabarcoding. Limnol. Oceanogr.: Methods 16, 209–221 (2018).
Lewin, H. A. et al. Earth Biogenome Project: sequencing life for the future of life. Proc. Natl Acad. Sci. USA 115, 4325–4333 (2018).
Moitinho-Silva, L., Nielsen, S., Amir, A. & Gonzalez, A. The sponge microbiome project. GigaScience 6, gix077 (2017).
Thompson, L. R., Sanders, J. G., McDonald, D. & Amir, A. A communal catalogue reveals Earth’s multiscale microbial diversity. Nature 551, 457–463 (2017).
Ainsworth, T. D., Krause, L., Bridge, T., Torda, G. & Raina, J. B. The coral core microbiome identifies rare bacterial taxa as ubiquitous endosymbionts. ISME J. 9, 2261–2274 (2015).
Ardura, A. et al. eDNA and specific primers for early detection of invasive species–a case study on the bivalve Rangia cuneata, currently spreading in Europe. Mar. Environ. Res. 112, 48–55 (2015).
Simmons, M., Tucker, A., Chadderton, W. L., Jerde, C. L. & Mahon, A. R. Active and passive environmental DNA surveillance of aquatic invasive species. Can. J. Fish. Aquat. Sci. 73, 76–83 (2015).
Baird, D. J. & Hajibabaei, M. Biomonitoring 2.0: a new paradigm in ecosystem assessment made possible by next-generation DNA sequencing. Mol. Ecol. 21, 2039–2044 (2012).
Bakker, J. et al. Environmental DNA reveals tropical shark diversity in contrasting levels of anthropogenic impact. Sci. Rep. 7, 16886 (2017).
Weltz, K. et al. Application of environmental DNA to detect an endangered marine skate species in the wild. PLoS ONE 12, e0178124 (2017).
Jeunen, G.-J. et al. Environmental DNA (eDNA) metabarcoding reveals strong discrimination among diverse marine habitats connected by water movement. Mol. Ecol. Resour. 19, 426–438 (2019).
Caplan, A. L., Parent, B., Shen, M. & Plunkett, C. No time to waste—the ethical challenges created by CRISPR. EMBO Rep. 16, 1421–1426 (2015).
A Research Review of Interventions to Increase the Persistence and Resilience of Coral Reefs (National Academies of Sciences, Engineering, and Medicine, 2018); https://doi.org/10.17226/25279
O’Leary, B. C. et al. Effective coverage targets for ocean protection. Conserv. Lett. 9, 398–404 (2016).
Laird, S. & Wynberg, R. Bioscience at a Crossroads: Implementing the Nagoya Protocol on Access and Benefit Sharing in a Time of Scientific, Technological and Industry Change (CBD, 2012).
Morgera, E. Fair and equitable benefit-sharing in a new treaty on marine biodiversity: a principled approach towards partnership building? Marit. Safe. Secur. Law J. 5, 48–77 (2018).
Muir, P. et al. The real cost of sequencing: scaling computation to keep pace with data generation. Genome Biol. 17, 53 (2016).
Greiber, T. An Explanatory Guide to The Nagoya Protocol on Access and Benefit-Sharing (IUCN, 2012).
Oldham, P., Hall, S. & Forero, O. Biological diversity in the patent system. PLoS ONE 8, e78737 (2013).
Hendriks, I. E. & Duarte, C. M. Allocation of effort and imbalances in biodiversity research. J. Exp. Mar. Biol. Ecol. 360, 15–20 (2008).
Stokstad, E. Norwegian billionaire funds deluxe deep ocean research ship. Science (19 November 2018); https://go.nature.com/2VeUDzA
Kyeremeh, K. et al. in Africa and the Sustainable Development Goals (eds Ramutsindela, M. & Mickler, D.) 257–266 (Springer International Publishing, 2020); https://doi.org/10.1007/978-3-030-14857-7_24
Mueller, M. R. Genetic Resources as Natural Information Implications for the Convention on Biological Diversity and Nagoya Protocol (Routledge, 2015).
Green, E. D., Rubin, E. M. & Olson, M. V. The future of DNA sequencing. Nat. News 550, 179 (2017).
Laird, S. & Wynberg, R. A Fact-Finding and Scoping Study on Digital Sequence Information on Genetic Resources in the Context of the Convention on Biological Diversity and the Nagoya Protocol 2–79 (Secretariat of CBD, 2018).
Carradec, Q. et al. A global ocean atlas of eukaryotic genes. Nat. Commun. 9, 373 (2018).
