Abstract
Egg yolk constitutes about a third of the structure of the chicken egg however, the molecular structure and physiological effects of egg yolk-derived lipid membranous vesicles are not clearly understood. In this study, for the first record, the egg yolk nanovesicles (vitellovesicles, VVs) were isolated, characterized, and used as a supplement for porcine embryo culture. Yolks of ten freshly oviposited eggs were filtered and ultracentrifuged at 100,000 × g for 3 h to obtain a pellet. Cryogenic transmission electron microscopy and nanoparticle tracking analysis of the pellet revealed bilipid membranous vesicles. Protein contents of the pellet were analyzed using tandem mass spectrometry and the miRNA content was also profiled through BGISEQ-500 sequencer. VVs were supplemented with the in vitro culture medium of day-7 hatched parthenogenetic blastocysts. After 2 days of blastocyst culture, the embryonic cell count was increased in VVs supplemented embryos in comparison to the non-supplemented embryos. TUNEL assay showed that apoptotic cells were increased in control groups when compared with the VVs supplemented group. Reduced glutathione was increased by 2.5 folds in the VVs supplemented group while reactive oxygen species were increased by 5.3 folds in control groups. Quantitative PCR analysis showed that VVs significantly increased the expression of lipid metabolism-associated genes (monoglyceride lipase and lipase E), anti-apoptotic gene (BCL2), and superoxide dismutase, while significantly reducing apoptotic gene (BAX). Culturing embryos on Matrigel basement membrane matrix indicated that VVs significantly enhanced embryo attachment and embryonic stem cell outgrowths compared to the non-supplemented group. This considers the first report to characterize the molecular bioactive cargo contents of egg yolk nanovesicles to show their embryotrophic effect on mammalian embryos. This effect might be attributed to the protein and miRNA cargo contents of VVs. VVs can be used for the formulation of in vitro culture medium for mammalian embryos including humans.
Similar content being viewed by others
Introduction
Egg yolk constitutes approximately one-third of the structure of the chicken egg and is enriched with essential metabolites and nutrients for chicken embryonic development1. The composition of egg yolk proteins and RNAs has been explored by several reports2,3,4,5. Egg yolk is enriched in proteins, cholesterol, and phospholipids and all of them are embedded inside the high-density and low-density lipoprotein micelles or granular particles6. Although many studies have provided a theoretical basis, research on ultramicroscopic and compositional compartments of yolk vesicles have not been performed so far.
Extracellular vesicles (EVs) including exosomes and microvesicles have been successfully identified and characterized in most of the biological fluids of humans, animals, and plants including plasma, milk, and saliva7,8,9,10. However, no work has been published on the isolation and characterization of egg yolk vesicles, to the best of our knowledge. Conversely, egg yolk lecithin was used to manufacture nanosized liposome vesicles for therapeutic use and drug delivery11,12,13.
The embryonic development of chicken embryos was used as a common paradigm to explore the roles of egg components on the early stages of embryonic development. Avian embryos undergo rapid cleavages during the first days of development, and this raises questions about the involvement of egg yolk components in enhancing embryonic development14. Pigs are considered a great tool for biomaterial production, organ xenotransplantation, and the development of biomedical models15. However, several challenges are facing to establish embryonic stem cells due to the low developmental competence of the in vitro–produced porcine embryos16,17. Pig embryonic stem cell research and the implantation process are important links to exploration and action18,19,20,21,22,23.
In the current work, we introduce the term “vitellovesicles” to describe chicken egg yolk lipid nanovesicles and outline ultramicroscopic features and contents of cargo protein and miRNA. Moreover, we studied the potential embryotrophic effects of these egg yolk vesicles on blastocyst culture, apoptosis, and oxidative stress in the porcine model.
Materials and methods
Chemicals and reagents
Chemicals and reagents were purchased from Sigma-Aldrich (St. Louis, MO, USA) unless otherwise specified.
Isolation and characterization of egg yolk vesicles
Egg yolk of freshly oviposited commercially purchased eggs (n = 10) were aseptically aspirated and diluted 10 times with phosphate-buffered saline (PBS), and then centrifuged at 16,000 × g for 30 min to remove the big protein particles. The clear supernatants were microfiltered through 0.4 µm and 0.2 µm microfilters. The filtrates were subjected to ultracentrifugation at 100,000 × g for 3 h using Optima XPN-100 Ultracentrifuge (Beckman Coulter, Indianapolis, IN, USA)24. VVs pellets were eluted in 50 µL of PBS solution. VVs were characterized through nanoparticle tracking analysis (NTA, NanoSight NS300, Malvern Instruments, Worcestershire, UK)25. 100-nm polystyrene particles (ThermoFisher Scientific, Waltham, MA, USA) were used for the device calibration. 50 µL of eluted VVs was diluted 20 × in 1 × PBS and was loaded into the NTA device. The machine was adjusted to read the mean particle size (diameter), particle concentration, and outlier removal. The reads were set to eleven different positions and two reading cycles per position in each sample. VVs were also visualized through transmission electron microscopy (TEM) and cryogenic TEM (Cryo-TEM)26,27. For TEM, 5 μL of eluted VVs was negatively stained with 2% uranyl acetate, moved onto 200-mesh copper grids, dried, and examined through an OMEGA-energy filtering TEM (ZEISS LEO 912, Carl Zeiss, Jena, Germany) at 120 kV. For Cryo-TEM, 3.5 µL of the eluted VVs was applied to glow-discharged Quantifoil R1.2/1.3 Cu 300 grids (Quantifoil, Jena, Germany) and were flash frozen in liquid ethane using Vitrobot mark IV (Thermo Fisher Scientific) set at 100% humidity and 4 °C for the preparation chamber and 5 s of blot time. Cryo-TEM micrographs were imaged on a Glacios microscope (Thermo Fisher Scientific) operated at an accelerating voltage of 200 kV with a 70-µm C2 aperture at an indicated magnification of 73 k × . A Falcon III direct electron detector (Thermo Fisher Scientific) in linear mode was used to acquire images of the samples with a 100-µm objective aperture. The EVs cargo contents of proteins were analyzed through proteomics as discussed below.
