Abstract
Hornets are the largest of the social wasps, and are important regulators of insect populations in their native ranges. Hornets are also very successful as invasive species, with often devastating economic, ecological and societal effects. Understanding why these wasps are such successful invaders is critical to managing future introductions and minimising impact on native biodiversity. Critical to the management toolkit is a comprehensive genomic resource for these insects. Here we provide the annotated genomes for two hornets, Vespa crabro and Vespa velutina. We compare their genomes with those of other social Hymenoptera, including the northern giant hornet Vespa mandarinia. The three hornet genomes show evidence of selection pressure on genes associated with reproduction, which might facilitate the transition into invasive ranges. Vespa crabro has experienced positive selection on the highest number of genes, including those putatively associated with molecular binding and olfactory systems. Caste-specific brain transcriptomic analysis also revealed 133 differentially expressed genes, some of which are associated with olfactory functions. This report provides a spring-board for advancing our understanding of the evolution and ecology of hornets, and opens up opportunities for using molecular methods in the future management of both native and invasive populations of these over-looked insects.
Similar content being viewed by others
Introduction
Insects are the most speciose and abundant organisms on the planet. This is an exciting time for insect research as we are enjoying an explosion in the sequencing of insect genomes1,2. At the time of writing, 3058 insect genomes have been published on International Nucleotide Sequence Database Collaboration (INSDC); although impressive, this represents only 0.1% of described insect species. Having a genome sequenced for a species opens up a wealth of research opportunities with benefits to evolutionary biologists, ecologists and conservation scientists3. Genomic resources provide clues to understand how and why species distributions are affected by anthropogenic actions, and to develop effective ways to track and manage populations. This is especially important for ecologically and economically important species, which provide critical ecosystem services on which planetary health depends (e.g., crop pest control, biodiversity and agriculture pollination); but it is also important for managing species which become problematic outside of their native ranges, as invasive species. The social insects (termites, ants, some bees and wasps) account for 75% of the insect biomass4; ant biomass alone equals 20% of human biomass5. It is not surprising, therefore, that this group of insects has amongst the greatest ecological and economic impact on natural and farmed ecosystems6. This, together with their fascinating social lives and relatively small genomes, has made the social insects popular choices for genome sequencing projects. Indeed, there are more species of the social Hymenoptera (ants, bees, wasps) with published genome sequences (National Center for Biotechnology Information (NCBI) RefSeq reference genomes or 158 species at the time of writing) than there are for the more highly speciose insect orders such as Coleoptera (beetles) (NCBI RefSeq reference genomes for 110 species at the time of writing). Amongst the insects, the number of hymenopteran genomes sequenced is second only to the Diptera1. Despite this focus, there remains a critical taxonomic bias7, with 89% of the sequenced hymenopteran genomes belonging to ants and bees, leaving a mere 11% Vespidae wasp species genomes on INSDC (including nine social wasp and hornet species; Supplementary Table ST1). The paucity of genomic resources for wasps is perplexing as they provide critical ecological services as regulators of arthropod populations, pollinators, seed dispersers and as a popular source of human nutrition in some parts of the world (e.g. Japan; China; South America)8. Moreover, the wasps include several of the world’s worst invasive species9,10,11. There is an urgent need to widen the genomic resources available for these important facets of our planet’s biodiversity.
Insect genome sequencing has been increasingly popular thanks to significant technological advances such as long-read sequencing12 and three-dimensional chromosomal mapping methods13. Substantial funding initiatives contribute to this genome production line, such as the Darwin Tree of Life project, which aims to generate high-quality genomes from all eukaryote species in Britain and Ireland14,15, the i5K Consortium which aims to sequence 5000 high-priority arthropod species, including the top 100 agricultural pests in the US16, and the Global Ant Genomic Alliance which is sequencing high-quality genomes for 200 ant species17. Social insect genomes improve our understanding of the evolution of social organisation7,18, e.g. by identifying loci involved in communication, such as gene family expansions of ant and bee odorant receptors and termite lipocalins19,20,21, as well as improving our understanding of their ecology and behaviour, e.g. on how bees adapt to living at high altitudes22. Furthermore, genomic data can provide insights into biodiversity trends associated with climate change and invasive species23,24,25.
There are around 1,200 species of social wasps, including relatively well-known Vespidae yellowjackets and hornets26, and lesser-known species of Stenogastrinae27 and Polistinae28. They display a huge variety of ecological and life-history characteristics; e.g. the colony size varies enormously, from species with less than 10 individuals in the society (e.g. as found in the Stenogastrine hover wasps) to those with tens of thousands of workers (e.g. Vespula species); some species have many reproductive queens (e.g. epiponine wasps, like Metapolybia) whilst others are monogynous (e.g. Vespa crabro); reproductive hierarchies can be regulated by conventions such as age or size29, aggression30 or pheromones31.
The first aculeate wasp genome was published in 2015 (Polistes canadensis32), closely followed by the European paper wasp Polistes dominula33. Currently there are genome sequences published for seven polistine wasp species and nine vespine wasps (including those presented in this paper; see Table 1). However, evolutionary analyses of these vespine genomes are lacking, and particularly so for the hornets, the Vespa genus. Such analyses are important for several reasons (Fig. 1). Vespa exhibit some of the most complex societies of the aculeate wasps, with colonies headed by one (or a few) queens who are morphologically distinct from their workers34. They constitute an important part of our planet’s natural capital as top predators and regulators of insect populations, pollinators and seed dispersers8. Several species of Vespa have been inadvertently introduced outside of the native range where their colony life span can be longer and they have become problematic as invasive species35; for instance the European hornet Vespa crabro has become established in the USA34. Some invasions threaten local fauna; e.g. the recent invasion of Vespa velutina from South East Asia threatens beekeeping in Europe36,37,38; Vespa mandarinia from South East Asia was introduced in several parts of North America where it poses a threat to local ecosystems and human health39. Given their ecological and economic importance and their history of invasion success, it is surprising that Vespa have not received more attention as subjects of genomic studies. It is therefore time to put Vespa genomes on the map and explore their potential.
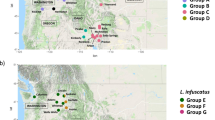
Biology of Vespa hornets. Comparing the key life-history traits of the three Vespa species analyzed in this study. Species column: Female adult morphology for Vespa crabro, Vespa velutina and Vespa mandarinia (photos taken from www.inaturalist.org, respectively from the following users: rainyang, Mиxaил Maлышeв, Kinmatsu Lin, all photos have CCBY-NC license). Distribution column: Known geographical distribution of species (from https://www.gbif.org/)40,41,42; the redder patches indicate higher occurrence records; their invasive distributions are circled. Biological information column: Descriptions of individual and nest traits and behaviours28,37,39,43. Diet column: All three Vespa species (left) prey on a diverse set of arthropod orders (right)8,44,45.
