Abstract
Genomic information on alfalfa adaptation to long-term grazing is useful for alfalfa genetic improvement. In this study, 14 alfalfa populations were collected from long-term grazing sites (> 25 years) across four soil zones in western Canada. Alfalfa cultivars released between 1926 and 1980 were used to compare degree of genetic variation of the 14 populations. Six agro-morphological and three nutritive value traits were evaluated from 2018 to 2020. The genotyping-by-sequencing (GBS) data of the alfalfa populations and environmental data were used for genotype-environment association (GEA). Both STRUCTURE and UPGMA based on 19,853 SNPs showed that the 14 alfalfa populations from long-term grazing sites had varying levels of parentages from alfalfa sub-species Medicago sativa and M. falcata. The linear regression of STRUCTURE membership probability on phenotypic data indicated genetic variations of forage dry matter yield, spring vigor and plant height were low, but genetic variations of regrowth, fall plant height, days to flower and crude protein were still high for the 14 alfalfa populations from long-term grazing sites. The GEA identified 31 SNPs associated with 13 candidate genes that were mainly associated with six environmental factors of. Candidate genes underlying environmental factors were associated with a variety of proteins, which were involved in plant responses to abiotic stresses, i.e., drought, cold and salinity-alkali stresses.
Similar content being viewed by others
Introduction
Cultivated alfalfa (Medicago sativa L.), a perennial auto-tetraploid (2n = 4x = 32) and cross-pollinated forage legume, is widely grown as forage due to its high yield and high adaptation in temperate regions of the world1. The cultivated alfalfa complex consists of three main sub-species: M. sativa subsp. sativa, M. sativa subsp. falcata, and M. sativa media2 with each sub-species displaying distinguishing agro-morphological traits2,3. Identification of genetic diversity and relationships within and among different alfalfa populations is of great importance for alfalfa genetic improvement. Especially, information on local adaption under long-term grazing is critical, as alfalfa stands are often affected by grazing4, and environmental conditions and their interactions3,5, which potentially results in genetic changes over time.
Long-term continuous grazing can deplete the alfalfa total non-structural carbohydrates (TNC) and N reserves in the root, and remove photosynthetically active leaf areas6,7, resulting in greater susceptibility to biotic and abiotic stresses, i.e., winter injuries and disease infections. A number of genomic studies demonstrated that plant genetic diversity decreased with increasing grazing intensities in several forage grasses, i.e., Stipa grandis8, Elymus nutans Griseb.9, blue grama (Bouteloua gracilis)10, and meadow fescue (Festuca pratensis Huds.)11. However, a few studies reported no evident genetic diversity change between grazed and ungrazed mountain rough fescue (Festuca campestris Rydb.) populations12, and Idaho fescue (Festuca idahoensis Elmer)13. The reason for no significant changes of genetic diversity in these two studies was that a high degree of gene flow occurred between the populations13. Recent studies indicated that the environmental heterogeneities alone can drive species to accumulate local adaptive loci14,15. Blanco-Pastor et al.16 identified 143 candidate genes of local adaptation primarily associated with environmental factors in diploid alfalfa populations. In western Canada, unique and highly variable environmental factors such as drought, heat, and extreme cold weather affect alfalfa survival and adaptation17,18,19,20,21. Lack of soil moisture during growing season is common in the regions, which can limit alfalfa persistence and productivity3,22,23.
In recent years, genome-wide single nucleotide polymorphisms (SNPs) by genotyping-by-sequencing (GBS) have been applied in the genetic diversity analysis of alfalfa including population structure analysis24. The GBS method makes it possible to generate high-density, genome-wide SNPs25. The abundant SNP markers are more informative than previously used molecular markers in the above-mentioned grass studies and have been successfully applied to identify the genetic structure and relationship in alfalfa breeding materials26,27. However, there are few genomic studies available in examining local adaptation of alfalfa populations under long-term grazing and varying environmental factors.
In this study, alfalfa populations with a minimum of 25 years of grazing history were collected from 14 ranch sites across four soil zones of Saskatchewan, Canada. It was hypothesized that: (1) there will be large phenotypic and genetic variations among the alfalfa populations adapted to long term grazing history; and (2) because of their stand age, the alfalfa populations from long-term grazing sites will be genetically related to one or more of commercial cultivars released from 1926 to 1980 in western Canada. However, each of the alfalfa populations will have unique adaptive loci due to long-term grazing effects and environmental heterogeneity at each ranch site. The objectives of this study were: (1) to identify the genetic relatedness of the 14 alfalfa populations with long-term grazing history inferred by 11 commercial alfalfa cultivars released from 1926 to 1980; and (2) to evaluate genetic and phenotypic variations of 14 alfalfa populations from long-term grazing sites.
Results
Population structure
In STRUCTURE analysis, the optimum sub-population number was determined using the largest delta K value and the change in LnP(K) variance. Both methods confirmed that the optimal sub-population number was K = 2 (Fig. 1A and B), dividing the 25 populations (251 genotypes) into two main genetic backgrounds of falcata and sativa sub-groups (Fig. 1C). Based on the analysis, alfalfa cultivar Anik was a yellow flowering falcata sub-species. Anchor was a sativa sub-species with the highest sativa parentage. The genetic make-up of the majority populations and cultivars were intermediate to the two cultivars. Based on the individual genotype distribution, there was also two main groups. In Group I, there were 10 genotypes from Anik, and two from Drylander representing falcata genetic background (Fig. 1C). Group II was composed of 98 genotypes from 10 commercial alfalfa cultivars, and 141 genotypes from the 14 alfalfa populations from long-term grazing sites (Fig. 1C).
Genetic backgrounds inferred by STRUCTURE using 19,853 SNPs of 14 alfalfa populations from long-term grazing sites sampled across four soil zone of Saskatchewan Canada and 11 alfalfa cultivars released between 1926 and 1980. (A): Support for change in LnP(K) variance; (B): Support for the largest Delta K value; (C): Two optimal clusters of 251 genotypes (K = 2).
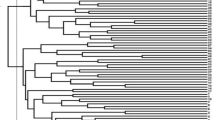
Based on the UPGMA analysis, the populations from five grazed sites (Fillmore, Rockhaven, Ceylon, Crooked River, and Val Marie) were related to the old Canadian cultivar Rambler (Fig. 2). The Duck Lake population was closely related to the cultivar Algonquin, which is variegated light purple flowering type (Table 1S). Eight populations from various grazing sites (Shellbrook, Gull Lake, Moose Jaw, Erwood, Pike Lake, MacDowall, Dalmeny, and Arcola) were associated with the cultivar Ladak, which is a purple-flowered type (Table 1S). The UPGMA analysis was also supported by flower color-based clustering except Fillmore population (Fig. 3). The 14 alfalfa populations from long-term grazing sites were sampled from the four soil zones of Saskatchewan, and when we analyzed them by soil zone there was an apparent genetic divergence of populations by each soil zone, indicating a genetic shift by growth environment (Fig. 4).
UPGMA tree based on Nei’s genetic distance among 25 alfalfa populations representing the 14 alfalfa populations from long-term grazing sites and 11 alfalfa cultivars released from 1926 to 1980; The original soil zone of 14 alfalfa populations from long-term grazing sites or the country and year of cultivars released are in the bracket. Bootstrap values (1000 replicates) are present for each node.
Discriminant analysis of principal component (DAPC) using the 19,853 SNPs of 141 alfalfa genotypes representing 14 alfalfa populations with long-term grazing history sampled across four soil zones of Saskatchewan, Canada. Each dot represents individual alfalfa genotype and the bottom right inset illustrates the legend for four soil zones.
Phenotypic variation
There were significant differences for the six agro-morphological traits among the 14 alfalfa populations from long-term grazing sites (p < 0.001) and among the four soil zones (p < 0.01) (Tables 1 and 2). The MacDowall population had the highest forage dry matter yield, spring vigor, plant height, regrowth yield and fall plant height among them (Table 1). The Gull Lake population required the fewest growing degree days (GDDs) to produce the first flower, although it had 90% sativa parentage based on the STRUCTURE membership analysis. The most GDDs were needed for Ceylon population (29% falcata parentage) (Table 1 and Fig. 1C). Of the 14 populations, five of them (MacDowall, Dalmeny, Fillmore, Arcola and Rockhaven) were the top yielding populations (Table 1), while the top five rapid regrowing populations included Erwood, and Gull Lake in addition to the highest yielding populations (Table 1). The five early fall dormant populations based on fall plant height were Ceylon, Val Marie, Pike Lake, Rockhaven and Arcola, with falcata parentages of 29%, 34%, 21%, 30%, and 24%, respectively (Table 1 and Fig. 1C). The later dormant populations were MacDowall, Dalmeny, Duck Lake, Moose Jaw and Gull Lake, with sativa percentages of 84%, 80%, 85%, 82% and 90% (Table 1 and Fig. 1C).
