Abstract
Enchytraeids (Annelida) are soil invertebrates with worldwide distribution that have served as ecotoxicology models for over 20 years. We present the first high-quality reference genome of Enchytraeus crypticus, assembled from a combination of Pacific Bioscience single-molecule real-time and Illumina sequencing platforms as a 525.2 Mbp genome (910 gapless scaffolds and 18,452 genes). We highlight isopenicillin, acquired by horizontal gene transfer and conferring antibiotic function. Significant gene family expansions associated with regeneration (long interspersed nuclear elements), the innate immune system (tripartite motif-containing protein) and response to stress (cytochrome P450) were identified. The ACE (Angiotensin-converting enzyme) — a homolog of ACE2, which is involved in the coronavirus SARS-CoV-2 cell entry — is also present in E. crypticus. There is an obvious potential of using E. crypticus as a model to study interactions between regeneration, the innate immune system and aging-dependent decline.
Similar content being viewed by others
Main
As Charles Darwin pointed out well over 100 years ago, ‘The plough is one of the most ancient and valuable of man’s inventions; but long before he existed the land was in fact regularly ploughed, and still continues to be thus ploughed by earthworms. It may be doubted whether there are many other animals which have played so important a part in the history of the world, as have these lowly organized creatures.’1. Without worms, it is likely that the earth’s soil would not be capable of sustaining the growth of food for humans and other plant-eating species. Annelid worms cover >22,000 species and are found worldwide in all types of habitats. Yet, knowledge of their genome was virtually absent until now. Sequencing big animals (e.g., gorillas2) has a large impact for conservation, but the importance of small species, often invisible at the naked-eye scale, is well known for their role in supporting life itself3,4.
Well-known model species with sequenced genomes include Drosophila melanogaster, Caenorhabditis elegans and Arabidopsis thaliana. However, very few ecotoxicology models have become genome model species, i.e., equipped with genomic-level endpoints in addition to phenotypic endpoints. The genome of Folsomia candida, a standard terrestrial ecotoxicology arthropod collembolan species5, was sequenced6 in 2017. This added on to the first aquatic ecotoxicology Daphnid model7. The species Daphnia pulex was sequenced8 in 2008; however, this is not the commonly tested Daphnia magna species. Recently among soil annelids, the Eisenia andrei genome has been sequenced9; this provides a high-quality assembly for a soil representative model that is also used in ecotoxicology similarly to Eisenia fetida. Other sequenced annelids include Helobdella robusta and Capitella teleta, but these are not ecotoxicology models.
The species sequenced in this study, Enchytraeus crypticus, is a soil invertebrate belonging to the phylum Annelida, class Clitellata, order Oligochaeta and family Enchytraeidae (Fig. 1). Enchytraeids are the most important organisms in many habitats, dominant both in biomass and abundance10 and ranging between 102 and 105 individuals/m2. They belong to the saprophagous mesofauna and play an important role in the degradation of organic matter. Contrary to many larger earthworms, which live in the humus or soil surface, enchytraeids inhabit the actual soil layer. Through their feeding activity, the soil assumes a fine-grained ‘crumb’ structure with an often higher stability than that of the bulk soil11. Enchytraeids are generally obligatory amphimictic hermaphrodites, but some species are able to reproduce by either parthenogenesis or self-fertilization. Most species reproduce sexually by means of egg and sperm production, cross-fertilization and cocoon deposition. E. crypticus can also reproduce via fragmentation: observations confirmed the regenerative ability in the posterior part (tail segments) after artificial amputation, whereas the anterior part was not able to regenerate12. One hypothesis is that autotomy can be used by this species as a self-defense mechanism in response to stress or injuries from physical or chemical stimuli, allowing detoxification and survival. Enchytraeus crypticus are probably diploid, although this has not been confirmed.
E. crypticus are soil invertebrates, belonging to the Oligochaete. Their size ranges from 6 to 9 mm, and they reproduce both sexually and asexually, carrying the cocoons with the embryos in the clitellum and releasing these when matured; they are semi-transparent, and the cocoons and other organelles can be visualized directly (e.g., under a binocular in the culture dishes). a, A photo in a natural habitat assembly. b, A cocoon with embryos. c, A cocoon at post-eggs stage (start of differentiation). d, A cocoon with juveniles. e, Juveniles from a hatched cocoon; f, An adult.
Because of their relevance and sensitivity, enchytraeids are standard models when it comes to evaluating the environmental risk of human-made compounds13 and have been used for >20 years for hazard assessment of chemicals. There are standardized protocols to assess survival and reproduction (ISO (International Standard Organization) and OECD (Organization for Economical Cooperation and Development))14, bioaccumulation15,16,17 and avoidance18 in enchytraeids, as well as vast arrays of other endpoints available.
There are few terrestrial environmental species with such a tool suite covering genotypic to phenotypic endpoints that are also ecotoxicological models. There has been impressive development in terms of molecular tools for E. crypticus, with a full transcriptome19 and suite of omics tools available at present; these include customized microarrays with a wide range of transcriptomics applications20,21,22,23,24,25,26, proteomics27, metabolomics28 and epigenetics29,30, with considerations of big data analysis and progress24. This ecotoxicology model species also includes phenotype-level endpoints for embryotoxicity31, full life cycle with hatching success, growth, maturation, survival, reproduction32,33,34,35, multigeneration36,37, full life span38, species interactions by using multispecies test systems39,40,41,42, histological tools43, oxidative stress biomarkers44,45,46,47 and cellular energy allocation48,49. The possibility of studying embryo development (and all life stages in the full life cycle test) in E. crypticus and its ability to reproduce via regeneration12 also represents some major opportunities. Hence, the progress toward sequencing the genome of this species will provide a major step forward in many related fields (e.g., for evolutionary studies and understanding the mechanisms underlying stress responses).
In this study, we present the first reference genome of E. crypticus, assembled from a combination of long and short reads produced on the Pacific Bioscience single-molecule real-time (SMRT) and Illumina sequencing platforms.
Results
De novo assembly and annotation of the E. crypticus genome
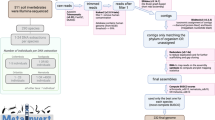
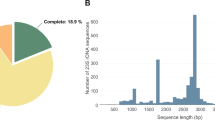
De novo assembly of the E. crypticus genome was done with 1.3 × 109 Illumina paired-end reads, 1.3 × 108 Illumina mate-pair reads and 1.2 × 106 PacBio long reads. These were assembled into 910 gapless scaffolds ≥1,000 nt long, for a total of 525.2 Mbp having an N50 of 1.2 Mbp and an L50 of 118 (see Table 1 for a summary). The largest scaffold had a sequence length of 5.7 Mbp. The GC content of the genome was 35.4%. Genome quality and completeness were checked via a benchmarking universal single copy orthologs (BUSCO) analysis: out of 954 metazoan genes, the method detected 856 (89.7%) complete single-copy orthologs and 41 (4.3%) complete but duplicated orthologs. There were 14 (1.5%) fragmented and 43 (4.5%) missing orthologs. Finally, 97.7% of the Illumina input reads and 80.6% of the PacBio reads mapped back on the genome.
Supported by experimental data (see Methods), the genome was predicted to contain a total of 18,452 gene models, accounting for 24.78% of the genome size and a gene density of 35 genes per mega base pair. We found 16,424 protein-coding genes, of which 82.8% were supported by public transcriptome data. The identified non-coding RNA genes consisted of 295 rRNA genes, 815 tRNA genes and 918 tRNA pseudogenes. A list of the predicted E. crypticus gene models is presented in Supplementary Table 1 (complemented by the genome, found in the Supplementary Data).
Repeated DNA segments comprised 39.03% of the genome and consisted, in decreasing order of abundance, of long interspersed nuclear elements (LINE), unclassified repeats, DNA transposons and simple repeats (Supplementary Table 2).
From the 7,540 genes with gene ontology annotation, the distribution showed a majority of genes involved in molecular functions, followed by biological processes and cellular components (Fig. 2a).
a, GO (Gene Ontology) functional distribution, including the top 50 per biological process, molecular function and cellular component. b, Ultrametric phylogenetic tree based on orthogroup analysis from this study. The indicative age of the root was derived from the TimeTree database divergence time of C. elegans to all members of the clade composed by F. candida, D. pulex and D. melanogaster. c, Comparative analysis via Venn diagram, displaying the number of shared gene families between Annelids (E. crypticus, E. fetida, E. andrei, H. robusta and C. teleta) as determined by OrthoFinder.
Most of the genes are involved in binding and catalytic activity within molecular functions, while for biological processes, metabolic and cellular processes are the most represented, followed by regulation, response to stimulus and signaling. Further detail on each gene ontology (GO) term can be found in Supplementary Fig. 1.