In vitro generation of porcine parthenogenetic embryos
Electric parthenogenetic activation of in vitro matured porcine oocytes was used to generate porcine embryos as showed in previous reports16,17,28. Briefly, ovaries were collected from a local slaughterhouse in Daejeon city, and cumulus-oocyte complexes (COCs) were aspirated. Good quality oocytes (homogenous ooplasm and surrounded by compact layers of cumulus cells) were microscopically selected and washed three times in N-2-hydroxyethylpiperazine-N′-ethane-sulphonic acid (HEPES) buffered Tyrode’s medium comprising of 0.05% polyvinyl alcohol (TLH-PVA). COCs were in vitro matured in 500 mL of a maturation medium in 4-well dishes (Nunc, ThermoFisher Scientific, Roskilde, Denmark). Maturation medium was composed of TCM-199 (Gibco, Waltham, MA, USA) supplemented with 10% (v/v) porcine follicular fluid, 0.6 mM cysteine, 0.91 mM sodium pyruvate, 10 ng/mL epidermal growth factor, 1 μg/mL insulin, 10 IU/mL hormones (human chorionic gonadotrophin and equine chorionic gonadotrophin (Daesung Microbiological Labs, Gyeonggi-do, Korea), and 75 μg/mL kanamycin. COCs were matured for 22 h in this medium and then were transferred to the same culture conditions without the presence of hormones for an additional 22 h. Mature COCs were subjected to hyaluronidase (0.6%), gently pipetted to remove cumulus cells and then washed in TLH-PVA and electrically activated inside a glass chamber of BTX Electro-Cell Manipulator 2001 (BTX Inc., San Diego, CA, USA) connected with two electrodes to transmit a single direct current pulse of 1.5 kV/cm for 60 μs. Activation medium comprised mannitol (0.28 M), calcium chloride (0.1 mM), HEPES (0.5 mM), and magnesium sulfate (0.1 mM). Parthenogenetically activated oocytes were cultured in 25 μL of porcine zygote medium-5 (IFP Co. Ltd., Yamagata, Japan) covered with mineral oil in a humidified atmosphere at 38.5 °C (5% O2, 5% CO2, and 90% N2) for 7 days. Blastocysts were collected and zona pellucida was removed after incubation with 0.1% pronase (w/v in PBS) for 30 s and zona-free blastocysts were used for the experiments.
Embryo attachment and embryonic stem cells outgrowth
Feeder-free culture condition for embryos attachment was used according to a previous report 29. The 4-well dishes (Nunc) were cooled and forty μL of microdrops of Matrigel® (BD Biosciences, San Jose, CA, USA) were placed in each well and then incubated for 30 min at 37 °C. Excess Matrigel was removed and 40 μL of culture medium were placed over the drop and covered with mineral oil. Culture medium comprised Dulbecco’s modified eagle medium/nutrient mixture F-12, fetal bovine serum (10% v/v), 0.1 mM β-mercaptoethanol, 1% nonessential amino acids (Invitrogen Corporation, Carlsbad, CA, USA), and 1% penicillin/streptomycin (100 U/mL penicillin and 100 µg/mL streptomycin). Zona-free blastocysts were washed with the culture medium before placed in the microdrops and cultured in a humidified atmosphere of 5% CO2 at 38.5 °C. On days 2–5 of culture, embryos were checked for attachment and outgrowths. VVs (1.6 × 108/ml) were supplemented with the in vitro culture medium (n = 160, 8 replicates of 10 blastocysts from each control and VVs-supplemented group), while control group was not supplemented with VVs.
TdT-mediated deoxyuridine triphosphate (dUTP-X) nick end labeling (TUNEL) assay
Detection of apoptosis was performed after labeling of DNA strand breaks by using in situ–cell death detection TUNEL assay Kit (Roche Holding AG, Basel, Switzerland). Embryos at the designed days were fixed in 4% paraformaldehyde, permeabilized in 0.1% TritonX, washed in PBS, and then placed in the enzyme (TdT) and a label (fluorescein-dUTP) solutions for 1 h at 37 °C. Vectashield antifade mounting medium was used for nuclear staining. Green fluorescence signals were captured, counted, and calculated using ImageJ software.
Measuring reduced glutathione and reactive oxygen species in embryos
Intracellular levels of reactive oxygen species (ROS) were measured after staining with 2′,7′-dichlorodihydrofluorescein diacetate (H2DCFDA; Invitrogen Corporation, Carlsbad, CA, USA). ROS accumulation was indicated as green, fluorescent signals. Intracellular levels of reduced glutathione (GSH) were measured after staining with CellTracker™ Blue including a blue fluorescent dye 4-chloromethyl-6,8-difluoro-7-hydroxycoumarin (CMF2HC). In each experiment, twenty-five embryos (n = 5 for 5 replicates) from each treatment group were used. Fluorescence signals were visualized through an epifluorescence microscope (Leica DM IRB; Leica Microsystems, Wetzlar, Germany) associated with ultraviolet filters (460 nm for ROS and 370 nm for GSH). Fluorescent images were processed through ImageJ (version 1.41; National Institutes of Health, Bethesda, MD, USA, https://imagej.nih.gov/ij/) to measure the fluorescence intensity after normalization to corresponding control group.
Relative quantitative polymerase chain reaction (RT-qPCR)
RNeasy Micro Kit (Qiagen GmbH, Hilden, Germany) was used to extract total RNA from the embryos (n = 5, 5 replicates). The concentration and quality of RNA was determined with NanoDrop 2000 (Thermo Scientific, Waltham, MA, USA). Reverse transcription was performed through using 2X RT Pre-Mix of QuantiNova Reverse Transcription Kit (Qiagen). Real time polymerase chain reaction (qPCR) was conducted by the CFX Connect Real-Time PCR system (Bio-Rad) and SYBR 2X Real-Time PCR Pre-Mix (BioFACT, Daejeon, Korea). Details about the primers, examined genes, and amplicon size are illustrated in Table 1. Quantification of the target transcripts were compared to those of housekeeping genes (ACTB-mRNA and U6-snRNA) through the ΔΔCt method.
Preparation of protein fraction of VVs and proteomic analysis
Protein isolation and proteomics were conducted according to Lee et al.26. Eluted VVs protein contents were measured through the bicinchoninic acid method. Sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) was used for protein fractionation. Gels were stained in NH4CO3 (10 mM) and acetonitrile (50%) for Coomassie Brilliant Blue staining30, then rinsed twice with distilled water and acetonitrile (100%). Gels were then dried in a speed vacuum evaporator and then treated with NH4HCO3 (100 mM) and dithiothreitol (10 mM) at 56 °C, before treated with 100 nM iodoacetamide to reduce alkylate S–S bridges. Gels were then washed in distilled water and then dried in a speed vacuum evaporator. Gels were kept for 12–16 h in trypsin (10 ng/mL) and NH4HCO3 (50 mM) at 37 °C for protein digestion. The resulting peptides were retrieved after treatment and suspension in 0.5% trifluoroacetic acid, purified, and concentrated through MGU30-C18 trapping columns (LC Packings). MS and MS/MS spectra were captured through LTQ-Velos ESI ion trap mass spectrometer (Thermo Scientific, Waltham, MA). The data-dependent mode was used for fragmentation of ten most abundant peaks from the full MS scan with 27% normalized collision energy. The eluted peptides were made to enter into the mass spectrometer at an electrospray voltage of 2.3 kV. The maximum ion injection times used were 100 ms for the MS scan and 100 ms for the MS/MS scans. The automatic gain control target settings were 5.0 × 104 for the MS scan mode and 1.0 × 105 for the MS/MS scan mode. The dynamic exclusion duration was set at 180 s and exclusion mass width 0.5 Da. The mass range for acquiring MS spectra was 150–2000 m/z. Protein identification were performed by MASCOT 2.4 with the false discovery rate at 1%, as a cutoff value. Relative quantitation of proteins was determined through the exponentially modified protein abundance index (emPAI) and were expressed as mol %. Protein functions and gene ontology (GO) were analyzed a web-based tool provided by DAVID (NIAID/NIH; https://david.ncifcrf.gov/)31,32. GO functional analysis was performed using the egg yolk vesicles proteomic data, 448 proteins accessions were process for functional analysis. DAVID database (https://david.ncifcrf.gov/) was used to convert the accessions into official names, and further used for GO functional annotation of the target genes. This was used for identification of biological process (BP), molecular function (MF), and cellular component (CC) and pathway. Bioinformatics tools were used for the construction of bubble plots (http://www.bioinformatics.com.cn/plot_basic_gopathway_enrichment_bubbleplot_081_en). Protein–protein interaction was performed through using String database. The generated network was further processed using String plugin in Cytoscape using confidence score cutoff (0.4).
