Abstract
To better clarify the causal effects between matrix metalloproteinases (MMPs) and estrogen-receptor (ER)-negative breast cancer (BC), we investigated the bidirectional causal relationship between MMPs and ER-negative BC by mendelian randomization (MR) analysis. Summary statistic data of five MMPs were extracted from European participants in 13 cohorts. Data of ER-negative BC collected from one of genome-wide association studies of European ancestry was used as experimental datasets and another four ER-negative BC datasets were used as validation sets. Inverse variance weighted method was used for main MR analysis and sensitivity analysis was also conducted. Serum level of MMP-1 has negative effect on ER-negative BC (odds ratio = 0.92, P = 0.0008) but the latter one was not the cause of the former one, which was supported by validation sets. No bidirectional causal effect was detected between the other four types of MMPs and ER-negative BC (P > 0.05). Sensitivity analysis indicated robustness of the above results without remarkable bias. To conclude, serum MMP-1 may be a protective factor against ER-negative BC. No reciprocal causality was found between the other kinds of MMPs and ER-negative BC. MMP-1 was indicated as a biomarker for risk of ER-negative BC.
Similar content being viewed by others
Introduction
According to the latest version of global cancer statistic GLOBOCAN published in 2020, female breast cancer (BC) was the most common solid malignancy, with approximately 2.3 million newly diagnosed cases (11.7%). It took the fourth place (6.9%) regarding cause of cancer-specific death of female patients globally1. Conventionally, BC includes four main subtypes. Luminal A, Luminal B, human epidermal growth factor receptor 2 (HER-2) positive, and triple-negative BC (TNBC)2. The classification is mainly dependent on status of estrogen receptor (ER) and HER-2, indicating different clinical managements to different subtypes of BC3. ER is an essential predictor for response of endocrine therapy and prognosis of BC patients. Regarding ER-positive BC, endocrine therapy could largely reduce recurrence and mortality rate4. Patients with ER-negative BC has a relatively more aggressive biological trait and a worse prognosis than ER-positive BC after endocrine therapy5. HER-2 positive BC and TNBC are two special types of ER-negative BC. The former one accounts for around 15% of BC patients and possesses aggressive clinical features and results in a poor prognosis until the appearance of anti-HER-2 monoclonal antibody (trastuzumab and pertuzumab, etc.), which lifts response rate and improvs survival6,7,8,9. Resistance and recurrence, however, usually occur in HER-2 positive tumor, especially for advanced and metastatic one10,11. Different from the other three subtypes, TNBC lacks of expression of neither ER or HER-2, rendering it the most aggressive and refractory BC subtype especially in younger patients12,13. Commonly-used endocrine treatment (tamoxifen and aromatase inhibitors) and targeting anti-HER-2 therapy trastuzumab are ineffective in patients with TNBC14. Thus, it is indispensable to find some other targets to improve the treatment response of ER-negative BC.
Matrix metalloproteinases (MMPs) are a family of zinc-dependent endopeptidases, a subgroup of metzincin superfamily. In 1962, MMPs were first reported by Charles Lapiere and Jerome Gross in tadpole undergoing metamorphosis15,16. Totally, MMP include 26 zinc-dependent endopeptidases, among which 23 genes of MMP have been identified in homo sapiens16,17. Based on molecular structure and substrate specificity, MMPs in vertebrates can be classified in 6 groups: collagenases (MMP-1, -8, -13, and -18), gelatinases (MMP-2 and -9), matrilysins (MMP-7 and -26), stromelysins (MMP-3, -10, and -11), and transmembrane MMPs (MT-MMPs, MMP-14, -15, -16, -17, -24, and -25), and other unspecific types (MMP-12, -19, -20, -21, -22, -23, -27, and -28)16. Functionally, MMPs are involved in both physiological (degradation of extracellular matrix, embryonic growth, reproduction, etc.)18 and pathological process (aneurysms, atherosclerosis, arthritis, fibrosis, nephritis, tissue ulcers, and cancer)19,20. In tumor, specifically, MMPs degraded proteins in basement membrane and extracellular matrix, eliminating barriers against cancer cell invasion, facilitating the process of cancer progression and metastasis21. Previous studies have found relationships between single nucleotide polymorphisms (SNP) of MMPs genes and solid malignancies including lung cancer, esophageal cancer, head and neck cancer, colorectal cancer, and BC22,23,24.
Previous systemic review and meta-analysis suggested that several types of MMPs were associated with BC25. Huang study found that SNP of MMP-9 rs3918242 was remarkably relevant with incidence of BC among the overall population and Asian population25, which was supported by Xu study, indicating that one of MMP-9 polymorphisms, rs3918242, may be a risk factor of BC26. Another meta-analysis published by Han et.al corroborated this conclusion, indicating that MMP-9–1562 C/T polymorphism was a risk factor of BC, especially in European population whereas MMP2 polymorphism MMP-2–1306 C/T polymorphism was a protecting factor for BC in Asian population27. In a study from Ou et.al, however, no correlation was found between MMP-2–1306 C/T and risk of BC28. Ren and Song study demonstrated that MMP-9 overexpression was associated with a poorer overall survival and Ren study did not find significant impact of MMP-2 on prognosis of BC patients29,30. Different from the above research, Chen er.al reported that MMP-2 expression was significantly associated with a poor survival and risk of lymph node metastasis31. For SNP of MMP-1, the results were also controversial. Sui study showed that SNP of MMP-1 rs1799750 was related to a reduction of risk of BC in both the overall population and Asian group32. On the contrary, MMP-1 1G/2G genotype and MMP-1 2G/2G genotype were significantly associated with metastasis of BC33.
The above studies not only failed to draw consistent conclusion, but also did not confirm a causal relationship between MMPs and risk of BC in that all meta-analysis were based on observational (case–control) studies. Moreover, few study focused on the association of MMPs specifically with ER-negative BC. The traditional epidemiological approach is vulnerable to confounders and reverse causality, causing conflicting evidence34,35. To neutralize the adverse effect of confounders and reverse causality, we used the Mendelian Randomization (MR) method36. MR uses genetic variants (SNP) strongly associated with certain type of exposure as instrumental variables (IVs) to predict causality of the given exposure on an outcome of interest37,38,39. Genetic variants are randomly assorted, hardly modified during meiosis and they are also unrelated with other confounders (socioeconomical and environmental factors). Thus, different outcomes between populations with and without these IVs can be attributed to this exposure39. The results of MR estimation are reliable to reflect life-long exposure and reduce the impact of confounding factors and reverse causation39,40,41,42. As an extension of basic MR analysis, bidirectional MR can further validate whether two phenotypes can interact as reciprocal causality43. Herein, we aimed to explore the bidirectional causal relationship between several types of MMPs and ER-negative BC via two-sample MR analysis to find new targets for BC treatment.