De novo assembly and annotation of the E. crypticus mitochondrial genome
Because the full genome assembly did not contain a scaffold representing an intact mitochondrial genome, a separate assembly was attempted by using the Illumina paired-end reads only and specialized software. The resulting mtDNA of E. crypticus has a length of 15,205 bp. When searching for this sequence in the main genome assembly, two scaffolds containing fragmented copies of the mitochondrial genome were identified and removed from the assembly. Annotation of the mitochondrial genome detected a replication origin, 22 tRNA genes, 2 rRNA genes and 13 protein-coding genes, for a total of 37 genes (see MT (Mitochondrial) scaffold in Supplementary Table 1). The gene order is identical to that reported for Lumbricus terrestris50, with the exception of a non-coding segment located between trnH and nad5 instead of separating trnR from trnH. A map of the annotated mitochondrial genome is available in Supplementary Fig. 2.
Gene family analysis and orthogroups
The comparison between E. crypticus and eight other relevant species assigned 218,791 genes to orthogroups (~85% (>80%)) (Supplementary Table 3). A phylogenetic tree based on the orthogroup analysis is shown in Fig. 2b. The number of shared orthogroups between the four annelid species is represented in the Venn diagram (Fig. 2c). One would expect larger overlap between E. fetida and E. andrei, but E. fetida data are derived from a poor-quality assembly, and hence results can change substantially once quality increases.
The list of significant expansions of gene families in E. crypticus, based on the z-scores, can be found in Supplementary Table 4 (see Supplementary Table 5 for the E. crypticus orthogroup protein description list). A total of 1,751 gene families were shared between E. crypticus and all the other eight species, with 104 being expanded in the E. crypticus genome (when including at least three more species in the comparison). The top 10 largest expansions (Supplementary Table 6) included long interspersed nuclear elements (LINEs) (129 genes), cytochrome P450 (44 genes), tripartite motif (TRIM) (26 genes), ankyrin (ANK; 19 genes), heat shock protein (16 genes), purple acid phosphatase (15 genes), paramyosin (11 genes), vitellogenin (VTG; 10 genes), caspase (10 genes) and hydroxy acid oxidase (HAO) (9 genes), besides several groups with unknown function (e.g., 54 genes in OG0000259). Other gene groups, such as inositol phosphate synthase (21 genes), potassium voltage-gated channel protein Shaw (11 genes) or nAChRbeta1:acetylcholine receptor subunit beta-like (10 genes) also showed high representation and are briefly integrated in the discussion.
With respect to lineage-specific gene families, we counted 307 orthogroups containing 1,370 genes unique for E. crypticus when compared to the eight selected species. An overview of the E. crypticus–specific orthogroups and their gene content can be found in Supplementary Table 7. Zinc fingers, one of the most abundant groups of proteins known for their wide range of molecular functions (transcriptional regulation, ubiquitin-mediated protein degradation, signal transduction, actin targeting, DNA repair, cell migration, etc.)51, were among the most represented. Another example included the sarcoplasmic calcium-binding protein, an invertebrate EF-hand calcium-buffering protein, suggested to have a similar function in muscle relaxation as vertebrate parvalbumin52.
Collinearity analysis
Intragenomic collinearity analysis detected 313 collinear genes in 25 syntenic blocks (see Supplementary Table 8 for a detailed list). Of those, one appears as an intra-scaffold palindrome on scaffold 15 (ANK) and another one as a tandem repeat in scaffold 129 (zinc finger) (Fig. 3).
Scaffolds were renumbered after ordering them by decreasing length for readability; only relevant scaffolds are shown. Gray bars represent scaffolds with no collinearity in the context shown. Scale in million bp. a, Inter-scaffold collinearity. b, Intra-scaffold collinearity; palindromes are drawn with blue links, tandem repeats with red links.
Biosynthetic gene clusters
We used antiSMASH (v5.1.2) to identify biosynthetic gene clusters in the E. crypticus genome. The tool reports only one multi-gene cluster as a chemical hybrid of type I polyketide synthase and non-ribosomal peptide synthetase. The two genes in this cluster were also identified as horizontal gene transfer (HGT) candidates (ECRY_011785-RA, ECRY_011786-RA and malonyl CoA-acyl carrier protein transacylase, from the fatty acid biosynthesis) although not confirmed as HGT.
Hox genes
Based on similarity with Uniprot and HomeoDB, we identified a total 160 homeobox genes in the E. crypticus genome. Of these, 38 are members of the ANTP/HOXL class, which is involved in embryonic development. This number is comparable to that found in the recently assembled high-quality genome of Metaphire vulgaris53, another annelid. Supplementary Fig. 3 shows the distribution of the homeobox genes over the known classes. A complete list of identified hox genes is presented in Supplementary Table 9. Manual assessment of synteny reveals that genes of the ANTP/HOXL class exist as multiple homologs located on several scaffolds. We do, however, notice a cluster of Hox1, Hox3, two Hox5 variants and a Hox7 gene on scaffold scf7180000023640.912933. A smaller cluster consisting of Hox1, Hox5 and Hox7 is present on another scaffold, scf7180000023512.337295. In both cases, the orientation is the same for all genes in the cluster.
HGT
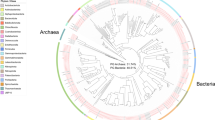
By calculation of h-scores, 105 HGT candidates were initially identified; 33 of them were rejected because of the absence of native neighbor genes and long read linkage. Based on their low metazoan bitscore, five genes were confirmed to have been the result of HGT. The remaining 67 HGT candidates were subjected to a phylogenetic test and resulted in an additional 27 confirmed HGT genes, for a total of 32 genes (Supplementary Table 10). The origin of the confirmed HGT genes is represented in Fig. 4. Bacterial origin is detected for 59.4% of the HGT genes, followed by plants and fungi for 25.0% and 12.5%, respectively, and finally Archaea for 3.1%. A Gene Ontology (GO) term enrichment analysis on the set of horizontally transferred genes yields 14 Biological Process (BP) terms and a single Molecular Function term (Supplementary Table 11).
a, Origin. b, Enriched Gene Ontology. Supplementary Table 11 contains the P values for the GO terms.
Discussion
Genome assembly
In this study, we produced the first high-quality genome for the oligochaete enchytraeid E. crypticus. The presented genome has good contiguity and completeness, as revealed by an N50 of 1.2 Mbp and an overall BUSCO score of 94.0%. The genome sequence, together with the currently over 18,000 identified genes, will allow exploration of the mechanisms underlying interactions with the worms’ environment and its potential toxicants, organ development/regeneration, adaptation and evolutionary aspects.
Genome size
With a 525-Mbp genome, it is interesting to realize the size differences compared to other annelids: larger than the clitellate H. robusta (235 Mbp) and the polychaete C. teleta (333 Mb), but smaller than E. andrei (1,300 Mbp), E. fetida (1,000 Mbp) and Mesenchytraeus solifugus (1,250 Mbp)54. In invertebrates, genome size differences have been found to correlate with, for example, life cycle duration55 and negatively with developmental rate; that is, species with multiple generations per year have smaller genomes (C-values) compared to species with one generation per year.
When comparing the two enchytraeid species, the ice worm M. solifugus and E. crypticus (1,250 versus 525 Mbp), the former’s twofold larger genome size can be due to fast mutational mechanisms or to natural selection. M. solifugus, a small and heavily pigmented enchytraeid, inhabits glacier areas in some of the coldest and highest UV radiation habitats on earth; it also has a much longer life span, living ~10 years, compared to ~1 year for E. crypticus38.
The enchytraeid family has an interesting trait regarding freeze tolerance: an RNAseq study in Enchytraeus albidus showed how the population from Greenland has specific transcriptional differences compared to the German population; both of these strains are freeze tolerant, but the Greenland population is extremely freeze tolerant56. The involved key processes are anion transport in the hemolymph, fatty acid metabolism, metabolism and transport of cryoprotective sugars as well as protection against oxidative stress, with peroxisome and toll-like receptor (TLR) signaling pathways being differentially expressed56. E. crypticus may be a well-adapted species for its life in the deeper soil layer, more buffered from variations compared to the upper layer, where other annelids, such as E. fetida and E. andrei (compost worms) or M. solifugus (ice worm), live. The fact that E. crypticus inhabits a less-variable environment than other worms may have reduced its gene bank source for adaptation (e.g., gene duplication) to cope with a changing environment.
Species-specific evolution and environment contribute to the end result of genome size and gene diversity. For instance, the E. crypticus genome (life span: 12 months; size: 6–9 mm; and genome size: 525 Mb) is twice as large as that for F. candida (life span: 5–8 months; size: 1–5 mm; and genome size: 220 Mb), a terrestrial arthropod, but the latter has ~10,000 more genes. Among other main differences, the arthropod F. candida is a parthenogenetic species, whereas the oligochaete E. crypticus mostly reproduces sexually, besides alternatives like regeneration. For the small crustacean Daphnia pulex (life span: 4–6 months; length: 1–5 mm; and genome size: ~200 Mb), gene duplication seems to be at the core of their evolutionary strategy8. Although there seems to be a trend, a larger number of genomes would be needed to allow such an analysis.