Among the four soil zones, the populations from the Black soil zone, the moistest region, had the highest forage yield, spring vigor, plant height, regrowth and fall plant height compared to the other three soil zones (Table 2). The populations from the Brown soil zone, the driest among four, had the lowest forage, spring vigor, plant height and regrowth (Table 2). The GDDs for days to flower was higher for the populations from two southern soil zones (Brown and Dark Brown soil zones) than the two northern zones (Black and Grey Wooded soil zones) (Table 2). Forage dry matter yield and regrowth were similar between populations from the Dark Brown and Grey Wooded soil zones (Table 2). The fall plant height was taller for the populations from the Black and Dark Brown soil zones than from the Brown and Grey Wooded soil zones (Table 2).
Significant variation existed among the 14 alfalfa populations from long-term grazing sites in each of the three nutritive value measurements (p < 0.01) and among the four soil zones (p < 0.001) (Tables 3 and 4). The concentrations of ADF ranged from 33.1 to 37.7%, and NDF ranged from 39.4 to 45.6%. The populations from Arcola and MacDowall had the highest concentrations of ADF and NDF, while populations from Crooked River had the lowest ADF and NDF concentrations (Table 3). The concentration of CP ranged from 16.2 to 18.6% among the 14 alfalfa populations from long-term grazing sites. The Crooked River population produced the highest CP concentration, followed by the Val Marie and Gull Lake populations (Table 3). Populations from the Black soil zone had the highest concentrations of ADF and NDF, while populations from the Brown and Grey Wooded soil zones had the lowest concentrations of ADF and NDF. The populations from Grey Wooded soil zones had the highest CP concentrations (Table 4).
We grouped genotypes of the 14 alfalfa populations from long-term grazing sites into sativa and falcata sub-groups based on the percent of sativa (or falcata) parentage in STRUCTURE analysis. If a genotype is made up of greater than 50% sativa parentage, and then it will be grouped into sativa cluster, and vice versa (Fig. 1C). The linear regressions of STRUCTURE based sativa parentage percent (membership probability) on phenotypic values of forage dry matter yield, spring vigor and plant height were not significant (Fig. 5), while the phenotypic values of fall plant height, regrowth, days to flower, NDF and CP were significantly associated with their STRUCTURE sativa membership probability (Figs. 5 and 6). Based on the UPGMA results, the sativa populations (Duck Lake, Shellbrook, Gull Lake, Moose Jaw, Erwood, Pike Lake, MacDowall, Dalmeny, and Arcola) had higher phenotypic values of spring vigor, plant height, forage dry matter yield, fall plant height, ADF and NDF but lower values of days to flower and CP as compared to the falcata populations (Fillmore, Rockhaven, Ceylon, Crooked River and Val Marie) (Figs. 7 and 8).
Genotype-environment association (GEA)
GEAs for soil nutrients
A total of seven significant GEAs were identified for soil K on chromosomes 1.3, 1.4, 2.4, 4.2, and 8.1 (Table 5). The correlations between soil K with allele frequencies at these SNPs ranged from 0.17 to 0.37. Of these, one SNP on chromosome 2.4 located at 28.5 Mb had the highest correlation of 0.37 and was related to MS.gene77592 encoding Cytochrome P450. Four significant GEAs were identified for soil P on chromosomes 1.4, 3.2 and 7.4 (Table 5). The correlations at these SNPs ranged between 0.24 and 0.33. The SNP on chromosome 3.2 located at 49.3 Mb had the highest correlation of 0.33 and was associated with MS.gene064294 encoding ABC transporter type 1, transmembrane domain superfamily. The SNP on chromosome 1.4 was related to MS.gene46469 encoding Leucine-rich repeat, cysteine-containing subtype. A total of four significant GEAs were identified for soil S on chromosomes 3.2 and 4.3 (Table 5). The correlations at these SNPs ranged from 0.22 to 0.24. One SNP on chromosome 3.2 at 42.9 Mb indicated the highest correlation of 0.24 and was related to MS.gene30119 encoding Armadillo-like helical. The three other SNPs on chromosome 4.3 located at 7.1 Mb had correlation of 0.22 and were related to MS.gene000431 encoding Ankyrin repeat-containing domain. Four significant GEAs were identified for soil pH on chromosomes 3.1, 4.3 and 7.4 (Table 5). The correlations at these SNPs ranged from 0.24 to 0.34 with a SNP on chromosome 3.1 at 74.9 Mb showing the highest correlation of 0.34. This SNP was associated with MS.gene049423 encoding Zinc finger, RING-type. In addition, the correlations at two other SNPs on chromosome 4.3 located at 7.1 Mb were 0.24. The SNP on chromosome 4.3 located at 7.1 Mb had correlation of 0.24 and was related to the same gene MS.gene000431 for soil pH (Table 5). Another SNP on chromosome 4.3 located at 7.1 Mb was related to MS.gene000426 encoding WD40 repeat (Table 5).
GEAs for summer and winter extreme temperature
A total of eight significant GEAs were identified for summer extreme temperature on chromosomes 1.1, 4.4, 5.2 and 8.1 (Table 5). The correlations at these SNPs with summer extreme temperature ranged from 0.14 to 0.38. Two SNPs on chromosome 4.4 at 10.0 Mb had the highest correlation of 0.38 and were related to same gene MS.gene28113 encoding FKBP-type peptidyl-prolyl cis–trans isomerase domain. The SNP on chromosome 4.4 at 8.7 Mb had correlation of 0.26 and was related to MS.gene28435 encoding MORN motif. Moreover, four significant GEAs were identified for winter extreme temperature on chromosomes 4.3, 7.1, and 8.1 (Table 5). The correlations at these SNPs ranged from 0.23 to 0.35. One major SNP on chromosome 7.1 at 62.1 Mb had the correlation of 0.35 and was related to MS.gene30010 encoding XPG/Rad2 endonuclease. The SNP on chromosome 4.3 at 7.1 Mb had a correlation of 0.23 and was related to the same gene MS.gene000426 for soil pH (Table 5). Two SNPs on chromosome 8.1 at 9.9 Mb had the same correlation of 0.27 and were related to MS.gene58730 encoding Alpha/Beta hydrolase fold (Table 5).
To estimate r2 values of SNPs within the same gene identified in GEA results, LD was analyzed using a total of 12,402 SNP markers. The r2 of LD across all chromosomes was calculated and is presented in Fig. 1S (see the Supplementary files). A rapid drop in r2 was observed with the increase in physical distance between 0 and 5 cM, and then r2 showed no decrease afterward (Fig. 1S). Meanwhile, five pairs of SNPs were found within same gene: two for soil K, one for soil S, one for winter extreme temperature, and one for summer extreme temperature (Fig. 2S). The physical distances for these SNP pairs ranged from 12 to 54 bp, and the r2 values were greater than 0.9 (Fig. 2S).
Discussion
Population structure
The STRUCTURE analysis confirmed two optimal clusters among 251 alfalfa genotypes representing the 14 alfalfa populations from long-term grazing sites and the 11 commercial alfalfa cultivars widely seeded from 1926 to 1980 in western Canada. The Cluster I included genotypes with a high percentage of falcata background, supported by cultivar Anik (known as falcata population) as an independent cluster with two genotypes from cultivar Drylander. M. falcata has yellow flowers and uncoiled seed pods, and is widely considered to be indigenous to Siberia28. The subspecies has also shown slow regrowth and creeping root-habit29,30, with good drought, extreme winter hardiness and grazing resistance3,31. The cultivars Rambler, Roamer and Rangelander have intermediate levels of falcata backgrounds, which is consistent with their pedigrees. For example, Rambler was recorded as having 45% of M. falcata parentage31. M. falcata was used as one of parents for artificial crossing with the cultivar Ladak (a purple flowered M. sativa) to develop creeping-rooted cultivars for grazing in western Canada, which resulted in Rambler32, Roamer33 and Drylander34. Rangelander was selected from a cross among Rambler, Roamer, Drylander and strains of M. falcata35. The Cluster II was composed of genotypes with high percentages of M. sativa background including cultivars of Algonquin, Anchor, Beaver, Grimm, Ladak, and Vernal. M. sativa is a purple-flowered form with more rapid growth, and a relatively more erect growth form than M. falcata3,22,36,37. The cultivar Anchor is considered as typical of the M. sativa subspecies, while Beaver alfalfa has 98% of M. sativa parentage31. Algonquin showed 50% of M. sativa parentage, since this cultivar was developed by open pollinating M. media (M. falcata × M. sativa) to the cultivar Rhizoma38, which had 50% parentage from M. falcata, and another 50% from Grimm (M. media)31.
