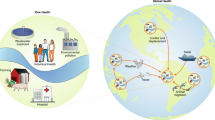
Rising levels of antimicrobial and pesticide resistance increasingly undermine human health and systems for biomass production, and emphasize the sustainability challenge of preserving organisms susceptible to these biocides. In this Review, we introduce key concepts and examine dynamics of biocide susceptibility that must be governed to address this challenge. We focus on the impact of biocides on the capacity of susceptible organisms to prevent spread of resistance, and we then review how biocide use affects a broader suite of ecosystem services. Finally, we introduce and assess the state of what we term the Anthropocene operating space of biocide susceptibility, a framework for assessing the potential of antibiotic and pesticide resistance to undermine key functions of human society. Based on current trends in antibiotic, insecticide and herbicide resistance, we conclude that the states of all six assessed variables are beyond safe zones, with three variables surpassed regionally or globally.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Ellis, E. C. Ecology in an anthropogenic biosphere. Ecol. Monogr. 85, 287–331 (2015).
Carroll, S. P. et al. Applying evolutionary biology to address global challenges. Science 346, 1245993 (2014).
Palumbi, S. R. Humans as the world’s greatest evolutionary force. Science 293, 1786–1790 (2001).
Millennium Ecosystem Assessment. Ecosystems and Human Well-being: Synthesis (Island, Washington DC, 2005).
Jørgensen, P. S., Wernli, D., Folke, C. & Carroll, S. P. Changing antibiotic resistance: sustainability transformation to a pro-microbial planet. Curr. Opin. Environ. Sustain 25, 66–76 (2017).
Wernli, D. et al. Mapping global policy discourse on antimicrobial resistance. BMJ Glob. Health 2, e000378 (2017).
Polasky, S., Carpenter, S. R., Folke, C. & Keeler, B. Decision-making under great uncertainty: environmental management in an era of global change. Trends Ecol. Evol. 26, 398–404 (2011).
IPBES/5/Inf/24: Update on the Classification of Nature’s Contributions to People by the Intergovernmental Science-Policy Platform on Biodiversity and Ecosystem Services (IPBES, 2017).
Zhang, W., Ricketts, T. H., Kremen, C., Carney, K. & Swinton, S. M. Ecosystem services and dis-services to agriculture. Ecol. Econ. 64, 253–260 (2007).
Fisher, B., Costanza, R., Kerry, T. & Morling, P. Defining and Classifying Ecosystem Services for Decision Making CSERGE Working Paper EDM 07-04 (CSERGE, 2007).
Saunders, M. E. Ecosystem services in agriculture: understanding the multifunctional role of invertebrates. Agric. For. Entomol. 20, 298–300 (2018).
Saunders, M. E., Peisley, R. K., Rader, R. & Luck, G. W. Pollinators, pests, and predators: recognizing ecological trade-offs in agroecosystems. Ambio 45, 4–14 (2016).
Saunders, M. E. & Luck, G. W. Limitations of the ecosystem services versus disservices dichotomy. Conserv. Biol. 30, 1363–1365 (2016).
Lynch, S. V. & Pedersen, O. The human intestinal microbiome in health and disease. N. Engl. J. Med. 375, 2369–2379 (2016).
Díaz, S. et al. The IPBES Conceptual Framework—connecting nature and people. Curr. Opin. Environ. Sustain 14, 1–16 (2015).
Daulaire, N., Bang, A., Tomson, G., Kalyango, J. N. & Cars, O. Universal access to effective antibiotics is essential for tackling antibiotic resistance. J. Law Med. Ethics 43, 17–21 (2015).
Creanza, N., Kolodny, O. & Feldman, M. W. Cultural evolutionary theory: how culture evolves and why it matters. Proc. Natl Acad. Sci. USA 114, 7782–7789 (2017).
Faith, D. P., Magallón, S., Hendry, A. P. & Donoghue, M. J. Future benefits from contemporary evosystem services: a response to Rudman et al. Trends Ecol. Evol. 32, 717–719 (2017).
Faith, D. P. et al. Evosystem services: an evolutionary perspective on the links between biodiversity and human well-being. Curr. Opin. Environ. Sustain 2, 66–74 (2010).
Rudman, S. M., Kreitzman, M., Chan, K. M. A. & Schluter, D. Evosystem services: rapid evolution and the provision of ecosystem services. Trends Ecol. Evol. 32, 403–415 (2017).