Here we compare the genomes of three ecologically, economically and societally important hornets—the European hornet Vespa crabro, the yellow-legged Asian hornet Vespa velutina, and the northern giant hornet Vespa mandarinia46. Two of these genomes (V. crabro and V. velutina) were sequenced and annotated for the purposes of this study. We compare the genome compositions of the three Vespa species with each other and contrast them with other social insect genomes (Aim 1). We are specifically interested in comparing lineages of insects that have evolved superorganismality independently47, to explore losses and gains of genes related to social organisation, such as chemoreceptors48,49. We identified genes undergoing rapid evolution and family expansions (Aim 2) and identified specific differences among the three Vespa species, such as duplication events among genes involved in communication. These provide insights into the ecological differences among the three study species, for instance by describing species-specific olfactory systems. We identified signs of positive selection in Vespa relative to other social insects and determined whether they were enriched for any specific functionalities. We found little evidence that genes under selection may be involved in caste determination. Finally, we examined differential gene expression among castes in one species, V. crabro (Aim 3), presenting the first transcriptomic analyses of castes in a superorganismal wasp.
Materials and methods
Sample collection
For Aim 1, four colonies of V. crabro were collected in September 2017 from four different locations in their native range of southern England (see Supplementary Table ST2). Individual workers, gynes (non-mated queens), males and queens were flash frozen on dry ice or collected straight into RNAlater and stored at – 80 °C thereafter. Samples from two colonies of V. velutina were collected in 2017 from Ventimiglia, Italy which is part of the invasive range50. Workers and gynes were collected alive and stored immediately in ethanol and − 20 °C thereafter.
DNA and RNA library preparation and sequencing
DNA was extracted from whole bodies of two males of V. crabro using Qiagen DNeasy blood and tissue kit (Catalogue number: 69504; Hilden, Germany) following the manufacturers’ protocol (details of this and all following protocols in Supplementary Information). One sample contributed to the two runs from a pair-end TruSeq Nano LT library (Cat #: FC-121-4001, Illumina, San Diego, CA, USA; INSDC accession numbers: SRR11213735 and SRR11213736) and the other to the Nextera mate-pair library (Cat #: FC-132-1001, Illumina; SRR11213734). Sequencing was performed at the Vienna BioCenter Core Facilities on an Illumina HiSeq 2500 sequencer.
DNA was extracted from one male of V. velutina using Qiagen MagAttract HMW DNA extraction kit (Cat # 67563, Illumina). It was sequenced by the Wellcome Sanger Institute as part of their 25 genomes initiative (https://www.sanger.ac.uk/collaboration/25-genomes-for-25-years/, INSDC accession ERS3567203), producing a combination of Pacific Biosciences CLR (PacBio Express Library Kit Cat # 102-088-900; on needle-sheared DNA), 10 × Genomics Chromium and Hi-C data.
RNA was extracted from a range of tissues (from workers, gynes, queens and larvae) for gene annotation purposes. For Aim 3, we also sequenced brain RNA from two different castes—gynes and workers—of V. crabro. Using gynes rather than mature queens avoids the effects of matedness, age, overwintering, and reproductive maturity (developed eggs) on gene expression. We focused on measuring variation in gene expression related to behaviour between castes47; thus, brains were dissected from individuals derived from three to four different nests (see Supplementary Table ST2), and total RNA was extracted from individuals following published methods51 (RNeasy Mini Kit Cat # 74104, Qiagen). RNA extractions were checked for quality using TapeStation. Three to four brains were then pooled for each caste, ensuring that no single pool had more than one wasp from the same nest; this generated a total of four worker pooled samples and five gyne pooled samples for sequencing, representing respectively four and five biological replicates. We sequenced these nine pooled samples (150 bp paired-end, Illumina NovaSeq platform, by Novogene) and obtained at least 22 million reads per sample (Supplementary Table ST2).
Genome assembly and annotation
We assembled and annotated two new Vespa genomes. The genome of V. crabro was assembled with SOAPdenovo_v2.0452,53 into contigs from one haploid male. The resulting assembly was screened for contaminants during INSDC submission (INSDC accession: JAITYU000000000). The genome of V. velutina (INSDC accession PRJEB46979) was assembled with the following process, palindromic read correction with pbclip, initial PacBio assembly generation with Falcon-unzip54, retained haplotig separation with purge_dups, Hi-C based scaffolding with SALSA2, Arrow polishing (from Pacific Biosciences, https://github.com/PacificBiosciences/GenomicConsensus), and short-read polishing using FreeBayes-called variants from 10× Genomics Chromium reads aligned with LongRanger. Chromosome-scale scaffolds confirmed by the Hi-C data have been named in order of size.
In order to compare genome compositions amongst Vespa and with other social insects (Aim 2), we annotated these two new genomes for structural and functional loci. We used RNAseq data combined with “ab initio” and comparative computational methods, including mapping wasp UniProt-derived proteins, to predict genes in the repeatmasked genomes of V. crabro and V. velutina (Supplementary Information; Supplementary Table ST3). We obtained genome and annotation files from all other species from INSDC (Supplementary Table ST4). We estimated the assemblies’ statistics using BUSCO v5.1.255 and QUAST v5.0.256. Additionally, we compared synteny between genomes (Supplementary Table ST5A) with D-genies using Minimap v.257,58. Finally, to annotate Transposable Elements (TE), we used RepeatModeler v2.0.459 to generate a model of TEs for the V. velutina and V. crabro genomes separately. Consensus sequences were clustered and filtered using cd-hit v4.6.860 and BBmap v39.0161 as recommended62. We then used RepeatMasker v4.1.063 to annotate TEs based on the model for that genome (Supplementary Table ST4.
Orthogroup exploration for positive selection and duplication events
In order to explore patterns of gene evolution across the species (Aim 2), we ran OrthoFinder 2.5.464 on the longest isoform of each protein, first on eight Hymenoptera species (Vespa crabro, Vespa velutina, Vespa mandarinia, Vespula germanica, Vespula pensylvanica, Vespula vulgaris, Apis mellifera, Solenopsis invicta; Fig. 2a), then on seven Hymenoptera species (without the ant, S. invicta). We based our duplication events exploration on the resulting listed multiple-copy orthologues for each species and each internal node.