Interestingly, both DAPC and genetic distance analyses showed no distinctive separation of the 14 alfalfa populations from long-term grazing sites. Initially, we expected that the 14 populations would be associated with one of the 11 cultivars used in the region at that time. This result might be caused by a genetic shift in response to regional environments39, and long-term grazing managements8,9,10,11,39. The 14 alfalfa populations from long-term grazing sites had at least 25 years of utilization by harvesting hay or grazing annually, which affected their population genetic diversity. However, the DAPC analysis on variability by soil zone showed a genetic shift of the 14 populations by soil zone. This might be attributed to interactive effects of climatic factors (precipitation and temperature), differential grazing pressure and soil properties such as texture, pH, and nutrient levels on genetic variation of the alfalfa populations. Blanco-Pastor et al.16 found a set of candidate genes associated with local environmental factors for diploid alfalfa populations, confirming local adaptation. In addition, long-term grazing activities may have also changed genetic structure. Although there are no reports of genetic shift of alfalfa populations under grazing, a few studies indicated that intensive grazing decreased genetic diversity in forage grasses, i.e., Stipa grandis8, Elymus nutans Griseb.9, blue grama (Bouteloua gracilis)10, and meadow fescue (Festuca pratensis Huds.)11.
When analyzed by populations based on the UPGMA analysis, five alfalfa populations from the 14 long-term grazing sites (Val Marie, Crooked River, Ceylon, Rockhaven and Fillmore) were genetically related to the cultivar Rambler. This cultivar is a variegated flower type with a higher level of parentage of M. falcata than Roamer and Rangelander32. This cultivar was superior in drought resistance and winter hardiness to Ladak and Grimm, but has slow recovery after cutting or grazing than either variety32. It was selected for improved drought tolerance, the climatic conditions often found in the Brown soil zone40. Thus, it would not be surprising to find this cultivar at the Ceylon, Val Marie sites, which are located in the Brown soil zone. The population from Duck Lake was genetically associated with the cultivar Algonquin. This cultivar is a variegated light purple type with high of parentage of M. sativa. Algonquin was developed by backcrossing plants resistant to bacterial wilt (Corynebacterium insidiosum (McCull) H. L. Jens) into the cultivar Rhizoma38. Eight alfalfa populations from the long-term grazing sites were genetically related to Ladak, which is a purple-flowered type which has a greater percent of M. sativa parentage. In the history of alfalfa cultivation under northern Great Plains conditions, Ladak not only outyielded Grimm but also had a longer stand persistence than other alfalfa cultivars at that time due to its excellent cold and drought tolerances41. M. sativa generally has higher forage yield, faster regrowth, and a relatively more erect growth form than M. falcata3,22,36,37. These populations were generally from the Black or Dark Brown soil zones where soil moisture is higher than the Brown soil zone.
Phenotypic variation
Significant differences were observed for six agro-morphological and three nutritive values traits among the 14 alfalfa populations with long-term grazing history. These traits were related to their relative levels of parentage to either falcata or sativa sub-species. For example, based on the genetic background inferred by UPGMA analysis, the mean forage dry matter yield, regrowth and yield-related traits (e.g., plant height, fall plant height and spring vigor), ADF and NDF of alfalfa populations with long-term grazing history in the sub cluster with Ladak and Algonquin were higher than those in the subcluster with Rambler, while days to flower and CP were lower. The subcluster with Ladak and Algonquin with a high percentage of M. sativa genome, while the subcluster with Rambler had a high percentage of falcata genome. The M. sativa genome has been considered to provide the genetic basis for high forage dry matter yield, rapid regrowth and low fall dormancy as compared with the M. falcata genome7,30,42,43. Interestingly, the alfalfa populations with a greater parentage from the M. falcata genome (Crooked River, Val Marie, Rockhaven, Fillmore and Ceylon) required longer GDDs and produced higher concentrations of CP and lower concentrations of ADF and NDF. The alfalfa cultivar AC Yellowhead, belonging to the falcata sub-species, has been shown to contain higher CP concentration to Beaver alfalfa cultivar, a M. sativa44 cultivar. Meanwhile, significant differences were also observed for the agro-morphological traits and nutritive values among the four soil zones. The alfalfa populations from the Black soil zone produced the highest forage dry matter yield, spring vigor, plant height, regrowth and fall plant height, while the populations from the Brown soil zone had the lowest values for agro-morphological traits. This is because that three of four alfalfa populations (Duck Lake, Arcola and MacDowall) in the Black soil zone were identified as having high percentages of the M. sativa genome based on the UPGMA analysis. By contrast, two of four alfalfa populations (Ceylon and Val Marie) in the Brown soil zone were identified as having high percentages of M. falcata genome.
The GDDs required for developing flowers for the 14 alfalfa populations from long-term grazing sites increased gradually in the order of Grey Wooded < Black < Dark Brown < Brown soil zones. This is an interesting regional adaptation trait as a reduced number of GDDs requirement for the populations from the Grey-Wooded and Black soil zones has an ecological significance for maintaining plant population and its genetic diversity by sexual reproduction. These regions have colder springs, and are at higher latitudes (longer days) than the Brown soil zone sites. Day-length and temperature are the two main environmental factors influencing flower development45. Alfalfa is a perennial, long-day plant, and its flowering is induced when the photoperiod exceeds a critical threshold value46. Lower accumulated temperature has been shown to delay flowering time for alfalfa47. Knowledge on environmental and genetic factors regulating flowering time can help farmers to balance forage quality and yield48.
The linear regression of STRUCTURE membership probability on phenotypic values indicated that there was lower genetic variation of first cut forage yield (hay yield) and yield-related traits (spring vigor and plant height) as sativa parentage percent increased. This result may be explained by recurrent phenotypic selection on forage dry matter yield and related traits that has been carried out in the past. However, the phenotypic variation of the traits regrowth, fall plant height, days to flower, NDF and CP were significantly associated with higher sativa percentage, which indicated that high genetic variation of these five traits existed in the 14 alfalfa populations with long-term grazing history. These findings may suggest that less phenotypic selection was carried out on these five traits in past breeding programs.
Genotype-environment association
The GEA analysis identified 31 SNPs highly associated with six environmental factors in the alfalfa populations from long-term grazing sites. Of the 31 SNPs identified, eight SNPs were associated with the summer extreme temperature, which is a critical factor affecting alfalfa production in western Canada49. In most cases, the summer extreme temperatures have been accompanied by drought stress in western Canada22,50,51,52. Historically, drought stress has occurred frequently in the Brown soil zone53, and drought could become more common under climate change54. Therefore, these genes may be useful for marker assisted breeding of climate resilient cultivars. Two major SNPs identified on chromosome 4.4 were linked with the putative candidate gene, MS.gene28113 (FKBP-type peptidyl-prolyl cis–trans isomerase domain) and MS.gene28435 (Membrane occupation and recognition nexus motif). It has been reported that the peptidyl prolyl cis–trans-Isomerase FKBP77 gene is involved in heat stress tolerance55. The PI monophosphate 5-kinase (PIP5K) enzyme containing membrane occupation and recognition nexus motif is involved in heat and drought-responses in plants56,57.
In western Canada, alfalfa dry matter yield is often limited by winter injury58, which can be caused by fall cutting or grazing, resulting low root organic matter20. M. falcata with strong winter hardiness has been used to breed alfalfa for improved winter survival31. Among the four SNPs associated with winter extreme temperature, the SNP on chromosome 4.3 was the most significant, which was linked to the putative candidate gene MS.gene000426 (WD40 repeat). The WD40 gene OsTTG1 regulates anthocyanin biosynthesis, which plays an important role in the growth of plants and is affected by temperature59,60,61,62,63.
Potassium (K) promotes alfalfa persistence and stand longevity64,65,66,67. Alfalfa plants deficient with K had low root organic reserves for new growth and maintenance67,68,69. Of seven significant SNPs, one major SNP on chromosome 2.4 is linked to the candidate gene MS.gene77592 (Cytochrome P450), which is involved in glucosinolate synthesis70,71,72. Glucosinolate plays an important role in plant response to abiotic stress73. Another two SNPs on chromosome 4.2. The putative candidate gene linked with these two markers is MS.gene043198 (S-phase kinase-associated protein 1). Plant SKP1-like family proteins have been found to associate with the regulation of plant alkaline tolerance and ABA sensitivity74. These two candidate genes identified here had no direct relationship with K stress in plants. This may be because the soils in our study were not K deficient based on soil tests.
Sulfur (S) has been reported to decrease the survival of alfalfa seedlings75. Four significant SNPs were identified for S on chromosome 3.2 and 4.3. Several repeat protein gene families have been identified in plants. The SNP on chromosome 3.2 is linked to the putative candidate gene MS.gene30119 (Armadillo-like helical). Some studies have reviewed the importance of the relationship between S assimilation and phytohormones76,77. Phytohormones are essential for plant acclimation and adaptation to environmental changes78. The other three SNPs on chromosome 4.3 were linked with MS.gene000431 (Ankyrin repeat-containing domain). The Ankyrin repeat domain C3HC4-Type RING finger genes play important roles in various physiological processes of plants79.