Gregory, A. C. et al. Marine DNA viral macro- and microdiversity from pole to pole. Cell 177, 1109–1123 (2019).
Angrist, M. & Cook-Deegan, R. Distributing the future: the weak justifications for keeping genomic databases secret and the challenges and opportunities in reverse engineering them. Appl. Transl. Genomics 3, 124–127 (2014).
Österblom, H. et al. Towards Ocean Equity (High Level Panel for a Sustainable Ocean Economy, 2020).
Heeks, R., Amalia, M., Kintu, R. & Shah, N. Inclusive Innovation: Definition, Conceptualisation and Future Research Priorities Development Informatics Working Paper no. 53 (Centre for Development Informatics, Institute for Development Policy and Management, SEED, University of Manchester, 2013).
Foster, C. & Heeks, R. Conceptualising inclusive innovation: modifying systems of innovation frameworks to understand diffusion of new technology to low-income consumers. Eur. J. Dev. Res. 25, 333–355 (2013).
Von Schomberg, R. in Responsible Innovation. Managing the Responsible Emergence of Science and Innovation in Society (eds Owen, R. et al.) 51–74 (Wiley, 2013).
Broggiato, A., Arnaud-Haond, S., Chiarolla, C. & Greiber, T. Fair and equitable sharing of benefits from the utilization of marine genetic resources in areas beyond national jurisdiction: bridging the gaps between science and policy. Mar. Policy 49, 176–185 (2014).
Blasiak, R., Jouffray, J.-B., Wabnitz, C. C. & Österblom, H. Scientists should disclose origin in marine gene patents. Trends Ecol. Evol. 34, 392–395 (2019).
Elbe, S. & Buckland-Merrett, G. Data, disease and diplomacy: GISAID’s innovative contribution to global health. Glob. Chall. 1, 33–46 (2017).
Deplazes-Zemp, A. et al. The Nagoya Protocol could backfire on the Global South. Nat. Ecol. Evol. 2, 917–919 (2018).
Vierros, M., Suttle, C. A., Harden-Davies, H. & Burton, G. Who owns the ocean? Policy issues surrounding marine genetic resources. Limnol. Oceanogr. Bull. 25, 29–35 (2016).
Halpern, B. S. et al. Recent pace of change in human impact on the world’s ocean. Sci. Rep. 9, 11609 (2019).
Sequence Read Archive (NCBI, 2020); https://go.nature.com/2yPdoSV
Wetterstrand, K. A. DNA Sequencing Costs: Data from the NHGRI Genome Sequencing Program (GSP) (NHGRI, 2020).
Wright, G., Rochette, J., Druel, E. & Gjerde, K. The Long and Winding Road Continues: Towards a New Agreement on High Seas Governance (IDDRI, 2016); https://go.nature.com/2KmgEYj
Acknowledgements
The author group was assembled under the auspices of the High-Level Panel for a Sustainable Ocean Economy, and provided with support to produce a paper titled ‘The ocean genome: conservation and the fair equitable and sustainable use of marine genetic resources’, on which this Review heavily draws. We thank J. Lokrantz (Azote) and J. B. Jouffray for their support with conceptualizing and designing figures.
Author information
Authors and Affiliations
Contributions
R.B., R.W., K.G.-C., S.T., N.M.B., A.V.M.C., J.D.S., C.M.D., M.J., A.R., K.S. and C.C.C.W. all contributed to the writing and reviewing of this manuscript.
Corresponding author
Ethics declarations
Competing interests
M.J. is founder of, shareholder in and consultant for ‘GyreOx Ltd’ which uses marine and terrestrial enzymes for the rapid production of complex molecules to target protein-protein interactions involved in disease. S.T. has participated in the BBNJ negotiations as a Pacific Islands Forum Adviser. All other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Blasiak, R., Wynberg, R., Grorud-Colvert, K. et al. The ocean genome and future prospects for conservation and equity. Nat Sustain 3, 588–596 (2020). https://doi.org/10.1038/s41893-020-0522-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41893-020-0522-9
This article is cited by
-
When will the BBNJ Agreement deliver results?
npj Ocean Sustainability (2024)
-
Making marine biotechnology work for people and nature
Nature Ecology & Evolution (2023)
-
Social equity is key to sustainable ocean governance
npj Ocean Sustainability (2022)
-
Seeing through sedimented waters: environmental DNA reduces the phantom diversity of sharks and rays in turbid marine habitats
BMC Ecology and Evolution (2021)
-
Our future in the Anthropocene biosphere
Ambio (2021)