Rudman, S. M., Kreitzman, M., Chan, K. M. A. & Schluter, D. Contemporary evosystem services: a reply to Faith et al. Trends Ecol. Evol. 32, 719–720 (2017).
Baquero, F. et al. Public health evolutionary biology of antimicrobial resistance: priorities for intervention. Evol. Appl 8, 223–239 (2015).
Powles, S. B. & Yu, Q. Evolution in action: plants resistant to herbicides. Annu. Rev. Plant Biol. 61, 317–347 (2010).
Davies, J. & Davies, D. Origins and evolution of antibiotic resistance. Microbiol. Mol. Biol. Rev. 74, 417–33 (2010).
Cousens, R. D. & Fournier-Level, A. Herbicide resistance costs: what are we actually measuring and why? Pest Manag. Sci. 74, 1539–1546 (2018).
Carpenter, S., Walker, B., Anderies, J. M. & Abel, N. From metaphor to measurement: resilience of what to what? Ecosystems 4, 765–781 (2001).
Sommer, F., Anderson, J. M., Bharti, R., Raes, J. & Rosenstiel, P. The resilience of the intestinal microbiota influences health and disease. Nat. Rev. Microbiol. 15, 630–638 (2017).
Lozupone, C. A., Stombaugh, J. I., Gordon, J. I., Jansson, J. K. & Knight, R. Diversity, stability and resilience of the human gut microbiota. Nature 489, 220–230 (2012).
Chen, Q.-L. et al. Do manure-borne or indigenous soil microorganisms influence the spread of antibiotic resistance genes in manured soil? Soil Biol. Biochem. 114, 229–237 (2017).
Griffiths, B. S. & Philippot, L. Insights into the resistance and resilience of the soil microbial community. FEMS Microbiol. Rev. 37, 112–129 (2013).
Mullineaux-Sanders, C., Suez, J., Elinav, E. & Frankel, G. Sieving through gut models of colonization resistance. Nat. Microbiol 3, 132–140 (2018).
Francino, M. P. Antibiotics and the human gut microbiome: dysbioses and accumulation of resistances. Front. Microbiol 6, 1543 (2016).
Andersson, D. I. & Hughes, D. Antibiotic resistance and its cost: is it possible to reverse resistance? Nat. Rev. Microbiol. 8, 260–271 (2010).
Waglechner, N. & Wright, G. D. Antibiotic resistance: it’s bad, but why isn’t it worse? BMC Biol. 15, 84 (2017).
Seekatz, A. M. et al. Recovery of the gut microbiome following fecal microbiota transplantation. MBio 5, e00893–14 (2014).
Caballero, S. et al. Cooperating commensals restore colonization resistance to vancomycin-resistant Enterococcus faecium. Cell Host Microbe 21, 592–602 (2017).
Baquero, F., Coque, T. M. & Cantón, R. Counteracting antibiotic resistance: breaking barriers among antibacterial strategies. Expert Opin. Ther. Targets 18, 851–861 (2014).
REX Consortium. Heterogeneity of selection and the evolution of resistance. Trends Ecol. Evol. 28, 110–118 (2013).
Wales, A. & Davies, R. Co-selection of resistance to antibiotics, biocides and heavy metals, and its relevance to foodborne pathogens. Antibiotics 4, 567–604 (2015).
Pal, C., Bengtsson-Palme, J., Kristiansson, E. & Larsson, D. G. J. Co-occurrence of resistance genes to antibiotics, biocides and metals reveals novel insights into their co-selection potential. BMC Genomics 16, 964 (2015).
Baker-Austin, C., Wright, M. S., Stepanauskas, R. & McArthur, J. V. Co-selection of antibiotic and metal resistance. Trends Microbiol. 14, 176–182 (2006).
Chang, Q., Wang, W., Regev‐Yochay, G., Lipsitch, M. & Hanage, W. P. Antibiotics in agriculture and the risk to human health: how worried should we be? Evol. Appl 8, 240–247 (2015).
Neve, P., Busi, R., Renton, M. & Vila‐Aiub, M. M. Expanding the eco‐evolutionary context of herbicide resistance research. Pest Manag. Sci. 70, 1385–1393 (2014).
Zhu, Y.-G. et al. Microbial mass movements. Science 357, 1098–1099 (2017).