Collinearity
As mentioned, the arthropod F. candida is a parthenogenetic species, whereas the oligochaete E. crypticus mostly reproduces sexually. E. crypticus showed 313 collinear genes in 25 syntenic blocks, much less than the collembolan F. candida with 883 genes in 55 syntenic blocks6. Gene collinearity has been associated with parthenogenic reproduction types; for example, the sexually reproducing collembolan Orchesella cincta does not show this pattern, and the parthenogenetic nematode Meloidogyne incognita has a mitotic cell division reproduction system57. Of the few intra-collinear genes in E. crypticus, zinc finger appears as a tandem repeat in scaffold 129 and is also found among the lineage-specific gene families, adding to its relevance. Zinc fingers, which can have many functions (e.g., binding DNA and RNA and being involved in transcriptional regulation), have also been found in F. candida in a palindrome6. In E. crypticus, scaffold 129 has 60 zinc finger genes in inter-collinearity with scaffold 52, besides the 7 intra-collinear tandem repeats.
Furthermore, ANK genes are present in an intra-scaffold palindrome, together with the protocadherin FAT4 and serine/threonine-protein kinase pak-1 (see the discussion section on ANK).
Expanded gene families and HGT
LINEs
Focusing on the gene family expansions, LINEs are a group of non–long terminal repeat retrotransposons that are widespread in the genome of many eukaryotes. These transposable elements are believed to contribute to the instability and evolution of genomes and are tightly regulated by DNA methylation, histone modifications and PIWI-interacting RNAs (piRNA). LINE2 has significant expansion (7.49%) in the earthworm E. andrei compared to other representative metazoan species: 2.52% in C. teleta, 3.90% in H. robusta, 0.00% in Macrostomum lignano and 0.84% in Apostichopus japonicus; the closely related species E. fetida also has a relatively high LINE2 proportion of ~4.10%9. With 5.04% LINE2, E. crypticus is closest to H. robusta and E. andrei, the top percentage species.
For E. andrei, the LINE2 transposable elements and gene families were functionally related to regeneration (e.g., epidermal growth factor receptor); thus, LINE2 is potentially involved in regulating genes involved in regeneration. E. crypticus is known to regenerate12, although only its posterior end, whereas E. andrei regenerates both. Regeneration has not yet been studied at the genomic level for E. crypticus.
Like regeneration, embryogenesis is a stage of high cell proliferation; this has been studied at the transcriptomic level in E. crypticus embryos when exposed to cadmium (Cd)22. The down-regulation of pms1, a gene coding for a protein involved in DNA mismatch repair, was observed, suggesting that Cd affects DNA synthesis and repair in E. crypticus embryos. Cd also induced the down-regulation of several genes involved in cell cycle/cell division, including cell division cycle proteins and cell division protein kinases. Injured E. fetida showed wound-induced transcriptional activation of early growth response protein 1 gene (EGR1)9; this could also be the case with E. crypticus. The epidermal growth factor receptor is a transmembrane receptor with tyrosine kinase activity that can regulate cell proliferation and differentiation. Hence, some of the mechanisms involved in regeneration also occur during embryogenesis, which is not surprising given the need for cell proliferation and differentiation in both events. Because earthworms are considered of great interest from the perspective of regenerative biology9, this can now be complemented by studies in E. crypticus; that is, the underlying mechanisms for regeneration58 can now also be studied in enchytraeids (E. crypticus), which have a shorter life cycle than E. andrei.
TRIM
Recent studies have revealed that TRIM proteins play key roles in innate antiviral immunity. TRIM, expanded in E. crypticus, is a protein super-family conserved in metazoans that expanded rapidly during vertebrate evolution. There are more members in humans (65) and mice (64) than in worms (~20) and flies (<10). Many TRIM proteins are induced by type I and II interferons, which are crucial for resistance to pathogens, and several are known to be required for the restriction of infection by lentiviruses59.
Type I interferon induction is a central event of the immune response against viral infection, relying on the recognition of pathogens by cellular pattern recognition receptors (PRRs), which then trigger several signaling cascades resulting in pro-inflammatory cytokines and interferon production60. TRIM proteins are essential and act as restriction factors or by modulating PRR signaling. TLRs and other PRRs are engaged by bacterial, viral or fungal components, which triggers the innate immune responses. Although TRIM genes clearly arise from a common ancestral gene, they evolved independently, having acquired species-specific functions59.
Invertebrates are exposed to a wide array of natural and anthropogenic threats with which the immune system has to deal. For instance, M. solifugus tolerate huge amounts of UV radiation compared to many other organisms to endure in the arctic ice and snow. Melanin synthesis, which gives M. solifugus its dark brown color, is known to be a central mechanism of innate immunity and a major response to various immune challenges, including UV. Part of the melanin synthesis pathway is catalyzed by the enzyme phenoloxidase; the phenoloxidase cascade produces melanin and induces multiple potent bioactive agents, such as peroxinectin and Reactive Oxigen Species (ROS), that aid in phagocytosis and cell adhesion. E. crypticus, which has a milky transparent dermis, would have to cope with UV in a different manner.
Other examples include exposure to nanomaterial (NM) contamination, which also activates the innate immune system via different mechanisms20,25,26. NM recognition can first occur upon interaction with surface receptors—typically innate immune PRRs61. As NMs enter a biological environment, they become covered with a corona of proteins, sugars or other compounds. The coronas can mask the NM surface and prevent immune recognition. The importance of protein corona composition for NM recognition was studied in coelomocytes by using coelomic proteins (native repertoire) of the earthworm E. fetida compared to FBS (non-native reference)62. Over time, silver (Ag) NMs can competitively acquire a biological identity native to the cells in situ, although significantly greater cellular accumulation is observed with coelomic protein corona complexes, with lysenin having a key role. On the basis of the genome sequence, we can now look for similarities between E. crypticus and E. fetida, and we find that lysenin is present only in E. fetida. This is a case of species-specific formation of biomolecular coronas and suggests that the use of representative species may need careful consideration in assessing the risks associated with NMs. With our knowledge of the genome, it also means that a protein with similar function in E. crypticus can potentially be identified and possibly linked to the same initiating event.
As mentioned, various organisms may activate similar but not identical mechanisms for the recognition of and response to NMs. For instance, plant PRRs, similar to animal TLR, recognize microbe- or pathogen-associated molecular patterns and trigger defense responses (e.g., ROS production, Mitogen Activated Protein (MAP) kinase activation and induction of defense genes). Worms (e.g., both E. fetida and E. crypticus) have a wide range of genes coding for extracellular recognition proteins (e.g., lectins, peptidoglycan-recognition proteins, lipopolysaccharide- and β1,3-glucan–binding proteins and fibrinogen-related proteins), and any of these are good candidates for similar function identified for lysenin, for example, in enchytraeids.
Research in innate and primed immunity in E. crypticus may open new horizons for developing strategies to prevent or combat infectious diseases, inflammatory conditions and autoimmune disorders. The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is a compelling case of innate immune hyperactivity63, causing acute respiratory distress syndrome. SARS-CoV-2 enters cells through the angiotensin-converting receptor (ACE) 2 (ACE-2) receptor, which is expressed in a small set of alveolar type 2 epithelial cells. The gene coding for ACE is present in the E. crypticus genome, further confirming the potential of this species for immunology studies. Transcriptomic studies showed the activation of the ACE gene in E. crypticus as a response to stress when exposed to TiO2 NMs21 and Ag NMs25. This activation was material specific; for example, the Joint Research Centre (JRC) reference TiO2 NMs NM105, NM104 and NM103 caused an ACE upregulation when exposed under UVB light, whereas exposure to a TiO2-Fe–doped library without UV caused its downregulation. Ag materials caused upregulation of ACE for exposure to AgNM and Ag-PVP coated25. As mentioned, it has been shown that NMs are handled as invaders by cells, like viruses, and can activate similar mechanisms64. A related observation is that SARS-CoV-2 disables interferons—‘strikingly depressed interferon activity and elevated chemokines in individuals whose disease became severe and critical’—hence, a dosage of synthetic interferons to both healthy and infected people might help tame the disease65. Although there is more to coronavirus disease than innate response, E. crypticus may be useful for studying fundamental mechanisms (see the earlier discussion on interferons associated with the TRIM).
Caspases
Another expanded family in E. crypticus includes caspases, a group of cysteine-based proteases that are essential not only during apoptosis but also for the immune system. The role of caspases in cell death has further revealed a caspase-driven compensatory proliferation, apoptosis-induced proliferation66, known to be involved for some forms of regeneration (as discussed above). Several NMs activate the NLRP3 inflammasome, inducing caspase-1 activation and the production of inflammatory IL-1β. Silica particles have been shown to induce caspase-1 activation and pulmonary inflammation.
In a parallel manner, a major known application of NMs is as antimicrobial agents (e.g., Ag and Cu NMs). The microbiota and immune functions are integrally linked; hence, studies should cover the impact on the interaction between bacteria and host immunity. The gut microbiome of E. crypticus has been shown to be altered when exposed to Cu materials67, shifting the communities to a decline in the relative abundance of Planctomycetes and an increase in Bacteroidetes, Firmicutes and Acidobacteria; antibiotic resistance genes in E. crypticus decreased significantly. The fungicide azoxystrobin also altered the abundance of core potential beneficial bacteria and increased the number and abundance of antibiotic resistance genes in the E. crypticus gut68, besides having a severe impact on survival and reproduction69.