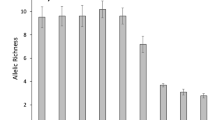
Vespa genomes statistics in Hymenoptera context. (a) Phylogenetic context and protein-coding gene content of the two new genomes (in bold) with Apis bee, Solenopsis ant, Polistes paper wasps, Vespula yellowjacket wasps. Branch lengths (unit: number of substitutions per site) are from species tree inference algorithm STAG (OrthoFinder). Left Bar Plot: Most Hymenoptera BUSCO genes are found as single copies (mauve) although a small number were duplicated (purple), fragmented genes (dark orange) or missing (light orange). Vespa crabro had a higher proportion of fragmented BUSCOs (Complete: 91.3% [Single-copy: 91.1%, Duplicated: 0.2%], Fragmented: 3.2%, Missing: 5.5%, n: 5991) than Vespa velutina (C: 96.3%[S: 96.1%, D: 0.2%], F: 0.9%, M: 2.8%, n: 5991) and Vespa mandarinia (C: 96.4%[S: 96.1%, D: 0.3%], F: 0.9%, M: 2.7%, n: 5991). Right Bar Plot: Total gene counts in each species in relation to OrthoFinder results. Out of 17,061 orthogroups, 163 are single-copy across the ten species (orange) and 1595 are found in multiple copies (green). Most of the protein-coding genes of each species are orthologous to one extend (Other, blue). (b) Vespa genomes composition: number of protein-coding genes, number of proteins, gene density (number of genes per 1000 bp), based on V. crabro and V. velutina EVM-consensus annotations, and based on V. mandarinia RefSeq annotation. All data in Supplementary Table ST4.
We kept 2685 resulting nearly-1:1 orthogroups (i.e. between one and three copies of the gene in any given species, and any given gene had to be present in a minimum of six species) for the six wasps and the bee. This is a more relaxed filtering method than strict single-copy orthologs, allowing more data points in our exploration49,65. We then aligned all species’ sequences using PRANK v.151120 (gffread -x, PRANK options -f = paml -F -codon66). We investigated evidence of positive selection being experienced in the past on a specific branch (branch-test; M0 (Model = 0, Nsites = 0); M2 (Model = 2, Nsites = 0)) with PAML v.l4.9j67 using codeml with the phylogenetic tree from OrthoFinder of six vespid species (three Vespa and three Vespula species). To gain more insight on specific loci, we then ran branch-site models, focusing on potential loci on specific branches (“foreground”) against other branches (“background”) using codeml’s dN/dS branch-site models of positive selection68. Briefly, we compared the ratio of non-synonymous mutations to synonymous mutations (ω) in nearly 1:1 single-copy orthogroups of six those vespid species and tested two models: neutral evolution in foreground branch (Model = 2, Nsites = 2; ω fixed at 1, null hypothesis) and positive selection in foreground branch (Model = 2, Nsites = 2; ω > 1, alternative hypothesis; Supplementary Table ST6). We ran likelihood ratio tests67 and further tested for association with significance of log-likelihood test with chi square tests. Significant loci having experienced positive selection are those orthogroups with adjusted P value (Benjamini-Hochberg) below 0.05. We then assigned a description to each orthogroup using BLASTp v.2.10.069 with A. mellifera as query to the nr database. Finally, we obtained GO terms from our gene subsets using TopGO v.2.38.170.
Differential gene expression analysis
For Aim 3, we employed Nextflow pipeline nf-core/rnaseq v.19.10.0.517071 to assess the quality of RNA reads with FastQC v0.11.8, to trim adapters with ‘TrimGalore!’ v0.6.4. We mapped the nine pooled samples (four gynes, five workers) against the genome with STAR vSTAR_2.6.1d72 and to obtain read count with featureCounts v1.6.473. We assessed the quality of our data with a Principal Components Analysis of normalised read count, which showed samples to cluster by caste (Supplementary Figure SF7). We then conducted a differential gene expression analysis between worker and gyne read count using DESeq2 v.1.26.074 (log normalization; alpha cutoff = 0.05; Bonferoni adjustment). Similar to the positive selection analysis, we used BLASTp and TopGO to explore results.
Results and Discussion
Aim 1: Vespa genomes statistics
We sequenced and assembled the draft genomes of V. crabro and V. velutina, using short-read and long-read data, respectively. We compare these assemblies to a publicly available assembly for V. mandarinia (Fig. 1), together with genomes from representatives of other aculeate wasps, ants and bees (Fig. 2a, Table 1). The Vespa genomes are similar in size (V. crabro: 211,313,510 bp, V. velutina: 193,994,974 bp, V. mandarinia: 247,710,421 bp; Table 1 and Supplementary Table ST4), and are within the expected range for Hymenoptera7. The degree to which the genome assemblies are fragmented reflects the sequencing technology used for each species. Those generated using short-read technology are more fragmented than those generated using long-read technology, and thus might influence our downstream report. The N50 for the short-read sequenced V. crabro was 34,273 bp, whilst that for the long-read sequences are significantly better (N50 for V. velutina was 9,190,824 bp; N50 for V. mandarinia was 2,778,186 bp; Table 1). All three Vespa species have low GC content (Table 1), as is characteristic of Hymenoptera7. V. crabro has 16,409 protein-coding genes and 19,597 annotated proteins; V. velutina has 12,928 protein-coding genes and 17,334 annotated proteins; the NCBI RefSeq annotation of V. mandarinia has 12,089 protein-coding genes and 27,185 annotated proteins (Fig. 2b); all of which are within known range for Hymenoptera (Supplementary Table ST4). The three Vespa assemblies have a high level of expected single-copy orthologous genes, as measured by the Hymenoptera BUSCO score (V. crabro: 91.1%, V. velutina: 96.1%, V. mandarinia: 96.1%; Fig. 2a and Supplementary Table ST4). These high BUSCO scores and the gene counts variation seen in the full-gene set barplot (Fig. 2a, Supplementary Table ST7) hint at lineage-specific differences. Indeed, there are only 14 genes common to the three Vespa species (in blue-green) whereas we counted 1,681 genes specific to the Vespula species (in grey). The genome of Vespa crabro contains a smaller proportion of TE (22%), compared to Vespa velutina (26%, Supplementary Table ST4; Supplementary Figure SF10). This range is larger than the 10% calculated in the Polistine lineage75. Both species’ TE landscape (Supplementary Figure SF8-SF9) fits within the expected range for insects. Interestingly, while all families appear to be larger in V. velutina, a larger proportion of Penelope elements are found in V. crabro (5.9%) compared to V. velutina (0.2%, which is within the range of recently published annotations of Polistes exclamans (0.16%) and Mischocyttarus mexicanus (0.02%)75). An extensive survey of Hymenoptera TEs will provide more insights in these variations.
In the pairwise assembly mapping between the three Vespa genomes, we find the highest contig similarity between V. mandarinia and V. velutina long-read genomes (88% of contigs are between 50 and 75% similar, Supplementary Table ST5A), while short-read V. crabro genome typically scores lower synteny score due to its fragmentation34,57. Interestingly, there is a large part of the V. crabro assembly that is not similar to the other two species’ assembly: over 5% of the V. crabro genome does not map to the V. mandarinia genome nor the V. velutina genome (i.e. no match in contig similarity; Supplementary Figures SF4–SF6), but these regions are small (less than 2000 bp) and not in coding regions (i.e. not in the annotation file). Their sequences are not similar to those of the closest model organism Apis mellifera (Supplementary Table ST5B); they are most probably representing some repetitive elements. We additionally find some evidence of a one-off diagonal inversion in V. mandarinia (length: 1,152,777 bp; on original strand: locus NW_023395844.1, start 3,701,405, end 5,336,819) when mapped to V. velutina (on original strand: locus SUPER_22, start 2,372,742, end 4,003,498; Supplementary Figure SF6). This region has 96 annotated V. mandarinia genes and seems to be only inverted between V. mandarinia and V. velutina, hinting towards a V. mandarinia-specific rearrangement. Further improvements in assembly contiguity for V. crabro as well as further V. mandarinia population-level resequencing are needed to quantify and qualify both the unique region covering 5% of V. crabro genome and this potential inversion in V. mandarinia.