Zhu, Y. G. et al. Human dissemination of genes and microorganisms in Earth’s critical zone. Glob. Chang. Biol 24, 1488–1499 (2018).
Elmqvist, T. et al. Response diversity, ecosystem change, and resilience. Front. Ecol. Environ. 1, 488–494 (2003).
Folke, C. et al. Regime shifts, resilience, and biodiversity in ecosystem management. Annu. Rev. Ecol. Evol. Syst. 35, 557–581 (2004).
Nyström, M., Folke, C. & Moberg, F. Coral reef disturbance and resilience in a human-dominated environment. Trends Ecol. Evol. 15, 413–417 (2000).
Spasojevic, M. J. et al. Scaling up the diversity–resilience relationship with trait databases and remote sensing data: the recovery of productivity after wildfire. Glob. Chang. Biol 22, 1421–1432 (2016).
Biggs, R. et al. Toward principles for enhancing the resilience of ecosystem services. Annu. Rev. Environ. Resour. 37, 421–448 (2012).
Ives, A. R. & Carpenter, S. R. Stability and diversity of ecosystems. Science 317, 58–62 (2007).
Rivera-Chávez, F. et al. Depletion of butyrate-producing Clostridia from the gut microbiota drives an aerobic luminal expansion of Salmonella. Cell Host Microbe 19, 443–454 (2016).
Manichanh, C. et al. Reshaping the gut microbiome with bacterial transplantation and antibiotic intake. Genome Res. 20, 1411–1419 (2010).
Rothschild, D. et al. Environment dominates over host genetics in shaping human gut microbiota. Nature 555, 210–215 (2018).
Wright, G. D. Antibiotic resistance in the environment: a link to the clinic? Curr. Opin. Microbiol. 13, 589–594 (2010).
Surette, M. & Wright, G. D. Lessons from the environmental antibiotic resistome. Annu. Rev. Microbiol. 71, 309–329 (2017).
Wybouw, N., Pauchet, Y., Heckel, D. G. & Van Leeuwen, T. Horizontal gene transfer contributes to the evolution of arthropod herbivory. Genome Biol. Evol 8, 1785–1801 (2016).
Ambrose, K. V., Koppenhöfer, A. M. & Belanger, F. C. Horizontal gene transfer of a bacterial insect toxin gene into the Epichloë fungal symbionts of grasses. Sci. Rep 4, 5562 (2014).
Van Kleunen, M. et al. Global exchange and accumulation of non-native plants. Nature 525, 100–103 (2015).
Stokstad, E. New crop pest takes Africa at lightning speed. Science 356, 473–474 (2017).
Multi-pronged Approach Key for Effectively Defeating Fall Armyworm in Africa (CIMMYT, 2017); https://www.cimmyt.org/press_release/multi-pronged-approach-key-for-effectively-defeating-fall-armyworm-in-africa/
Tay, W. T. et al. Mitochondrial DNA and trade data support multiple origins of Helicoverpa armigera (Lepidoptera, Noctuidae) in Brazil. Sci. Rep 7, 45302 (2017).
Nagoshi, R. N. et al. Comparative molecular analyses of invasive fall armyworm in Togo reveal strong similarities to populations from the eastern United States and the Greater Antilles. PLoS One 12, e0181982 (2017).
Watt, S. Biotype of Australia’s Russian Wheat Aphid Populations Now Known (GRDC, 2017); https://grdc.com.au/news-and-media/news-and-media-releases/south/2017/11/biotype-of-australias-russian-wheat-aphid-populations-now-known
Berrazeg, M. et al. New Dehli metallo-beta-lactamase around the world: an eReview using Google Maps. Euro Surveill 19, 20809 (2014).
Campos, J., Cristino, L., Peixe, L. & Antunes, P. MCR-1 in multidrug-resistant and copper-tolerant clinically relevant Salmonella 1,4,[5],12:i:- and S. Rissen clones in Portugal, 2011 to 2015. Euro Surveill. 21, 30270 (2016).
Munoz-Price, L. S. et al. Clinical epidemiology of the global expansion of Klebsiella pneumoniae carbapenemases. Lancet Infect. Dis. 13, 785–796 (2013).
Porse, A., Schou, T. S., Munck, C., Ellabaan, M. M. H. & Sommer, M. O. A. Biochemical mechanisms determine the functional compatibility of heterologous genes. Nat. Commun. 9, 522 (2018).