Cytochromes P450 (CYPs)
When it comes to stress response, cytochromes P450 (CYPs), a superfamily of enzymes found across most species, are important for hormone biosynthesis and the clearance of various compounds, oxidizing steroids, fatty acids, and xenobiotics. CYPs are expanded in E. crypticus; their expansion in other species such as Mesobuthus martensii (scorpion)70 and F. candida6 has been associated with their survival in hazardous environments and linked to feeding on phytotoxins from herbivorous insects or larva. Hence, they could be an important mechanism of adaptation to an environment (here, soil) where toxic compounds persist and accumulate in decaying soil organic matter; these toxic compounds can include plant anti-herbivory toxins, lignocellulose breakdown products and feeding deterrents.
Heat shock proteins (HSPs)
The expanded HSP family are highly conserved proteins produced by cells in response to stress. For example, the HSP70 group consists of both constitutive and stress-induced HSPs as studied in E. crypticus29. One of the essential roles of HSPs under ‘normal’ conditions is to promote proper embryonic and postnatal development of multiple organ systems, particularly the nervous system71. A study covering embryo development and transcriptomics in response to Cd exposure showed no activation of HSPs in E. crypticus22, except for SSB1; this is in line with the findings72 that loss of SSB1 (combined with SSB2) impairs embryogenesis72. HSP70 induction as a response to stress has been shown in E. crypticus, for example, an increase after multigenerational exposure to copper and a turn-off when transferred to clean media30.
Purple acid phosphatases (PAPs)
PAPs are metalloenzymes that catalyze the hydrolysis of phosphomonoesters and amide substrates. PAPs are highly conserved within eukaryotic species, although varying substantially between plants and animals73. Functional studies indicate that PAPs have flexible mechanisms. They are well known in mammals for their involvement in bone metabolism, and functions include iron transport and generation of ROS as an immune response. For plants, a speculated function is the mobilization/scavenging of inorganic phosphate from organophosphates in the soil. The fact that the PAP family is expanded in E. crypticus could be related to its soil habitat and the need to extract and manage its broad range of metallic elements, as well as the response to stress. PAP seemed to be a good candidate for HGT from plants, although we did not find evidence supporting this in our analysis. For instance, in F. candida there was HGT from the arbuscular mycorrhizal fungus (AMF) Rhizophagus irregularis, which facilitates its grazing by F. candida6. In return, these AMFs benefit from spreading and inoculation to other plants, and plants benefit from phosphorus uptake from AMFs: it is a tritrophic mutualism, contributing to soil health. F. candida was also the first animal discovered with penicillin biosynthesis genes in its genome6,74; the isopenicillin N synthase gene is now also found in the E. crypticus HGT gene list. This suggests that these organisms have evolved to be well adapted in their soil habitat and have been able to develop antibiotic capacity in their microbe-dominated environment.
HAO
HAO (glycolate oxidase) 1 (HAO1) is a protein in the peroxisome encoded by the HAO1 gene in humans. HAO1 belongs to the superfamily of the alpha HAO enzymes. HAO1 catalyzes the flavin mononucleotide–mediated oxidation of glycolate to glyoxylate and glyoxylate to oxalate with reduction of oxygen to hydrogen peroxide; hence, it is central in the toxicity of ethylene glycol poisoning. The gene is primarily expressed in the liver and pancreas. Why HAO is expanded in E. crypticus is not obvious, but it could be for detoxifying functions, because response to stress seems to be prioritized in these organisms.
VTG
This is the major egg yolk precursor protein, which provides protein- and lipid-rich nutrients for developing embryos. The response of VTG to endocrine disruptive chemicals has been well studied in fish, where males can express the VTG gene in a dose-dependent manner. Invertebrates also possess an endocrine system75 and VTG-like proteins, although this is poorly understood. The roles of VTG and its derived yolk proteins lipovitellin and phosvitin include host innate immune defense with various functions76. VTG could play a role in response to stress and innate immunity in E. crypticus, although further studies are needed to clarify this.
Paramyosin
Paramyosin has been found in invertebrate muscles, and it would make sense that their expansion in E. crypticus, a soil worm, relates to the movement and burrowing function requirement for strong muscle contraction. Paramyosin is also a prominent antigen in human cysticercosis and may have a role as a modulator of the host immune response.
Potassium voltage-gated channel protein Shaw
Potassium voltage-gated channel protein Shaw, the Kv3 family, is highly represented in E. crypticus, and these proteins are important in shaping action potentials and in neuronal excitability and plasticity. In animal cells, the K+ channels are involved in neural signaling and generation of the cardiac rhythm, act as effectors in signal transduction pathways involving G protein-coupled receptors and may have a role in targeted cell lysis. Some K+ channels open in response to depolarization of the plasma membrane, others to hyperpolarization or an increase in intracellular calcium concentration; some can be regulated by binding of a transmitter with intracellular kinases or regulated by GTP-binding proteins or other second messengers.
Acetylcholine receptor
The acetylcholine (ACh) receptor (subunit beta-like 1) was well represented in E. crypticus. It binds ACh and responds by a change in conformation, which leads to opening of an ion-conducting channel across the plasma membrane. ACh and γ-aminobutyric acid (also present in E. crypticus) are among the group of neurotransmitters described in vertebrate and invertebrate nervous systems: ACh is a major excitatory transmitter, and GABA is a major inhibitory transmitter, both at the neuromuscular synapses and in the central nervous system. Several pesticides/insecticides (e.g., dimethoate) are designed to target the ACh pathway, and the impacts have been studied in regard to avoidance behavior, a relevant ecological trait for organisms to escape contaminated environments. Studies with E. crypticus showed an association between lack of avoidance behavior because of boric acid and an increase in the γ-aminobutyric acid receptor-associated protein, whereas acetylcholinesterase did not seem to be affected77. Non-avoidance to phenmedipham, however, seems to be associated with acetylcholinesterase inhibition in E. albidus45,78.
Inositol phosphate synthase
The expanded enzyme inositol phosphate synthase, which catalyzes the conversion of D-glucose 6-phosphate into 1D-myo-inositol 3-phosphate, is important for the production of inositol-containing compounds, including phospholipids (important for cell membrane formation and integrity), cell signaling and membrane trafficking. Mechanisms of cold adaptation or acclimation have been associated with changes in the membrane phospholipid composition, gradually undergoing a transition from liquid–crystalline to gel phase. The properties of membranes of E. albidus from seven populations (polar to temperate) have been studied, showing that the composition of phospholipid fatty acids varied significantly but that the ‘optimal’ fluidity of the membrane was apparently kept79. The accumulation of glucose, a cryoprotectant, has been observed, and glucose could possibly have a putative role in the fluidity of membranes.
ANK
ANK, also expanded, are a family of proteins that serve as adaptor proteins linking membrane proteins to the underlying cytoskeleton80. This is required to maintain the integrity of plasma membranes and to anchor specific ion channels, ion exchangers and ion transporters in the plasma membrane. Hence, both inositol phosphate synthase and ANK play a role in membrane integrity; this seems to be an important feature for E. crypticus, given the gene families’ expansion. In addition, ANK is required for the polarized distribution of many membrane proteins, including the Na+/K+ ATPase, the voltage-gated Na+ channel and the Na+/Ca2+ exchanger; hence, this must be an important regulation, because the K+ voltage-gated channel protein Shaw was also observed expanded. The ANK genes may have been transferred from plants and fungi to E. crypticus via HGT (see Supplementary Table 10, from Phytophthora megakarya and Planctomycetes bacterium). As mentioned above, ANK genes are present in an intra-scaffold palindrome, together with the protocadherin FAT4, a calcium-dependent cell adhesion protein playing a role in maintaining cell polarity. Other genes in the palindrome include serine/threonine-protein kinase pak-1 (which has important roles in cytoskeleton dynamics and cell adhesion), Ras-related protein Rab-5C (cell transporter; e.g., vesicular traffic). Hence, this hairpin gene structure with proximity between ANK, FAT4, protein kinases, etc., is not random and must aid in repairing and keeping a key function. One could argue the importance of these ionic stabilizers for their role in the observed plasticity of enchytraeids to survive in aquatic biotopes—many often living in marine interstitial environments, where the level of salts is much higher than terrestrial soil. Studies have shown that the presence of low levels of salinity (15–20‰ NaCl) clearly improves the reproduction of E. albidus81, a species often found at coastal shores among algae and in large abundances. Taking advantage of this asset from a toxicological perspective, an aquatic test has even been developed for enchytraeids82, and exposure to stressors in water is possible during a short period of time83,84,85, allowing researchers to screen effects via an aquatic exposure route.
Challenges and future research applications
From the genome, a potential advance involves the possibility to confirm hypotheses and underlying mechanisms of response to stressors, often a missing link in ecotoxicology. This is feasible by using gene knock-out or gene knock-down (silencing) techniques that have been successfully demonstrated in other invertebrates, including CRISPR–Cas986, transcriptional activator-like effector nucleases87 and RNA interference88. Such development and creation of proof-of-concepts will have direct impact for regulation, for example, in Registration, Evaluation, Authorisation and Restriction of Chemicals for chemicals and developing adverse outcome pathways89, where the causality between transcriptomics and impacts on the phenotype remains one of the main sources of uncertainty for their wider usage and implementation.