Phosphorus (P) is a vital macronutrient that plays important roles in energy transfer, respiration, enzyme activation, photosynthesis, and root elongation80. Low soil P availability is a major limiting factor in old alfalfa stands, which was verified by our soil nutrient analyses at these long-term sites. Four significant SNPs were identified for P in our study with a major SNP on chromosome 3.2 located at 49.3 Mb. The putative candidate gene linked with this marker is MS.gene064294 (ABC transporter type 1, transmembrane domain superfamily). ATPase binding cassette (ABC) transport G subfamily plays a crucial role in the transportation of biological molecules across the membrane. Overexpression of L.albABCG29 in white lupin hairy root enhanced P accumulation in cluster roots under low P and improved plant growth81. Another SNP on chromosome 1.4 located at 45.0 Mb had a correlation of 0.25, and the putative candidate gene linked with this marker is MS.gene46469 (Leucine-rich repeat, cysteine-containing subtype).
Alfalfa plants are sensitive to acid soil82. The collected sites of the 14 alfalfa populations from long-term grazing sites had soil pH > 7. Four significant SNPs were identified for soil pH with a major SNP on chromosome 3.1. The putative candidate gene linked with this marker is MS.gene049423 (Zinc finger, RING-type). NCA1 gene encoding RING-type zinc finger domain regulates other protein activities in response to alkaline stress83. Two different SNPs on chromosome 4.3 were linked to the putative candidate genes MS.gene000431 (Ankyrin repeat-containing domain) and MS.gene000426 (WD40 repeat), respectively. The WD40 related proteins are involved in plant resistance to alkali stress84. The genes encoding ankyrin repeat domain has been reported to be associated with salinity-alkali stresses85. Our results showed that the LD distance within the alfalfa populations with long-term grazing history was larger (r2 reduced to 0.2 after 3 cM) than previous studies24,86,87. One main reason is that the alfalfa populations with long-term grazing history in our study were from genetically related alfalfa cultivars, and then low chromosome recombination events occurred within these cultivars.
This study identified 13 candidate genes associated with six environmental factors, reflecting adaptation of the 14 alfalfa populations from long-term grazing sites to their given environments across different soil zones. Most of these genes play an important role in plant tolerances to biotic and abiotic stresses, such as heat, drought, cold, salt and diseases. In western Canada, extreme low temperature17,18,19,20,21, and frequent drought stresses5,22,37,50,51,52,88 can significantly reduce alfalfa forage yield and stand persistence, acting as a natural selection for local adaptation. Inevitably, the majority of original alfalfa plants at the 14 long-term grazing sites might have died because of lack of adaption to interaction of grazing and environmental conditions (i.e., extreme drought in summer, extreme low temperature in winter or diseases). Similar to the results in our study, a number of studies reported candidate loci responsible for adaptation to climate gradients in diploid alfalfa16, M. truncatula16,39,89,90 and perennial ryegrass91. In addition, Humphries et al.92 demonstrated that collecting alfalfa germplasms that have evolved to survive in extreme environments with low rainfall, high temperature, and winter freezing would be one option to breed alfalfa adapted to the warming climates around the world. Therefore, the collected alfalfa populations with higher forage dry matter in our study can be a valuable genetic resource for future alfalfa breeding. Meanwhile, the 13 candidate genes involved in regulation of alfalfa plants’ response to environmental factors need to be validated in the future experiments, and then these genes can be used for marker-assisted selection for alfalfa breeding and genetic improvement.
Methods
Plant materials and soil sampling
In spring 2016, 14 alfalfa-grass mixed stands with a minimum of 25 years of grazing history were selected across Brown, Dark Brown, Black and Grey Wooded soil zones of Saskatchewan, Canada (Table 6; Fig. 9). Our research team obtained permissions to collect cultivated alfalfa plants from the 14 private ranchers prior to this study. The plant materials are not wild plants or endangered species. Experimental research and field studies on plants including the collection of plant material, complied with relevant institutional, national, and international guidelines and legislation. The Brown soil zone is the most arid region characterized by annual precipitation in the range of 275–325 mm. It has the highest evapotranspiration among the four soil zones. The Dark Brown soil zone is characterized by annual precipitation in the range of 325–375 mm. It has a moderate evapotranspiration. The Black soil zone is characterized by annual precipitation in the range of 400–475 mm. The Grey Wooded soil zone is characterized by annual precipitation ranging from 300 to 500 mm with lower evapotranspiration than in the Black soil zone40. At all grazing sites, alfalfa plants were grown in a grass-alfalfa mixture, which were grazed at least once a year over 25 years. At each site, 30 mature alfalfa plants were dug and stored in a plastic container with 10 cm water before transplanting to a spaced nursery at Saskatoon, Saskatchewan for agronomic traits evaluation. The distance between any two individual plants was 30 m to minimize sampling similar clones. Meanwhile, 30 soil samples at a depth of 30 cm were collected near the sampled plants at each site and then bulked by site for analyses of nitrogen (NO3-N), phosphorus, potassium, sulfur (SO4-S), and soil pH and electrical conductivity (E.C.) at ALS Laboratory Group (Saskatoon, Saskatchewan).
Locations of 14 long-term grazing sites for alfalfa plant selection during this study. (Sites names 1–14 are: Crooked River, Shellbrook, and Erwood; MacDowall, Duck Lake, Rockhaven and Arcola; Dalmeny, Pike Lake, and Fillmore; Ceylon, Gull Lake, Moose Jaw, and Val Marie). Map was adapted from Canadian Soil Information Service (https://sis.agr.gc.ca/cansis/index.html).
Reference cultivars for genetic diversity study
Ten random alfalfa plants from each of the 14 alfalfa populations from long-term grazing sites were selected for the genetic diversity study. To identify population structure of the 14 alfalfa to the Canadian commercial alfalfa cultivars released in western Canada from 1926 to 1980, 11 commercial alfalfa cultivars (Table 1S) were also included in the study. The seeds of the 11 alfalfa cultivars were germinated on top of a double layer of filter paper (VWR 413) in 9 cm plastic Petri-dishes. Then, 10 random germinated seeds per cultivar were transplanted into a 1/8-gallon pot (11 × 0.9 cm, Sun Gro Horticulture) filled with propagation soil mix containing 70–80% sphagnum peat moss, vermiculite and dolomite (SS#3, Sun Gro Horticulture Ltd.) at the College of Agriculture and Bioresources greenhouse at the University of Saskatchewan. The greenhouse has an 18-h photoperiod supplemented with high pressure sodium halogen lamps to a total of 490–550 μMs−1 m−2 PAR. The air temperature was 26 °C and 20 °C, for day/night, respectively. Relative humidity was maintained at 67%.
Field experimental design
Four hundred twenty alfalfa plants collected from the 14 long-term grazing sites were propagated by stem cuttings. Then, the cloned plants were transplanted to a 1/4-gallon pot (13 × 12 cm, Sun Gro Horticulture) with propagation soil mix containing 70–80% sphagnum peat moss, vermiculite and dolomite (SS#3, Sun Gro Horticulture Ltd.) at the College of Agriculture and Bio-resources greenhouse in the University of Saskatchewan. A field spaced nursery was established on June 6, 2017, at the Agriculture and Agri-Food Canada (AAFC) Saskatoon Research Farm, Saskatoon, SK. The experimental design was a nested randomized complete block design (RCBD) with two blocks. There were 840 plots in the nursery (14 alfalfa populations × 30 genotypes population−1 × 2 blocks). Spacing between rows and plants within a row was 1 m. Weed control was done using a triple rototiller (KS-190, Tram sales Ltd, AB, Canada) in early spring in addition to hand weeding around individual plants. The soil was a Sutherland clay loam (Dark Brown Chernozem, Typic Haploloroll)93. Weather data for Saskatoon from 2018 to 2020 was obtained from Environment Canada (https://climate.weather.gc.ca). The average monthly air temperature and precipitation at the study site are shown in Fig. 3S. Growing season (May to September) air temperature during the three study years was near the long-term mean except for a higher temperature in May 2018, and lower temperatures in September 2018, May and August 2019. Rainfall was lower than normal during June and July in 2018. Almost no rainfall was recorded in May 2019. There was a higher than normal rainfall in June 2020.