Small RNA sequencing and bioinformatic analysis
Transcriptome library construction and sequencing
The total RNA of VVs was isolated from three random samples as mentioned above and integrity and concentration of collected small RNAs (sRNAs) were evaluated through Agilent 2100 Bioanalyzer (Agilent RNA 6000 Nano Kit, Agilent Technologies, Santa Clara, CA, USA). sRNA sequencing libraries were executed following the BGISEQ-500 Small RNA unique molecular identifier (UMI) Library Construction Protocol. Each 1 ng of sRNA sample was adaptor-ligated in consequential steps: connection of the 3´-adaptor system at 70 °C for 2 min and 25 °C for 2 h, then UMI-labeled primers were added at 65 °C for 2 min, ramp to 4 °C at a rate of 0.3 °C/s. Finally, a 5´-adaptor system was added at 70 °C for 2 min, 25 °C for 1 h. After adaptors ligation, the first strand master mix was mixed with SuperScript II (Invitrogen Corporation) for reverse transcription at 42 °C for 1 h followed by enzyme inactivation at 70 °C for 15 min. cDNA was amplified through PCR using primers and PCR master mix in the following cycling conditions: initial 95 °C / 3 min, 20 cycles of 98 °C / 20 s, 56 °C / 15 s, 72 °C / 15 s, and final extension at 72 °C /10 min. PAGE gel electrophoresis was used to purify the final PCR products and then were dissolved in an elution buffer. The resulting PCR product was heat-denatured and circularized by the split oligo sequence to be single-stranded circle DNAs (ssCir DNA) that were considered as the final library. Constructed libraries were analyzed through the Agilent 2100 Bioanalyzer and then were amplified with phi29 to make DNA nanoballs (DNBs) that contain > 300 copies per ball. The DNBs were transferred into a patterned nanoarray and single end 50 bases reads were generated during sequencing by synthesis. BGISEQ-500 sequencer was used for the RNA-Seq libraries and base-calling was conducted by its accompanied software (version #0.3.8.111).
Sequencing data analysis
RNA-seq libraries were checked for quality through FastQC v0.11.5 software (https://www.bioinformatics.babraham.ac.uk/projects/fastqc/). The raw reads that contain the sequence of the primers/adaptors, high content (10%) of unknown bases (N), and low-quality reads (≤ 5 bases) were identified as ‘dirty’ reads and were excluded to reduce data noise. The remaining “clean” reads were stored in the FASTQ format and mapped to reference Gallus genome (NCBI: bGalGal1.mat.broiler.GRCg7b), and small RNA sequences from miRBase (version 22.1, https://www.mirbase.org/) with mirDeep2 software (miRDeep2.0.0.8, https://github.com/Drmirdeep/miRDeep2). Read numbers of known and predicted novel miRNAs and small RNAs of each sample were counted Using miRDeep2.pl script. Moreover, the ortholog miRNAs from human, pig, rat, and mouse were explored and compared with the resultant chicken miRNAs.
Target genes and GO functional analysis
Using (http://www.mirdb.org/mirdb/ontology.html) database, target genes based on ontology for each ortholog were identified. Target genes with score ≥ 99 were considered for the analysis. Only 25 orthologs were considered for the matured miRNA, and 4 orthologs for Novel mi-RNA. Other miRNA orthologs that have target genes with less score were not considered. DAVID database was further used for GO functional annotation of the target genes. This was used for identification of biological process (BP), molecular function (MF), and cellular component (CC) and pathway. Bioinformatics tools were used for the construction of bubble plots (http://www.bioinformatics.com.cn/plot_basic_gopathway_enrichment_bubbleplot_081_en).
Statistical analyses
Samples homogeneity and normality of variance distribution were validated through Lieven’s test and Kolmogorov–Smirnov test, respectively. Values were presented as mean ± standard error of means (SEM) and analyzed through an unpaired Student’s t-test or with univariate analysis of variance (ANOVA) followed by Tukey’s multiple comparison test. Statistical analyses were conducted with GraphPad Prism 5 (GraphPad Software Inc., San Diego, CA, USA, https://www.graphpad.com/) and differences were considered significant when p < 0.05 or p < 0.01. was used for statistical analyses.
Results
Isolation, characteristics, and protein cargo contents of VVs
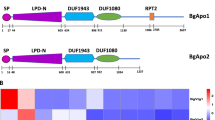
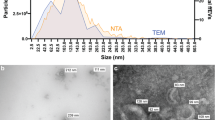
NTA results showed the existence of 1.6 × 1010 particles/mL with mean particle size 158.6 ± 20.5 nm in the isolated VVs. Furthermore, Cryo-TEM images indicated the presence of lipid bilayer vesicles (Fig. 1). The protein contents of isolated VVs were analyzed using proteomics and 498 protein reads were identified (Supplementary Table 1), of which, the top twenty proteins accounted around 83.5% of the total proteins of VVs, including proteins associated with different cellular functions such as protein (albumin, vitellogenins, ovalbumin, fibrinogen, angiotensinogen, transthyretin, and insulin-like growth factor binding protein) and lipid metabolism (apovitellenin-1, apolipoprotein A–I, apolipoprotein B, and lipocalin) in addition to the protective immune proteins (lysozyme C, ovoinhibitor, and complementary system) and immunoglobulins (Table 2, and Supplementary Figures S1–S3). Moreover, the entire proteins showed involvement in different cellular components and biological processes such as protein and lipid metabolism and signaling as well as the exosomal structures (Fig. 2). Furthermore, oxidative defense proteins were also expressed in VVs such as thioredoxin domain-containing protein (accession # A0A1D5PLH7). Importantly, proteomics of VVs revealed exosomes-related tetraspanins and pentaspan transmembrane glycoprotein such as CD81 (accession # F1NW06), heat shock protein family A (Hsp70) member 8 (HSPA8; accession # A0A1D5PFJ6), cystatin C (accession #A0A1L1S0C3), annexin A6 (accession #A0A1D5PY67), and prominin or CD133 (accession # A0A3Q2U8I8) (Supplementary Table S1).
