Abstract
Histone methyltransferase SETD2 plays a critical role in maintaining genomic integrity and stability. Here, we investigated the characteristics of SETD2 somatic mutation in the cancer genome atlas pan-cancer cohort. Our data revealed that, compared with SETD2 nonmutant patients, SETD2 mutant patients had higher tumor mutation burden and microsatellite instability. In addition, the transcriptions of most genes related to immune activities were upregulated in patients with SETD2 mutant tumors. Further examination of cancer patients treated with immune checkpoint inhibitors suggested SETD2 mutation was associated with favorable clinical outcomes. These results have implication for the personalization of cancer immunotherapy.
Similar content being viewed by others
Immune checkpoint inhibitors (ICIs) targeting programmed cell death protein-1 (PD-1), programmed cell death ligand 1 (PD-L1), and cytotoxic T-lymphocyte-associated antigen 4 (CTLA-4) can significantly improve the overall survival (OS) in cancer patients1. However, most patients cannot benefit from immunotherapy and reliable biomarkers are warranted2. Although the US Food and Drug Administration (FDA) has approved the application of PD-L1, defective mismatch repair or microsatellite instability high (dMMR/MSI-H), and tumor mutation burden (TMB) in clinical practice, we and others have shown these biomarkers are imperfect2,3.
Histone methyltransferase SETD2, the sole human gene responsible for the trimethylation of histone H3 at lysine 36 (H3K36me3), plays a critical role in maintaining genomic integrity and stability by several distinct pathways4. Pfister et al. found SETD2 was necessary for homologous recombination repair5, depletion of SETD2 shows MSI and an increased spontaneous mutation frequency, characteristic of dMMR cells6. SETD2 also provides an alternative mechanism that leads to DNA damage repair through interacting with p53 tumor suppressor7. Moreover, SETD2 can directly change the chromatin accessibility, which will generate RNA processing defects8. It is estimated that mRNA processing defects occur in 25% of expressed gene across the whole genome when SETD2 is mutant8. We speculate the mutation of SETD2 results in the enrichment of tumor mutation-specific neo-antigens in the cell surface, the immune system will recognize and attack these cells with the help of ICIs. The unique features of SETD2 mutation makes it a potential biomarker for cancer immunotherapy. Accordingly, with accumulated data that are publicly available, here we conducted a comprehensive analysis to examine the characteristics of SETD2 mutation and its association with the efficacy of immunotherapy.
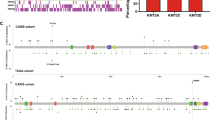
We first examined the prevalence of SETD2 somatic mutations in the cancer genome atlas (TCGA) pan-cancer cohort. Of all 10,427 patients, 451 (4.33%) harbored SETD2 mutations (Fig. 1a). SETD2 mutations occurred in a small subset of most tumor types, and the mutant frequencies differed significantly across various tumors (P < 0.001). Totally, 569 SETD2 mutations were identified, 375 (65.9%) were missense mutations, 193 (33.9%) were truncating mutations, and 1 (0.2%) was inframe mutation (Fig. 1b). These mutations occurred in a dispersed manner throughout the whole sequence (Fig. 1b) and 3D protein structure (Fig. 1c).
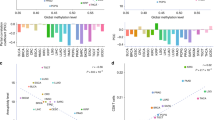
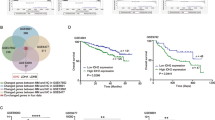
a The prevalence of SETD2 mutations across tumors. b The subtypes and distributions of SETD2 somatic mutations. X-axis, amino acid; Y-axis, numbers of SETD2 mutations; green box, SET domain (1561–1667); red box, WW domain (2391–2420); blue box, SRI domain (2466–2558); green dot, missense mutation; black dot, truncating mutation; orange dot, inframe mutation. c Location of variants on the 3D protein structure of SETD2. Purple, mutated amino acid. d Tumor mutation burden (TMB) in SETD2 nonmutant cancer and different subtypes of SETD2 mutant cancer. Each gray dot represents one patient, black line represents the median TMB and its interquartile ranges. e The prevalence of SETD2 mutation and median TMB in multiple tumors. Red line, fitted curve; HNC, head and neck cancer; LUAD, lung adenocarcinoma; LUSC, lung squamous cell carcinoma; KIRC, kidney renal clear cell carcinoma; KIRP, kidney renal papillary cell carcinoma. f MSIsensor scores in SETD2 nonmutant cancer and different subtypes of SETD2 mutant cancer. g MSI MANTIS scores in SETD2 nonmutant cancer and different subtypes of SETD2 mutant cancer. h The mutant frequencies of MSH2, MSH6, MLH1, and PMS2 in SETD2 mutant and nonmutant cancer. i Overall survival (OS) analysis stratified by SETD2 mutation status in the whole TCGA cohort. j Disease-free survival (DFS) analysis stratified by SETD2 mutation status in TCGA. k Disease-specific survival (DSS) analysis stratified by SETD2 mutation status in TCGA. l Progress-free survival (PFS) analysis stratified by SETD2 mutation status in TCGA. NS, P > 0.05; *P < 0.05; ***P < 0.001.