Agro-morphological trait measurement
Six agro-morphological traits were measured on individual genotypes for three consecutive summers from 2018 to 2020, in addition to nutritive value analysis in 2018 and 2019. Spring vigor (SV) was visually scored for each genotype using a 1–5 scale (poor to good) on the basis of spring growth, plant size and leafiness. Days to flower (GDD) was recorded as the date of first flower emerged on each genotype. Plant height (PH) for each genotype was measured from the soil surface to the tip of the stem by stretching the stems upwards. For forage dry matter yield (DMY) determination, each genotype was harvested, and dried at 60 °C for 48 h in a forced-air oven and was weighed in grams. For regrowth yield (RY) determination, each genotype was harvested, and dried at 60 °C for 48 h in a forced-air oven and was weighed in grams. Fall plant height (FPH) was measured from the soil surface to the tip of the stem by stretching the stems upwards for each genotype. The flowering date was expressed as growing degree days (GDDs) using a base temperature of 5 °C94.
Forage nutritive value
For forage nutritive value determinations, plants at full bloom stage before the first cut were dried at 60 °C for 48 h in a forced-air oven. The dried plant samples were ground in a Wiley mill (Thomas-Wiley, Philadelphia, PA, USA) and then passed through a 1-mm mesh screen (Cyclone Mill, UDY Mfg., Fort Collins, Colorado, USA). Ground samples were stored in filter bags (Nasco Whirl–Pak, USA) prior to crude protein (CP), neutral detergent fiber (NDF) and acid detergent fiber (ADF) determinations. The values of three nutritive value traits were determined using a FOSS XDS rapid content analyzer (Foss, Denmark).
Phenotypic data analysis
For each phenotypic trait, a linear mixed model was used to perform analysis of variance (ANOVA) using JMP (JMP 15.2.1. SAS Institute Inc., Cary, NC). Soil zone, year, population (soil zone) and genotype (population, soil zone), soil zone × year, were treated as fixed effects in the model. The block was treated as a random effect. For each of six agro-morphological and three nutritive values traits, if the ANOVA indicated a significant difference at p ≤ 0.05 level, means were separated using the studentized Tukey multi-treatment method. Degrees of freedom were calculated using Satterthwaite’s method.
Genotype-environment associations (GEA) were determined using the redundancy analysis (RDA) procedure using the R package “vegan 2.5-7 version” with the rda function95,96. The long-term (1986–2015) lowest and highest air temperatures and total precipitation (from May to September) for GEA analysis were obtained from Canada Weather Stats (available at https://www.weatherstats.ca/) and Environment Canada (available at https://climate.weather.gc.ca/). The soil pH, nitrogen (NO3-N), phosphorus (P), potassium (K), sulfur (SO4-S), and electrical conductivity (E.C.) from each long-term grazing site were also added into the GEA40. The electrical conductivity was removed as it was highly correlated (\(\left| {\text{r}} \right|\) > 0.7) with soil pH, because the RDA method for GEA is a regression-based method subject to problems when using highly correlated predictors as described by Dormann et al.97. Two alfalfa populations from Dalmeny and Pike Lake were excluded from the GEA due to no soil test information.
Individual genetic data were considered as response matrix Y, and a set of environmental predictors were used as explanatory matrix X. Genotypes were coded using the values 0, 1 or 2 that correspond to homozygote for the most frequent alleles, and heterozygote and homozygote for the less frequent alleles. RDA aims at constructing a matrix Y′ of fitted genetic values estimated from the regression of each locus by the environmental predictors and at performing principal component analysis on the matrix Y′98. The number of axes used (K) is determined by looking at the amount of information retained on the different axes of the RDA. A Mahalanobis distance D is then computed for each locus to identify loci showing extreme D values compared to the rest of the SNPs. A Mahalanobis distance is a multidimensional generalization of the idea of measuring how many standard deviations is a point from an average point. Computation of the Mahalanobis distance uses the covRob function of R package “ROBUST”99. Mahalanobis distances are distributed as a chi-squared distribution with K degrees of freedom after correcting with the genomic inflation factor100. Inflation factors are constant values that are used to rescale chi-square statistics in order to limit inflation due to diverse confounding factors101. Inflation factors are constant values that are used to rescale chi-square statistics to limit inflation due to diverse confounding factors101. Multicollinearity between environmental predictors was assessed using the variance inflation (VIF) and since all predictors showed VIF < 5 none were excluded102. We then adjusted the resulting p-values for the false discovery rate (FDR) by computing q-values with the “qvalue” R package103. A locus is considered as an outlier if its q-value is < 0.01.
DNA extraction and library establishment
Fresh young leaves of 10 random alfalfa genotypes from the 14 alfalfa populations from long-term grazing sites (except 11 genotypes for Shellbrook) and 10 genotypes from the 11 commercial alfalfa cultivars were collected into 1.5 ml centrifuge tubes and flash frozen in liquid nitrogen. Then the tubes were immediately stored in a − 80 \(\mathrm{^\circ{\rm C} }\) freezer before the leaf samples were dried in a freeze-drier (Freezone 6, Labconco, USA) at − 52 °C and < 0.09 Pa for 72 h. One hundred milligram of dried leaf samples were ground individually with a sterilized plastic stick. The genomic DNA was extracted using DNeasy Plant Mini Kit (Qiagen, Toronto, ON, Canada). The Quant-iT PicoGreen dsDNA Assay Kit (Invitrogen, Carisbad, CA, USA) was used to quantify the concentration of the DNA samples. The DNA concentration was adjusted to 20 ng μl−1, and 100 ng DNA was digested by ApeKI (New England Biolabs, Ipswitch, MA). Then, the individual DNAs were ligated with a unique barcode adapter and a common adapter (Eurofins Genomics, Louisville, KY, USA) for 96 genotypes per plate using the protocol from Elshire et al.25. The ligated DNA fragments were purified by Qiagen PCR cleanup kit (Qiagen, Toronto, ON, Canada). Following the purification, PCR primers (Eurofins Genomics, Louisville, KY, USA) and KAPA HiFi HotStart Library Amp Kit (Roche, Laval, QC, Canada) were added through PCR amplification. The amplicon fragments were dual-side size selected using 0.57 × and 0.78 × (Average fragment size 400 bp, approximate size distribution 220–900 bp) using Sera Mag Select (Cytiva, Marlborough, MA, USA). An aliquot of each sample was run on an Agilent 2100 Bioanalyzer (Agilent Technologies 2013) for evaluation of fragment sizes. The three libraries of 96 barcoded samples were sequenced with paired-ends of 125 bp in three lanes using an Illumina HiSeq 2500 at the National Research Council of Canada, Saskatoon, Canada.
SNP calling
The quality of pooled, raw reads was assessed using FastQC software104. The raw reads were then demultiplexed using Sabre105. After demultiplexing, the raw reads of individual samples were cleaned by removing adapters and low-quality sequences using Trimmomatic v.0.36 based on the default setting of paired-end mode, phred 33 and threads 6106. The M. sativa genome sequence107 was used as a reference genome for alignment of the reads using Burrows-Wheeler Aligner v0.7.17 software108. SNP calling was performed using Freebayes v1.2.0109 with the following settings: minimum coverage 1, minimum alternate fraction 0.1, minimum alternate count 5, genotype variant threshold 26, minimum base quality 13, pooled continuous, best 1 alleles, mapping quality 30, ploidy 2, other settings, default. The minor allele frequency, extent of missing SNP data and distribution of SNPs on the reference genome were calculated. The SNPs with Phred-scaled quality (QUAL) score ≥ 1260 were kept for imputation using BEAGLE software110. After imputation, 19,853 high quality SNPs were obtained and used in further analysis.
Genetic diversity analysis
The Bayesian method implemented in the program STRUCTURE is a common method applied to detect population genetic structure111,112,113,114. This program uses a Bayesian clustering approach to define clusters containing groups of individuals whose genotypes maximize Hardy–Weinberg and linkage equilibrium111. A discriminant analysis of principal components (DAPC)115 using the “adegenet” package for R116,117 was conducted to identify clusters of genetically related individuals. The Bayesian Information Criterion (BIC) was used for selecting the optimal number of clusters (K) based on the elbow method115. The find.clusters function was used to detect the number of clusters among the populations. It uses K-means clustering which decomposes the total variance of a variable into between-group and within-group components. The optimum number of sub-populations has the lowest associated BIC. A cross validation function Xval.dapc was used to confirm the correct number of principal components (PC) to be retained.
The genetic diversity analysis was based on 19,853 SNP markers obtained from the SNP calling procedure. Three types of diversity analysis were performed at individual genotype level. First, genetic structure of 251 alfalfa genotypes was assessed using a model-based Bayesian method implemented in the program STRUCTURE version 2.3.4111. Sub-population numbers (K) ranging from 1 to 10 were evaluated, repeating each analysis 20 times. Modelling decisions included correlated allele frequencies and use of an admixture model. A burn-in of 30,000 iterations and 30,000 iterations of the Markov Chain Monte Carlo (MCMC) were used. The most likely number of sub-populations was inferred with the delta K method118 implemented in the STRUCTURE HARVESTER program119. The CLUMPP program120 was used to generate a consensus ancestry estimate from the 20 independent runs for each K and ancestry bar plots were generated with the “pophelper” package for R117,121.
