Abstract
Improved molecular understanding is needed for rational treatment of diffuse intrinsic pontine gliomas (DIPG). Here, using multi-focal paired tumor and germline exome DNA and RNA sequencing, we uncovered phosphatase and tensin homolog (PTEN) loss as a clonal mutation in the case of a 6-year-old boy with a diffuse intrinsic pontine glioma, and incorporated copy number alteration analyses to provide a more detailed understanding of clonal evolution in diffuse intrinsic pontine gliomas. As well, using the PedcBioPortal, we found alterations in PTEN in 16 of 326 (4.9%) cases of pediatric high-grade glioma (3 of 154 (1.9%) brainstem) for which full sequencing data was available. Our data strengthens the association with PTEN loss in diffuse intrinsic pontine gliomas and provides further argument for the inclusion of PTEN in future targeted sequencing panels for pediatric diffuse intrinsic pontine gliomas and for the development and optimization of mTOR/PI3K inhibitors with optimal central nervous system penetration.
Similar content being viewed by others
Introduction
Brainstem tumors comprise 10–15% of central nervous system tumors in children.1, 2 Of these, diffuse intrinsic pontine gliomas (DIPG) are the most common,1 and the most aggressive. Despite improvements in diagnostic accuracy and multimodal treatments, prognosis for patients with DIPG remains dismal with a median survival of around 1 year.3, 4
Improved molecular understanding is needed for rational treatment of DIPG. Pediatric DIPG has been shown to demonstrate intratumoral spatial histologic heterogeneity.5, 6 DIPG appears to be a molecularly homogenous tumor as compared to adult glioblastoma multiforme (GBM).7, 8 Deep sequencing of multiple sites within tumors has revealed key mutations, which are inferred to be driving events as they are conserved throughout all samples in tumors with multi-focal analysis. Activating point mutations (H3K27M) in the genes encoding the histone H3 histone family member 3A (H3F3A) or histone cluster 1 H3 family member B (HIST1H3B) proteins are now understood to be early oncogenic events in DIPG.7,8,9 These mutations are frequently accompanied by driving mutations in genes encoding activin A receptor type 1 (ACVR1, one of the bone morphogenetic protein type I receptors), tumor protein p53 (TP53), or phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha (PIK3CA).6, 10, 11 Here, using multi-focal paired tumor and germline exome DNA and RNA sequencing, we uncovered phosphatase and tensin homolog (PTEN) loss as a clonal mutation in the case of a 6-year-old boy with DIPG, and incorporated copy number alteration analyses to provide a more detailed understanding of clonal evolution in a DIPG.
Case report
A 5-year-old boy presented with repetitive episodes of stumbling and tripping, as well as bilateral ptosis. Magnetic resonance imaging (MRI) of the brain revealed a 4.6 cm, diffusely infiltrating pontine mass. Surgical resection was not recommended due to the location and size of the lesion, and he was treated on a Children’s Oncology Group phase I trial (NCT01922076) with AZD1775 (Wee1 kinase inhibitor) concurrently during radiation. Two months following chemo-radiotherapy, the patient underwent convection-enhanced delivery of 8-H9 antibody (NCT01502917). An interval MRI 1 month later was concerning for progressive disease with recurrence of prior symptoms and development of new motor deficits. He was then treated off-trial for six cycles with the histone deacetylase inhibitor panobinostat, based on pre-clinical data showing its efficacy in H3K27M-mutated DIPG.12 Approximately 13 months following his initial diagnosis, the patient died from progressive disease. Consent for research autopsy was obtained and research autopsy was performed within hours of patient’s death.