Another important future direction from the genome will be the study of the epigenome, representing one of the major regulators of observed effects and its environmental linkage. Although epigenetics has received vast attention for some species, this is not the case for invertebrates, and even less for environmentally relevant species. With the availability of the complete genome of E. crypticus comes the possibility of applying cheaper, more-feasible and/or more-targeted epigenetic genotyping tools, such as Methylated DNA Immunoprecipitation Sequencing (MeDIP-seq), Methyl-CpG-binding domain sequencing (MBD-seq) and epigenetic microarrays. These tools can be used to study how the genome is accessed in different cell types and during development and differentiation. These tools can also provide valuable information on how the organism is reacting on a molecular level to environmental changes. For example, we will be able to learn much more about innate immune memory and priming, information also relevant for humans.
Conclusions
The first high-quality draft genome for E. crypticus was sequenced and assembled, resulting in a 525-Mbp genome, with currently >18,000 identified genes, good contiguity and completeness. Evidence suggests that E. crypticus may be a well-adapted species in its environment, but its genome adaptation and evolution can now be explored. Expanded gene families showed that the genome evolved to respond to stress (CYP) and to develop the innate immune system (TRIM), which are often activated via connected mechanisms. Its capacity to regenerate is a very interesting asset (LINEs), and although it has been found to be inversely related with the evolution of the innate immune system, successful regeneration requires adequate immune response. There is an obvious potential for using E. crypticus as a model to study interactions between regeneration, innate immune system and its aging-dependent decline. Last, the possibility of studying embryo development, a stage of high cell proliferation like regeneration, and the ability to link genes to phenotypic effects represent major advantages of working with E. crypticus. Available transcriptomics studies have linked response to stress to genome features. The potential for future research now includes hypothesis confirmation via gene knock-out and epigenetics.
Methods
Organisms
E. crypticus (Oligochaeta: Enchytraeidae) cultures are kept in the laboratory for many years33. Cultures consist of agar media plates prepared with a salt solution of CaCl2, MgSO4, KCl and NaHCO3, fed ad libitum with oatmeal and maintained in the laboratory under controlled conditions at 18 °C and a photoperiod of 16:8 (light:dark).
Sample preparation and sequencing
E. crypticus were collected from the cultures; six pools (replicates) of 300 adult organisms (with similar size and well-developed clitellum) were rinsed in ultra-pure water and frozen in liquid nitrogen. The samples were stored at −80 °C, until further analysis.
DNA extraction
To obtain high quality and long fragments (150 kbp) of DNA, genomic DNA was extracted from the six pools of E. crypticus adults by using Qiagen Blood & Cell Culture DNA Midi kit (Qiagen). Each sample was diluted in 90 µl of Tris-EDTA (TE) buffer, pH 8.0. The quantity and purity of the isolated DNA were measured spectrophotometrically with a nanodrop (NanoDrop ND-1000 spectrophotometer), and its quality and fragment sizes were analyzed on 1% weight/volume (w/v) agarose gel, by pulse-field gel electrophoresis (CHEF-DR II pulse-field electrophoresis system). After the confirmation of DNA quality, the six samples of E. crypticus were pooled.
Short-read library preparations and sequencing
High-molecular-weight genomic DNA of E. crypticus was quantified by using picogreen (Invitrogen), and its integrity was checked on a 1% (w/v) E-gel (Invitrogen). For the construction of a paired-end library, 2.5 µg of genomic DNA was sheared by using a Covaris S2 sonicator (Covaris), aiming for 500-bp fragments. The fragmented DNA was used to build a PCR-free sequencing library with the NEBNext Ultra II DNA library prep kit for Illumina (New England BioLabs) by using TruSeq adapters. For the construction of a mate-pair library, 1 µg of intact genomic DNA was used with the Nextera mate pair library prep kit (Illumina). Both sequencing libraries were size-selected on a 2% (w/v) agarose E-gel. Fragments ranging from 700 to 800 bp were cut out of the gel and purified with the Zymoclean gel recovery kit (Zymo Research). Library quality was checked on an Agilent Bioanalyzer by using a high-sensitivity chip (Agilent Technologies), and concentration was measured via qPCR according to the qPCR Quantification Protocol (Illumina). Paired-end and mate-pair libraries were pooled with a 75:25 percent ratio and sequenced for 2 × 150 cycles in two lanes on a HiSeq3000 (Illumina).
Long-read sequencing library preparations and sequencing
High-molecular-weight DNA (no shearing required) was purified with 0.6× AMPure PB beads (Beckman Coulter) and eluted in Elution Buffer (EB) buffer. Quantification was done on a Qubit (Thermo Fisher Scientific) by using the ‘Broad Range Kit’. A sequencing library was prepared by using the SMRTbell express template prep kit (PacBio) according to the ‘Procedure & Checklist - Preparing > 15 kb Libraries Using SMRTBell Template Preparation Kit’ (PacBio). Library fragments in the 15–50-kb range were size-selected on a Blue Pippin (Sage Science) by using a 0.75% dye-free agarose gel cassette according to the ‘0.75% DF Marker S1 high-pass 15–20 kb protocol’. Library quality was checked on a Fragment Analyzer (Agilent) by using the ‘High Sensitivity Large Fragment 50 kb Kit’. Final yield was measured on a Qubit by using the ‘High Sensitivity Kit’. The library was sequenced on a PacBio Sequel I (PacBio) by using the V2.1 Binding kit, Chemistry kit and three SMRT cells. The loading type was ‘diffusion’ with an on-plate concentration of 6 pM. There was no pre-extension, and data were collected as a 10-h movie.
De novo genome assembly
For Illumina mate-pair reads, junction adapter removal and read categorization were done by using nxtrim (v0.4.3, Illumina). The resulting true mate-pair reads were used in combination with unmodified Illumina paired-end reads and PacBio long reads for a hybrid de novo genome assembly with MaSuRCA (v3.2.8)90 by using default settings. The quality and integrity of the assembly were assessed with Quast (v4.6.2)91 and BUSCO (v4.0.6)92.
The mitochondrial genome was assembled separately by using the Illumina paired-end reads with NOVOPlasty (v2.7.2)93. Scaffolds in the genome assembly consisting of fragments of the mitochondrial genome were removed.
All sequencing reads and the E. crypticus genome (including the mitochondrial genome) were submitted at the European Nucleotide Archive as study PRJEB41884. The de novo genome assembly is available via accession number GCA_905160935.
Genome annotation
Annotation of the assembled genome was done iteratively by using Maker (v2.31.10)94. Repetitive sequences were masked with RepeatMasker (v4.0.7) and RepeatModeler (v1.0.11)95,96. A first round of annotation with biological evidence was done by using publicly available transcriptome data from E. crypticus (National Center for Biotechnology Information accession GALF01), cDNA data and protein sequences from the related organism E. albidus collected from the National Center for Biotechnology Information (NCBI), as well as all proteins from Uniprot/Swissprot. Next, several ab initio gene predictors were trained and applied in succession to improve the accuracy of identified features on the genome: two training and application rounds of SNAP (v2006–07–28)97 and a single training and application round of Augustus (v3.2.3)98 and GeneMark (v4.38)99. Transfer RNA genes were identified by using trnaScan-SE (v1.3.1)100, and ribosomal RNA genes were detected by using barrnap (v0.9)101. Protein domain information and GO terms were added by using Interproscan (v5.29)102. Finally, a putative function was assigned to the predicted genes by blastp comparison (E-value threshold: 10−6) of the derived protein sequences with the Uniprot/Swissprot database. The mitochondrial genome was annotated by using MITOS (v1.0.1)103. For details on the bioinformatics settings, please see Supplementary Table 12.
Gene family analysis and orthogroups
Identification of homologous protein coding genes between E. crypticus and eight other relevant species was performed by using the protein sequences of the selected species and OrthoFinder (v2.4.0), with default settings to infer orthogroups104. These groups contain all genes descending from a single gene in the last common ancestor of the species whose genes are being analyzed. The eight species (Supplementary Table 13) included in the comparison were selected for being relevant models (C. elegans (Nematoda) and D. melanogaster (Insecta)), ecotoxicology models (F. candida (Collembola)6 and D. pulex (Crustacea)8) and phylogenetically closely related annelids (C. teleta (Polychaete, a marine polichaete), H. robusta (Clitellata, Hirudinida, a freshwater leech), E. andrei (Clitellata) and E. fetida (Clitellata)105). M. solifugus, a glacier enchytraeid species9, could not be included because its genome sequence information is not available106.
To identify significant expansions and contractions of gene families in E. crypticus, the z-score method70 was used. For each gene family (orthogroup), represented by at least three species besides E. crypticus, a z-score was calculated for each species as: (the gene number for the species – the average gene number of the family from all species)/the standard deviation of gene numbers of the family from all species. The families with z-scores ≥2 were considered as significantly expanded, whereas families with a z-score ≤−2 were considered contracted.