In TCGA cohort, higher TMB was observed in patients with SETD2 mutant cancer (median, 5.9; interquartile range, 1.8–29.2) than those in patients with SETD2 nonmutant disease (1.5, 0.7–3.3; P < 0.0001). Moreover, TMB was significant different among SETD2 truncating mutant cancer (2.2,1.–10.2), SETD2 missense mutant cancer (9.4, 2.3–39.5), and cancer with multiple mutations (118.1, 21.2–270.7; Fig. 1d). TMB stratified by SETD2 mutation status in different tumors were presented in Supplemental Fig. 1a. In colorectal cancer, further analysis revealed that TMB in non-MSI SETD2 mutant tumors (4.9, 2.8–132.2) was significantly higher than TMB in non-MSI SETD2 nonmutant tumors (2.5, 2.0–3.3; P < 0.0001). Interestingly, we observed a significant correlation between the frequencies of SETD2 mutation and median tumor mutation burdens across multiple tumor types (correlation coefficient, 0.62; P = 0.005; Fig. 1e).
MSIsensor is an effective and efficient tool for deriving MSI status9. MSIsensor scores in patients with SETD2 mutant cancer (0.12; 0.01–0.84) were significantly higher than the scores in patient with SETD2 nonmutant cancer (0.05, 0.00–0.31; P < 0.0001; Fig. 1f). There was no correlation between the frequency of SETD2 mutation and median MSIsensor scores (correlation coefficient, 0.10; P = 0.71). The associations between MSIsensor scores and SETD2 mutation in different tumors were presented in Supplemental Fig. 1b. To further validate the association between SETD2 mutation and MSI status, we also examined the MSI MANTIS10 scores in patients with SETD2 mutant cancer (0.32, 0.30–0.34) and patients with SETD2 nonmutant cancer (0.31, 0.29–0.33; P < 0.0001). Of note, the scores showed no differences among various subtypes of SETD2 mutation (Fig. 1g). MSH2, MSH6, MLH1, and PMS2 played critical roles during the mismatch repair (MMR) process11,12, the mutation in any of these four MMR genes might cause MSI-H. Here, we investigated the co-occurrence patterns of these four MMR mutant genes and SETD2 mutation (Fig. 1h). Compared with patients with SETD2 nonmutant cancer, patients with SETD2 mutant cancer harbored more MMR mutant genes (MSH6, 1.24% vs.14.38%; MSH2, 0.98% vs.11.73%; MLH1, 1.06% vs.8.41%; PMS2, 0.98% vs.9.73%; P < 0.0001 for all four genes).
Next, we investigated the correlations between SETD2 mutation and various immune signatures, including 28 tumor-infiltrating lymphocytes, 24 immunoinhibitors, 45 immunostimulators, 21 major histocompatibility complex molecules, 40 chemokines, and 18 chemokine receptors, in kidney renal clear cell carcinoma (KIRC, n = 43), colon adenocarcinoma (COAD, n = 41), lung adenocarcinoma (LUAD, n = 30), bladder urothelial carcinoma (BLCA, n = 27), and uterine corpus endometrial carcinoma (UCEC, n = 22), five tumors with over 20 SETD2 mutant cases in TCGA cohort (Fig. 2). Compared with SETD2 nonmutant samples, most immune-related genes were upregulated in SETD2 mutant samples, and many showed statistically significant. These results suggested the immune system was more active in SETD2 mutant cancer, which might be recognized as immunologically “hot” tumor. Moreover, our data provided strong evidence that cancer epigenetic driver mutations could shape tumor immune phenotype.
a The correlation between the frequencies of SETD2 mutation and objective response rates across multiple tumors. Red line, fitted curve; HNC, head and neck cancer. b Kaplan–Meier survival analysis stratified by SETD2 mutation status in 2734 cancer patients treated with immune checkpoint inhibitors. CI, confidence interval; HR, hazard ratio. c Multivariate analysis of the association between SETD2 mutation and overall survival.