Identification of unique loci associated with long-term grazing
The 19,853 SNPs, with minor allele frequency (MAF) below 5% removed, were used for genotype-environment association for 122 genotypes representing 12 alfalfa populations sampled from the long-term grazing sites. The SnpEff software122 was used to annotate all SNPs based on the M. sativa reference genome database. The candidate genes were localized by retrieving the positions of significant SNP markers in the annotation database of M. sativa genome function107.
Linkage disequilibrium
Pairwise Pearson correlation (r2) between pairs of SNPs were performed to assess the linkage disequilibrium (LD) across all chromosomes. A total of 12,402 SNPs were used in the LD analysis. The LD decay over physical distance was determined as the mean distance under the LD threshold of r2 = 0.2. The R package “sommer” (version 4.2.0) was used to conduct LD analysis123.
Data availability
The datasets generated during the current study are available from Sequence Read Archive (SRA) database under NCBI [SRA accession PRJNA900199].
References
Annicchiarico, P., Barrett, B., Brummer, E. C., Julier, B. & Marshall, A. H. Achievements and challenges in improving temperate perennial forage legumes. Crit. Rev. Plant Sci. 34, 327–380. https://doi.org/10.1080/07352689.2014.898462 (2015).
Small, E. & Brookes, B. S. Taxonomic circumscription and identification in the Medicago sativa-falcata (alfalfa) continuum. Econ. Bot. 38, 83–96. https://doi.org/10.1007/BF02904419 (1984).
Boe, A. et al. Breeding alfalfa for semiarid regions in the northern Great Plains: History and additional genetic evaluations of novel germplasm. Agronomy 10, 1686. https://doi.org/10.3390/agronomy10111686 (2020).
Smith, S., Bouton, J. & Hoveland, C. Alfalfa persistence and regrowth potential under continuous grazing. Agron. J. 81, 960–965. https://doi.org/10.2134/agronj1989.00021962008100060023x (1989).
Jefferson, P. & Cutforth, H. Sward age and weather effects on alfalfa yield at a semi-arid location in southwestern Saskatchewan. Can. J. Plant Sci. 77, 595–599. https://doi.org/10.4141/P96-110 (1997).
Berdahl, J., Wilton, A., Lorenz, R. & Frank, A. Alfalfa survival and vigor in rangeland grazed by sheep. Rangeland Ecol. Manag./J. Range Manag. Arch. 39, 59–62. https://doi.org/10.2307/3899688 (1986).
Smith, S. R., Bouton, J. H., Singh, A. & McCaughey, W. P. Development and evaluation of grazing-tolerant alfalfa cultivars: A review. Can. J. Plant Sci. 80, 503–512. https://doi.org/10.4141/p99-048 (2000).
Shan, D., Zhao, M., Han, B. & Han, G. Examining the genetic diversity of Stipa grandis under various grazing pressures. Acta Ecol. Sin. 26, 3175–3182. https://doi.org/10.1016/S1872-2032(06)60048-6 (2006).
Ma, D. et al. Plant genetic diversity and grazing management on the Qinghai-Tibetan Plateau: A case study of a dominant native wheatgrass (Elymus nutans). Biochem. Syst. Ecol. 56, 16–23. https://doi.org/10.1016/j.bse.2014.04.014 (2014).
Aguado-Santacruz, G. A. et al. Genetic variability of Bouteloua gracilis populations differing in forage production at the southernmost part of the North American Graminetum. Plant Ecol. 170, 287–299. https://doi.org/10.1023/B:VEGE.0000021706.12328.61 (2004).
Kölliker, R., Stadelmann, F., Reidy, B. & Nösberger, J. Fertilization and defoliation frequency affect genetic diversity of Festuca pratensis Huds. in permanent grasslands. Mol. Ecol. 7, 1557–1567. https://doi.org/10.1046/j.1365-294x.1998.00486.x (1998).
Fu, Y., Thompson, D., Willms, W. & Mackay, M. Long-term grazing effects on genetic variability in Mountain rough fescue. Rangel. Ecol. Manage. 58, 637–642. https://doi.org/10.2111/05-032R2.1 (2005).
Matlaga, D. & Karoly, K. Long-term grazing effects on genetic variation in Idaho fescue. Rangel. Ecol. Manage. 57, 275–279. https://doi.org/10.2111/1551-5028(2004)057[0275:LGEOGV]2.0.CO;2 (2004).
Forester, B. R., Lasky, J. R., Wagner, H. H. & Urban, D. L. Comparing methods for detecting multilocus adaptation with multivariate genotype–environment associations. Mol. Ecol. 27, 2215–2233. https://doi.org/10.1111/mec.14584 (2018).
Forester, B. R., Jones, M. R., Joost, S., Landguth, E. L. & Lasky, J. R. Detecting spatial genetic signatures of local adaptation in heterogeneous landscapes. Mol. Ecol. 25, 104–120. https://doi.org/10.1111/mec.13476 (2016).
Blanco-Pastor, J. L. et al. Annual and perennial Medicago show signatures of parallel adaptation to climate and soil in highly conserved genes. Mol. Ecol. 30, 4448–4465. https://doi.org/10.1111/mec.16061 (2021).
Bélanger, G. et al. Winter damage to perennial forage crops in eastern Canada: Causes, mitigation, and prediction. Can. J. Plant Sci. 86, 33–47. https://doi.org/10.4141/P04-171 (2006).
Castonguay, Y. & Nadeau, P. Enzymatic control of soluble carbohydrate accumulation in cold-acclimated crowns of alfalfa. Crop Sci. 38, 1183–1189. https://doi.org/10.2135/cropsci1998.0011183X003800050012x (1998).
Castonguay, Y., Nadeau, P., Lechasseur, P. & Chouinard, L. Differential accumulation of carbohydrates in alfalfa cultivars of contrasting winterhardiness. Crop Sci. 35, 509–516. https://doi.org/10.2135/cropsci1995.0011183X003500020038x (1995).
Castonguay, Y., Laberge, S., Brummer, E. C. & Volenec, J. J. Alfalfa winter hardiness: A research retrospective and integrated perspective. Adv. Agron. 90, 203–265. https://doi.org/10.1016/S0065-2113(06)90006-6 (2006).
Reynolds, J. H. Carbohydrate trends in alfalfa (Medicago sativa L.) roots under several forage harvest schedules1. Crop Sci. 11, 103–106. https://doi.org/10.2135/cropsci1971.0011183X001100010036x (1971).
Misar, C. G., Xu, L., Gates, R. N., Boe, A. & Johnson, P. S. Stand persistence and forage yield of 11 alfalfa (Medicago sativa) populations in semiarid rangeland. Rangel. Ecol. Manage. 68, 79–85. https://doi.org/10.1016/j.rama.2014.12.012 (2015).
Kang, Y. et al. System responses to long-term drought and re-watering of two contrasting alfalfa varieties. Plant J. 68, 871–889. https://doi.org/10.1111/j.1365-313X.2011.04738.x (2011).
Sakiroglu, M. & Brummer, E. C. Identification of loci controlling forage yield and nutritive value in diploid alfalfa using GBS-GWAS. Theor. Appl. Genet. 130, 261–268. https://doi.org/10.1007/s00122-016-2782-3 (2017).
Elshire, R. J. et al. A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PLoS ONE 6, e19379. https://doi.org/10.1371/journal.pone.0019379 (2011).
Annicchiarico, P. et al. Accuracy of genomic selection for alfalfa biomass yield in different reference populations. BMC Genom. 16, 1020. https://doi.org/10.1186/s12864-015-2212-y (2015).
Jia, C. et al. Genomic prediction for 25 agronomic and quality traits in alfalfa (Medicago sativa). Front. Plant Sci. 9, 1220–1220. https://doi.org/10.3389/fpls.2018.01220 (2018).
Oakley, R. A. & Garver, S. Medicago falcata, a yellow-flowered alfalfa https://books.google.ca/books?hl=en&lr=&id=WtVKKmrQ6iUC&oi=fnd&pg=PA2&dq=Medicago+falcata,+a+yellow-flowered+alfalfa&ots=M0nuIYY6Eq&sig=m0u63836PYKbGeMeYrU-zj54SU4&redir_esc=y#v=onepage&q=Medicago%20falcata%2C%20a%20yellow-flowered%20alfalfa&f=false (1917).
Riday, H. & Brummer, E. C. Heterosis of agronomic traits in alfalfa. Crop Sci. 42, 1081–1087. https://doi.org/10.2135/cropsci2002.1081 (2002).
Riday, H. & Brummer, E. C. Morphological variation of Medicago sativa subsp. falcata genotypes and their hybrid progeny. Euphytica 138, 1–12. https://doi.org/10.1023/B:EUPH.0000047049.43566.1a (2004).