The involved brainstem was sectioned into six cross-sections (superior to inferior, Fig. 1). Each cross-section was separated into six quadrants, of which one quadrant (e.g., “A1”) from five of the cross-sections was flash frozen and sent for sequencing. Normal brain samples from cerebellum and frontal lobe were also flash frozen and sequenced as germline samples. Five exome libraries and five poly-A transcriptomes were generated from unique flash frozen tumor tissue and one exome and two poly-A transcriptomes were generated from germline samples (normal cerebellum and frontal lobe). Whole exome (paired tumor and germline DNA) and transcriptome (tumor RNA) sequencing was performed on all samples according to previously published methodology in the Michigan Center for Translational Pathology through PEDS-MIONCOSEQ. The PEDS-MIONCOSEQ study was approved by the University of Michigan Institutional Review Board and adheres to the Clinical Laboratory Improvement Amendments.13
Diffuse intrinsic pontine glioma cross-sections at autopsy and MRI. a Histopathological specimens, showing six equal cross-sections of tumor A–F progressing from superior to inferior sections, with one of six quadrants in each cross-section used for sequencing. b Brain MRI (T1 plus contrast) axial images, progressing from superior to inferior sections, demonstrating primarily unilateral necrosis and corresponding contrast enhancement
Whole exome sequencing revealed 61 somatic mutations distributed throughout the five tumor sites (Fig. 2a), including point mutations in HIST1H3B (p.K27M), ACVR1 (p.G328V), and PTEN (p.Q97X), and copy number loss of PTEN in all five samples. Somatic mutations were additionally found across all five sites in glycoprotein VI (GP6, p.R272W) and adenosine monophosphate deaminase 1 (AMPD1, p.R350C), though these mutations are of uncertain significance. We integrated these and the other (sub-clonal) mutations alongside copy number variants (CNVs) (Supplementary Fig. 1), which significantly informed a representative DIPG tumor evolution map (Fig. 2b). Several genes were found to have corresponding CNV and RNA level changes, including nine genes with copy number gain and overexpression and seven genes with copy number loss and reduced expression, including PTEN and multiple genes of unclear significance in DIPG pathogenesis (Fig. 2c). Of these genes that exhibited changes in copy number and expression, the most notable alterations in expression were overexpression of cyclin-dependent kinase 18 (CDK18), and reduced expression of glutamic-oxaloacetic transaminase 1 (GOT1) and proprotein convertase subtilisin/kexin type 9 (PCSK9). While there is no mention of PCSK9 in the context of oncogenesis in the literature, overexpression of CDK18 has been seen in association with breast cancer and reduced expression of GOT1 has recently been demonstrated to be associated with poorer outcomes in glioblastoma.14,15,16 Histopathologic analysis of contralateral quadrants from each cross-section revealed pleomorphic cells infiltrating through the pons. No microvascular proliferation or tumor necrosis was observed and the histology was consistent with a diffuse infiltrative glioma with focal anaplastic changes. Tumor cells showed some expression of glial fibrillary acidic protein. An immunostain for Ki-67 showed marked heterogeneity from region to region (Supplementary Fig. 2).
Tumor sequencing and copy number analysis reveal clonal events and tumor evolution. a Overall, 61 mutations were noted in the five sequenced tumor samples with five mutations seen in all samples, and previously identified cancer consensus mutations in red (“A1” corresponds to “A1” quadrant in Fig. 1a, etc.) b Tumor evolutionary map displays distance corresponding to degree of mutational and copy number alterations. HIST1H3B, ACVR1, and PTEN are conserved in all five specimens. c Candidate pathway genes with conserved CNVs and corresponding expression changes by RNA sequencing
Discussion
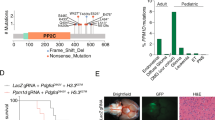
PTEN is an established negative regulator of PI3K signaling and tumor suppressor in many human cancers.17,18,19,20 While previously reported in adult high-grade glioma, it has not been regarded as a main driving lesion in pediatric high-grade glioma or DIPG.10, 21 Interestingly, one study of DIPG specimens obtained at autopsy utilized comparative genomic hybridization to demonstrate hemizygous loss of chromosome 10q, which includes PTEN, in 6 out of 11 samples, as well as one instance of homozygous loss.6 From papers with sequencing data, one found hemizygous deletion of PTEN in 4 out of 48 (8%) DIPG tumors, and no cases with mutation or homozygous loss; another identified a PTEN SNV in 1 of 26 (4%) samples.22 Using the PedcBioPortal, which compiled multiple sequencing datasets of human pediatric high-grade glioma and DIPG10, 22,23,24,25 with additional samples deposited by Dr. Chris Jones (EGAS00001001436), we found alterations in PTEN in 16 of 326 (4.9%) cases for which full sequencing is available (3 of 154 (1.9%) brainstem). As well, we found homozygous deletion of PTEN in 7 of 834 (0.8%) in which DNA copy number was available (3 of 242 (1.2%) brainstem). The PI3K/mechanistic target of rapamycin (mTOR)/AKT/PTEN pathway is an attractive target for children with DIPG showing PTEN loss or PI3K activation. The mTOR protein is downstream of PI3K and in vitro data from multiple tumor types with PIK3CA mutations demonstrates sensitivity to mTOR inhibition with the mTOR inhibitor everolimus, which exhibits good CNS penetration.26, 27 Sapanisertib, an agent that inhibits both mTOR complexes 1 and 2, has been shown to be effective against in vitro and murine models of DIPG.28 Recently agents have been explored to dually target PI3K and mTOR, which have shown efficacy in GBM pre-clinically.29,30,31,32
The prognosis for pediatric DIPG remains bleak despite improved, biopsy-guided multi-modal treatment approaches. It is becoming increasingly evident that a deeper molecular understanding of each individual tumor is needed to guide therapeutic decisions. While our data confirms a relative homogeneity of DIPG in terms of recurrent lesions, our use of copy number analysis and RNA sequencing helps clarify distinct tumor evolution and behavior by region.