To investigate whether these distinct characterisitics of SETD2 mutation could translate into cancer prognosis, we compared the OS (P = 0.38, Fig. 1i), disease-free survival (P = 0.53, Fig. 1j), disease-specific survival (P = 0.76, Fig. 1k), and progress-free survival (P = 0.96, Fig. 1l) between patients with SETD2 mutant cancer and patients with SETD2 nonmutant cancer. The prognosis and survival for cancer patients in TCGA cohort were independent of SETD2 mutant status.
Previous studies suggested that copy number alteration (CNA) of SETD2 contributed to the nucleosome stabilization, coordination of DNA repair, and suppression of replication stress13. Hence, we examined the features of cancer patients with CNA of SETD2. Totally, 75 patients (0.68%) with SETD2 CNA were identified in TCGA cohort. The frequencies of the SETD2 CNA acorss different tumors were shown in Supplemental Fig. 2a. The CNA of SETD2 was not associated with TMB (P = 0.73, Supplemental Fig. 2b), MSI MANTIS scores (P = 0.32, Supplemental Fig. 2c), MSIsensor scores (P = 0.71, Supplemental Fig. 2d), and OS (P = 0.63, Supplemental Fig. 2e). It should be noted that, due to the limited number of SETD2 CNA patients included in the TCGA cohort, further investigations are needed to confirm these results.
Our previous study including 22,165 patients treated with PD-1/PD-L1 blockade monotherapy from 160 trials demonstrated the objective response rates (ORRs) in various tumors1. With the frequencies of SETD2 mutations extracted from TCGA, we found that there was a significant correlation between the prevalences of SETD2 mutations and ORRs (correlation coefficient, 0.72; P = 0.001, Fig. 3a).
For survival analysis, a total of 2734 patients from eight studies were included (Table 1). SETD2 mutation was associated with significantly better OS (hazard ratio (HR), 0.55; 95% confidence interval (CI), 0.46–0.65; P < 0.0001; Fig. 3b). This association remained robust after adjusting for confounding factors, including age, sex, cancer type, treatment strategy, and TMB (Fig. 3c), suggesting SETD2 mutation was not a prognostic, but a predictive biomarker for cancer immunotherapy.
Due to the success of POPLAR and OAK, two multicenter randomized controlled trials conducted in patients with non-small cell lung cancer, FDA granted the application of atezolizumab in clinical practice14. Here, we specifically examined the association between SETD2 mutation and various clinicopathological characteristics in patients enrolled in POPLAR and OAK. As shown in Table 2, more PD-L1-positive tumors and higher TMBs were discovered in patients with SETD2 mutant cancer.
In summary, our data reveal that SETD2 mutation is correlated with higher tumor mutation burden and MSI, and more immune activities in cancer. Moreover, SETD2 mutation status is a potential biomarker in predicting the clinical outcomes in patients treated with ICIs.
Methods
Study design
Our study was deemed exempt from institutional board approval and patient informed consent because all data are deidentified and publicly available. The nonsynonymous mutations were defined as frameshift, missense, nonsense, splice site, nonstop, and translation start site changes. Truncating mutations were defined as nonsense, nonstop, frameshift deletion, frameshift insertion, and splice site. Inframe mutations included inframe deletion and inframe insertion.
TCGA data
TCGA database included sequencing and clinicopathological data from patients with over 30 types of tumors. All data included for prevalence analysis of SETD2 mutations and CNA, subtype analysis, 3D protein structure, mutation counts, MSIsensor score, MSI MANTIS score, and survival analysis were queried and downloaded from the cBioPortal for Cancer Genomics database (https://www.cbioportal.org)15. To study the association between SETD2 mutation and immune characteristics, KIRC, COAD, LUAD, BLCA, and UCEC data obtained from TCGA were analyzed using TISIDB (http://cis.hku.hk/TISIDB)16, a database integrated multiple types of data resources in onco-immunology.