Barnes, D. Alfalfa Germplasm in the United States: Genetic Vulnerability, Use, Improvement, and Maintenance https://handle.nal.usda.gov/10113/CAT78694267 (1977).
Heinrichs, D. H. & Bolton, J. L. Rambler Alfalfa https://publications.gc.ca/collections/collection_2015/aac-aafc/A53-1030-1958-eng.pdf (1958).
Heinrichs, D. Roamer alfalfa. Can. J. Plant Sci. 47, 220–221. https://doi.org/10.4141/cjps67-040 (1967).
Heinrichs, D. H. Drylander alfalfa. Can. J. Plant Sci. 51, 430–432. https://doi.org/10.4141/cjps71-084 (1971).
Heinrichs, D. H., Lawrence, T. & McElgunn, J. D. Rangelander Alfalfa. Can. J. Plant Sci. 59, 491–492. https://doi.org/10.4141/cjps79-076 (1979).
Iversen, C., Meijer, G. & Langer, R. Types and varieties of lucerne. Lucerne Crop 1967, 74–84 (1967).
Jung, G. A. & Larson, K. L. Cold, drought, and heat tolerance. Alfalfa Sci. Technol. 15, 185–209. https://doi.org/10.2134/agronmonogr15.c9 (1972).
Baenziger, H. Algonquin alfalfa. Can. J. Plant Sci. 55, 1093–1094. https://doi.org/10.4141/cjps75-173 (1975).
Yoder, J. B. et al. Genomic signature of adaptation to climate in medicago truncatula. Genetics 196, 1263–1275. https://doi.org/10.1534/genetics.113.159319 (2014).
Willms, W., Adams, B. & McKenzie, R. Overview: Anthropogenic changes of Canadian grasslands. Arthropods Can. Grassl. 2, 1–22. https://doi.org/10.3752/9780968932155.ch1 (2011).
Garver, S. Alfalfa in South Dakota: Twenty-One Years of Research at the Redfield Station https://openprairie.sdstate.edu/agexperimentsta_bulletins/383/ (1946).
Katepa-Mupondwa, F., Singh, A., Smith, S. Jr. & McCaughey, W. Grazing tolerance of alfalfa (Medicago spp.) under continuous and rotational stocking systems in pure stands and in mixture with meadow bromegrass (Bromus riparius Rehm. Syn. B. biebersteinii Roem & Schult). Can. J. Plant Sci. 82, 337–347. https://doi.org/10.4141/P00-017 (2002).
Van Keuren, R. & Marten, G. Pasture production and utilization. Alfalfa Sci. Technol. 15, 641–658. https://doi.org/10.2134/agronmonogr29.c16 (1972).
McLeod, J., Muri, R., Jefferson, P., Bittman, S. & McCartney, D. Yellowhead alfalfa. Can. J. Plant Sci. 89, 653–655. https://doi.org/10.4141/CJPS08224 (2009).
Bhatt, J. Growth and flowering of cotton (Gossypium hirsutum L.) as affected by daylength and temperature. J. Agric. Sci. 89, 583–587. https://doi.org/10.1017/S0021859600061360 (1977).
Bula, R. & Massengale, M. Environmental physiology. Alfalfa Sci. Technol. 15, 167–184. https://doi.org/10.2134/agronmonogr15.c8 (1972).
Nelson, C. & Smith, D. Growth of birdsfoot trefoil and alfalfa. IV. carbohydrate reserve levels and growth analysis under two temperature regimes1. Crop Sci. 9, 589–591. https://doi.org/10.2135/cropsci1969.0011183X000900050022x (1969).
Arzani, H. et al. Phenological effects on forage quality of five grass species. J. Range Manag. 57, 624–629. https://doi.org/10.2111/1551-5028(2004)057[0624:PEOFQO]2.0.CO;2 (2004).
Ren, L., Bennett, J. A., Coulman, B., Liu, J. & Biligetu, B. Forage yield trend of alfalfa cultivars in the Canadian prairies and its relation to environmental factors and harvest management. Grass Forage Sci. 76, 390–399. https://doi.org/10.1111/gfs.12513 (2021).
Berdahl, J., Wilton, A. & Frank, A. Survival and agronomic performance of 25 alfalfa cultivars and strains interseeded into rangeland. Rangel. Ecol. Manage./J. Range Manage. Arch. 42, 312–316. https://doi.org/10.2307/3899501 (1989).
Bittman, S., Waddington, J. & McCartney, D. Performance of alfalfa strains grown in mixture with smooth bromegrass as affected by management. Can. J. Plant Sci. 71, 1029–1037. https://doi.org/10.4141/cjps91-145 (1991).
McKenzie, J. S., Paquin, R. & Duke, S. H. Cold and heat tolerance. Alfalfa Alfalfa Improvement 29, 259–302. https://doi.org/10.2134/agronmonogr29.c8 (1988).
Cutforth, H., Jones, K. & Lang, T. Agroclimate of the Brown soil zone of southwestern Saskatchewan research branch. Agric. Can. Publ. 1993, 251 (1993).
Schindler, D. W. & Donahue, W. F. An impending water crisis in Canada’s western prairie provinces. Proc. Natl. Acad. Sci. 103, 7210–7216. https://doi.org/10.1073/pnas.0601568103 (2006).
Kurek, I., Aviezer, K., Erel, N., Herman, E. & Breiman, A. The wheat peptidyl prolyl cis-trans-isomerase FKBP77 is heat induced and developmentally regulated. Plant Physiol. 119, 693–704. https://doi.org/10.1104/pp.119.2.693 (1999).
Mikami, K., Katagiri, T., Iuchi, S., Yamaguchi-Shinozaki, K. & Shinozaki, K. A gene encoding phosphatidylinositol-4-phosphate 5-kinase is induced by water stress and abscisic acid in Arabidopsis thaliana. Plant J. 15, 563–568. https://doi.org/10.1046/j.1365-313X.1998.00227.x (1998).
Liu, H., Sun, Z., Hu, L. & Yue, Z. Genome-wide identification of PIP5K in wheat and its relationship with anther male sterility induced by high temperature. BMC Plant Biol. 21, 598. https://doi.org/10.1186/s12870-021-03363-1 (2021).
Ouellet, C. Winter hardiness and survival of forage crops in Canada. Can. J. Plant Sci. 56, 679–689. https://doi.org/10.4141/cjps76-108 (1976).
Yang, X. et al. OsTTG1, a WD40 repeat gene, regulates anthocyanin biosynthesis in rice. Plant J. 107, 198–214. https://doi.org/10.1111/tpj.15285 (2021).
Nemesio-Gorriz, M. et al. Identification of Norway spruce MYB-bHLH-WDR transcription factor complex members linked to regulation of the flavonoid pathway. Front. Plant Sci. 8, 305. https://doi.org/10.3389/fpls.2017.00305 (2017).
Shvarts, M., Borochov, A. & Weiss, D. Low temperature enhances petunia flower pigmentation and induces chalcone synthase gene expression. Physiol. Plant. 99, 67–72. https://doi.org/10.1111/j.1399-3054.1997.tb03432.x (1997).
Islam, M. S., Jalaluddin, M., Garner, J. O., Yoshimoto, M. & Yamakawa, O. Artificial shading and temperature influence on anthocyanin compositions in sweetpotato leaves. HortScience 40, 176–180. https://doi.org/10.21273/HORTSCI.40.1.176 (2005).
Movahed, N. et al. The grapevine VviPrx31 peroxidase as a candidate gene involved in anthocyanin degradation in ripening berries under high temperature. J. Plant. Res. 129, 513–526. https://doi.org/10.1007/s10265-016-0786-3 (2016).
Wang, L., Attoe, O. & Truog, E. Effect of lime and fertility levels on the chemical composition and winter survival of alfalfa 1. Agron. J. 45, 381–384. https://doi.org/10.2134/agronj1953.00021962004500080010x (1953).
Smith, D. Effects of potassium topdressing a low fertility silt loam soil on alfalfa herbage yields and composition and on soil K values 1. Agron. J. 67, 60–64. https://doi.org/10.2134/agronj1975.00021962006700010016x (1975).
Collins, M., Lang, D. & Kelling, K. Effects of phosphorus, potassium, and sulfur on alfalfa nitrogen-fixation under field conditions1. Agron. J. 78, 959–963. https://doi.org/10.2134/agronj1986.00021962007800060005x (1986).
Berg, W. K., Lissbrant, S., Cunningham, S. M., Brouder, S. M. & Volenec, J. J. Phosphorus and potassium effects on taproot C and N reserve pools and long-term persistence of alfalfa (Medicago sativa L.). Plant Sci. 272, 301–308. https://doi.org/10.1016/j.plantsci.2018.02.026 (2018).