A phylogenetic tree was built on the basis of the orthogroup analysis, and the age of the root was derived from the TimeTree database50.
To identify lineage-specific gene families of E. crypticus among this group of eight species, the orthogroups that contain zero genes in all but E. crypticus were filtered out.
Collinearity analysis
Intragenomic collinearity and synteny of the E. crypticus genes were investigated by using MCScanX107.
HGT
HGT analysis was performed as described in ref. 6. In short, all E. crypticus transcripts were blasted against (i) a protein database containing all metazoan proteins (excluding annelida) and (ii) a protein database containing all non-metazoan proteins. All protein sequence databases were downloaded from Uniprot. Then, an h score was calculated as the best non-metazoan blast hit bitscore – the best metazoan blast hit bitscore. A gene with an h score ≥30 and best non-metazoan bitscore ≥100 is considered an HGT candidate. Furthermore, the HGT candidate gene under investigation must have native genes on the same scaffold, and a linkage to neighboring native genes must be confirmed by the presence of mapped PacBio long reads. When linkage and neighbor requirements were fulfilled, candidates were considered confirmed as HGT when the best metazoan bitscore was <50. When linkage was fulfilled but the best metazoan bitscore was ≥50, we performed a phylogenetic validation. To this end, the HGT candidate was blasted against several protein databases: (i) metazoa excluding annelida, (ii) annelida, (iii) plants, (iv) bacteria, (v) archaea and (vi) protists. For each HGT candidate, the top five blast hits in each database were collected and aligned with the HGT by using muscle (v3.8.31). The sequence alignment was trimmed with trimal (v1.4), and a phylogenetic tree was derived by using phyml (v3.3) with the approximate likelihood ratio test (aLRT) method. The trees were inspected, and the HGT candidate was confirmed when the smallest clade containing the HGT candidate did not contain any metazoan sequences.
Hox genes
To identify and assess the organization of Hox genes, we compared the complete HomeoDB108 set of homeodomain proteins to the E. crypticus protein-coding genes via blastp.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
All data are available and will be provided upon request if necessary. The de novo genome assembly is available via accession number GCA_905160935.
References
Darwin, C. The Formation of Vegetable Mould through the Action of Worms: with Observations of Their Habits. (John Murray, 1881).
Gordon, D. et al. Long-read sequence assembly of the gorilla genome. Science 352, aae0344 (2016).
Eisenhauer, N., Bonn, A. & Guerra, C. A. Recognizing the quiet extinction of invertebrates. Nat. Commun. 10, 1–3 (2019).
Wilson, E. O. The little things that run the world (the importance and conservation of invertebrates). Conserv. Biol. 1, 344–346 (1987) http://www.jstor.org/stable/2386020
OECD. Test No.232: Collembolan Reproduction Test in Soil, OECD Guidelines for Testing of Chemicals, Section 2—. Available at https://doi.org/10.1787/9789264264601-en (OECD Publishing, Paris, France, 2016).
Faddeeva-Vakhrusheva, A. et al. Coping with living in the soil: the genome of the parthenogenetic springtail Folsomia candida. BMC Genomics 18, 493 (2017).
OECD. Test No. 211: Daphnia magna Reproduction Test, OECD Guidelines for Testing of Chemicals, Section 2—. Available at https://doi.org/10.1787/9789264185203-en. (OECD Publishing, Paris, France, 2012).
Colbourne, J. K. et al. The ecoresponsive genome of Daphnia pulex. Science 331, 555–561 (2011).
Shao, Y. et al. Genome and single-cell RNA-sequencing of the earthworm Eisenia andrei identifies cellular mechanisms underlying regeneration. Nat. Commun. 11, 1–15 (2020).
Pelosi, C. & Römbke, J. Enchytraeids as bioindicators of land use and management. Appl. Soil Ecol. 123, 775–779 (2018).
Jänsch, S., Amorim, M. J. & Römbke, J. Identification of the ecological requirements of important terrestrial ecotoxicological test species. Env. Rev. 83, 51–83 (2005).
Gonçalves, M. F. M., Gomes, S. I. L., Soares, A. M. V. M. & Amorim, M. J. B. Enchytraeus crypticus (Oligochaeta) is able to regenerate—considerations for a standard ecotoxicological species. Appl. Soil Ecol. 107, 320–323 (2016).
OECD. Test No. 220: Enchytraeid Reproduction Test, Guidelines for the Testing of Chemicals, Section 2. Available at https://doi.org/10.1787/9789264264472-en (OECD Publishing, Paris, France, 2016).
Amorim, M. J. B., Sousa, J. P., Nogueira, A. J. A. & Soares, A. M. V. M. Comparison of chronic toxicity of Lindane (γ-HCH) to Enchytraeus albidus in two soil types: the influence of soil pH. Pedobiologia 43, 635–640 (1999).
OECD. Test No. 317: Bioaccumulation in Terrestrial Oligochaetes, OECD Guidelines for the Testing of Chemicals, Section 3. Available at https://doi.org/10.1787/9789264090934-en (OECD Publishing, Paris, France, 2010).
Amorim, M. J. B., Sousa, J. P., Nogueira, A. J. & Soares, A. M. V. M. Bioaccumulation and elimination of 14C-lindane by Enchytraeus albidus in artificial (OECD) and a natural soil. Chemosphere 49, 323–329 (2002).
Amorim, M. J. B., Sousa, J. P., Nogueira, A. J. A. & Soares, A. M. V. M. Bioavailability and toxicokinetics of 14C-lindane (γ-HCH) in the enchytraeid Enchytraeus albidus in two soil types: the aging effect. Arch. Environ. Contam. Toxicol. 43, 221–228 (2002).
ISO. Soil Quality—Avoidance Test for Testing the Quality of Soils and Effects of Chemicals on Behaviour—Part 1: Test with Earthworms (Eisenia fetida and Eisenia andrei). Available at https://www.iso.org/obp/ui/#iso:std:38402:en (2008).
Castro-Ferreira, M. P. et al. Transcriptome assembly and microarray construction for Enchytraeus crypticus, a model oligochaete to assess stress response mechanisms derived from soil conditions. BMC Genomics 15, 302 (2014).
Gomes, S. I. L. et al. High-throughput tool to discriminate effects of NMs (Cu-NPs, Cu-nanowires, CuNO3, and Cu salt aged): transcriptomics in Enchytraeus crypticus. Nanotoxicology 12, 325–340 (2018).
Gomes, S. I. L., Roca, C. P., von der Kammer, F., Scott-Fordsmand, J. J. & Amorim, M. J. B. Mechanisms of (photo)toxicity of TiO 2 nanomaterials (NM103, NM104, NM105): using high-throughput gene expression in Enchytraeus crypticus. Nanoscale 10, 21960–21970 (2018).
Gomes, S. et al. High-throughput gene expression in soil invertebrate embryos – mechanisms of Cd toxicity in Enchytraeus crypticus. Chemosphere 212, 87–94 (2018).
Gomes, S., Roca, C., Scott-Fordsmand, J. & Amorim, M. Identifying conserved UV exposure genes and mechanisms. Sci. Rep. 8, 8605 (2018).
Roca, C. P., Gomes, S. I. L., Amorim, M. J. B. & Scott-Fordsmand, J. J. A novel normalization approach unveils blind spots in gene expression profiling. Sci. Rep. 7, 42460 (2017).
Gomes, S. I. L., Roca, C. P. P., Scott-Fordsmand, J. J. & Amorim, M. J. B. High-throughput transcriptomics reveals uniquely affected pathways: AgNPs, PVP-coated AgNPs and Ag NM300K case studies. Environ. Sci. Nano 4, 929–937 (2017).
Gomes, S. I. L., Roca, C. P., Scott-Fordsmand, J. J. & Amorim, M. J. B. High-throughput transcriptomics: insights into the pathways involved in (nano) nickel toxicity in a key invertebrate test species. Environ. Pollut. 245, 131–140 (2019).
Maria, V. L., Licha, D., Scott-Fordsmand, J. J., Huber, C. G. & Amorim, M. J. B. The proteome of Enchytraeus crypticus—exposure to CuO nanomaterial and CuCl2—in pursue of a mechanistic interpretation. Proteomics 18, 1–6 (2018).
Maria, V. L. et al. The Enchytraeus crypticus stress metabolome–CuO NM case study. Nanotoxicology 12, 766–780 (2018).
Noordhoek, J. W. et al. Exploring DNA methylation patterns in copper exposed Folsomia candida and Enchytraeus crypticus. Pedobiologia 66, 52–57 (2017).
Bicho, R. C., Roelofs, D., Mariën, J., Scott-Fordsmand, J. J. & Amorim, M. J. B. Epigenetic effects of (nano) materials in environmental species – Cu case study in Enchytraeus crypticus. Environ. Int. 136, 105447 (2020).
Gonçalves, M. F. M. et al. Development of an embryotoxicity test for Enchytraeus crypticus – the effect of Cd. Chemosphere 139, 386–392 (2015).