Aim 2: evidence of gene duplication and site-specific positive selection in Vespa
Gene family expansions are thought to be associated with novel functions78 and invertebrate invasion79. Target gene families include those involved in communication, odorant binding49 and caste differentiation80. We thus compared the number of multiple copies of orthogroups between the three Vespa genomes. We focus on 16,913 orthogroups present in at least one species among ten chosen Hymenoptera representatives. Between lineages, we find more gene duplication events along the Vespa branch (25%; 5632 out of 22,976 events associated with 16,913 orthogroups) than along the Vespula branch (4%; 990 events; Supplementary Figure SF3). V. crabro has 1,187 duplicated orthogroups (7% of 16,913 orthogroups), including the highest proportion of species-specific gene family expansions: 77% of duplicated genes have two or more copies unique to this species (911 out of 1187). V. mandarinia has the highest number of duplicated orthogroups (2,177; 13% of 16,913 orthogroups), including 65% species-specific gene family expansions. V. velutina has 881 duplicated orthogroups (5% of 16,913 orthogroups), including 75% species-specific gene family expansions (Fig. 3a, Supplementary Table ST7). There are 28 genes that are duplicated in all three Vespa, some of which could be associated with the odorant or nervous systems (see notable examples of best BLASTp hits in Fig. 3b). V. crabro-specific gene duplications include sequences similar to notable Hymenoptera proteins. Those of particular interest include ten proteins from zinc finger family, a gene family known to also be duplicated in incipiently social Ceratina bees80; odorant receptors which are commonly found in lineage-specific expansions in Hymenoptera evolution49 and predicted to be duplicated in 80 invasive insect species81, and transposable element derived proteins which are involved in DNA-binding transcription factor activities and are thought to be associated with regulation of phenotypes via genome architecture variation82 (Supplementary Table ST8).
Comparative analyses of gene and protein evolution in vespine wasps. (a) Number of orthogroups with two or more copies, colour-coded by ancestral state (yellow: ancestral; grey: species-specific). Vespa mandarinia has the highest number of duplicated genes. All wasps have a high proportion of species-specific duplications. (b) Illustrative example of orthogroups that are duplicated in all three Vespa species, with associated Hymenoptera gene description after BLAST nr. (c) Orthogroups clustered by Euclidean distances of dN/dS categories (orthogroups in columns, blue represents positive selection in branch-site models) and by rows (species). There is very little overlap between species; overlapping areas with genes of interest are highlighted. Number of orthogroups that have experienced positive selection (dN/dS > 1, FDR 0.05) in Vespa and Vespula branches. Vespa crabro has the highest number of genes with a dN/dS ratio higher than 1 (n = 104). (d) Overlap of GO terms of orthogroups having experienced positive selection. Most of the species have a unique large set of GO terms, as seen in the tall, coloured bars on the far right-hand side of the graph (Fisher classic, unadjusted P value, see ST26–ST31.
Selection pressures associated with social phenotypic plasticity impact the molecular evolution of proteins83. We thus explored the rate of protein evolution (dN/dS) for each single-copy orthogroup in the six Vespa and Vespula species (branch models: Supplementary Tables ST8–ST16; branch-site models: Supplementary Tables ST17-ST24). In branch-site models, V. crabro has 104 genes that showed evidence of having experienced positive selection (FDR 0.05, Fig. 3c, Supplementary Table ST17). From the best BLAST hits, notable loci include four zinc finger proteins (e.g. XP_006560154.1 zinc finger CCHC domain-containing protein 4)—a family known for its binding sites being more variable in social bees compared to solitary bees84 and known for being included in an ant social supergene85; an intraflagellar transport protein (XP_006560607.2)—known to be associated with insect olfactory sensilia used for communication86. In V. velutina, 57 genes showed evidence of positive selection (Supplementary Table ST21). Notable loci included: ELMO domain-containing protein C (XP_395400.7), found to be upregulated in a incipiently social bee when experimentally forced to increased brood provisioning87; and senecionine N-oxygenase (XP_006571261.2), which is involved in detoxification and found upregulated in pesticide-sprayed beetle88. In Vespa mandarinia, 31 genes showed evidence of positive selection (Supplementary Table ST19) including deformed epidermal autoregulatory factor 1 (XP_395757.3), a transcription factor associated with honey bee egg laying behaviour89. We found only one locus (OG0002464) that had experienced positive selection across all three Vespa species: the sequence matches in BLAST Apis cerana’s transmembrane protein-like (Fig. 3c). One locus—Spidroin-2-like precursor protein (NP_001314894.1)—was under selection in both V. crabro and V. mandarinia; this gene could be playing a role during pupa development90. Zinc finger CCHC domain-containing protein 4 (XP_006560154.1) was also under selection in both V. crabro and V. velutina.
When we compared these results with Vespula species, we found similar range of genes with evidence of positive selection (Supplementary Tables ST18, ST20, ST22). However, there was no overlap of genes with dN/dS > 1 between the six vespine wasp species nor distinct vespine chromosomal hotspot for positive selection (Fig. 3c in blue).
We next tested for GO term enrichment among the genes under positive selection in each Vespa branch-site model (Supplementary Tables ST25–ST30, Fig. 3d). There was no significant enrichment at the most stringent level (FDR 0.05). However, examination of terms with the lowest raw P values (Praw < 0.05) revealed functions associated with reproduction and morphological innovations: oogenesis (GO:0048477, hallmark of eusocial species91) for V. crabro; hippo signalling (GO:0035329, involved in termite soldier morphological differentiation92) for V. mandarinia; signalling pathway (GO:0007224, part of the hedgehog signalling pathway associated with beetle horn morphological innovation93) for V. velutina. We additionally found species overlaps in GO terms, namely: cell signalling and reproduction in Vespa species; immunity in Vespula species.
In summary, the Vespa lineage may have experienced positive selection on genes related to communication, reproduction, production/regulation of alternative phenotypes. Our comparative analyses on orthologous genes consistently highlighted V. crabro, with the highest number of genes under positive selection and associated GO Terms, and with the highest proportion of species-specific duplicated genes in the Vespa lineage. Thus, we explored gene expression between phenotypes in this species, for our last Aim.