Dutil, L. et al. Ceftiofur resistance in Salmonella enterica serovar Heidelberg from chicken meat and humans, Canada. Emerg. Infect. Dis. 16, 48–54 (2010).
Surveillance Bulletin: Reductions in Antimicrobial Use and Resistance: Preliminary Evidence of the Effect of the Canadian Chicken Industry’s Elimination of Use of Antimicrobials of Very High Importance to Human Medicine (CIPARS, 2016).
Overdevest, I. et al. Extended-spectrum β-lactamase genes of Escherichia coli in chicken meat and humans, the Netherlands. Emerg. Infect. Dis. 17, 1216–1222 (2011).
Tamang, M. D. et al. Prevalence and molecular characterization of CTX-M β-lactamase-producing Escherichia coli isolated from healthy swine and cattle. Foodborne Pathog. Dis. 10, 13–20 (2013).
Davis, M. A. et al. Recent emergence of Escherichia coli with cephalosporin resistance conferred by bla CTX-M on Washington State dairy farms. Appl. Environ. Microbiol. 81, 4403–4410 (2015).
Cormier, A. C. et al. Extended-spectrum-cephalosporin resistance genes in Escherichia coli from beef cattle. Antimicrob. Agents Chemother. 60, 1162–1163 (2016).
2015 NARMS Integrated Report (NARMS, 2017).
Bennett, E. M., Peterson, G. D. & Gordon, L. J. Understanding relationships among multiple ecosystem services. Ecol. Lett 12, 1394–1404 (2009).
Raudsepp-Hearne, C., Peterson, G. D. & Bennett, E. M. Ecosystem service bundles for analyzing tradeoffs in diverse landscapes. Proc. Natl Acad. Sci. USA 107, 5242–5247 (2010).
Renard, D., Rhemtulla, J. M. & Bennett, E. M. Historical dynamics in ecosystem service bundles. Proc. Natl Acad. Sci. USA 112, 13411–13416 (2015).
Spake, R. et al. Unpacking ecosystem service bundles: towards predictive mapping of synergies and trade-offs between ecosystem services. Glob. Environ. Chang 47, 37–50 (2017).
Mie, A. et al. Human health implications of organic food and organic agriculture. Environ. Health 16, 88 (2017).
Bahram, M. et al. Structure and function of the global topsoil microbiome. Nature 560, 233–237 (2018).
Lessa, F. C. et al. Burden of Clostridium difficile infection in the United States. N. Engl. J. Med. 372, 825–834 (2015).
Bohnhoff, M., Drake, B. L. & Miller, C. P. Effect of streptomycin on susceptibility of intestinal tract to experimental Salmonella infection. Proc. Soc. Exp. Biol. Med. 86, 132–137 (1954).
Wadolkowski, E. A., Burris, J. A. & O’Brien, A. D. Mouse model for colonization and disease caused by enterohemorrhagic Escherichia coli O157: H7. Infect. Immun. 58, 2438–2445 (1990).
Wadolkowski, E. A., Laux, D. C. & Cohen, P. S. Colonization of the streptomycin-treated mouse large intestine by a human fecal Escherichia coli strain: role of growth in mucus. Infect. Immun. 56, 1030–1035 (1988).
Desneux, N., Decourtye, A. & Delpuech, J.-M. The sublethal effects of pesticides on beneficial arthropods. Annu. Rev. Entomol. 52, 81–106 (2007).
Wilson, L. J., Bauer, L. R. & Lally, D. A. Effect of early season insecticide use on predators and outbreaks of spider mites (Acari: Tetranychidae) in cotton. Bull. Entomol. Res. 88, 477–488 (1998).
Romeis, J., Meissle, M. & Bigler, F. Transgenic crops expressing Bacillus thuringiensis toxins and biological control. Nat. Biotechnol. 24, 63–71 (2006).
Naranjo, S. E. Impacts of Bt transgenic cotton on integrated pest management. J. Agric. Food Chem. 59, 5842–5851 (2010).
Ellsworth, P. C. & Martinez-Carrillo, J. L. IPM for Bemisia tabaci: a case study from North America. Crop Prot. 20, 853–869 (2001).
Gould, F., Kennedy, G. G. & Johnson, M. T. Effects of natural enemies on the rate of herbivore adaptation to resistant host plants. Entomol. Exp. Appl 58, 1–14 (1991).