Isolation and characterization of VVs. VVs were isolated by the targeted nanofiltration method and were characterized using NanoSight nanoparticle tracking analysis and showed an average diameter of 158.6 ± 20.5 nm with a concentration of 1.1 × 108 particles/mL (dilution factor is 20 ×) (A). VVs were visualized by transmission electron microscopy (B), and cryo-transmission electron microscope (C) and showed bilipid vesicles (yellow arrows). The scale bar = 200 nm.
GO functional analysis and protein–protein interaction (PPI)
Using the converted accession IDs, the proteins were utilized for the functional analysis from DAVID. The GO-BP analysis shows that, the genes are highly enriched in negative regulation of peptidase activity (GO:0,010,466), and negative regulation of endopeptidase activity (GO:0,010,951). GO-CC are enriched in extracellular region (GO:0,005,576), and extracellular region part (GO:0,044,421). While GO-MF from our data shows enrichment in endopeptidase and serine type endopeptidase inhibitor activity (GO:0,004,866), and peptidase regulator activity (GO:0,061,134) as shown in Supplementary Figure SF1. Pathway analysis was found to be significantly enriched in phagosome (gga04145) and regulation of actin cytoskeleton (gga04810) among others as shown in (Supplementary Figure SF2). The ClueGo functional analysis from the Cytoscape shows the highly enriched BP, CC, and MF from the PPI network. The enriched BP include blood coagulation, peptidase activity, and homotypic cell–cell adhesion among others. While highly enriched CC include extracellular space, and cell adhesion molecule binding among others. MF include plasma lipoprotein particles, and extracellular exosome as shown in (Supplementary Figure SF1). pathway analysis was performed by integrating the KEGG, reactome and WikiPathway shows phagosome and glycolysis/gluconeogenesis as highly enriched pathways as shown in (Supplementary Figure SF3).
MicroRNA contents of VVs
The miRNA proportion was around 4% of the whole detected RNAs (Supplementary Figure SF4). Among the 25,653,000 raw tags, we clarified around 14,138,572 reads of average base pair length 19 bp and after exclusion of all reads less than 17 bp, we identified 10,479 mapped miRNAs according to miRBase 22.1 database (Supplemental Tables S2 and S3). Among the identified siRNAs, around 71 mature miRNA and 186 novel mature miRNAs were detected in VVs (Fig. 3). Twenty miRNAs were expressed in all analyzed samples, while still there are some variations in the detected sequences in the analyzed samples. Among these commonly expressed miRNAs, the top 10 miRNAs by reads count were gga-miR-92-3p, gga-miR-21-5p, gga-miR-199-5p, gga-miR-22-5p, gga-miR-7, gga-miR-31-5p, gga-miR-181a-5p, gga-miR-429-5p, gga-miR-1416-5p, and gga-miR-26a-5p, that showed similar ortholog with pigs miRNA library except for the gga-miR-1416-5p that showed no similarity with the pig or with other species. These top 10 miRNA occupy around 66.31 ± 5.2% from the total reads of the mature miRNAs in VVs. Moreover, from these mature miRNAs, 21 miRNAs were found to affect the embryonic and stem cell development in different mechanisms such as embryo implantation and viability as illustrated in Table 3. Interestingly, these miRNAs are of similar ortholog with pigs (Sus scrofa) and humans (Homo sapiens) according to miRBase 22.1 database. The known miRNAs in Homo sapiens, (hsa, 2,656), Gallus (gga, 1,235), and Sus scrofa (ssc, 457) (Supplementary Tables S4–S6). Interestingly, data indicated the presence of 186 novel miRNA, and of these, seven are matched to the sequenced database of porcine, human, and mouse species. Additionally, 32 novel miRNA were similar to the human miRNA database, three were similar to mouse (Mus musculus, mmu) sequences, and two were similar to rat (Rattus norvegicus, rno) sequence (Supplementary Table S7).
miRNAs target genes and GO functional analysis
Using the 25 miRNAs from the matured miRNA ortholog. Total of 306 target genes were identified. hsa-miR-30a-5p has the highest number target genes. For the novel miRNA data, 71 target genes were identified from the 4 miRNAs ortholog with hsa-miR-23a-3p has the highest number of target genes. For the matured miRNA data, the GO-BP analysis shows that, the target genes are highly enriched in nervous system development (GO:0,007,399), and positive regulation of cell metabolic process (GO:0,009,893). GO-CC are enriched in cytosol (GO:0,005,829), and nucleoplasm (GO:0,005,654). While GO-MF from our data shows enrichment in sequence-specific double-stranded DNA binding (GO:1,990,837), and double-stranded DNA binding (GO:0,007,399) as shown in Supplementary Figure SF5. On the other hand, the novel miRNA GO-BP was found highly enriched in phosphate-containing compound metabolic process (GO:0,006,796). The GO-CC is enriched in chromatin (GO:0,000,785). While the GO-MF is found to be significantly enriched phosphoric ester hydrolase activity (GO:0,042,578) among others as shown in Supplementary Figure SF6.
Effect of VVs on embryo attachment and outgrowths
VVs supplementation for 1.5 day increased embryonic cell number in comparison to the whole egg yolk supplementation of either 10% or 5% and the control non-supplemented group (Fig. 4). Moreover, VVs significantly reduced the apoptosis incidence in day-2 cultured embryos (Fig. 5). Embryo development and outgrowths were daily checked for five subsequent days in VVs supplemented or control groups and VVs increased the embryonic attachment, embryonic cell outgrowths, and cell numbers, and reduced the apoptotic cells (Fig. 6).
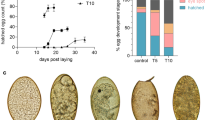
The effects of egg yolk and egg yolk vesicles (VVs) on porcine embryo development. (A) Day-7 zona-free embryos (n = 20, 3 replicates) were cultured in micro drops of culture medium in a humidified atmosphere of 5% CO2 for 48 h. The control group was cultured in a plain culture medium and the groups were supplemented with egg yolk 10% and 5% and VVs of 2.6 × 106 particles/mL. Scale bar = 100 µm. All groups were imaged in a bright field before staining with 4′,6-diamidino-2-phenylindole stain. (B) Graph shows the cell number in the groups, and values were compared using ANOVA followed by Tukey’s test to determine the difference among the groups. Values (mean ± SD) denoted by asterisks (*, **, and ***) were considered statistically different (p < 0.05).
Effects of VVs on porcine embryo apoptosis. (A) Day-7 zona-free embryos (n = 20, 3 replicates) were cultured on in micro drops of culture medium in a humidified atmosphere of 5% CO2 for 48 h. VVs of 2.6 × 106 particles/mL were supplemented to treatment group and compared with the control non-supplemented group. All groups were examined through TUNEL assay and contrasted with 4′,6-diamidino-2-phenylindole stain. Apoptotic nuclei showed green fluorescence signals (yellow arrows). Scale bar = 100 µm. (B) The graph shows the apoptosis percentage in the two groups, and values were compared using Student’s t-test. Value (mean ± SD) denoted by an asterisk (*) was considered statistically different (p < 0.05).