Results
Causal effect of serum MMP levels on ER-negative BC
Ten types of MMPs (MMP-1, MMP-2, MMP-3, MMP-7, MMP-9, MMP-10, MMP-12, MMP-14, MMP-16, MMP-17) were available in the open GWAS summary datasets. After excluded MMPs with less than five significant associated SNPs (P < 5 × 10−8), five SNPs (MMP-1, MMP-3, MMP-7, MMP-10, MMP-12) were selected for following analysis. After extracting IVs of the five MMPs, we input these IVs into Phenoscanner database to remove SNPs associated with confounding factors. No confounder-related SNPs had been found for MMP-1, MMP-7, and MMP-12 whereas each of two SNPs (rs17360661, rs2267373 for MMP-3; rs3129886 and rs601338 for MMP-10) were found to be associated with BCs (cause of death; body weight; long-standing illness; alcohol intake). The number of ultimately selected LD-independent SNPs for data harmonization were 17, seven, nine, nine, and 15 for MMP-1, MMP-3, MMP-7, MMP-10, and MMP-12, respectively. F-statistic for SNPs of all the five MMPs were greater than the threshold of 10, suggesting strong IVs, which reducing bias of IVs estimates (Supplementary Tables S1, S2, S3, S4, S5). During harmonization, two, three, and one palindromic SNP(s) significantly associated with MMP-1, MMP-3, and MMP-12 were removed for following MR analysis. No palindromic SNPs were removed for MMP-7 and MMP-10.
The first dataset of BC (GWAS ID: ieu-a-1128) was used as experimental set to explore the causal effect of MMPs on ER-negative BC. Genetically elevated serum MMP-1 level were causally associated with a low risk of ER-negative BC (OR = 0.92, 95% confidence interval [CI]: 0.88–0.97, P = 0.0008), which was validated in the other three datasets as suggestive associations (ieu-a-1135: OR = 0.93, 95% CI: 0.87–0.99, P = 0.03; ieu-a-1136: OR = 0.92, 95% CI: 0.86–1.00, P = 0.049; ieu-a-1166: OR = 0.92, 95% CI: 0.85–1.00, P = 0.047) (Table 1). Genetically elevated serum MMP-3 level were not causally associated with risk of ER-negative BC (OR = 1.01, 95%CI: 0.93–1.09, P = 0.88), which was supported by the results of the other four datasets (ieu-a-1135: OR = 0.99, 95%CI: 0.91–1.08; P = 0.88; ieu-a-1136: OR = 0.99, 95% CI: 0.86–1.14, P = 0.884; ieu-a-1137: OR = 1.08, 95%CI: 0.97–1.20, P = 0.17; ieu-a-1166: OR = 1.00, 95%CI: 0.89–1.12, P = 1.00)(Table 2). Genetically elevated serum MMP-7 level were not causally associated with risk of ER-negative BC (OR = 1.05, 95%CI: 0.97–1.14, P = 0.24), which was supported by the results of the other four datasets (ieu-a-1135: OR = 0.98, 95%CI: 0.89–1.07; P = 0.60; ieu-a-1136: OR = 1.10, 95% CI: 0.96–1.26, P = 0.19; ieu-a-1137: OR = 1.22, 95%CI: 0.99–1.50, P = 0.06; ieu-a-1166: OR = 1.08, 95%CI: 0.94–1.25, P = 0.28) (Table 3). Genetically elevated serum MMP-10 level were not causally associated with risk of ER-negative BC (OR = 1.00, 95%CI: 0.95–1.06, P = 0.86), which was supported by the results of the other four datasets (ieu-a-1135: OR = 0.99, 95%CI: 0.93–1.06; P = 0.86; ieu-a-1136: OR = 1.00, 95% CI: 0.89–1.13, P = 1.00; ieu-a-1137: OR = 1.05, 95%CI: 0.93–1.19, P = 0.43; ieu-a-1166: OR = 1.00, 95%CI: 0.89–1.13, P = 0.98) (Table 4). Genetically elevated serum MMP-12 level were not causally associated with risk of ER-negative BC (OR = 1.02, 95%CI: 0.96–1.07, P = 0.56), which was supported by the results of the other three datasets (ieu-a-1136: OR = 0.98, 95% CI: 0.91–1.05, P = 0.59; ieu-a-1137: OR = 0.95, 95%CI: 0.85–1.06, P = 0.35; ieu-a-1166: OR = 0.99, 95%CI: 0.92–1.07, P = 0.86) except GWAS ieu-a-1135 (OR = 1.07, 95%CI: 1.00–1.13, P = 0.04) (Table 5). All the results above were calculated by IVW method. MR-Egger analysis did not suggest any directional pleiotropy for the IVs of all types of MMPs (P for intercept > 0.1 in both experimental and validation datasets). MR-PRESSO global test did not detect any outliers and pleiotropy, either. For heterogeneity analysis, Cochran's Q test did not detect the heterogeneity in MMP-1 (P > 0.10), MMP-7 (P > 0.05), MMP-10 (P > 0.10), and MMP-12 (P > 0.10) whereas data of MMP-3 (P < 0.01) were significantly heterogenous. Both of the result of MR-Egger and IVW method were consistent in heterogeneity analysis.
The results of leave-one-out sensitivity analysis showed that no SNPs with potential effect on the pooled result in analysis of experimental datasets Figs. 1, 2, 3, 4, 5. Scatter plots and funnel plots for analysis of MMP and BC in both experimental and validation sets are presented in Supplementary Figure (Figs. S1, S2, S3, S4, S5 for scatter plots, S6–S10 for forest plots, and S11–S15 for funnel plots).
Leave-one-out plots for analysis of causal effect of MMP-1 on ER-negative BC. (a) Associations between MMP-1 and ER-negative BC (experimental set: ieu-a-1128); (b) Associations between MMP-1 and ER-negative BC (Validation set 1: ieu-a-1135); (c) Associations between MMP-1 and ER-negative BC (Validation set 2: ieu-a-1136); (d) Associations between MMP-1 and ER-negative BC (Validation set 3: ieu-a-1137); (e) Associations between MMP-1 and ER-negative BC (Validation set 4: ieu-a-1166). MMP, matrix metalloproteinases; ER-negative BC, estrogen receptor-negative breast cancer.
Leave-one-out plots for analysis of causal effect of MMP-3 on ER-negative BC.(a) Associations between MMP-3 and ER-negative BC (experimental set: ieu-a-1128); (b) Associations between MMP-3 and ER-negative BC (Validation set 1: ieu-a-1135); (c) Associations between MMP-3 and ER-negative BC (Validation set 3: ieu-a-1136); (d) Associations between MMP-3 and ER-negative BC (Validation set 3: ieu-a-1137); (e) Associations between MMP-3 and ER-negative BC (Validation set 4: ieu-a-1166), MMP, matrix metalloproteinases; ER-negative BC, estrogen receptor-negative breast cancer.