In conclusion, we report PTEN loss as an early event in whole exome and transcriptome multi-focal sequencing of a case of DIPG. Our data strengthens its association with DIPG, and provides further argument for the inclusion of PTEN in future targeted sequencing panels for pediatric DIPG and for the development and optimization of mTOR/PI3K inhibitors with optimal CNS penetration.
Code availability
Genomic data will be deposited according to established practices of the Michigan Center for Translational Pathology.
References
Ostrom, Q. T. et al. Alex’s Lemonade stand foundation infant and childhood primary brain and central nervous system tumors diagnosed in the United States in 2007-2011. Neuro. Oncol. 16, x1–x36 (2015).
Vanan, M. I. & Eisenstat, D. D. DIPG in Children - what can we learn from the past? Front. Oncol. 5, 237 (2015).
Gokce-Samar, Z. et al. Pre-radiation chemotherapy improves survival in pediatric diffuse intrinsic pontine gliomas. Child’s Nerv. Syst. 32, 1415–1423 (2016).
Veldhuijzen van Zanten, S. E. et al. A twenty-year review of diagnosing and treating children with diffuse intrinsic pontine glioma in The Netherlands. Exp. Rev. Anticancer Ther. 15, 157–164 (2015).
Buczkowicz, P., Bartels, U., Bouffet, E., Becher, O. & Hawkins, C. Histopathological spectrum of paediatric diffuse intrinsic pontine glioma: diagnostic and therapeutic implications. Acta Neuropathol. 128, 573–581 (2014).
Warren, K. E. et al. Genomic aberrations in pediatric diffuse intrinsic pontine gliomas. Neuro. Oncol. 14, 326–332 (2012).
Nikbakht, H. et al. Spatial and temporal homogeneity of driver mutations in diffuse intrinsic pontine glioma. Nat. Commun. 7, 11185 (2016).
Hoffman, L. M. et al. Spatial genomic heterogeneity in diffuse intrinsic pontine and midline high-grade glioma: implications for diagnostic biopsy and targeted therapeutics. Acta Neuropathol. Commun. 4, 1 (2016).
Castel, D. et al. Histone H3F3A and HIST1H3B K27M mutations define two subgroups of diffuse intrinsic pontine gliomas with different prognosis and phenotypes. Acta Neuropathol. 130, 815–827 (2015).
Wu, G. et al. The genomic landscape of diffuse intrinsic pontine glioma and pediatric non-brainstem high-grade glioma. Nat. Genet. 46, 444–450 (2014).
Puget, S. et al. Biopsy in a series of 130 pediatric diffuse intrinsic Pontine gliomas. Child’s Nerv. Syst. 31, 1773–1780 (2015).
Grasso, C. S. et al. Functionally defined therapeutic targets in diffuse intrinsic pontine glioma. Nat. Med. 21, 555–559 (2015).
Mody, R. J. et al. Integrative clinical sequencing in the management of refractory or relapsed cancer in youth. JAMA 314, 913–925 (2015).
Valladares, A. et al. Genetic expression profiles and chromosomal alterations in sporadic breast cancer in Mexican women. Cancer Genet. Cytogenet. 170, 147–151 (2006).
Barone, G. et al. Human CDK18 promotes replication stress signaling and genome stability. Nucleic Acids Res. 44, 8772–8785 (2016).
Panosyan, E. H., Lin, H. J., Koster, J. & Lasky, J. L. 3rd In search of druggable targets for GBM amino acid metabolism. BMC Cancer 17, 162 (2017).
Wang, Y., Liu, Y., Du, Y., Yin, W. & Lu, J. The predictive role of phosphatase and tensin homolog (PTEN) loss, phosphoinositol-3 (PI3) kinase (PIK3CA) mutation, and PI3K pathway activation in sensitivity to trastuzumab in HER2-positive breast cancer: a meta-analysis. Curr. Med. Res. Opin. 29, 633–642 (2013).
Ren, Y. et al. PTEN activation sensitizes breast cancer to PI3-kinase inhibitor through the beta-catenin signaling pathway. Oncol. Rep. 28, 943–948 (2012).
Chalhoub, N. & Baker, S. J. PTEN and the PI3-kinase pathway in cancer. Annu. Rev. Pathol. 4, 127–150 (2009).
DeGraffenried, L. A. et al. Reduced PTEN expression in breast cancer cells confers susceptibility to inhibitors of the PI3 kinase/Akt pathway. Ann. Oncol. 15, 1510–1516 (2004).
Pollack, I. F. et al. Rarity of PTEN deletions and EGFR amplification in malignant gliomas of childhood: results from the Children’s Cancer Group 945 cohort. J. Neurosurg. 105, 418–424 (2006).