Effects of VVs on porcine embryo attachment and development. (A) Day-7 zona-free embryos (n = 20, 3 replicates) were cultured on Matrigel-coated dishes in micro drops of culture medium in a humidified atmosphere of 5% CO2 for 72 h. The control group was cultured in a plain culture medium and the VVs groups was supplemented with VVs of 2.6 × 106 particles/mL. All groups were imaged in a bright field before staining with TUNEL assay and contrasted with 4′,6-diamidino-2-phenylindole stain. Scale bar = 100 µm. (B, C, and D) Graphs show the attachment percentage, the cell number in the groups, and the apoptosis percentages, respectively. Values (mean ± SD) were compared using Student’s t-test. Values denoted by an asterisk (*) were considered statistically different (p < 0.05).
Effect of VVs on oxidative stress and gene expression of embryos
Supplementation of VVs with the embryo culture medium significantly increased GSH contents and reduced ROS generation (Fig. 7). Furthermore, the expressions of BCL2, SOD1, LIPE, and MGL in the embryos supplemented with VVs were significantly increased (Fig. 8). Conversely, the expression of BAX and BAX/BCL2 ratio were significantly reduced with the supplementation of VVs.
Effect of VVs on antioxidative status of embryos. (A) Reduced glutathione (GSH) was detected with CMF2HC stain and expressed as blue fluorescence, while reactive oxygen species (ROS) were stained with H2DCFDA and expressed as green fluorescence (B). (C and D) graphs show the comparison between the fluorescence intensity and values (mean ± SD) compared with Student’s t-test. Values denoted by an asterisk (*) were considered statistically different (p < 0.05).
Relative quantitative analysis (RT-qPCR) of mRNA transcripts expressed in the embryos treated with VVs. Five blastocysts from each group of four replicates were used for qPCR analysis. The means were normalized to the control group and expressed as arbitrary units (fold change). Data are expressed as mean ± SD and the difference between the two groups was compared using Student’s t-test. Values denoted by an asterisk (*) are considered statistically significant (p < 0.05). House-keeping genes ACTB and U6 were used for quantification of the target transcripts through the ΔΔCt method.
Discussion
For the first time, we successfully isolated and characterized the protein and miRNA contents of bilipid-layer nanovesicles of the chicken egg yolk “vitellovesicles (VVs)” and revealed their embryotrophic effects on porcine embryos. VVs contain exosome and microvesicle markers such as CD81, prominin, annexin A6, cystatin C, albumin, and Hsp7033,34,35,36. VVs contain several proteins and mature miRNAs that are essential for supporting early embryonic development.
Albumin from human and bovine sources has been used extensively for manufacturing embryo culture media in different animal species as a source of protein37,38,39. In addition, some other proteins were found and possess some embryotrophic effects and facilitate the immune and metabolic activities of the embryos such as immunoglobulin, apolipoprotein B, and IGF-binding proteins40. Moreover, VVs contain bone morphogenetic protein 2, retinol-binding protein, and exosome cargo such as fibronectin and CD81 that can support embryonic development41,42,43,44.
Some contents of VVs were linked to lipid metabolism such as apovitellenin (a major very-low-density lipoprotein component)45 and apolipoproteins46. Pre-implantation mammalian embryos were proved to actively utilize, synthesize, and store lipids as lipid droplets47 and therefore, activation of lipases will trigger the mobilization of accumulated lipids in response to different metabolic requirements48,49. Our results indicated an increase in MGLL and LIPE, hormone-sensitive type mRNA expression that subsequently promote lipid metabolism and enhance cell proliferation, survival, and developmental competence of embryos16,50,51,52,53.
The redox defense is also expressed in the cargo contents of VVs. Thioredoxin is a protein disulfide oxidoreductase that transfers two electrons and two protons to oxidize two cysteine thiol groups to a disulfide and a dithiol54. Moreover, the presence of ovotransferrin, which possesses antioxidant activity in addition to its role in iron metabolism55, explains the improved oxidative defense (increased GSH and SOD expression) and reduced ROS in the current results. Additionally, glutathione peroxidase 3 was also found in VVs as a major player in hydrogen peroxide catabolic process and oxidative stress defense56. Moreover, the increase in the antiapoptotic BCL2 mRNA expression and decrease in the apoptotic Bax expression and Bax/Bcl2 ratio explain the reduced apoptosis in the embryos supplemented by VVs. Taken together, the proteomics data indicate the involvement of the VVs protein in inhibiting peptidases, and endopeptidases, and the regulation of actin cytoskeleton which all protects the cells against the apoptosis induction57,58,59.
Tetraspanins and transgelin were also expressed in the cargo of VVs and this protein is a TGFβ signaling pathway that is essential in stem cell differentiation60 and importantly required for trophoblast implantation by promoting actin polymerization61, which can interpret the improvement in embryo attachment after supplementation of VVs.
The mature miRNAs present in VVs are of interest because we were able to define 21 of them as potential target mRNAs, which are essential for embryo development, cell metabolism, and differentiation of stem cells (References 64–87 in Table 3). Interestingly, unlike results of Wade et al.5 who detected only four miRNAs such as gga-miR-30c-5p, gga-miR-99a-5p, gga-miR-92-3p, and gga-miR-100-5p, we found 10 miRNAs expressed in VVs such gga-miR-92-3p, gga-miR-22-5p, gga-miR-26a-5p, gga-miR-100-5p, gga-miR-10a-5p, gga-miR-99a-5p, gga-miR-30c-5p, gga-let-7f.-5p, gga-miR-10b-5p, and gga-miR-204 (Table 3). Although, we found some miRNAs that are commonly expressed in the profile of the analyzed samples (Fig. 3), there is still some variability that might be due to the individual variations5 or the sequencing method62.
Interestingly, the ultrastructure of egg yolk LDL and HDL micelles were recently evaluated through TEM, cryogenic TEM, and the phosphorus-31 nuclear magnetic resonance and yielded small rounded or irregular particles of 17 to 200 nm in size63,64,65,66,67.
Conclusion
We provide a detailed method for the isolation and molecular cargo contents of proteins and miRNA in addition to the functional characterization of egg yolk vesicles and their embryotrophic effects on the porcine embryo model. These VVs can be used for the formulation of an in vitro–culture medium for supporting different mammalian embryos development and could be engineered and used as a vehicle for nucleic acids for therapeutic purposes.
Data availability
The data supporting the findings of this study are included in the manuscript. Proteomics data generated during the current study are available in ProteomeXchange dataset PXD038679 [doi:@10.25345/C5RV0D524]. RNA sequencing dataset generated during the current study is available in the NCBI GEO (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE219218).