Leave-one-out plots for analysis of causal effect of MMP-7 on ER-negative BC. (a) Associations between MMP-7 and ER-negative BC (experimental set: ieu-a-1128); (b) Associations between MMP-7 and ER-negative BC (Validation set 1: ieu-a-1135); (c) Associations between MMP-7 and ER-negative BC (Validation set 3: ieu-a-1136); (d) Associations between MMP-7 and ER-negative BC (Validation set 3: ieu-a-1137); (e) Associations between MMP-7 and ER-negative BC (Validation set 4: ieu-a-1166), MMP, matrix metalloproteinases; ER-negative BC, estrogen receptor-negative breast cancer.
Leave-one-out plots for analysis of causal effect of MMP-10 on ER-negative BC. (a) Associations between MMP-10 and ER-negative BC (experimental set: ieu-a-1128); (b) Associations between MMP-10 and ER-negative BC (Validation set 1: ieu-a-1135); (c) Associations between MMP-10 and ER-negative BC (Validation set 3: ieu-a-1136); (d) Associations between MMP-10 and ER-negative BC (Validation set 3: ieu-a-1137); (e) Associations between MMP-10 and ER-negative BC (Validation set 4: ieu-a-1166), MMP, matrix metalloproteinases; ER-negative BC, estrogen receptor-negative breast cancer.
Leave-one-out plots for analysis of causal effect of MMP-12 on ER-negative BC. (a) Associations between MMP-12 and ER-negative BC (experimental set: ieu-a-1128); (b) Associations between MMP-12 and ER-negative BC (Validation set 1: ieu-a-1135); (c) Associations between MMP-12 and ER-negative BC (Validation set 3: ieu-a-1136); (d) Associations between MMP-12 and ER-negative BC (Validation set 3: ieu-a-1137); (e) Associations between MMP-12 and ER-negative BC (Validation set 4: ieu-a-1166). MMP, matrix metalloproteinases; ER-negative BC, estrogen receptor-negative breast cancer.
Causal effect of on ER-negative BC on serum MMP levels
To evaluate reverse causation effects, we planned to use the above five GWAS summary data of ER-negative BC. In the five BC GWAS data, no SNP potentially associated with confounders was removed. For the five GWAS summary datasets of ER-negative BC, the first one (GWAS ID: ieu-a-1128) has 40 exposure-associated SNPs, the second one (GWAS ID: ieu-a-1135) has 14 exposure-associated SNPs, The third one (GWAS ID: ieu-a-1136) has seven SNPs, the fourth one (GWAS ID: ieu-a-1137) has only 2 significantly related SNPs, and the last one (GWAS ID: ieu- a-1166) has eight SNPs. Because the number of selected SNP in the fourth dataset (GWAS ID: ieu-a-1137) was less than five, we used the other four datasets to investigate potential causal effect of ER-negative BC on serum level of the five MMPs. Using IVW method, neither of the results derived from these datasets indicated causality from ER-negative BC to the serum level of the five kinds of MMPs (For MMP-1, GWAS ID: ieu-a-1128: P= 0.63; GWAS ID: ieu-a-1135: P=0.87; GWAS ID: ieu-a-1136: P= 0.61; GWAS ID: ieu-a-1166: P= 0.89; For MMP-3, GWAS ID: ieu-a-1128: P= 0.95; GWAS ID: ieu-a-1135: P=0.45; GWAS ID: ieu-a-1136: P= 0.84; GWAS ID: ieu-a-1166: P= 0.88; For MMP-7, GWAS ID: ieu-a-1128: P= 0.38; GWAS ID: ieu-a-1135: P=0.24; GWAS ID: ieu-a-1136: P= 0.90; GWAS ID: ieu-a-1166: P= 0.65; For MMP-10, GWAS ID: ieu-a-1128: P= 0.74; GWAS ID: ieu-a-1135: P=0.94; GWAS ID: ieu-a-1136: P= 0.71; GWAS ID: ieu-a-1166: P= 0.59; For MMP-12, GWAS ID: ieu-a-1128: P= 0.98; GWAS ID: ieu-a-1135: P=0.50; GWAS ID: ieu-a-1136: P= 0.80; GWAS ID: ieu-a-1166: P= 0.36). These results were also supported by the other four methods (MR-Egger, Weighted median, simple mode, and weighted mode). No pleiotropy (P>0.4) or outlier was detected by sensitivity analysis throughout the analysis (Tables 6, 7, 8, 9, 10). No remarkable heterogeneity was found either by MR-Egger or IVW methods for analysis of causal effect of BC on MMP-1/-7/-10/-12 (P>0.05) but MMP-3 (In GWAS data ieu-a-1128: MR Egger: P=0.03; IVW method: P=0.01) (Table 7). For analysis between ER-negative BC (GWAS data ieu-a-1128) and the five types of MMPs, three SNPs were removed for being palindromic with intermediate allele frequencies: rs2735846, rs62116991, and rs191981806. As a result, the results were derived from the remaining 37 SNPs. Leave-one-out plots indicated that no SNP in all four GWAS summary datasets of ER-negative BC had great impact on the MR analysis (Figs. 6, 7, 8, 9, 10). F-statistic for SNPs of all the four datasets of ER-negative BC were greater than the threshold of 10, suggesting strong IVs, which reducing bias of IVs estimates (Supplementary Tables S6, S7, S8, S9, the F statistics for analysis with the other four types of MMPs were the same as the analysis with MMP-1).
Leave-one-out plots for analysis of causal effect of ER-negative BC on MMP-1. (a) Associations between ER-negative BC (ieu-a-1128) and MMP-1; (b) Associations between ER-negative BC (ieu-a-1135) and MMP-1; (c) Associations between ER-negative BC (ieu-a-1136) and MMP-1; (d) Associations between ER-negative BC (ieu-a-1166) and MMP-1, MMP, matrix metalloproteinases; ER-negative BC, estrogen receptor-negative breast cancer.
Leave-one-out plots for analysis of causal effect of ER-negative BC on MMP-3. (a) Associations between ER-negative BC (ieu-a-1128) and MMP-3; (b) Associations between ER-negative BC (ieu-a-1135) and MMP-3; (c) Associations between ER-negative BC (ieu-a-1136) and MMP-3; (d) Associations between ER-negative BC (ieu-a-1166) and MMP-3. MMP, matrix metalloproteinases; ER-negative BC, estrogen receptor-negative breast cancer.
Leave-one-out plots for analysis of causal effect of ER-negative BC on MMP-7. (a) Associations between ER-negative BC (ieu-a-1128) and MMP-7; (b) Associations between ER-negative BC (ieu-a-1135) and MMP-7; (c) Associations between ER-negative BC (ieu-a-1136) and MMP-7; (d) Associations between ER-negative BC (ieu-a-1166) and MMP-7.MMP, matrix metalloproteinases; ER-negative BC, estrogen receptor-negative breast cancer.
Leave-one-out plots for analysis of causal effect of ER-negative BC on MMP-10. (a) Associations between ER-negative BC (ieu-a-1128) and MMP-10; (b) Associations between ER-negative BC (ieu-a-1135) and MMP-10; (c) Associations between ER-negative BC (ieu-a-1136) and MMP-10; (d) Associations between ER-negative BC (ieu-a-1166) and MMP-10.MMP, matrix metalloproteinases; ER-negative BC, estrogen receptor-negative breast cancer.