Aim 3: caste-associated differential gene expression in Vespa crabro
Analyses of the molecular basis of caste differentiation in the brains of social insects has provided important insights into how division of labour in group living societies evolves94,95. However, to date these analyses in wasps have been limited to species from simple societies. Here we provide the first analysis of brain transcriptomes among castes of a superorganismal wasp. We extracted RNA from 40 individual brains of V. crabro. These were sequenced as five pooled samples of gynes (i.e. each sample contains brains from four individuals) and four pooled samples of workers (i.e. each sample contains brains from three individuals). We detected 1171 differentially expressed genes between workers and gynes (FDR-adjusted P value < 0.05; no log2 fold change filter; 648 upregulated, 523 downregulated, Fig. 4a). After filtering for log2 fold change 1.5, 133 genes remained: 63 were upregulated and 70 were downregulated genes in gynes relative to workers (Fig. 4b, Supplementary Table ST31). These included sequences with BLAST match to Apis mellifera’s zinc finger CCCH domain-containing protein 13—also found in V. mandarinia’s queen transcriptome96—, and odorant receptor proteins, recurrent in all hymenopteran olfactory repertoires, including hornets97. Caste-enriched GO terms for the differentially expressed genes (FDR < 0.05, Supplementary Table ST32) included pheromone-related activity (GO:0016229, also upregulated in experimentally-isolated bumblebee brains98), as well as oxidoreductase activity (GO:0016491) involved in reprogramming cell metabolism and previous found to be caste-biased in social Hymenoptera32,99.
Caste-specific differentially expressed genes in Vespa crabro brain transcriptomes. (a) Differential expression analysis based on negative binomial distribution of read counts of 1171 genes from 4 workers and 5 non-mated queens (gynes) of Vespa crabro. Read counts from DESeq2 results are filtered by FDR adjusted P value < 0.05 and cluster by castes. Genes are colour-coded in the right-hand side column by stricter filtering (absolute log fold change above 1.5), resulting in 63 genes upregulated in gynes (green) and 70 downregulated (orange). (b) Top Gene Ontology Terms enriched in Vespa crabro’s differentially expressed genes (n = 133 DEG, FDR < 0.05, absolute log fold change = 1.5). 12 Molecular Function GO terms (red) are significantly enriched in the DEG.
We examined whether any of the differentially expressed genes in V. crabro had experienced positive selection or duplication events. A single locus was found in both the dataset of duplicated genes (duplicated in OrthoFinder analysis; aim 2) and differentially expressed between castes (Vcabro1a000853P1), which shares sequence similarity with heterogeneous nuclear ribonucleoprotein H in Apis mellifera (Supplementary Table ST31). This locus has previously been identified as caste-biased in Cape honey bees, where it is implicated in alternative splicing leading to worker ovary activation100. It is also known in several bee species to recognise epigenetic modifications and is part of the post-transcriptional RNA modification machinery101.
Conclusion
In this study, we sequenced, assembled and annotated two new Vespa genomes; we also generated gene annotation files, available for the community. We then conducted an analysis of gene evolution for vespine wasps, comparing with other social Hymenoptera. We detected high levels of gene duplications among genes associated with reproduction, communication and production of phenotypic variation in the Vespa species; for instance, two genes related to odorant receptors are duplicated in V. velutina. Furthermore, we found 104 genes under positive selection in V. crabro, including some associated with reproduction such as oogenesis. Finally, we provide the first analyses of caste-specific brain gene expression for a superorganismal wasp, comparing transcriptomes derived from adult workers and unmated female reproductives (gynes).
Our analyses of duplications, positive selection and differentially expressed genes detected associations with the olfactory system. Communication is behaviourally key to all social insects for reproduction102, colony health103 and species’ survival, including invasive species104. This is supported by evolutionary variation in chemosensory repertoire105 in social and invasive insects, for instance, loss of gustatory receptors in caterpillar pests, and duplicated chemosensory genes in invasive insects. Genes belonging to the zinc finger family were also found repeatedly in our analyses. This is one of the largest families of transcription factors, and so may not be so surprising. However, we suggest that this is a biologically interesting result: zinc finger domains found to be under selection in the Vespa lineage (duplicated, dN/dS, differentially expressed) may have experienced lineage-specific expansions and may be related to new gene regulatory functions106,107. Future work should focus on conducting functional tests, e.g. CRISPR to demonstrate functionality.
Accordingly, the genomic resources and analyses we provide secures the hornets a place in the ever-growing world of genome sequencing and analyses, and adds significantly to what remains an underrepresented group of insects in the genomics world—the aculeate wasps7. As an example of the exponential growth of genomics datasets leading to population genetics and pangenome analyses, since our analyses, the Darwin Tree of Life initiative made available a new Vespa crabro assembly. These new datasets will improve the scope of comparative analyses in the Hymenoptera by, for instance, providing a more complete survey of non-coding regions and olfactory receptors related to the evolution of social organisation in insects. Finally, our datasets hint at candidate genes with important gene functions—namely those involved in regulation of reproduction and communication, and open up new opportunities to explore the molecular mechanisms underpinning key ecological and physiological traits, such as those associated with invasive species and venom components in invasive Vespula species76.
Future work should focus on population genetic studies to further explore selection pressures associated with social evolution and anthropogenic impacts (e.g. high FST regions indicating directional selection and Tajima’s D indicating balancing selection within species, especially in the context of comparison between invasive and native ranges). Furthermore, it would be interesting to conduct a comprehensive TE analysis across wasps to assess potential functions related to phenotypic plasticity, for instance when comparing native and invasive species108,109,110. Finally, we encourage single-cell brain expression analyses111, and analysis of multiple tissues from across the diversity of phenotypes112 to further explore differential gene expression in Vespa crabro. Such approaches will help build a better understanding of these ecological and economically important insects.
Data availability
The datasets generated during and analysed during the current study are available in the NCBI repository, SRR11213734, SRR11213735, SRR11213736, JAITYU000000000, SRX9350949, SRX9350950, SRX9350951, SRX9350952, SRX9350953, SRX9350956, SRX9350958, SRX9350959, SRX9350960, ERS3567203, PRJEB46979, SRR22217221, SRR22217222. The annotations are available on Github: https://github.com/EmelineFavreau/Vespa-Genomes-Analyses/tree/master/input/gff.
References
Hotaling, S. et al. Long-reads are revolutionizing 20 years of insect genome sequencing. Genome Biol. Evol. https://doi.org/10.1093/gbe/evab138 (2021).
Li, F. et al. Insect genomes: Progress and challenges. Insect Mol. Biol. 0, (2019).
Ungerer, M. C., Johnson, L. C. & Herman, M. A. Ecological genomics: Understanding gene and genome function in the natural environment. Heredity 100, 178–183 (2008).
Hölldobler, B. & Wilson, E. O. The Ants (Harvard Univ. Press, 1990).