Santos, F. C. F., Gomes, S. I. L., Scott-Fordsmand, J. J. & Amorim, M. J. B. Hazard assessment of nickel nanoparticles in soil—the use of a full life cycle test with Enchytraeus crypticus. Environ. Toxicol. Chem. 36, 2934–2941 (2017).
Bicho, R. C., Santos, F. C. F., Gonçalves, M. F. M., Soares, A. M. V. M. & Amorim, M. J. B. Enchytraeid Reproduction TestPLUS: hatching, growth and full life cycle test—an optional multi-endpoint test with Enchytraeus crypticus. Ecotoxicology 24, 1053–1063 (2015).
Bicho, R., Santos, F., Scott-Fordsmand, J. & Amorim, M. Effects of copper oxide nanomaterials (CuONMs) are life stage dependent – full life cycle in Enchytraeus crypticus. Environ. Pollut. 224, 117–124 (2017).
Bicho, R. C., Ribeiro, T., Rodrigues, N. P., Scott-Fordsmand, J. J. & Amorim, M. J. B. B. Effects of Ag nanomaterials (NM300K) and Ag salt (AgNO3) can be discriminated in a full life cycle long term test with Enchytraeus crypticus. J. Hazard. Mater. 318, 608–614 (2016).
Bicho, R., Santos, F., Scott-Fordsmand, J. & Amorim, M. Multigenerational effects of copper nanomaterials (CuONMs) are different of those of CuCl2: exposure in the soil invertebrate Enchytraeus crypticus. Sci. Rep. 7, 1–7 (2017).
Ribeiro, M. J., Scott-Fordsmand, J. J. & Amorim, M. J. B. Multigenerational exposure to cobalt (CoCl2) and WCCo nanoparticles in Enchytraeus crypticus. Nanotoxicology 13, 751–760 (2019).
Gonçalves, M. F. M., Gomes, S. I. L., Scott-Fordsmand, J. J. & Amorim, M. J. B. Shorter lifetime of a soil invertebrate species when exposed to copper oxide nanoparticles in a full lifespan exposure test. Sci. Rep. 7, 1–8 (2017).
Mendes, L. A., Amorim, M. J. B. & Scott-Fordsmand, J. J. Interactions of soil species exposed to CuO NMs are different from Cu salt: a multispecies test. Environ. Sci. Technol. 52, 4413–4421 (2018).
Mendes, L. A., Amorim, M. J. B. & Scott-Fordsmand, J. J. Assessing the toxicity of safer by design CuO surface-modifications using terrestrial multispecies assays. Sci. Total Environ. 678, 457–465 (2019).
Menezes-Oliveira, V. B. B., Scott-Fordsmand, J. J., Soares, A. M. V. M. & Amorim, M. J. B. Development of ecosystems to climate change and the interaction with pollution—unpredictable changes in community structures. Appl. Soil Ecol. 75, 24–32 (2014).
Menezes-Oliveira, V. B., Scott-Fordsmand, J. J., Soares, A. M. V. M. & Amorim, M. J. B. Effects of temperature and copper pollution on soil community—extreme temperature events can lead to community extinction: effect of temperature and copper pollution on soil community. Environ. Toxicol. Chem. 32, 2678–2685 (2013).
Bicho, R. C., Faustino, A. M. R., Rêma, A., Scott-Fordsmand, J. J. & Amorim, M. J. B. Confirmatory assays for transient changes of omics in soil invertebrates – copper materials in a multigenerational exposure. J. Hazard. Mater. 402, 123500 (2021).
Howcroft, C. F., Amorim, M. J. B., Gravato, C., Guilhermino, L. & Soares, A. M. V. M. Effects of natural and chemical stressors on Enchytraeus albidus: can oxidative stress parameters be used as fast screening tools for the assessment of different stress impacts in soils? Environ. Int. 35, 318–324 (2009).
Howcroft, C. F. et al. Biochemical characterization of cholinesterases in Enchytraeus albidus and assessment of in vivo and in vitro effects of different soil properties, copper and phenmedipham. Ecotoxicology 20, 119–130 (2011).
Maria, V. L. et al. Silver (nano)materials cause genotoxicity in Enchytraeus crypticus, as determined by the comet assay. Environ. Toxicol. Chem. 37, 184–191 (2017).
Gomes, S. I. L. et al. Effect of Cu-nanoparticles versus one Cu-salt: analysis of stress biomarkers response in Enchytraeus albidus (Oligochaeta). Nanotoxicology 6, 134–143 (2012).
Gomes, S. I. L., Soares, A. M. V. M. & Amorim, M. J. B. Changes in cellular energy allocation in Enchytraeus crypticus exposed to copper and silver—linkage to effects at higher level (reproduction). Environ. Sci. Pollut. Res. Int. 22, 14241–14247 (2015).
Gomes, S., Scott-Fordsmand, J. & Amorim, M. Cellular energy allocation to assess the impact of nanomaterials on soil invertebrates (Enchytraeids): the effect of Cu and Ag. Int. J. Environ. Res. Public Health 12, 6858–6878 (2015).
Kumar, S., Stecher, G., Suleski, M. & Hedges, S. B. TimeTree: a resource for timelines, timetrees, and divergence times. Mol. Biol. Evol 34, 1812–1819 (2017).
Cassandri, M. et al. Zinc-finger proteins in health and disease. Cell Death Discov. 3, 17071 (2017).
White, A. J. et al. Characterization of sarcoplasmic calcium binding protein (SCP) variants from freshwater crayfish Procambarus clarkii. Comp. Biochem. Physiol. B 160, 8–14 (2011).
Jin, F. et al. High-quality genome assembly of Metaphire vulgaris. PeerJ 8, 1–16 (2020).
Iyer, R. G., Valle Rogers, D., Levine, M., Winchell, C. J. & Weisblat, D. A. Reproductive differences among species, and between individuals and cohorts, in the leech genus Helobdella (Lophotrochozoa; Annelida; Clitellata; Hirudinida; Glossiphoniidae), with implications for reproductive resource allocation in hermaphrodites. PLoS One 14, 1–25 (2019).
Gregory T. R. Genome size evolution in animals. In The Evolution of the Genome (ed. Gregory, T. R.) 3–87 (Elsevier, 2005).
de Boer, T. E., Roelofs, D., Vooijs, R., Holmstrup, M. & Amorim, M. J. B. Population-specific transcriptional differences associated with freeze tolerance in a terrestrial worm. Ecol. Evol. 8, 3774–3786 (2018).
Abad, P. et al. Genome sequence of the metazoan plant-parasitic nematode Meloidogyne incognita. Nat. Biotechnol. 26, 909–915 (2008).
Zhao, A., Qin, H. & Fu, X. What determines the regenerative capacity in animals? Bioscience 66, 735–746 (2016).
Long, Ö., Nicole, M. & Badre, D. TRIM family proteins and their emerging roles in innate immunity. Nat. Rev. Immunnol. 8, 849–860 (2008).
García-Sastre, A. & Miorin, L. Intrinsic cellular defenses (TRIMS) in modulating viral infection and immunity. Encycl. Immunobiol. 4, 235–242 (2016).
Boraschi, D. et al. Addressing nanomaterial immunosafety by evaluating innate immunity across living species. Small 16, e2000598 (2020).
Hayashi, Y. et al. Species differences take shape at nanoparticles: protein corona made of the native repertoire assists cellular interaction. Environ. Sci. Technol. 47, 14367–14375 (2013).
Vardhana, S. A. & Wolchok, J. D. The many faces of the anti-COVID immune response. J. Exp. Med. 217, 1–10 (2020).
Hayashi, Y. et al. Earthworms and humans in vitro: characterizing evolutionarily conserved stress and immune responses to silver nanoparticles. Environ. Sci. Technol. 46, 4166–4173 (2012).
Wadman, M. Can interferons stop COVID-19 before it takes hold? Science 369, 125–126 (2020).
Fogarty, C. E. & Bergmann, A. Killers creating new life: caspases drive apoptosis-induced proliferation in tissue repair and disease. Cell Death Differ. 24, 1390–1400 (2017).
Ma, J., Chen, Q. L., O’Connor, P. & Sheng, G. D. Does soil CuO nanoparticles pollution alter the gut microbiota and resistome of Enchytraeus crypticus? Environ. Pollut. 256, 113463 (2020).
Zhang, Q. et al. The fungicide azoxystrobin perturbs the gut microbiota community and enriches antibiotic resistance genes in Enchytraeus crypticus. Environ. Int. 131, 104965 (2019).
Gomes, S. I. L., Ammendola, A., Casini, S. & Amorim, M. J. B. Toxicity of fungicides to terrestrial non-target fauna – formulated products versus active ingredients (azoxystrobin, cyproconazole, prothioconazole, tebuconazole) – a case study with Enchytraeus crypticus (Oligochaeta). Sci Total Env. 754, 142098 (2020).
Cao, Z. et al. The genome of Mesobuthus martensii reveals a unique adaptation model of arthropods. Nat. Commun. 4, 2602 (2013).