Leave-one-out plots for analysis of causal effect of ER-negative BC on MMP-12. (a) Associations between ER-negative BC (ieu-a-1128) and MMP-12; (b) Associations between ER-negative BC (ieu-a-1135) and MMP-12; (c) Associations between ER-negative BC (ieu-a-1136) and MMP-12; (d) Associations between ER-negative BC (ieu-a-1166) and MMP-12, MMP, matrix metalloproteinases; ER-negative BC, estrogen receptor-negative breast cancer.
Discussion
In our study, we found that serum MMP-1 is a protective factor for ER-negative BC. In other words, a reduction of serum MMP-1 concentration had causal effect on the risk of ER-negative BC. In the opposite direction, no causal effect was found from ER-negative BC to the serum MMP-1 level. Different from MMP-1, no mutual causal relationship between the other four types of MMPs (MMP-3, MMP-7, MMP-10, and MMP-12) and ER-negative MMPs.
To the best of our knowledge, our study is the first study reporting the bidirectional causality between serum MMP level and BC. MMP could stimulate tumor cell migration, invasion, and metastasis through proteolysis of the extracellular matrix16,44. MMP-1 is a kind of interstitial collagenase which is capable of degrading type I, II, and III collagens. Previous studies have found that exosomal MMP-1 in circulation and MMP-1 expressed on BC cells empowered BCs (especially for TNBC) the potential of distal metastasis (brain, lung, etc.) and led to a poor disease-free survival45,46. One study found that MMP-1 expression was significantly higher in TNBC tissue than in ER-positive and HER-2-positive BC tissue. And MMP-1 expression was also enriched in metastatic BC tissue than in non-metastatic BC tissue47. Studies also reported that MMP-1 or their specific polymorphisms contributed to initiation and progression of BC but the association between MMP-1 level and overall survival was still controversial48,49,50,51. What’s more, certain studies even found that specific genetic variants of MMP-1 did not affect the risk of BC52,53,54. Different from the above results, one study suggested that serum MMP-1 level was significantly lower in BC patients than in healthy controls (P < 0.0001) and patients with a lower serum concentration of MMP-1 had a remarkably shorter 5-year survival55. And another study even demonstrated that stromal expression of MMP-1 was an independent prognostic factor for a longer overall survival (Hazard ratio = 0.528, P = 0.042)56. Nevertheless, few studies focused on association between circulating/serum MMP-1 and each subtype of BC. In our study, serum MMP-1 level had causal effect on ER-negative BC and a high level of MMP-1 serum level caused a lower risk of ER-negative BC, suggesting a protective role of MMP-1 in ER-negative BC. The result was derived not only from IVW method, but also from weighted median and weighted mode methods. In our study, all results were based on IVW method. Moreover, our results were considered robust as selected GWAS summary data of MMP-1 and ER-negative BC had a large sample size. Different types of sensitivity analysis also corroborated the strength and power of our results. According to result of MR analysis of causal effect of ER-negative BC on MMP-1, no positive result was found. This suggested that low serum level of MMP-1 caused ER-negative BC instead of that the latter one resulted in reduction of MMP-1 level. According to the status quo of the research of MMP-1 in breast cancer, inconsistent conclusions could be found in these studies mentioned in our discussion. Firstly, some studies only indicated that MMP-1 promoted carcinogenesis and metastasis of BC though whether all subtypes of BC could be empowered by MMP-1 was unclear. Secondly, most of MMP-related studies focused on tumoral or histological MMP expression. Instead, our study investigated relationship between specifically serum MMP molecules and ER-negative BC. Whether same result could happen in serum MMP should be further discussed. Lastly, studies have suggested that MMP-1 has several genetic variants (polymorphisms) and different variants could impact on prognosis of each subtype of BC in different ways57. In one study from the US in which most of the patients were from Hispanic and non-Hispanic white, investigators found that not all polymorphisms of serum MMP-1 were associated with prognosis of BC and different gene sequences could cause different clinical outcomes. MMP-1 rs17293761 TT genotype was not a risk factor for more advanced breast tumor57. Hence, our study not only corroborated research results in studies believing that MMP-1 was a protective factor but also put forward a new possibility of relationship between serum MMP-1 and ER-negative BC. Regarding that research of relationship between serum MMP-1 and BC was still lacking and the potential mechanism of this phenomenon was unclear, it is worth being furtherly explored to validate this result.
For the other four types of MMPs, we did not find any causality between each of them and ER-negative BC. For MMP-3 (Stromelysin-1) and MMP-10 (stromelysin-2), both of them degrade extracellular matrix (ECM) proteins including aggrecan, collagen types III and IV, and fibronectin58. The former one is not only expressed in cancer cells, but also in normal cells (endothelial cells, epithelial cells, macrophages, and stromal fibroblasts) while the latter one is merely detected in abnormal tissue including acute or chronic injury and cancer59,60. One study suggested that serum level of MMP-10 was significantly higher in BC patients than that in healthy control (P < 0.001). Median serum of MMP-3 was significantly higher in advanced BC (stage III and IV) than that in early-stage BC (stage I) (P = 0.018)61. Another study drew a different conclusion, suggesting that expression of MMP-10 was lower in BC tissue compared with adjacent normal tissue62. Also in the aforementioned study published by Dr. Martha L Slattery from Utah, USA, the clinical significance of MMP-3 were investigated57. Results showed that MMP-3 was associated with breast cancer risk only in part of Native Americann, with merely borderline significance (P = 0.06). For relationship between MMP polymorphism and tumor prognosis, two genetic variants of MMP-3 could drastically increased risk of tumor progression and distant metastasis. Nevertheless, these results were based on mixed population in which Hispanic and Native Amerivans predominated. Whether the results could be applied in other ethnicities should be further explored. For association between MMP-3 and prognosis of breast cancer, one study indicated that MMP-3 did not impact on overall survival but a higher level of cellular expression of MMP-3 had a significantly poorer metastasis-free survival63. Up till now, studies on relationship between MMP-10 and prognosis of breast cancer was not available. As a type of matrilysin, MMP-7 disrupts the structure of and degrade casein, collagen, elastin, fibronectin, gelatins, laminin, and proteoglycans64. Amongst, collagen IV, laminin, and proteoglycan are the major components of basement membrane65. Thus, the biological process of MMP-7 plays a crucial role in local invasion, lymph-node, and distal metastasis of cancer cells66. Studies have shown that serum MMP-7 was higher in BC patients compared with control group67. Another study found that BC patients with bone metastasis had a higher serum level of MMP-7, suggesting a potential circulating biomarker for BC progression towards bone metastasis68. In one study from Xi’an Jiaotong University, researchers illustrated that MMP-7 expression was higher in tissue from advanced breast cancer (larger focus, lymphatic metastasis, and distant metastasis) and patients with positive MMP-7 expression had a poorer 5-year survival (P = 0.046)69. On the contrary, another study reported that serum level of MMP-7 did not correlate with risk of breast tumor and it did not reduce after the removal of the tumor70. Currently, few study has reported positive result and conclusion for association or causal relatoinship between tissue/serum level of MMP-12 with BC. Above all, no consensus has been made on causal effect between these four types of MMP and BC. More intense investigtions in this field should be performed.