Carrière, Y. et al. Large-scale, spatially-explicit test of the refuge strategy for delaying insecticide resistance. Proc. Natl Acad. Sci. USA 109, 775–780 (2012).
Liu, X. et al. Natural enemies delay insect resistance to Bt crops. PLoS One 9, e90366 (2014).
Klein, A.-M. et al. Importance of pollinators in changing landscapes for world crops. Proc. R. Soc. Lond. B 274, 303–313 (2007).
Olotu, M. I. & Gujar, G. T. Many-fold less than the field recommended concentrations of neonicotinoids and malathion affect foraging of honeybee in three important crops in India. ENTOMON 41, 47–60 (2016).
Assessment Report on Pollinators, Pollination and Food Production (IPBES, 2016); https://www.ipbes.net/assessment-reports/pollinators
Hokkanen, H. M. T., Menzler-Hokkanen, I. & Keva, M. Long-term yield trends of insect-pollinated crops vary regionally and are linked to neonicotinoid use, landscape complexity, and availability of pollinators. Arthropod. Plant. Interact 11, 449–461 (2017).
Feld, L. et al. Pesticide side effects in an agricultural soil ecosystem as measured by amoA expression quantification and bacterial diversity changes. PLoS One 10, e0126080 (2015).
National Research Council. The Impact of Genetically Engineered Crops on Farm Sustainability in the United States (National Academies Press, Washington DC, 2010).
Schlatter, D. C., Yin, C., Hulbert, S., Burke, I. & Paulitz, T. Impacts of repeated glyphosate use on wheat-associated bacteria are small and depend on glyphosate use history. Appl. Environ. Microbiol. 83, e01354–17 (2017).
Langdon, A., Crook, N. & Dantas, G. The effects of antibiotics on the microbiome throughout development and alternative approaches for therapeutic modulation. Genome Med 8, 39 (2016).
Barzegari, A., Saeedi, N. & Saei, A. A. Shrinkage of the human core microbiome and a proposal for launching microbiome biobanks. Future Microbiol. 9, 639–656 (2014).
Cho, I. & Blaser, M. J. The human microbiome: at the interface of health and disease. Nat. Rev. Genet. 13, 260–270 (2012).
Clemente, J. C., Ursell, L. K., Parfrey, L. W. & Knight, R. The impact of the gut microbiota on human health: an integrative view. Cell 148, 1258–1270 (2012).
Riiser, A. The human microbiome, asthma, and allergy. Allergy, Asthma Clin. Immunol. 11, 35 (2015).
Von Hertzen, L. et al. Helsinki alert of biodiversity and health. Ann. Med. 47, 218–225 (2015).
Martinez, J. L. & Olivares, J. in Antimicrobial Resistance in the Environment (eds Keen, P. L. & Montforts, M. H. M. M.) 151–172 (Wiley, Hoboken, 2012).
Berendonk, T. U. et al. Tackling antibiotic resistance: the environmental framework. Nat. Rev. Microbiol. 13, 310–317 (2015).
Lerner, A., Jeremias, P. & Matthias, T. The world incidence and prevalence of autoimmune diseases is increasing. Int. J. Celiac Dis 3, 151–155 (2015).
Stefka, A. T. et al. Commensal bacteria protect against food allergen sensitization. Proc. Natl Acad. Sci. USA 111, 13145–13150 (2014).
Tito, R. Y. et al. Insights from characterizing extinct human gut microbiomes. PLoS One 7, e51146 (2012).
Zaneveld, J. R., McMinds, R. & Vega Thurber, R. Stress and stability: applying the Anna Karenina principle to animal microbiomes. Nat. Microbiol 2, 17121 (2017).
Steffen, W., Broadgate, W., Deutsch, L., Gaffney, O. & Ludwig, C. The trajectory of the Anthropocene: the Great Acceleration. Anthr. Rev 2, 81–98 (2015).
Rockström, J. et al. Planetary boundaries: exploring the safe operating space for humanity. Ecol. Soc. 14, 32 (2009).
Steffen, W. et al. Planetary boundaries: guiding human development on a changing planet. Science 347, 1259855 (2015).
Shen, Z., Wang, Y., Shen, Y., Shen, J. & Wu, C. Early emergence of mcr-1 in Escherichia coli from food-producing animals. Lancet Infect. Dis. 16, 293 (2016).
Ventola, C. L. The antibiotic resistance crisis: part 1: causes and threats. P T 40, 277–83 (2015).