Schultheiss, P. et al. The abundance, biomass, and distribution of ants on Earth. Proc. Natl. Acad. Sci. USA 119, e2201550119 (2022).
Elizalde, L. et al. The ecosystem services provided by social insects: Traits, management tools and knowledge gaps. Biol. Rev. Camb. Philos. Soc. 95, 1418–1441 (2020).
Branstetter, M. G. et al. Genomes of the Hymenoptera. Curr. Opin. Insect Sci. 25, 65–75 (2018).
Brock, R. E., Cini, A. & Sumner, S. Ecosystem services provided by aculeate wasps. Biol. Rev. Camb. Philos. Soc. 96, 1645–1675 (2021).
Beggs, J. R. et al. Ecological effects and management of invasive alien Vespidae. Biocontrol 56, 505–526 (2011).
Kenis, M. et al. Ecological effects of invasive alien insects. Biol. Invasions 11, 21–45 (2009).
Spradbery, J. P. Wasps. An account of the biology and natural history of social and solitary wasps, with particular reference to those of the British Isles (Sidgwick & Jackson, 1973).
Amarasinghe, S. L. et al. Opportunities and challenges in long-read sequencing data analysis. Genome Biol. 21, 30 (2020).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Blaxter, M. et al. Why sequence all eukaryotes? Proc. Natl. Acad. Sci. 119, e2115636118 (2022).
The Darwin Tree of Life Project Consortium et al. Sequence locally, think globally: The Darwin Tree of Life Project. Proc. Natl. Acad. Sci. 119, e2115642118 (2022).
i5K Consortium. The i5K Initiative: advancing arthropod genomics for knowledge, human health, agriculture, and the environment. J Hered 104, 595–600 (2013).
Boomsma, J. J. et al. The global ant genomics Alliance (GAGA). Myrmecol. News 25, 61–66 (2017).
Favreau, E., Martínez-Ruiz, C., Rodrigues Santiago, L., Hammond, R. L. & Wurm, Y. Genes and genomic processes underpinning the social lives of ants. Curr. Opin. Insect Sci. 25, 83–90 (2018).
Shell, W. A. et al. Sociality sculpts similar patterns of molecular evolution in two independently evolved lineages of eusocial bees. Commun. Biol. 4, 253 (2021).
Shigenobu, S. et al. Genomic and transcriptomic analyses of the subterranean termite Reticulitermes speratus: Gene duplication facilitates social evolution. Proc. Natl. Acad. Sci. USA 119, e2110361119 (2022).
Zhou, X. et al. Phylogenetic and transcriptomic analysis of chemosensory receptors in a pair of divergent ant species reveals sex-specific signatures of odor coding. PLoS Genet. 8, e1002930 (2012).
Sun, C. et al. Genus-wide characterization of bumblebee genomes provides insights into their evolution and variation in ecological and behavioral traits. Mol. Biol. Evol. 38, 486–501 (2021).
Ometto, L. et al. Linking genomics and ecology to investigate the complex evolution of an invasive Drosophila pest. Genome Biol. Evol. 5, 745–757 (2013).
Wu, N. et al. Fall webworm genomes yield insights into rapid adaptation of invasive species. Nat. Ecol. Evol. 3, 105–115 (2019).
North, H. L., McGaughran, A. & Jiggins, C. D. Insights into invasive species from whole-genome resequencing. Mol. Ecol. 30, 6289–6308 (2021).
Carpenter, J. M. & Kojima, J.-I. Checklist of the species in the subfamily Vespinae (Insecta: Hymenoptera: Vespidae). Nat. Hist. Bull. Ibaraki Univ. 1, 51–92 (1997).
Carpenter, J. M. & Kojima, J.-I. Checklist of the species in the subfamily stenogastrinae (Hymenoptera: Vespidae). J. N. Y. Entomol. Soc. 104, 21–36 (1996).
Ross, K. G. & Matthews, R. W. The Social Biology of Wasps (Cornell University Press, 1991).
Taylor, B. A., Cini, A., Cervo, R., Reuter, M. & Sumner, S. Queen succession conflict in the paper wasp Polistes dominula is mitigated by age-based convention. Behav. Ecol. 31, 992–1002 (2020).
Patalano, S. et al. Self-organization of plasticity and specialization in a primitively social insect. Cell Syst. 13, 768-779.e4 (2022).
Oi, C. A. et al. Dual effect of wasp queen pheromone in regulating insect sociality. Curr. Biol. 25, 1638–1640 (2015).
Patalano, S. et al. Molecular signatures of plastic phenotypes in two eusocial insect species with simple societies. Proc. Natl. Acad. Sci. 112, 13970–13975 (2015).
Standage, D. S. et al. Genome, transcriptome and methylome sequencing of a primitively eusocial wasp reveal a greatly reduced methylation system in a social insect. Mol. Ecol. 25, 1769–1784 (2016).
Perrard, A., Pickett, K., Villemant, C., Kojima, J. & Carpenter, J. Phylogeny of hornets: A total evidence approach (Hymenoptera, Vespidae, Vespinae, Vespa). J. Hymenopt. Res. 32, 1–15 (2013).
Wilson Rankin, E. E. Emerging patterns in social wasp invasions. Curr. Opin. Insect Sci. 46, 72–77 (2021).
Cini, A. et al. Competition between the native and the introduced hornets Vespa crabro and Vespa velutina: A comparison of potentially relevant life-history traits: Competition between native and alien hornets. Ecol. Entomol. 43, 351–362 (2018).
Monceau, K., Bonnard, O. & Thiéry, D. Vespa velutina: A new invasive predator of honeybees in Europe. J. Pest Sci. 87, 1–16 (2014).
Cappa, F., Cini, A., Bortolotti, L., Poidatz, J. & Cervo, R. Hornets and honey bees: A coevolutionary arms race between ancient adaptations and new invasive threats. Insects 12, 1037 (2021).
Alaniz, A. J., Carvajal, M. A. & Vergara, P. M. Giants are coming? Predicting the potential spread and impacts of the giant Asian hornet (Vespa mandarinia, Hymenoptera:Vespidae) in the USA. Pest Manag. Sci. 77, 104–112 (2021).
Vespa crabro Linnaeus in GBIF Secretariat (2022). GBIF Backbone Taxonomy. Checklist dataset https://doi.org/10.15468/39omei accessed via GBIF.org on 2022–03–16.
Vespa velutina Lepeletier, 1836 in GBIF Secretariat (2022). GBIF Backbone Taxonomy. Checklist dataset https://doi.org/10.15468/39omei accessed via GBIF.org on 2022–03–16.
Vespa mandarinia Smith, 1852 in GBIF Secretariat (2022). GBIF Backbone Taxonomy. Checklist dataset https://doi.org/10.15468/39omei accessed via GBIF.org on 2022–03–16.
Matsuura, M. & Yamane, S. Biology of the vespine wasps (Springer, 1990).