Data analysis of patients with immunotherapy
We searched “immune checkpoint blockade clinical trials” across all tumor types on ClinicalTrials.gov for status as completed. The treatment strategies were classified as anti-PD-L1 (avelumab, atezolizumab, and durvalumab), anti-PD-1 (nivolumab, pembrolizumab, and cemiplimab), and anti-CTLA-4 (ipilimumab and tremelimumab), in each tumor type. Then, we conducted systematic search of PubMed database for potential trials in November 2020. Two investigators (M.L. and B.Z.) independently screened the full texts were checked for their eligibility. Any discrepancy was resolved by discussion. The selection criteria were prespecified. To be eligible, studies had to meet the following standards: (1) population: clinical trials including over 30 adult patients with solid tumor; (2) intervention: at least one arm in the trial was treated with ICIs irrespective the dosage and duration of the treatment; and (3) outcomes: reported information regarding SETD2 mutation status and OS. In addition, the reference lists of all trials fulfilling the eligibility criteria were also checked for possible relevant studies. When multiple publications of the same study appeared, only the most recent and/or most complete reporting study were included. We retrospectively collected clinical data of cancer patients samples from three melanoma studies17,18,19, two lung cancer trials20,21,22, one renal cancer datasets23, and two cohorts, including multiple tumors24,25. After removing patients samples without survival information, a total of 2734 patients treated with ICIs were included in this study.
Statistics
Survival analysis was analyzed by Kaplan–Meier method and compared using log-rank test. It was censored at the last date that the patient was not dead. HR was calculated by Cox proportional hazards model and 95% CI was reported. Median OS time and 95% CI were presented where relevant. Spearman’s ρ correlation coefficient was calculated. The relations between various clinical characteristics and SETD2 mutation were evaluated with χ2 test, Student’s t test, or Fisher’s exact test depending on the context. Two-sided P < 0.05 was considered statistically significant. All statistical analysis was conducted by MedCalc 18.2.1 (MedCalc Software, Belgium).
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author upon reasonable request.
Code availability
The code that supports the findings of this study are available from the corresponding author upon reasonable request.
References
Zhao, B., Zhao, H. & Zhao, J. Efficacy of PD-1/PD-L1 blockade monotherapy in clinical trials. Ther. Adv. Med. Oncol. 12, 1758835920937612 (2020).
Shen, X. & Zhao, B. Efficacy of PD-1 or PD-L1 inhibitors and PD-L1 expression status in cancer: meta-analysis. BMJ Clin. Res. Ed. 362, k3529 (2018).
Hegde, P. S. & Chen, D. S. Top 10 challenges in cancer immunotherapy. Immunity 52, 17–35 (2020).
Fahey, C. C. & Davis, I. J. SETting the stage for cancer development: SETD2 and the consequences of lost methylation. Cold Spring Harb. Perspect. Med. 7 (2017).
Pfister, S. X. et al. SETD2-dependent histone H3K36 trimethylation is required for homologous recombination repair and genome stability. Cell Rep. 7, 2006–2018 (2014).
Li, F. et al. The histone mark H3K36me3 regulates human DNA mismatch repair through its interaction with MutSalpha. Cell 153, 590–600 (2013).
Carvalho, S. et al. SETD2 is required for DNA double-strand break repair and activation of the p53-mediated checkpoint. Elife 3, e02482 (2014).
Simon, J. M. et al. Variation in chromatin accessibility in human kidney cancer links H3K36 methyltransferase loss with widespread RNA processing defects. Genome Res. 24, 241–250 (2014).
Niu, B. et al. MSIsensor: microsatellite instability detection using paired tumor-normal sequence data. Bioinformatics 30, 1015–1016 (2014).
Kautto, E. A. et al. Performance evaluation for rapid detection of pan-cancer microsatellite instability with MANTIS. Oncotarget 8, 7452–7463 (2017).
Tamura, K. et al. Genetic and genomic basis of the mismatch repair system involved in Lynch syndrome. Int. J. Clin. Oncol. 24, 999–1011 (2019).