Despite the originality and a robust result of our study, some limitations are necessary to be stated: (1) Individuals of this study are from European Ancestry, results derived from selected SNPs could not directly extend to other ethnic groups; (2) Temporarily the GWAS summary data did not contain sufficient IVs to complete analysis for other types of MMPs so that MR analysis between these MMPs and BC could not be conducted; (3) Number of SNPs for the five MMPs were relatively small, especially for MMP-3, MMP-7, and MMP-10, a larger GWAS with more eligible IVs is needed to increase the power of MR analysis.
Conclusions
To conclude, a low level of serum MMP-1 has a causal effect on a high risk of ER-negative BC in European population. In reverse analysis, no causal effect was found from ER-negative BC on the level of serum MMP-1. No evidence supported any causality between MMP-3, -7, -10, -12 and ER-negative BC in European ancestry. More intense research ought to be carried on to validate the serum MMPs as potential biomarkers and therapeutic targets in ER-negative BC.
Methods
Study design
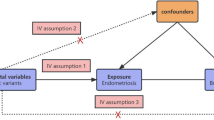
Selection of IVs from genetic variants in this MR analysis strictly meet the three stringent assumptions of MR: (1) as selected IVs, the genetic variants is remarkably associated with the exposure; (2) the genetic variants is not associated with any confounding factors; (3) genetic variants could only indirectly affect the outcome via the exposure, not directly affecting or any other pathways (Fig. 11)71. In our study, we selected summary-level data of 5 kinds of MMPs (MMP-1,-3,-7,-10,-12, containing not less than 5 SNPs) and ER-negative BC from open database of published genome-wide association studies (GWASs) summary dataset (https://gwas.mrcieu.ac.uk/)72,73. We firstly collected genetic variants for each type of MMP in order to determine the causality from MMP to ER-negative BC. Then we collected genetic variants robustly associated with ER-negative BC to validate the reverse causality from BC to MMPs. This is the main goal of our study. The design of bidirectional MR study is overviewed in Fig. 12.
Data sources and SNP selection for MMPs
Genetic variants of 5 kinds of MMPs (MMP-1, MMP-3, MMP-7, MMP-10, and MMP-12) were obtained from a meta-analysis of GWASs including 21,758 individuals from 13 cohorts of European ancestry74. All the five kinds of MMPs passed quality control and were normalized with rank-based inverse normal transformation and/or standardized to unit variance in order to control unrelated variables among cohorts. Genetic associations between 20.3 million genetic variants (SNPs) and log-transformed MMPs were adjusted for population structure (age, sex, smoking status, oral contraceptive usage, blood cell counts, etc.) and study-specific parameters (OLINK plate, storage time, MDS component, etc.) 74. To meet the first assumption of MR, we extracted the IVs of the five types of MMPs at genome-wide significance (5 × 10−8). 17 SNPs were significantly associated with MMP-1; 12 SNPs were significantly related with MMP-3; seven SNPs were significantly associated with MMP-7; eleven SNPs were significantly associated with MMP-10; and 15 SNPs were remarkably associated with MMP-12. Meanwhile, a linkage disequilibrium (LD) test was conducted on these SNPs to clump SNPs for independence. All SNPs were strongly and independently (R2 < 0.01 within 5 Mb) predicted MMP level from the published GWAS. Subsequently, we input all the SNPs significantly associated with MMPs into Phenoscanner database (V2) (http://www.phenoscanner.medschl.cam.ac.uk/) to determine if any SNPs were associated with confounders (P < 5 × 10−8)75,76. Resulted SNPs would be deleted to reduce the possibility of pleiotropic effect.
Data sources and SNP selection for ER-negative BC
Summary-level data on ER-negative BC were extracted from a GWAS of 127,442 individuals of European ancestry from Breast Cancer Association Consortium (BCAC), combined with Discovery, Biology and Risk of Inherited Variants in Breast Cancer Consortium (DRIVE), iCOGS project, and data from other GWAS meta-analysis77. This data would be used as experimental dataset to explore potential causal effect between MMPs and ER-negative BC. Then we used the other four datasets as validation datasets to prove the conclusion draught from the experimental datasets. The four datasets were all derived from European Ancestry (OncoArray1, case: 9655, control: 45494; iCOGS, case: 7333, control: 42892; GWAS meta-analysis1, case: 4480, control: 17588; GWAS meta-analysis2: case: 3611, control: 18084)77,78. Similar to SNP selection for MMPs, potential SNPs correlated with confounders would be removed by using Phenoscanner database (P<5×10−8).To further evaluate robustness of selected SNPs, statistics F and R2 were used in both the process of SNP selection for MMP and ER-negative BC. F statistic stands for the precision and magnitude of the genetic effect on the trait. The Eq. (1) is:
N stands for sample size of a certain GWAS and R2 is the proportion of the variance of the trait caused by genetic variants (SNPs). The Eq. (2) is:
EAF is short for “Effect Allele Frequency” (EAF) of the SNP and β is the estimated effect of SNP on the trait. SNPs with F less than ten would be removed and SNPs with F larger than 10 were robust to prove the validity of selected SNPs
Bidirectional mendelian randomizasion analysis
Bidirectional two-sample MR was performed by using the R pacakge “TwosampleMR”. Information of SNPs, β value (created by log-transformation of odds ratios [ORs]), standard error, P-value, and EAF value of selected exposure instrument were necessary for this package to harmonize exposure and outcome data to investigate direction of causality between MMPs and ER-negative BC by using summary association data. In our study we did not look for proxies to replace SNPs that were not available in the outcome datasets. During data harmonization, we should ensure that all selected SNPs were derived from the same allele no matter in exposure or outcome data. For palindromic SNPs, however, they were too difficult to be recognized whether the SNPs were from the same allele because the sequence were same on both strands. As a result, palindromic SNPs were removed to eliminate the ambiguity as to whether exposure and outcome GWAS infer the same effect allele43. In the core process of MR analysis, we measured Wald ratio (i.e, βoutcome/βexposure) for each SNP and then summarized these SNP-sepcific Wald ratio via inverse-variance-weighted (IVW) method which estimated causal effects of genetically predicted exposure on outcome81,82. We demonstrated the estimate effects in ORs for binary outcome (ER- negative BC) and in β for continuous outcome (MMP level). To explore the direction of causality from MMP to BC, OR was elaborated as risk for ER-negative BC (outcome) per unit increase in serum level of certain type of MMP. Other methods in MR anaysis include: MR-Egger, weighted median, simple mode, and weighted mode. A series of sensitivity analysis were performed, consisting of weighted median (WM) method, MR-Egger, and Mendelian Randomization Pleiotropy RESidual Sum and Outlier (MR-PRESSO). WM method reckons the causal effect by selecting median MR estimate for condition in which multiple genetic variants are invalid or present pleiotropy83. MR-Egger method is robust not only to provide a consistent estimate of causal effect, but also to evaluate horizontal pleiotropy of IVs and a non-zero intercept suggesting that the IVW estimate is biased84,85. MR-PRESSO is capable of detecting and correcting any potentially pleiotropic outliers (SNPs) for all reported results to avoid bias86. Heterogeneity was quantified by the Cochran Q statistics and I2 statistics, in which larger I2 indicates increasing heterogeneity87. Furthermore, “leave-one-out analysis” was also conducted by removing each SNPs to test the stability and reliability of the MR results. By virtue of multiple testing in our analysis, Bonferroni correction was used to modify the significant level for multiple tests. Thus we considered P-values below (0.05/25=0.002) as strong evidence of associations. Results with P-values between 0.002 and 0.05 were regarded as suggestive associations43. All statistical analysis were two-sided. All analysis was conducted using R software (4.2.0) with R package of “TwosampleMR” (version 0.5.6), “MRPRESSO” (version 1.0). Reporting follows the STROBE-MR statement.