Taylor, K. R. et al. Recurrent activating ACVR1 mutations in diffuse intrinsic pontine glioma. Nat. Genet. 46, 457–461 (2014).
Buczkowicz, P. et al. Genomic analysis of diffuse intrinsic pontine gliomas identifies three molecular subgroups and recurrent activating ACVR1 mutations. Nat. Genet. 46, 451–456 (2014).
Fontebasso, A. M. et al. Recurrent somatic mutations in ACVR1 in pediatric midline high-grade astrocytoma. Nat. Genet. 46, 462–466 (2014).
Wu, G. et al. Somatic histone H3 alterations in pediatric diffuse intrinsic pontine gliomas and non-brainstem glioblastomas. Nat. Genet. 44, 251–253 (2012).
Weigelt, B., Warne, P. H. & Downward, J. PIK3CA mutation, but not PTEN loss of function, determines the sensitivity of breast cancer cells to mTOR inhibitory drugs. Oncogene 30, 3222–3233 (2011).
Li, X. et al. PI3K/Akt/mTOR signaling pathway and targeted therapy for glioblastoma. Oncotarget 7, 33440–33450 (2016).
Miyahara, H. et al. The dual mTOR kinase inhibitor TAK228 inhibits tumorigenicity and enhances radiosensitization in diffuse intrinsic pontine glioma. Cancer Lett. 400, 110–116 (2017).
Matsuda, S., Nakanishi, A., Wada, Y. & Kitagishi, Y. Roles of PI3K/AKT/PTEN pathway as a target for pharmaceutical therapy. Open Med. Chem. J. 7, 23–29 (2013).
Malemud, C. J. The PI3K/Akt/PTEN/mTOR pathway: a fruitful target for inducing cell death in rheumatoid arthritis? Future Med. Chem. 7, 1137–1147 (2015).
Dienstmann, R., Rodon, J., Serra, V. & Tabernero, J. Picking the point of inhibition: a comparative review of PI3K/AKT/mTOR pathway inhibitors. Mol. Cancer Ther. 13, 1021–1031 (2014).
Gil del Alcazar, C. R. et al. Inhibition of DNA double-strand break repair by the dual PI3K/mTOR inhibitor NVP-BEZ235 as a strategy for radiosensitization of glioblastoma. Clin. Cancer Res. 20, 1235–1248 (2014).
Acknowledgements
The authors thank the patient, his family, and the Chad Tough Foundation for their gift to the field and to the future fight against DIPG. Additionally, the authors thank the Michigan Center for Translational Pathology for whole exome and transcriptome tumor sequencing analysis through the PEDS-MIONCOSEQ program. C.K. is supported by the National Institutes of Health/National Institute of Neurological Disorders and Stroke (NIH/NINDS) grant K08-NS099427-01. R.M. is a Hyundai Hope on Wheels Scholar. Funds from the University of Michigan Pediatric Brain Cancer Research Initiative and the UM Chad Tough Fund were used to support this work. The PEDS-MIONCOSEQ study was supported by grant 1UM1HG006508 from the National Institutes of Health Clinical Sequencing Exploratory Research Award (PI: A.C., Co-I: R.M.). Brain tumor research in the Castro–Lowenstein Lab is supported by NIH/NINDS grants R37-NS094804, R01-NS074387, and R21-NS091555 to M.G.C.; NIH/NINDS grants R01-NS061107, R01-NS076991, and R01-NS082311 to P.R.L.; NIH/NINDS grant RO1-EB022563, Leah’s Happy Hearts, University of Michigan Comprehensive Cancer Center, and The Phase One Foundation to M.G.C. and P.R.L.
Author information
Authors and Affiliations
Contributions
Drafting of the manuscript: C.K, Z.F., and S.S. Tumor sequencing: K.K., X.C., A.C., and R.M. Coordination of clinical care and specimen procurement and analysis: D.Z., S.V., S.H.-J., H.G., K.M., L.F., P.L.R., M.L., V.O., M.G.C., and P.R.L.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Koschmann, C., Farooqui, Z., Kasaian, K. et al. Multi-focal sequencing of a diffuse intrinsic pontine glioma establishes PTEN loss as an early event. npj Precision Onc 1, 32 (2017). https://doi.org/10.1038/s41698-017-0033-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41698-017-0033-y
This article is cited by
-
Convergent evolution and multi-wave clonal invasion in H3 K27-altered diffuse midline gliomas treated with a PDGFR inhibitor
Acta Neuropathologica Communications (2022)
-
Pharmaco-proteogenomic profiling of pediatric diffuse midline glioma to inform future treatment strategies
Oncogene (2022)
-
Targeting and Therapeutic Monitoring of H3K27M-Mutant Glioma
Current Oncology Reports (2020)