Abbreviations
- COC:
-
Cumulus-oocyte complex
- GSH:
-
Reduced glutathione
- HEPES:
-
N-2-hydroxyethylpiperazine-N′-ethane-sulphonic acid
- IVM:
-
In vitro maturation
- miRNA:
-
Micro-RNA
- mRNA:
-
Messenger RNA
- NTA:
-
Nanoparticles tracking analysis
- PBS:
-
Phosphate-buffered saline
- PCR:
-
Polymerase chain reaction
- PPI:
-
Protein–protein interaction
- PVA:
-
Polyvinyl alcohol
- RNA-seq:
-
RNA sequencing
- ROS:
-
Reactive oxygen species
- sRNAs:
-
Small RNAs
- ssCir DNA:
-
Single-stranded circle DNAs
- TEM:
-
Transmission electron microscopy
- TUNEL:
-
TdT-mediated deoxyuridine triphosphate (dUTP-X) nick end labeling
- VVs:
-
Vitellovesicles
References
Schechtman, A. M. & Knight, P. F. Transfer of proteins from the yolk to the chick embryo. Nature 176(4486), 786–787 (1955).
Zhu, W., Zhang, J., He, K., Geng, Z. & Chen, X. Proteomic analysis of fertilized egg yolk proteins during embryonic development. Poult. Sci. 99(5), 2775–2784 (2020).
Mann, K. & Mann, M. The chicken egg yolk plasma and granule proteomes. Proteomics 8(1), 178–191 (2008).
Laca, A., Paredes, B., Rendueles, M. & Díaz, M. Egg yolk granules: Separation, characteristics and applications in food industry. LWT Food Sci. Technol. 59(1), 1–5 (2014).
Wade, B., Cummins, M., Keyburn, A. & Crowley, T. M. Isolation and detection of microRNA from the egg of chickens. BMC Res. Not. https://doi.org/10.1186/s13104-016-2084-5 (2016).
Abeyrathne, E. D. N. S., Nam, K.-C., Huang, X. & Ahn, D. U. Egg yolk lipids: Separation, characterization, and utilization. Food Sci. Biotechnol. 31(10), 1243–1256 (2022).
Babaker, M. A. et al. The therapeutic potential of milk extracellular vesicles on colorectal cancer. Int. J. Mol. Sci. 23(12), 6812 (2022).
Théry, C. et al. Minimal information for studies of extracellular vesicles 2018 (MISEV2018): A position statement of the international society for extracellular vesicles and update of the MISEV2014 guidelines. J. Extracell. Vesicles https://doi.org/10.1080/20013078.2018.1461450 (2018).
Ruf, A., Oberkofler, L., Robatzek, S. & Weiberg, A. Spotlight on plant RNA-containing extracellular vesicles. Curr. Opin. Plant Biol. https://doi.org/10.1016/j.pbi.2022.102272 (2022).
Alzahrani, F. A. & Saadeldin, I. M. Role of Exosomes in biological communication systems (Springer, 2021).
Kondratowicz, A. et al. Egg yolk extracts as potential liposomes shell material: Composition compared with vesicles characteristics. J. Food Sci. 83(10), 2527–2535 (2018).
Lupi, G. A. et al. Development of low-cost cage-like particles to formulate veterinary vaccines. Vet. Immunol. Immunopathol. https://doi.org/10.1016/j.vetimm.2022.110460 (2022).
Tang, Z. et al. nanovector assembled from natural egg yolk lipids for tumor-targeted delivery of therapeutics. ACS Appl. Mater. Interfaces 12(7), 7984–7994 (2020).
Hamburger, V. & Hamilton, H. L. A series of normal stages in the development of the chick embryo. Dev. Dyn. 195(4), 231–272 (1992).
Rafat, M. et al. Bioengineered corneal tissue for minimally invasive vision restoration in advanced keratoconus in two clinical cohorts. Nat. Biotechnol. https://doi.org/10.1038/s41587-022-01408-w (2022).
Fang, X. et al. Oviduct epithelial cells-derived extracellular vesicles improve preimplantation developmental competence of in vitro produced porcine parthenogenetic and cloned embryos. Mol. Reprod. Dev. 89(1), 54–65 (2021).
Tanga, B. M. et al. MiRNA-155 inhibition enhances porcine embryo preimplantation developmental competence by upregulating ZEB2 and downregulating ATF4. Theriogenology 183, 90–97 (2022).
Brevini, T. A., Vassena, R., Francisci, C. & Gandolfi, F. Role of adenosine triphosphate, active mitochondria, and microtubules in the acquisition of developmental competence of parthenogenetically activated pig oocytes. Biol. Reprod. 72(5), 1218–1223 (2005).
Kim, H. S. et al. Isolation and initial culture of porcine inner cell masses derived from in vitro-produced blastocysts. Zygote (Cambridge, England). 15(1), 55–63 (2007).
Miyoshi, K., Taguchi, Y., Sendai, Y., Hoshi, H. & Sato, E. Establishment of a porcine cell line from in vitro-produced blastocysts and transfer of the cells into enucleated oocytes. Biol. Reprod. 62(6), 1640–1646 (2000).
Yang, H. et al. Generation of genetically modified mice by oocyte injection of androgenetic haploid embryonic stem cells. Cell 149(3), 605–617 (2012).
Liu, Q. et al. Karyotype characterization of in vivo- and in vitro-derived porcine parthenogenetic cell lines. PLoS ONE 9(5), e97974 (2014).
Fang, X. et al. Vitamin C enhances porcine cloned embryo development and improves the derivation of embryonic stem-like cells. Reprod. Biol. 22(2), 100632 (2022).
Abumaghaid, M. M., Abdelazim, A. M., Belali, T. M., Alhujaily, M. & Saadeldin, I. M. Shuttle transfer of mRNA transcripts via extracellular vesicles from male reproductive tract cells to the cumulus-oocyte complex in rabbits (oryctolagus cuniculus). Front. Vet. Sci. 9, 81 (2022).
Mehdiani, A. et al. An innovative method for exosome quantification and size measurement. J. V. Exp. 95, e50974 (2015).
Lee, H. et al. Streptococcus equi-derived extracellular vesicles as a vaccine candidate against streptococcus equi infection. Vet. Microbiol. 259, 109165. https://doi.org/10.1016/j.vetmic.2021.109165 (2021).
Saadeldin, I. M., Kim, S. J., Choi, Y. B. & Lee, B. C. Improvement of cloned embryos development by co-culturing with parthenotes: A possible role of exosomes/microvesicles for embryos paracrine communication. Cell. Reprogram. 16(3), 223–234 (2014).
Saadeldin, I. M., Kim, S. J., Choi, Y. B. & Lee, B. C. Post-maturation zona perforation improves porcine parthenogenetic trophoblast culture. Placenta 35(4), 286–288 (2014).
Saadeldin, I. M., Kim, S. J. & Lee, B. C. Blastomeres aggregation as an efficient alternative for trophoblast culture from porcine parthenogenetic embryos. Dev. Growth Differ. 57(5), 362–368 (2015).
Lee, S.-Y. et al. Analysis of the extracellular proteome of colistin-resistant Korean acinetobacter baumannii strains. ACS Omega 5(11), 5713–5720 (2020).