Gelsomino, F., Barbolini, M., Spallanzani, A., Pugliese, G. & Cascinu, S. The evolving role of microsatellite instability in colorectal cancer: a review. Cancer Treat. Rev. 51, 19–26 (2016).
Kanu, N. et al. SETD2 loss-of-function promotes renal cancer branched evolution through replication stress and impaired DNA repair. Oncogene 34, 5699–5708 (2015).
Weinstock, C. et al. U.S. Food and Drug Administration approval summary: atezolizumab for metastatic non-small cell lung cancer. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 23, 4534–4539 (2017).
Gao, J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 6, pl1 (2013).
Ru, B. et al. TISIDB: an integrated repository portal for tumor-immune system interactions. Bioinformatics 35, 4200–4202 (2019).
Van Allen, E. M. et al. Genomic correlates of response to CTLA-4 blockade in metastatic melanoma. Science 350, 207–211 (2015).
Hugo, W. et al. Genomic and transcriptomic features of response to Anti-PD-1 therapy in metastatic melanoma. Cell 165, 35–44 (2016).
Riaz, N. et al. Tumor and microenvironment evolution during immunotherapy with nivolumab. Cell 171, 934–949.e916 (2017).
Fehrenbacher, L. et al. Atezolizumab versus docetaxel for patients with previously treated non-small-cell lung cancer (POPLAR): a multicentre, open-label, phase 2 randomised controlled trial. Lancet 387, 1837–1846 (2016).
Gandara, D. R. et al. Blood-based tumor mutational burden as a predictor of clinical benefit in non-small-cell lung cancer patients treated with atezolizumab. Nat. Med. 24, 1441–1448 (2018).
Rittmeyer, A. et al. Atezolizumab versus docetaxel in patients with previously treated non-small-cell lung cancer (OAK): a phase 3, open-label, multicentre randomised controlled trial. Lancet 389, 255–265 (2017).
Miao, D. & Margolis, C. A. Genomic correlates of response to immune checkpoint therapies in clear cell renal cell carcinoma. Science 359, 801–806 (2018).
Miao, D. et al. Genomic correlates of response to immune checkpoint blockade in microsatellite-stable solid tumors. Nat. Genet. 50, 1271–1281 (2018).
Samstein, R. M. et al. Tumor mutational load predicts survival after immunotherapy across multiple cancer types. Nat. Genet. 51, 202–206 (2019).
Acknowledgements
The authors thank Dr. Chenbin Chen from The Second Affiliated Hospital and Yuying Children’s Hospital of Wenzhou Medical University for statistical analysis and technique support. This work was funded by National Natural Science Foundation of China (Grant No. 31571417) and Natural Science Foundation of Zhejiang Province (Grant No. LZ21H160006).
Author information
Authors and Affiliations
Contributions
M.Lu and B.Z. contributed equally to this work, and should be considered as co-first authors. B.Z. and X.S. conceived and designed the study. M.Lu., B.Z., M.Liu, L.W., Y.L., Y.Z., and X.S. developed the protocol and performed the data analysis. B.Z., M.Lu, and X.S. collected data. B.Z., M.Lu, and X.S. wrote the manuscript. B.Z. and X.S. supervised this work. All of the authors discussed and commented the study. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lu, M., Zhao, B., Liu, M. et al. Pan-cancer analysis of SETD2 mutation and its association with the efficacy of immunotherapy. npj Precis. Onc. 5, 51 (2021). https://doi.org/10.1038/s41698-021-00193-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41698-021-00193-0
This article is cited by
-
RUNX3 pathway signature predicts clinical benefits of immune checkpoint inhibition plus tyrosine kinase inhibition in advanced renal cell carcinoma
BMC Urology (2024)
-
Systematic pan-cancer analysis identifies SLC31A1 as a biomarker in multiple tumor types
BMC Medical Genomics (2023)
-
Prognostic Mutational Signatures of NSCLC Patients treated with chemotherapy, immunotherapy and chemoimmunotherapy
npj Precision Oncology (2023)
-
Multi-omics identification of an immunogenic cell death-related signature for clear cell renal cell carcinoma in the context of 3P medicine and based on a 101-combination machine learning computational framework
EPMA Journal (2023)
-
ANK2 as a novel predictive biomarker for immune checkpoint inhibitors and its correlation with antitumor immunity in lung adenocarcinoma
BMC Pulmonary Medicine (2022)