Hersh, A. L., Newland, J. G., Beekmann, S. E., Polgreen, P. M. & Gilbert, D. N. Unmet medical need in infectious diseases. Clin. Infect. Dis. 54, 1677–1678 (2012).
Eades, C., Hughes, S., Heard, K. & Moore, L. S. Antimicrobial therapies for Gram-positive infections. Pharm. J. 9, 20203363 (2017).
Aksoy, D. Y. & Unal, S. New antimicrobial agents for the treatment of Gram-positive bacterial infections. Clin. Microbiol. Infect. 14, 411–420 (2008).
Johnson, A. P. et al. Mandatory surveillance of methicillin-resistant Staphylococcus aureus (MRSA) bacteraemia in England: the first 10 years. J. Antimicrob. Chemother 67, 802–809 (2012).
Edgeworth, J. D. Has decolonization played a central role in the decline in UK methicillin-resistant Staphylococcus aureus transmission? A focus on evidence from intensive care. J. Antimicrob. Chemother. 66, 41–47 (2011).
Sparks, T. C. & Nauen, R. IRAC: mode of action classification and insecticide resistance management. Pestic. Biochem. Physiol. 121, 122–128 (2015).
Heap, I. & Duke, S. O. Overview of glyphosate‐resistant weeds worldwide. Pest Manag. Sci. 74, 1040–1049 (2017).
Tabashnik, B. E. & Carrière, Y. Surge in insect resistance to transgenic crops and prospects for sustainability. Nat. Biotechnol. 35, 926–935 (2017).
Gual, M. A. & Norgaard, R. B. Bridging ecological and social systems coevolution: a review and proposal. Ecol. Econ. 69, 707–717 (2010).
Rockwood, L. L. et al. (eds) Foundations of Environmental Sustainability: The Coevolution of Science and Policy (Oxford Univ. Press, Oxford, 2009).
Bernhardt, E. S., Rosi, E. J. & Gessner, M. O. Synthetic chemicals as agents of global change. Front. Ecol. Environ. 15, 84–90 (2017).
Pesticides Use Database (FAO, accessed 4 October 2017); http://www.fao.org/faostat/en/#data/RP
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer, New York, 2016) .
Acknowledgements
This is a product of the Living with Resistance project supported by the National Socio-Environmental Synthesis Center (SESYNC) under funding received from the National Science Foundation DBI-1639145 and led by P.S.J. and S.P.C. We thank everyone who contributed to the four project meetings in 2016 and 2017. P.S.J. acknowledges funding from the Carlsberg foundation CF14-1050 and CF15-0988, FORMAS 2016-00451 and the Erling-Persson Family programme.
Author information
Authors and Affiliations
Consortia
Contributions
P.S.J. conceived the manuscript on the basis of four workshops where all authors contributed in person or virtually. P.S.J. wrote the first draft of the manuscript with primary contributions from Y.C., S.D., R.R.D., G.E., G.L., H.M.S. and D.W. P.S.J., Y.C., E.Y.K., D.W. and D.J. performed the assessment of the Anthropocene operating space. P.S.J. and F.K. designed the figures. M.S. and P.S.J. conceived the feedback loops of Fig. 4. All authors commented on and contributed to the writing of the manuscript.
Ethics declarations
Competing interests
DowDuPont and Monsanto did not provide funding to support this work, but may be affected by the publication of this paper and have funded other work by Y.C. All other authors have no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Living with Resistance project. Antibiotic and pesticide susceptibility and the Anthropocene operating space. Nat Sustain 1, 632–641 (2018). https://doi.org/10.1038/s41893-018-0164-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41893-018-0164-3
This article is cited by
-
Effects of environmentally relevant concentrations of oxytetracycline and sulfadiazine on the bacterial communities, antibiotic resistance genes, and functional genes are different between maize rhizosphere and bulk soil
Environmental Science and Pollution Research (2024)
-
Sustainable nano-pesticide platform based on Pyrethrins II for prevention and control Monochamus alternatus
Journal of Nanobiotechnology (2022)
-
Long-term field evaluation and large-scale application of a Metarhizium anisopliae strain for controlling major rice pests
Journal of Pest Science (2021)
-
Resilience: Now more than ever
Ambio (2021)
-
Interference competition and predation between invasive and native herbivores in maize
Journal of Pest Science (2021)