Data availability
All data generated or analysed during this study are included in this published article and its Supplementary information files.
Abbreviations
- BC:
-
Breast cancer
- BCAC:
-
Breast cancer association consortium
- CI:
-
Confidence interval
- EAF:
-
Effect allele frequency
- ER:
-
Estrogen receptor
- GWASs:
-
Genome-wide association studies
- HER-2:
-
Human epidermal growth factor receptor 2
- IVW:
-
Inverse-variance-weighted
- IVs:
-
Instrumental variables
- LD:
-
Linkage disequilibrium
- MMP:
-
Matrix metalloproteinases
- MR:
-
Mendelian randomization
- MR-PRESSO:
-
Mendelian randomization pleiotropy RESidual sum and outlier
- OR:
-
Odds ratio
- SNP:
-
Single nucleotide polymorphisms
- TNBC:
-
Triple-negative breast cancer
References
Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71(3), 209–249 (2021).
Voduc, K. D. et al. Breast cancer subtypes and the risk of local and regional relapse. J. Clin. Oncol. 28(10), 1684–1691 (2010).
Belete, A. M., Aynalem, Y. A., Gemeda, B. N., Demelew, T. M. & Shiferaw, W. S. The effect of estrogen receptor status on survival in breast cancer patients in ethiopia retrospective cohort study. Breast Cancer 14, 153–161 (2022).
Davies, C. et al. Relevance of breast cancer hormone receptors and other factors to the efficacy of adjuvant tamoxifen: Patient-level meta-analysis of randomised trials. Lancet 378(9793), 771–784 (2011).
Colditz, G. A., Rosner, B. A., Chen, W. Y., Holmes, M. D. & Hankinson, S. E. Risk factors for breast cancer according to estrogen and progesterone receptor status. J. Natl. Cancer Inst. 96(3), 218–228 (2004).
Goddard, K. A. et al. HER2 evaluation and its impact on breast cancer treatment decisions. Public Health Genom. 15(1), 1–10 (2012).
Howlader, N. et al. US incidence of breast cancer subtypes defined by joint hormone receptor and HER2 status. J. Natl. Cancer Inst. https://doi.org/10.1093/jnci/dju055 (2014).
Jerusalem, G., Lancellotti, P. & Kim, S. B. HER2+ breast cancer treatment and cardiotoxicity: Monitoring and management. Breast Cancer Res. Treat. 177(2), 237–250 (2019).
Asif, H. M., Sultana, S., Ahmed, S., Akhtar, N. & Tariq, M. HER-2 positive breast cancer - a mini-review. Asian Pac. J. Cancer Prev. 17(4), 1609–1615 (2016).
Parakh, S., Gan, H. K. & Scott, A. M. Sensitization of cancers resistant to HER2 antibodies. Crit. Rev. Oncog. 25(3), 175–207 (2020).
Bredin, P., Walshe, J. M. & Denduluri, N. Systemic therapy for metastatic HER2-positive breast cancer. Semin. Oncol. 47(5), 259–269 (2020).
Carey, L. A. et al. Race, breast cancer subtypes, and survival in the Carolina breast cancer study. JAMA 295(21), 2492–2502 (2006).
Siddharth, S. & Sharma, D. Racial disparity and triple-negative breast cancer in African-American women: A multifaceted affair between obesity, biology, and socioeconomic determinants. Cancers 10(12), 514 (2018).
de Jong, V. M. T. et al. Prognostic value of stromal tumor-infiltrating lymphocytes in young, node-negative, triple-negative breast cancer patients who did not receive (neo) adjuvant systemic therapy. J. Clin. Oncol. 40(21), 2361–2374 (2022).
Gross, J. & Lapiere, C. M. Collagenolytic activity in amphibian tissues: A tissue culture assay. Proc. Natl. Acad. Sci. U. S. A. 48(6), 1014–1022 (1962).
Visse, R. & Nagase, H. Matrix metalloproteinases and tissue inhibitors of metalloproteinases: Structure, function, and biochemistry. Circ. Res. 92(8), 827–839 (2003).
Lohi, J., Wilson, C. L., Roby, J. D. & Parks, W. C. Epilysin, a novel human matrix metalloproteinase (MMP-28) expressed in testis and keratinocytes and in response to injury. J. Biol. Chem. 276(13), 10134–10144 (2001).
Sternlicht, M. D. & Werb, Z. How matrix metalloproteinases regulate cell behavior. Annu. Rev. Cell Dev. Biol. 17, 463–516 (2001).
Malemud, C. J. Matrix metalloproteinases (MMPs) in health and disease: An overview. Front. Biosci. 11, 1696–1701 (2006).
Lemaître, V. & D’Armiento, J. Matrix metalloproteinases in development and disease. Birth Defects Res. C. Embryo. Today 78(1), 1–10 (2006).
Deryugina, E. I. & Quigley, J. P. Tumor angiogenesis: MMP-mediated induction of intravasation- and metastasis-sustaining neovasculature. Matrix. Biol. 44–46, 94–112 (2015).
Peng, B. et al. Meta-analysis of association between matrix metalloproteinases 2, 7 and 9 promoter polymorphisms and cancer risk. Mutagenesis 25(4), 371–379 (2010).
Peng, B. et al. Polymorphisms in the promoter regions of matrix metalloproteinases 1 and 3 and cancer risk: A meta-analysis of 50 case-control studies. Mutagenesis 25(1), 41–48 (2010).
McColgan, P. & Sharma, P. Polymorphisms of matrix metalloproteinases 1, 2, 3 and 9 and susceptibility to lung, breast and colorectal cancer in over 30,000 subjects. Int. J. Cancer 125(6), 1473–1478 (2009).
Huang, C. et al. Epidemiological evidence between variants in matrix metalloproteinases-2, -7, and -9 and cancer risk. Front. Oncol. 12, 856831 (2022).