Rome, Q. et al. Not just honeybees: predatory habits of Vespa velutina (Hymenoptera: Vespidae) in France. Ann. Soc. Entomol. Fr. 57, 1–11 (2021).
Verdasca, M. J. et al. A metabarcoding tool to detect predation of the honeybee Apis mellifera and other wild insects by the invasive Vespa velutina. J. Pest Sci. 95, 997–1007 (2022).
Childers, A. K., Geib, S. M., Smith, T., Foster, L. J. & Korlach, J. Asian giant hornet, Vespa mandarinia, genome assembly. (2020) https://doi.org/10.15482/USDA.ADC/1519179.
Boomsma, J. J. & Gawne, R. Superorganismality and caste differentiation as points of no return: how the major evolutionary transitions were lost in translation: Superorganisms, eusociality and major transitions. Biol. Rev. 93, 28–54 (2018).
Jongepier, E. et al. Convergent loss of chemoreceptors across independent origins of slave-making in ants. Mol. Biol. Evol. 39, msab305 (2022).
Legan, A. W., Jernigan, C. M., Miller, S. E., Fuchs, M. F. & Sheehan, M. J. Expansion and accelerated evolution of 9-exon odorant receptors in polistes paper wasps. Mol. Biol. Evol. 38, 3832–3846 (2021).
Cappa, F., Cini, A., Pepiciello, I., Petrocelli, I. & Cervo, R. Female body size, weight and fat storage rather than nestmateship determine male attraction in the invasive yellow-legged hornet Vespa velutina nigrithorax. Ethol. Ecol. Evol. 31, 73–85 (2019).
Taylor, B. A., Cini, A., Wyatt, C. D. R., Reuter, M. & Sumner, S. The molecular basis of socially mediated phenotypic plasticity in a eusocial paper wasp. Nat. Commun. 12, 775 (2021).
Luo, R. et al. SOAPdenovo2: An empirically improved memory-efficient short-read de novo assembler. Gigascience 1, 18 (2012).
Dohm, J. C. et al. The genome of the recently domesticated crop plant sugar beet (Beta vulgaris). Nature 505, 546–549 (2014).
Chin, C.-S. et al. Phased diploid genome assembly with single-molecule real-time sequencing. Nat. Methods 13, 1050–1054 (2016).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: Assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Gurevich, A., Saveliev, V., Vyahhi, N. & Tesler, G. QUAST: Quality assessment tool for genome assemblies. Bioinformatics 29, 1072–1075 (2013).
Cabanettes, F. & Klopp, C. D-GENIES: Dot plot large genomes in an interactive, efficient and simple way. PeerJ 6, e4958 (2018).
Li, H. Minimap2: Pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Flynn, J. M. et al. RepeatModeler2 for automated genomic discovery of transposable element families. Proc. Natl. Acad. Sci. 117, 9451–9457 (2020).
Li, W. & Godzik, A. Cd-hit: A fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22, 1658–1659 (2006).
Bushnell, B. BBMap: A fast, accurate, splice-aware aligner (2014).
Goubert, C. et al. A beginner’s guide to manual curation of transposable elements. Mob. DNA 13, 7 (2022).
Smit, A. F., Hubley, R., & Green, PR. RepeatMasker. (2013).
Emms, D. M. & Kelly, S. OrthoFinder: Phylogenetic orthology inference for comparative genomics. Genome Biol. 20, 1–14 (2019).
Wyatt, C. D. R. et al. Genetic toolkit for sociality predicts castes across the spectrum of social complexity in wasps. bioRxiv Preprint at https://doi.org/10.1101/2020.12.08.407056 (2020).
Löytynoja, A. & Goldman, N. A model of evolution and structure for multiple sequence alignment. Philos. Trans. R. Soc. Lond. B Biol. Sci. 363, 3913–3919 (2008).
Yang, Z. PAML: A program package for phylogenetic analysis by maximum likelihood. Comput. Appl. Biosci. 13, 555–556 (1997).
Zhang, J. Evaluation of an improved branch-site likelihood method for detecting positive selection at the molecular level. Mol. Biol. Evol. 22, 2472–2479 (2005).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Alexa, A., & Rahnenfuhrer, J. topGO: Enrichment Analysis for Gene Ontology. R package version 2.50.0. (2022).
Patel, H. et al. nf-core/rnaseq: nf-core/rnaseq v3.8.1: Plastered Magnesium Mongoose. (2022).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: An efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biology, 15, 550 (2014).
Miller, S. E., Legan, A. W., Uy, F. M. K. & Sheehan, M. J. Highly contiguous genome assemblies of the Guinea Paper Wasp (Polistes exclamans) and Mischocyttarus mexicanus. Genome Biol. Evol. 14, evac110 (2022).
Harrop, T. W. R. et al. High-quality assemblies for three invasive social wasps from the Vespula genus. G3 Genes Genomes Genet. 10, 3479–3488 (2020).
Miller, S. E. et al. Evolutionary dynamics of recent selection on cognitive abilities. Proc. Natl. Acad. Sci. 117, 3045–3052 (2020).
Long, M., VanKuren, N. W., Chen, S. & Vibranovski, M. D. New gene evolution: Little did we know. Annu. Rev. Genet. 47, 307–333 (2013).
Makino, T. & Kawata, M. Invasive invertebrates associated with highly duplicated gene content. Mol. Ecol. 28, 1652–1663 (2019).
Rehan, S. M. et al. Conserved genes underlie phenotypic plasticity in an incipiently social bee. Genome Biol. Evol. 10, 2749–2758 (2018).
Huang, C. et al. InvasionDB: A genome and gene database of invasive alien species. J. Integr. Agric. 20, 191–200 (2021).
Rubenstein, D. R. et al. Coevolution of genome architecture and social behavior. Trends Ecol. Evol. 34, 844–855 (2019).
Zhao, F. et al. Molecular evolution of bumble bee vitellogenin and vitellogenin-like genes. Ecol. Evol. 11, 8983–8992 (2021).
Kapheim, K. M. et al. Genomic signatures of evolutionary transitions from solitary to group living. Science (2015).
Purcell, J., Lagunas-Robles, G., Rabeling, C., Borowiec, M. L. & Brelsford, A. The maintenance of polymorphism in an ancient social supergene. Mol. Ecol. 30, 6246–6258 (2021).
Schmidt, H. R. & Benton, R. Molecular mechanisms of olfactory detection in insects: Beyond receptors. Open Biol 10, 200252 (2020).
Séguret, A. C. Ageing and the costs of reproduction: Insights from Euglossa viridissima, an orchid bee on the cusp of sociality (Universitäts- und Landesbibliothek Sachsen-Anhalt, 2021).
Bastarache, P. et al. Transcriptomics-based approach identifies spinosad-associated targets in the Colorado Potato Beetle, Leptinotarsa decemlineata. Insects 11, 820 (2020).