Saadeldin, I. M. et al. Thermotolerance and plasticity of camel somatic cells exposed to acute and chronic heat stress. J. Adv. Res. 22, 105–118 (2020).
Sherman, B. T. et al. DAVID: A web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res. 50(W1), W216–W221 (2022).
Pammi Guru, K. T., Sreeja, J. S., Dharmapal, D., Sengupta, S. & Basu, P. K. Novel gold nanoparticle-based quick small-exosome isolation technique from serum sample at a low centrifugal force. Nanomaterials 12(10), 1660 (2022).
Kang, M., Kim, S. & Ko, J. Roles of CD133 in microvesicle formation and oncoprotein trafficking in colon cancer. FASEB J. 33(3), 4248–4260 (2018).
Veschi, E. A. et al. Localization of annexin A6 in matrix vesicles during physiological mineralization. Int. J. Mol. Sci. 21(4), 1367 (2020).
Ghidoni, R. et al. Cystatin C is released in association with exosomes: A new tool of neuronal communication which is unbalanced in Alzheimer’s disease. Neurobiol. Aging 32(8), 1435–1442 (2011).
Bungum, M., Humaidan, P. & Bungum, L. Recombinant human albumin as protein source in culture media used for IVF: A prospective randomized study. Reprod. Biomed. Online 4(3), 233–236 (2002).
Saadeldin, I. M., Kim, B., Lee, B. & Jang, G. Effect of different culture media on the temporal gene expression in the bovine developing embryos. Theriogenology 75(6), 995–1004 (2011).
Morbeck, D. E. et al. Composition of protein supplements used for human embryo culture. J. Assist. Reprod. Genet. 31(12), 1703–1711 (2014).
Dyrlund, T. F. et al. Unconditioned commercial embryo culture media contain a large variety of non-declared proteins: A comprehensive proteomics analysis. Hum. Reprod. 29(11), 2421–2430 (2014).
Larson, R. C., Ignotz, G. G. & Currie, W. B. Effect of fibronectin on early embryo development in cows. Reproduction 96(1), 289–297 (1992).
Kam, K. T. et al. Retinoic acid synthesis and functions in early embryonic development. Cell Biosci. 2(1), 14 (2012).
Schubert, M. & Gibert, Y. Retinoids in embryonic development. Biomolecules 10(9), 1278 (2020).
Halloran, D., Durbano, H. W. & Nohe, A. Bone morphogenetic protein-2 in development and bone homeostasis. J. Dev. Biol. 8(3), 19 (2020).
MacLachlan, I., Steyrer, E., Hermetter, A., Nimpf, J. & Schneider, W. J. Molecular characterization of quail apolipoprotein very-low-density lipoprotein II: Disulphide-bond-mediated dimerization is not essential for inhibition of lipoprotein lipase. Biochem. J. 317(2), 599–604 (1996).
Ramasamy, I. Recent advances in physiological lipoprotein metabolism. Clin. Chem. Lab. Med. (CCLM). https://doi.org/10.1515/cclm-2013-0358 (2014).
Aizawa, R. et al. Synthesis and maintenance of lipid droplets are essential for mouse preimplantation embryonic development. Development https://doi.org/10.1242/dev.181925 (2019).
Zimmermann, R. et al. Fat mobilization in adipose tissue is promoted by adipose triglyceride lipase. Science 306(5700), 1383–1386 (2004).
Olzmann, J. A. & Carvalho, P. Dynamics and functions of lipid droplets. Nat. Rev. Mol. Cell Biol. 20(3), 137–155 (2018).
Torres, V. et al. Resveratrol-cyclodextrin complex affects the expression of genes associated with lipid metabolism in bovine in vitro produced embryos. Reprod. Dom. Anim. 53(4), 850–858 (2018).
Agirregoitia, E. et al. Dynamic of expression and localization of cannabinoid-degrading enzymes FAAH and MGLL in relation to CB1 during meiotic maturation of human oocytes. Cell Tissue Res. 365(2), 393–401 (2016).
Hu, R. et al. MiR-202–3p determines embryo viability during mid-blastula transition. Front. Cell Dev. Biol. https://doi.org/10.3389/fcell.2022.897826 (2022).
Mau, K. H. T. et al. Dynamic enlargement and mobilization of lipid droplets in pluripotent cells coordinate morphogenesis during mouse peri-implantation development. Nat. Commun. https://doi.org/10.1038/s41467-022-31323-2 (2022).
Holmgren, A. Thioredoxin structure and mechanism: Conformational changes on oxidation of the active-site sulfhydryls to a disulfide. Structure 3(3), 239–243 (1995).
Wu, J. & Acero-Lopez, A. Ovotransferrin: Structure, bioactivities, and preparation. Food Res. Int. 46(2), 480–487 (2012).
Ufer,. The roles of glutathione peroxidases during embryo development. Front. Mol. Neurosci. https://doi.org/10.3389/fnmol.2011.00012 (2011).
Desouza, M., Gunning, P. W. & Stehn, J. R. The actin cytoskeleton as a sensor and mediator of apoptosis. BioArchitecture 2(3), 75–87 (2012).
Zhivotovsky, B., Burgess, D. H. & Orrenius, S. Proteases in apoptosis. Experientia 52(10–11), 968–978 (1996).
Egger, L. et al. Serine proteases mediate apoptosis-like cell death and phagocytosis under caspase-inhibiting conditions. Cell Death Differ. 10(10), 1188–1203 (2003).
Elsafadi, M. et al. Transgelin is a TGFβ-inducible gene that regulates osteoblastic and adipogenic differentiation of human skeletal stem cells through actin cytoskeleston organization. Cell Death Dis. 7(8), e2321 (2016).
Liang, X. et al. Transgelin 2 is required for embryo implantation by promoting actin polymerization. FASEB J. 33(4), 5667–5675 (2019).
Shi, H. et al. Bias in RNA-seq library preparation: Current challenges and solutions. Biomed. Res. Int. 2021, 1–11 (2021).
Wakamatsu, H., Handa, A. & Chiba, K. Observations using Phosphorus-31 nuclear magnetic resonance (31P-NMR) of structural changes in freeze-thawed hen egg yolk. Food Chem. 244, 169–176 (2018).
Zhou, M., Hu, Q., Wang, T., Xue, J. & Luo, Y. Characterization of high density lipoprotein from egg yolk and its ability to form nanocomplexes with chitosan as natural delivery vehicles. Food Hydrocoll. 77, 204–211 (2018).
Anton, M. High-density lipoproteins (HDL) or lipovitellin fraction (Bioactive Egg Compounds, 2007).
Bellairs, R. The structure of the yolk of the hen’s egg as studied by electron microscopy. J. Biophys. Biochem. Cytol. 11(1), 207–225 (1961).
Anton, M. et al. Chemical and structural characterisation of low-density lipoproteins purified from hen egg yolk. Food Chem. 83(2), 175–183 (2003).