Xu, T., Zhang, S., Qiu, D., Li, X. & Fan, Y. Association between matrix metalloproteinase 9 polymorphisms and breast cancer risk: An updated meta-analysis and trial sequential analysis. Gene 759, 144972 (2020).
Han, M. et al. Associations of MMP-2 −1306 C/T and MMP-9 −1562 C/T polymorphisms with breast cancer risk among different populations: A meta-analysis. Genes Genom. 39, 331–340 (2017).
Ou, Y. X. & Bi, R. Meta-analysis on the relationship between the SNP of MMP-2-1306 C>T and susceptibility to breast cancer. Eur. Rev. Med. Pharmacol. Sci. 24(3), 1264–1270 (2020).
Ren, F. et al. Overexpression of MMP family members functions as prognostic biomarker for breast cancer patients: A systematic review and meta-analysis. PLoS ONE 10(8), e0135544 (2015).
Song, J., Su, H., Zhou, Y. Y. & Guo, L. L. Prognostic value of matrix metalloproteinase 9 expression in breast cancer patients: A meta-analysis. Asian Pac. J. Cancer. Prev. 14(3), 1615–1621 (2013).
Chen, Y., Wang, X., Chen, G., Dong, C. & Zhang, D. The impact of matrix metalloproteinase 2 on prognosis and clinicopathology of breast cancer patients: A systematic meta-analysis. PLoS ONE 10(3), e0121404 (2015).
Sui, J., Huang, J. & Zhang, Y. The MMP-1 gene rs1799750 polymorphism is associated with breast cancer risk. Genet. Test. Mol. Biomarkers 25(7), 496–503 (2021).
Liu, D. et al. Association between polymorphisms in the promoter regions of matrix metalloproteinases (MMPs) and risk of cancer metastasis: A meta-analysis. PLoS ONE 7(2), e31251 (2012).
Hill, H. A. et al. A longitudinal analysis of predictors of quitting smoking among participants in a self-help intervention trial. Addict. Behav. 19(2), 159–173 (1994).
Lee, Y. H., Bae, S. C. & Song, G. G. Hepatitis B virus (HBV) reactivation in rheumatic patients with hepatitis core antigen (HBV occult carriers) undergoing anti-tumor necrosis factor therapy. Clin. Exp. Rheumatol. 31(1), 118–121 (2013).
Chen, Y. C. et al. Assessing causality between childhood adiposity and early puberty: A bidirectional mendelian randomization and longitudinal study. Metabolism 100, 153961 (2019).
Bowden, J. & Holmes, M. V. Meta-analysis and mendelian randomization: A review. Res Synth Methods 10(4), 486–496 (2019).
Burgess, S., Small, D. S. & Thompson, S. G. A review of instrumental variable estimators for mendelian randomization. Stat. Methods. Med. Res. 26(5), 2333–2355 (2017).
Emdin, C. A., Khera, A. V. & Kathiresan, S. Mendelian randomization. JAMA 318(19), 1925–1926 (2017).
Yang, Z., Yu, R., Deng, W. & Wang, W. Genetic evidence for the causal association between programmed death-ligand 1 and lung cancer. J Cancer Res. Clin. Oncol. 147(11), 3279–3288 (2021).
Chen, D. et al. Assessing causality between osteoarthritis with urate levels and gout: A bidirectional Mendelian randomization study. Osteoarthr. Cartil. 30(4), 551–558 (2022).
Davey Smith, G. & Hemani, G. Mendelian randomization: Genetic anchors for causal inference in epidemiological studies. Hum. Mol. Genet. 23(R1), R89-98 (2014).
Wang, Q., Shi, Q., Lu, J., Wang, Z. & Hou, J. Causal relationships between inflammatory factors and multiple myeloma: A bidirectional Mendelian randomization study. Int. J. Cancer 151(1750), 1759 (2022).
Brinckerhoff, C. E., Rutter, J. L. & Benbow, U. Interstitial collagenases as markers of tumor progression. Clin. Cancer. Res. 6(12), 4823–4830 (2000).
Zhu, Y. et al. Exosomal MMP-1 transfers metastasis potential in triple-negative breast cancer through PAR1-mediated EMT. Breast Cancer Res. Treat. 193(1), 65–81 (2022).
Harati, R., Hafezi, S., Mabondzo, A. & Tlili, A. Silencing miR-202-3p increases MMP-1 and promotes a brain invasive phenotype in metastatic breast cancer cells. PLoS ONE 15(10), e0239292 (2020).
Wang, Q. M., Lv, L., Tang, Y., Zhang, L. & Wang, L. F. MMP-1 is overexpressed in triple-negative breast cancer tissues and the knockdown of MMP-1 expression inhibits tumor cell malignant behaviors in vitro. Oncol. Lett. 17(2), 1732–1740 (2019).
Balkhi, S., Mashayekhi, F., Salehzadeh, A. & Saedi, H. S. Matrix metalloproteinase (MMP)-1 and MMP-3 gene variations affect MMP-1 and -3 serum concentration and associates with breast cancer. Mol. Biol. Rep. 47(12), 9637–9644 (2020).
Hughes, S. et al. Matrix metalloproteinase single-nucleotide polymorphisms and haplotypes predict breast cancer progression. Clin. Cancer. Res. 13(22 Pt 1), 6673–6680 (2007).
Padala, C. et al. Synergistic effect of collagenase-1 (MMP1), stromelysin-1 (MMP3) and gelatinase-B (MMP9) gene polymorphisms in breast cancer. PLoS ONE 12(9), e0184448 (2017).
Boström, P. et al. MMP-1 expression has an independent prognostic value in breast cancer. BMC Cancer 11, 348 (2011).
Hsiao, C. L. et al. The Association of matrix metalloproteinase-1 promoter polymorphisms with breast cancer. In Vivo 32(3), 487–491 (2018).
Białkowska, K. et al. Polymorphisms in MMP-1, MMP-2, MMP-7, MMP-13 and MT2A do not contribute to breast, lung and colon cancer risk in polish population. Hered. Cancer Clin. Pract. 18, 16 (2020).
Zhou, P. et al. Current evidence on the relationship between four polymorphisms in the matrix metalloproteinases (MMP) gene and breast cancer risk: A meta-analysis. Breast Cancer Res. Treat. 127(3), 813–818 (2011).
Kulić, A., Dedić Plavetić, N., Vrbanec, J. & Sirotković-Skerlev, M. Low serum MMP-1 in breast cancer: A negative prognostic factor?. Biomarkers 17(5), 416–421 (2012).
Kim, G. E. et al. Expression of matrix metalloproteinases and their inhibitors in different immunohistochemical-based molecular subtypes of breast cancer. BMC Cancer 14, 959 (2014).
Slattery, M. L. et al. Matrix metalloproteinase genes are associated with breast cancer risk and survival: The breast cancer health disparities study. PLoS ONE 8(5), e63165 (2013).
Mirastschijski, U. et al. Novel specific human and mouse stromelysin-1 (MMP-3) and stromelysin-2 (MMP-10) antibodies for biochemical and immunohistochemical analyses. Wound Repair Regen. 27(4), 309–323 (2019).