Jones, B. M. et al. Individual differences in honey bee behavior enabled by plasticity in brain gene regulatory networks. Elife 9, e62850 (2020).
Sehnal, F. & Akai, H. Insect silk glands: their types, development and function, and effects of environmental factors and morphogenetic hormones on them. Int. J. Insect Morphol. Embryol. 19, 79–132 (1990).
Holman, L., Helanterä, H., Trontti, K. & Mikheyev, A. S. Comparative transcriptomics of social insect queen pheromones. Nat. Commun. 10, 1593 (2019).
Yaguchi, H., Suzuki, R., Matsunami, M., Shigenobu, S. & Maekawa, K. Transcriptomic changes during caste development through social interactions in the termite Zootermopsis nevadensis. Ecol. Evol. 9, 3446–3456 (2019).
Kijimoto, T. & Moczek, A. P. Hedgehog signaling enables nutrition-responsive inhibition of an alternative morph in a polyphenic beetle. Proc. Natl. Acad. Sci. USA 113, 5982–5987 (2016).
Toth, A. L. & Rehan, S. M. Molecular evolution of insect sociality: An eco-evo-devo perspective. Annu. Rev. Entomol. 62, 419–442 (2017).
Sumner, S. Determining the molecular basis of sociality in insects: Progress, prospects and potential in sociogenomics. Ann. Zool. Fenn. 43, 423–442 (2006).
Patnaik, B. B. et al. Transcriptome profile of the Asian Giant Hornet (Vespa mandarinia) using Illumina HiSeq 4000 sequencing: De novo assembly, functional annotation, and discovery of SSR markers. Int. J. Genom. Proteomics 2016, 4169587 (2016).
Couto, A., Mitra, A., Thiéry, D., Marion-Poll, F. & Sandoz, J.-C. Hornets have it: A conserved olfactory subsystem for social recognition in Hymenoptera?. Front. Neuroanat. 11, 48 (2017).
Wang, Z. Y. et al. Isolation disrupts social interactions and destabilizes brain development in bumblebees. Curr. Biol. 32, 2754-2764.e5 (2022).
Honeybee Genome Sequencing Consortium. Insights into social insects from the genome of the honeybee Apis mellifera. Nature 443, 931–949 (2006).
Jarosch, A., Stolle, E., Crewe, R. M. & Moritz, R. F. A. Alternative splicing of a single transcription factor drives selfish reproductive behavior in honeybee workers (Apis mellifera). Proc. Natl. Acad. Sci. USA 108, 15282–15287 (2011).
Bataglia, L., Simões, Z. L. P. & Nunes, F. M. F. Active genic machinery for epigenetic RNA modifications in bees. Insect Mol. Biol. 30, 566–579 (2021).
Wittwer, B. et al. Solitary bees reduce investment in communication compared with their social relatives. Proc. Natl. Acad. Sci. 114, 6569–6574 (2017).
Cremer, S., Armitage, S. A. O. & Schmid-Hempel, P. Social immunity. Curr. Biol. 17, R693–R702 (2007).
Smith, C. D. et al. Draft genome of the globally widespread and invasive Argentine ant (Linepithema humile). Proc. Natl. Acad. Sci. 108, 5673–5678 (2011).
Zhou, X. et al. Chemoreceptor evolution in hymenoptera and its implications for the evolution of eusociality. Genome Biol. Evol. 7, 2407–2416 (2015).
Dogantzis, K. A. et al. Insects with similar social complexity show convergent patterns of adaptive molecular evolution. Sci. Rep. 8, 1–8 (2018).
Chung, H.-R., Löhr, U. & Jäckle, H. Lineage-specific expansion of the zinc finger associated domain ZAD. Mol. Biol. Evol. 24, 1934–1943 (2007).
Stapley, J., Santure, A. W. & Dennis, S. R. Transposable elements as agents of rapid adaptation may explain the genetic paradox of invasive species. Mol. Ecol. 24, 2241–2252 (2015).
Schrader, L. et al. Transposable element islands facilitate adaptation to novel environments in an invasive species. Nat. Commun. 5, 5495 (2014).
Su, Y., Huang, Q., Wang, Z. & Wang, T. High genetic and epigenetic variation of transposable elements: Potential drivers to rapid adaptive evolution for the noxious invasive weed Mikania micrantha. Ecol. Evol. 11, 13501–13517 (2021).
Traniello, I. M. et al. Single-cell dissection of a collective behaviour in honeybees. bioRxiv (2022).
McKenzie, S. K., Oxley, P. R. & Kronauer, D. J. C. Comparative genomics and transcriptomics in ants provide new insights into the evolution and function of odorant binding and chemosensory proteins. BMC Genomics 15, 718 (2014).
Acknowledgements
We thank J. Lock, M. Coleman, P. Kennedy, E. Bell, S. Moreno for assistance in locating and/or collecting samples used in this study. The V. mandarinia genome assembly was generated as part of the U.S. Department of Agriculture, Agricultural Research Service (USDA-ARS) Ag100Pest Initiative. We thank the Ag100Pest Team and their collaborators for allowing us to include the assembly in our analysis. We thank the Sumner Lab and M. Blaxter for their helpful comments on earlier drafts. This work was funded by Grants from the UK’s Natural Environment Research Council awarded to S.S. (NE/M012913/2; NE/S011218/1), a Marie Sklodowska-Curie Action Individual fellowship awarded to A.C. (706208-SocParPhenoEvo); F.C.F. and R.G. were supported by the Plataforma de Recursos Biomoleculares y Bioinformáticos PT 13/0001/0021 from ISCIII, a platform co-funded by the European Regional Development Fund (FEDER), from the Spanish Ministry of Science and Innovation to the EMBL partnership, the Centro de Excelencia Severo Ochoa and the CERCA Programme/Generalitat de Catalunya. J.S. was supported by UCL’s Department of Genetics, Evolution and Environment (Harold and Olga Fox Fund). This research was funded in whole, or in part, by the Wellcome Trust Grants 206194 and 218328. We thank our colleagues in the Sanger Institute’s Scientific Operations, Genome Assembly and Genome Reference Informatics teams for extraction, sequencing, assembly and curation and the Director’s Office for support.
Author information
Authors and Affiliations
Contributions
E.F. designed data analyses, analysed the data and wrote the paper; A.C. collected data and wrote the paper; D.Taylor. designed the analysis and collected data; D.Thiery, R.L.B., F.C. and R.C. collected data; N.S.P. performed experiments; F.C.F., M.A.B., R.G., E.P., J.S., R.M., C.D.R.W. and D.M. analysed data; A.B.S analysed data supported by J.C.D., H.H. oversaw experimental work and data analysis; S.S. conceived the project, designed analyses and wrote the paper. All authors approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Favreau, E., Cini, A., Taylor, D. et al. Putting hornets on the genomic map. Sci Rep 13, 6232 (2023). https://doi.org/10.1038/s41598-023-31932-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-31932-x
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.