Fang, F. et al. MicroRNAs secreted by human embryos could be potential biomarkers for clinical outcomes of assisted reproductive technology. J. Adv. Res. 31, 25–34 (2021).
Zhou, F. et al. miRNA-122-5p stimulates the proliferation and DNA synthesis and inhibits the early apoptosis of human spermatogonial stem cells by targeting CBL and competing with lncRNA CASC7. Aging 12(24), 25528–25546 (2020).
Krawczynski, K., Najmula, J., Bauersachs, S. & Kaczmarek, M. M. MicroRNAome of porcine conceptuses and trophoblasts: expression profile of microRNAs and their potential to regulate genes crucial for establishment of pregnancy1. Biol. Reprod. https://doi.org/10.1095/biolreprod.114.123588 (2015).
de Alcântara-Neto, A. S. et al. Oviductal extracellular vesicles enhance porcine in vitro embryo development by modulating the embryonic transcriptome. Biomolecules 12(9), 1300 (2022).
Rosenbluth, E. M., Shelton, D. N., Wells, L. M., Sparks, A. E. & Van Voorhis, B. J. Human embryos secrete microRNAs into culture media–a potential biomarker for implantation. Fertil Steril. 101(5), 1493–1500 (2014).
Acuña-González, R. J. et al. MiR-191–5p is upregulated in culture media of implanted human embryo on day fifth of development. Reprod. Biol. Endocrinol. https://doi.org/10.1186/s12958-021-00786-1 (2021).
Nagosa, S. et al. microRNA-184 Induces a commitment switch to epidermal differentiation. Stem Cell Rep. 9(6), 1991–2004 (2017).
Zhang, J. et al. MiR-202-5p is a novel germ plasm-specific microRNA in zebrafish. Sci. Rep. 7(1), 7055 (2017).
Li, Z., Jia, J., Gou, J., Zhao, X. & Yi, T. MicroRNA-451 plays a role in murine embryo implantation through targeting Ankrd46, as implicated by a microarray-based analysis. Fertil. Steril. 103(3), 834–44.e4 (2015).
Krawczynski, K., Bauersachs, S., Reliszko, Z. P., Graf, A. & Kaczmarek, M. M. Expression of microRNAs and isomiRs in the porcine endometrium: Implications for gene regulation at the maternal-conceptus interface. BMC Genomics 16, 906 (2015).
Lin, X. et al. Bovine embryo-secreted microRNA-30c Is a potential non-invasive biomarker for hampered preimplantation developmental competence. Front. Genet. 10, 315 (2019).
Veno, M. T. et al. Cortical morphogenesis during embryonic development is regulated by miR-34c and miR-204. Front. Mol. Neurosci. 10, 31 (2017).
Chen, X., Liu, J., Feng, W.-K., Wu, X. & Chen, S.-Y. MiR-125b protects against ethanol-induced apoptosis in neural crest cells and mouse embryos by targeting Bak 1 and PUMA. Exp. Neurol. 271, 104–111 (2015).
Kim, K.-H. et al. The miR-125 family is an important regulator of the expression and maintenance of maternal effect genes during preimplantational embryo development. Open Biol. 6(11), 160181 (2016).
Emanueli, C. et al. miR-125b promotes early germ layer specification through Lin28/let-7d and preferential differentiation of mesoderm in human embryonic stem cells. PLoS ONE 7(4), e36121 (2012).
Szuszkiewicz, J., Nitkiewicz, A., Drzewiecka, K. & Kaczmarek, M. M. miR-26a-5p and miR-125b-5p affect trophoblast genes and cell functions important during early pregnancy. Biol. Reprod. 107(2), 590–604 (2022).
Zhang, W. et al. MiRNA-128 regulates the proliferation and neurogenesis of neural precursors by targeting PCM1 in the developing cortex. Elife 5, e11324 (2016).
Wu, H. M. et al. Extracellular vesicle-associated MicroRNA-138–5p regulates embryo implantation and early pregnancy by adjusting GPR124. Pharmaceutics 14(6), 1172 (2022).
Revel, A., Achache, H., Stevens, J., Smith, Y. & Reich, R. MicroRNAs are associated with human embryo implantation defects. Hum. Reprod. 26(10), 2830–2840 (2011).
Kataruka, S. et al. Physiologically relevant miRNAs in mammalian oocytes are rare and highly abundant. EMBO Rep. https://doi.org/10.15252/embr.202153514 (2021).
Sun, L., Li, H., Xu, X., Xiao, G. & Luo, C. MicroRNA-20a is essential for normal embryogenesis by targeting vsx1 mRNA in fish. RNA Biol. 12(6), 615–627 (2015).
Guo, L. et al. gga-mir-133a-3p regulates myoblasts proliferation and differentiation by targeting PRRX1. Front. Genet. https://doi.org/10.3389/fgene.2018.00577 (2018).
Gad, A. et al. Small-extracellular vesicles and their microRNA cargo from porcine follicular fluids: The potential association with oocyte quality. J. Anim. Sci. Biotechnol. 13(1), 82 (2022).
Tripurani, S. K., Lee, K. B., Wee, G., Smith, G. W. & Yao, J. MicroRNA-196a regulates bovine newborn ovary homeobox gene (NOBOX) expression during early embryogenesis. BMC Dev Biol. 11, 25 (2011).
Hornstein, E. et al. The microRNA miR-196 acts upstream of Hoxb8 and Shh in limb development. Nature 438(7068), 671–674 (2005).
Acknowledgements
We thank the staff members of the Daejeon metropolitan slaughterhouse for providing us with porcine ovaries. We thank Mrs. Hyeonsoon Kim from Okcheon Poultry Farm, and Dr. Jongung Yoon (DVM) from FarmShield Co. Ltd. For providing the fresh eggs.
Funding
This work was supported by the Ministry of Science and ICT through the National Research Foundation of Korea (NRF) (Grant numbers: 2021R1A2C2009294 and 2022R1I1A1A01065412) the Brain Pool program (Grant number: 2021H1D3A2A02040098).
Author information
Authors and Affiliations
Contributions
Saadeldin I.M.: Conceptualization, Methodology, Software, Data curation, Writing—Original draft preparation, Funding acquisition; T.B.M.: Methodology, Data curation. B.S.: Methodology, Data curation; S.C.: Methodology, Data curation; M.A.Y.: Methodology, Data curation; K.H.: Methodology; C.D.: Methodology Y.S.H.: Methodology, Data curation; K.S.I.: Methodology, Data curation; L.S.: Methodology, Data curation, Investigation; C.J.: Conceptualization, Methodology, Software, Data curation, Writing—Original draft preparation, Funding acquisition.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Saadeldin, I.M., Tanga, B.M., Bang, S. et al. Isolation, characterization, proteome, miRNAome, and the embryotrophic effects of chicken egg yolk nanovesicles (vitellovesicles). Sci Rep 13, 4204 (2023). https://doi.org/10.1038/s41598-023-31012-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-31012-0
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.