Fang, S. et al. Polymorphisms in the MMP1 and MMP3 promoter and non-small cell lung carcinoma in North China. Carcinogenesis 26(2), 481–486 (2005).
McMahan, R. S. et al. Stromelysin-2 (MMP10) moderates inflammation by controlling macrophage activation. J. Immunol. 197(3), 899–909 (2016).
Piskór, B. M. et al. Plasma level of MMP-10 may be a prognostic marker in early stages of breast cancer. J. Clin. Med. 9(12), 4122 (2020).
Benson, C. S., Babu, S. D., Radhakrishna, S., Selvamurugan, N. & Ravi Sankar, B. Expression of matrix metalloproteinases in human breast cancer tissues. Dis. Markers. 34(6), 395–405 (2013).
Mehner, C. et al. Tumor cell expression of MMP3 as a prognostic factor for poor survival in pancreatic, pulmonary, and mammary carcinoma. Genes. Cancer. 6(11–12), 480–489 (2015).
Basu, S., Thorat, R. & Dalal, S. N. MMP7 is required to mediate cell invasion and tumor formation upon Plakophilin3 loss. PLoS ONE 10(4), e0123979 (2015).
Paulsson, M. Basement membrane proteins: Structure, assembly, and cellular interactions. Crit. Rev. Biochem. Mol. Biol. 27(1–2), 93–127 (1992).
Chang, J. & Chaudhuri, O. Beyond proteases: Basement membrane mechanics and cancer in vasion. J. Cell. Biol. 218(8), 2456–2469 (2019).
Piskór, B. M. et al. Plasma concentrations of Matrilysins MMP-7 and MMP-26 as diagnostic biomarkers in breast cancer. J. Clin. Med. 10(7), 1436 (2021).
Voorzanger-Rousselot, N. et al. Association of 12 serum biochemical markers of angiogenesis, tumour invasion and bone turnover with bone metastases from breast cancer: A crossectional and longitudinal evaluation. Br. J. Cancer. 95(4), 506–514 (2006).
Cao, P. L. et al. Expressions of FOXC1 and MMP-7 in molecular subtypes of breast cancer and their association with clinicopathological characteristics. Zhejiang Da Xue Xue Bao Yi Xue Ban 43(4), 406–412 (2014).
Katunina, A. I. et al. Matrix metalloproteinases 2, 7, and 9 in tumors and sera of patients with breast cancer. Bull. Exp. Biol. Med. 151(3), 359–362 (2011).
Lawlor, D. A., Harbord, R. M., Sterne, J. A., Timpson, N. & Davey Smith, G. Mendelian randomization: Using genes as instruments for making causal inferences in epidemiology. Stat. Med. 27(8), 1133–1163 (2008).
Ben Elsworth, M. L. et al. The MRC IEU OpenGWAS data infrastructure. bioRxiv 08(10), 244293v1 (2020).
Hemani, G. et al. The MR-Base platform supports systematic causal inference across the human phenome. Elife 7, e34408 (2018).
Folkersen, L. et al. Genomic and drug target evaluation of 90 cardiovascular proteins in 30,931 individuals. Nat. Metab. 2(10), 1135–1148 (2020).
Staley, J. R. et al. PhenoScanner: A database of human genotype-phenotype associations. Bioinformatics 32(20), 3207–3209 (2016).
Kamat, M. A. et al. PhenoScanner V2: An expanded tool for searching human genotype-phenotype associations. Bioinformatics 35(22), 4851–4853 (2019).
Michailidou, K. et al. Association analysis identifies 65 new breast cancer risk loci. Nature 551(7678), 92–94 (2017).
Michailidou, K. et al. Genome-wide association analysis of more than 120,000 individuals identifies 15 new susceptibility loci for breast cancer. Nat. Genet. 47(4), 373–380 (2015).
Palmer, T. M. et al. Using multiple genetic variants as instrumental variables for modifiable risk factors. Stat. Methods. Med. Res. 21(3), 223–242 (2012).
D., F. Introduction to quantitative genetics. Prentice. Hall. (1996).
Bowden, J. et al. A framework for the investigation of pleiotropy in two-sample summary data Mendelian randomization. Stat. Med. 36(11), 1783–1802 (2017).
Burgess, S., Bowden, J., Fall, T., Ingelsson, E. & Thompson, S. G. Sensitivity analyses for robust causal inference from mendelian randomization analyses with multiple genetic variants. Epidemiology 28(1), 30–42 (2017).
Bowden, J., Davey Smith, G., Haycock, P. C. & Burgess, S. Consistent estimation in mendelian randomization with some invalid instruments using a weighted median estimator. Genet. Epidemiol. 40(4), 304–314 (2016).
Burgess, S. & Thompson, S. G. Interpreting findings from mendelian randomization using the MR-Egger method. Eur. J. Epidemiol. 32(5), 377–389 (2017).
Bowden, J., Davey Smith, G. & Burgess, S. Mendelian randomization with invalid instruments: Effect estimation and bias detection through Egger regression. Int. J. Epidemiol. 44(2), 512–525 (2015).
Verbanck, M., Chen, C. Y., Neale, B. & Do, R. Detection of widespread horizontal pleiotropy in causal relationships inferred from Mendelian randomization between complex traits and diseases. Nat. Genet. 50(5), 693–698 (2018).
Bowden, J. et al. Assessing the suitability of summary data for two-sample Mendelian randomization analyses using MR-Egger regression: The role of the I2 statistic. Int. J. Epidemiol. 45(6), 1961–1974 (2016).
Acknowledgements
We deeply appreciate all the patients and volunteers for their participation in this MR study. We deeply acknowledge all the investigators from all related GWAS consortia and technicians from Ieu Open GWAS Project for their great contribution in data curating, collection, and summary in GWAS summary database (https://gwas.mrcieu.ac.uk/). We sincerely thank to all scientists from all related fields for establishing several important open source software projects related to this dataset and making the GWAS data publicly available. Last but not the least, we also deeply thank the team of Phenoscanner led by Dr. James Staley for design, development, and production of this database of human genotype-phenotype associations.
Funding
This work was supported by CAMS Innovation Fund for Medical Sciences (CIFMS) (2021-I2M-1-061 and 2021-1-I2M-003 and 2018-I2M-3-001), CAMS Clinical and Translational Medicine Research Funds (2019XK320006), and Beijing Natural Science Foundation (7192158).
Author information
Authors and Affiliations
Contributions
Z.Z. and Q.C. mainly designed and performed analysis, and wrote the manuscript; M.Z. and C.W. performed statistical analysis and verified data; X.L. supervised the entire project. All authors have read, carefully discussed, provided critical feedback on intellectual content, and approved the submission of final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhao, Z., Cao, Q., Zhu, M. et al. Causal relationships between serum matrix metalloproteinases and estrogen receptor-negative breast cancer: a bidirectional mendelian randomization study. Sci Rep 13, 7849 (2023). https://doi.org/10.1038/s41598-023-34200-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-34200-0
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.