Miller, D. J. & Fort, P. E. Heat shock proteins regulatory role in neurodevelopment. Front. Neurosci. 12, 1–15 (2018).
Pfeifer, M. et al. SSB1/SSB2 proteins safeguard B cell development by protecting the genomes of B cell precursors. J. Immunol. 202, 3423–3433 (2019).
Schenk, G., Mitić, N. Š., Hanson, G. R. & Comba, P. Purple acid phosphatase: a journey into the function and mechanism of a colorful enzyme. Coord. Chem. Rev. 257, 473–482 (2013).
Roelofs, D. et al. A functional isopenicillin N synthase in an animal genome. Mol. Biol. Evol. 30, 541–548 (2013).
Oehlmann, J. & Schulte-Oehlmann, U. Endocrine disruption in invertebrates. Pure Appl. Chem. 75, 2207–2218 (2003).
Sun, C. & Zhang, S. Immune-relevant and antioxidant activities of vitellogenin and yolk proteins in fish. Nutrients 7, 8818–8829 (2015).
Bicho, R. C., Gomes, S. I. L., Soares, A. M. V. M. & Amorim, M. J. B. Non-avoidance behaviour in enchytraeids to boric acid is related to the GABAergic mechanism. Environ. Sci. Pollut. Res. 22, 6898–6903 (2014).
Amorim, M. J. B., Novais, S., Römbke, J. & Soares, A. M. V. M. Enchytraeus albidus (Enchytraeidae): a test organism in a standardised avoidance test? Effects of different chemical substances. Environ. Int. 34, 363–371 (2008).
Fisker, K. V. et al. Membrane properties of Enchytraeus albidus originating from contrasting environments: a comparative analysis. J. Comp. Physiol. B 185, 389–400 (2015).
Cunha, S. R. & Mohler, P. J. Ankyrin protein networks in membrane formation and stabilization. J. Cell. Mol. Med. 13, 4364–4376 (2009).
Silva, A. L. P., Amorim, M. J. B. & Holmstrup, M. Salinity changes impact of hazardous chemicals in Enchytraeus albidus. Environ. Toxicol. Chem. 34, 2159–2166 (2015).
Rombke, J. & Kcnacker, T. Aquatic toxicty test for enchytraeids. Hydrobiologia 180, 235–242 (1989).
Gomes, S. I. L., Caputo, G., Pinna, N., Scott-Fordsmand, J. J. & Amorim, M. J. B. Effect of 10 different TiO2 and ZrO2 (nano)materials on the soil invertebrate Enchytraeus crypticus. Environ. Toxicol. Chem. 34, 2409–2416 (2015).
Bicho, R. C., Soares, A. M. V. M., Nogueira, H. I. S. & Amorim, M. J. B. Effects of europium polyoxometalate encapsulated in silica nanoparticles (nanocarriers) in soil invertebrates. J. Nanoparticle Res. 18, 360 (2016).
Rodrigues, N. P., Scott-Fordsmand, J. J. & Amorim, M. J. B. Novel understanding of toxicity in a life cycle perspective – the mechanisms that lead to population effect – the case of Ag (nano)materials. Environ. Pollut. 262, 114277 (2020).
Hiruta, C., Kakui, K., Tollefsen, K. E. & Iguchi, T. Targeted gene disruption by use of CRISPR/Cas9 ribonucleoprotein complexes in the water flea Daphnia pulex. Genes Cells 23, 494–502 (2018).
Bannister, S. et al. TALENs mediate efficient and heritable mutation of endogenous genes in the marine annelid Platynereis dumerilii. Genetics 197, 77–89 (2014).
Zhu, F., Xu, J., Palli, R., Ferguson, J. & Palli, S. R. Ingested RNA interference for managing the populations of the Colorado potato beetle, Leptinotarsa decemlineata. Pest Manag. Sci. 67, 175–182 (2011).
Spinu, N., Cronin, M. T. D., Enoch, S. J., Madden, J. C. & Worth, A. P. Quantitative adverse outcome pathway (qAOP) models for toxicity prediction. Arch. Toxicol. 94, 1497–1510 (2020).
Zimin, A. V. et al. The MaSuRCA genome assembler. Bioinformatics 29, 2669–2677 (2013).
Gurevich, A., Saveliev, V., Vyahhi, N. & Tesler, G. QUAST: quality assessment tool for genome assemblies. Bioinformatics 29, 1072–1075 (2013).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Dierckxsens, N., Mardulyn, P. & Smits, G. NOVOPlasty: de novo assembly of organelle genomes from whole genome data. Nucleic Acids Res. 45, e18 (2016).
Cantarel, B. L. et al. MAKER: an easy-to-use annotation pipeline designed for emerging model organism genomes. Genome Res. 18, 188–196 (2008).
Smit, A. & Hubley, R. RepeatModeler Open-1.0. Available at https://www.repeatmasker.org/RepeatModeler/ (2008).
Smit, A., Hubley, R. & Green, P. RepeatMasker Open-4.0. Available at https://www.repeatmasker.org/ (2013).
Korf, I. Gene finding in novel genomes. BMC Bioinformatics 5, 59 (2004).
Stanke, M. et al. AUGUSTUS: ab initio prediction of alternative transcripts. Nucleic Acids Res. 34, W435–W439 (2006).
Lomsadze, A., Ter-Hovhannisyan, V., Chernoff, Y. O. & Borodovsky, M. Gene identification in novel eukaryotic genomes by self-training algorithm. Nucleic Acids Res. 33, 6494–6506 (2005).
Lowe, T. M. & Eddy, S. R. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 25, 955–964 (1997).
Seemann, T. Barrnap 0.9: Rapid Ribosomal RNA Prediction. Available at https://github.com/tseemann/barrnap (2013).
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics 30, 1236–1240 (2014).
Bernt, M. et al. MITOS: improved de novo metazoan mitochondrial genome annotation. Mol. Phylogenet. Evol. 69, 313–319 (2013).
Emms, D. M. & Kelly, S. OrthoFinder: solving fundamental biases in whole genome comparisons dramatically improves orthogroup inference accuracy. Genome Biol. 16, 1–14 (2015).
Zwarycz, A. S., Nossa, C. W., Putnam, N. H. & Ryan, J. F. Timing and scope of genomic expansion within Annelida: evidence from homeoboxes in the genome of the earthworm Eisenia fetida. Genome Biol. Evol. 8, 271–281 (2015).
Pacific Biosciences - PacBio. Webinar: Small Bodies, Big Genomes: Overcoming Large DNA Input Requirements for Long-Read Genome Assembly. Available at https://youtu.be/4-BGuflprw0 (2019).
Wang, Y. et al. MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 40, e49 (2012).
Zhong, Y.-F., Butts, T. & Holland, P. W. H. HomeoDB: a database of homeobox gene diversity. Evol. Dev. 10, 516–518 (2008).
Acknowledgements
This study received funding from the European Commission within NanoInformaTIX (H2020-NMBP-14-2018, No. 814426), BIORIMA (H2020-NMBP-12-2017 GA No. 760928) and NANORIGO (H2020-NMBP-13-2018, GA No. 814530) projects, FCT (Fundação para a Ciência e Tecnologia)/MCTES (Ministério da Ciência e Tecnologia do Ensino Superior) for financial support of project BEAUTY: Big gEnome wide Applications for an ecotoxicology soil model – a knowledge base to Unravel mechanisms (nanopesTY_cides) (PTDC/CTA-AMB/3970/2020) and CESAM (UIDP/50017/2020+UIDB/50017/2020). S.I.L.G. is funded by national funds (OE), through FCT – Fundação para a Ciência e a Tecnologia, I.P., in the scope of the framework contract foreseen in the numbers 4, 5 and 6 of the article 23, of the Decree-Law 57/2016, of August 29, changed by Law 57/2017, of July 19.
Author information
Authors and Affiliations
Contributions
M.J.B.A. was responsible for conceptualization, data curation, resources and writing, including the original draft, review and editing. Y.G. performed genome analysis, data curation and writing, including review and editing. S.I.L.G. was responsible for investigation, methodology, review and editing. F.V.N. was involved in conceptualization and writing, including review and editing. J.J.S.-F was involved in conceptualization, data curation, resources and writing, including review and editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Lab Animal thanks Steven Siciliano, Avril Coghlan and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–3
Supplementary Tables 1–13
Excel workbook with 13 sheets of supplemental tables
Supplementary Data
Genome file
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Amorim, M.J.B., Gansemans, Y., Gomes, S.I.L. et al. Annelid genomes: Enchytraeus crypticus, a soil model for the innate (and primed) immune system. Lab Anim 50, 285–294 (2021). https://doi.org/10.1038/s41684-021-00831-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41684-021-00831-x
This article is cited by
-
Fipronil Risk Assessment in Tropical Soils: Relevant Concentrations Cause High Risk to the Edaphic Community
Water, Air, & Soil Pollution (2024)
-
Automated detection and quantification of Enchytraeus crypticus (Oligochaeta: Enchytraeidae) in tropical artificial soil using image analysis
Environmental Monitoring and Assessment (2022)
-
Turning fifty
Lab Animal (2021)