Counce, P. A., Bouton, J. H. & Brown, R. H. Screening and Characterizing alfalfa for persistence under mowing and continuous grazing1. Crop Sci. https://doi.org/10.2135/cropsci1984.0011183X002400020017x (1984).
Teixeira, E. I., Moot, D. J. & Mickelbart, M. V. Seasonal patterns of root C and N reserves of lucerne crops (Medicago sativa L.) grown in a temperate climate were affected by defoliation regime. Eur. J. Agron. 26, 10–20. https://doi.org/10.1016/j.eja.2006.08.010 (2007).
Hull, A. K., Vij, R. & Celenza, J. L. Arabidopsis cytochrome P450s that catalyze the first step of tryptophan-dependent indole-3-acetic acid biosynthesis. Proc. Natl. Acad. Sci. 97, 2379–2384. https://doi.org/10.1073/pnas.040569997 (2000).
Zhao, Y. et al. Trp-dependent auxin biosynthesis in Arabidopsis: Involvement of cytochrome P450s CYP79B2 and CYP79B3. Genes Dev. 16, 3100–3112. https://doi.org/10.1101/gad.1035402 (2002).
Wittstock, U. & Halkier, B. A. Glucosinolate research in the Arabidopsis era. Trends Plant Sci. 7, 263–270. https://doi.org/10.1016/S1360-1385(02)02273-2 (2002).
Del-Carmen-Martínez-Ballesta, M., Moreno, D. A. & Carvajal, M. The physiological importance of glucosinolates on plant response to abiotic stress in Brassica. Int. J. Mol. Sci. 14, 11607–11625. https://doi.org/10.3390/ijms140611607 (2013).
Liu, A. et al. GsSKP21, a Glycine soja S-phase kinase-associated protein, mediates the regulation of plant alkaline tolerance and ABA sensitivity. Plant Mol. Biol. 87, 111–124. https://doi.org/10.1007/s11103-014-0264-z (2015).
Seim, E. C., Caldwell, A. C. & Rehm, G. W. Sulfur response by alfalfa (Medicago sativa L.) on a sulfur-deficient soil1. Agron. J. 61, 368–371. https://doi.org/10.2134/agronj1969.00021962006100030009x (1969).
Maruyama-Nakashita, A., Nakamura, Y., Watanabe-Takahashi, A., Yamaya, T. & Takahashi, H. Induction of SULTR1; 1 sulfate transporter in Arabidopsis roots involves protein phosphorylation/dephosphorylation circuit for transcriptional regulation. Plant Cell Physiol. 45, 340–345. https://doi.org/10.1093/pcp/pch029 (2004).
Kopriva, S. Regulation of sulfate assimilation in Arabidopsis and beyond. Ann. Bot. 97, 479–495. https://doi.org/10.1093/aob/mcl006 (2006).
Peleg, Z. & Blumwald, E. Hormone balance and abiotic stress tolerance in crop plants. Curr. Opin. Plant Biol. 14, 290–295. https://doi.org/10.1016/j.pbi.2011.02.001 (2011).
Yuan, X. et al. Global analysis of ankyrin repeat domain C3HC4-type RING finger gene family in plants. PLoS ONE 8, e58003. https://doi.org/10.1371/journal.pone.0058003 (2013).
Elhaissoufi, W. et al. Phosphate solubilizing rhizobacteria could have a stronger influence on wheat root traits and aboveground physiology than rhizosphere P solubilization. Front. Plant Sci. 11, 979. https://doi.org/10.3389/fpls.2020.00979 (2020).
Aslam, M. M. et al. Identification of ABC transporter G subfamily in white lupin and functional characterization of L.albABGC29 in phosphorus use. BMC Genom. 22, 723. https://doi.org/10.1186/s12864-021-08015-0 (2021).
El-Kherbawy, M., Angle, J., Heggo, A. & Chaney, R. Soil pH, rhizobia, and vesicular-arbuscular mycorrhizae inoculation effects on growth and heavy metal uptake of alfalfa (Medicago sativa L.). Biol. Fertil. Soils 8, 61–65. https://doi.org/10.1007/BF00260517 (1989).
Li, J. et al. A chaperone function of no catalase activity1 is required to maintain catalase activity and for multiple stress responses in arabidopsis. Plant Cell 27, 908–925. https://doi.org/10.1105/tpc.114.135095 (2015).
Zhao, Z. et al. Physiological and TMT-based proteomic analysis of oat early seedlings in response to alkali stress. J. Proteom. 193, 10–26. https://doi.org/10.1016/j.jprot.2018.12.018 (2019).
Chang, Y. et al. Genome-wide identification and characterization of ACBP gene family in Populus reveal salinity alkali-responsive profiles. J. For. Res. https://doi.org/10.1007/s11676-022-01485-2 (2022).
Li, X. et al. Development of an alfalfa SNP array and its use to evaluate patterns of population structure and linkage disequilibrium. PLoS ONE 9, e84329. https://doi.org/10.1371/journal.pone.0084329 (2014).
Zhang, F. et al. Whole-genome sequencing on 220 alfalfa (Medicago sativa L.) accessions identified loci associated with fall dormancy. bioRxiv https://doi.org/10.21203/rs.3.rs-1006624/v1 (2021).
Bonner, D. M. Comparative Water Relations and Drought Tolerance Among Alfalfa Cultivars http://hdl.handle.net/1993/753 (1997).
Guerrero, J., Andrello, M., Burgarella, C. & Manel, S. Soil environment is a key driver of adaptation in Medicago truncatula: new insights from landscape genomics. New Phytol. 219, 378–390. https://doi.org/10.1111/nph.15171 (2018).
Burgarella, C. et al. Adaptation to climate through flowering phenology: A case study in Medicago truncatula. Mol. Ecol. 25, 3397–3415. https://doi.org/10.1111/mec.13683 (2016).
Blanco-Pastor, J. L. et al. Canonical correlations reveal adaptive loci and phenotypic responses to climate in perennial ryegrass. Mol. Ecol. Resour. 21, 849–870. https://doi.org/10.1111/1755-0998.13289 (2021).
Humphries, A. W. et al. Characterization and pre-breeding of diverse alfalfa wild relatives originating from drought-stressed environments. Crop Sci. 61, 69–88. https://doi.org/10.1002/csc2.20274 (2021).
Soil Classification Working Group. The Canadian System of Soil Classification, (3rd edition). https://sis.agr.gc.ca/cansis/taxa/cssc3/index.html (1998).
Lipka, A. E. et al. Accelerating the switchgrass (Panicum virgatum L.) breeding cycle using genomic selection approaches. PLoS ONE 9, e112227. https://doi.org/10.1371/journal.pone.0112227 (2014).
Oksanen, J. et al. Vegan community ecology package: Ordination methods, diversity analysis and other functions for community and vegetation ecologists. R package ver, 2–3. https://doi.org/10.1111/j.1654-1103.2003.tb02228.x (2015).
Borcard, D., Gillet, F. & Legendre, P. Numerical Ecology with R 688 (Springer, 2011). https://doi.org/10.1007/978-1-4419-7976-6.
Dormann, C. F. et al. Collinearity: a review of methods to deal with it and a simulation study evaluating their performance. Ecography 36, 27–46. https://doi.org/10.1111/j.1600-0587.2012.07348.x (2013).
Legendre, P. & Legendre, L. Numerical Ecology (Elsevier, 2012).
Wang, J. et al. Package ‘robust’, https://cran.r-project.org/web/packages/robust/index.html (2022).
Luu, K., Bazin, E. & Blum, M. G. pcadapt: An R package to perform genome scans for selection based on principal component analysis. Mol. Ecol. Resour. 17, 67–77. https://doi.org/10.1111/1755-0998.12592 (2017).
François, O., Martins, H., Caye, K. & Schoville, S. D. Controlling false discoveries in genome scans for selection. Mol. Ecol. 25, 454–469. https://doi.org/10.1111/mec.13513 (2016).
Zuur, A. F., Ieno, E. N. & Elphick, C. S. A protocol for data exploration to avoid common statistical problems. Methods Ecol. Evol. 1, 3–14. https://doi.org/10.1111/j.2041-210X.2009.00001.x (2010).
Storey, J. D., Bass, A. J., Dabney, A. & Robinson, D. qvalue: Q-value Estimation for False Discovery Rate Control https://github.com/StoreyLab/qvalue (2015).
Schmieder, R. & Edwards, R. Quality control and preprocessing of metagenomic datasets. Bioinformatics 27, 863–864. https://doi.org/10.1093/bioinformatics/btr026 (2011).
Sabre-barcode-demultiplexing. Sabre—A Barcode Demultiplexing and Trimming Tool for FastQ Files https://github.com/najoshi/sabre (2013).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinform. (Oxf., Engl.) 30, 2114–2120. https://doi.org/10.1093/bioinformatics/btu170 (2014).
Chen, H. et al. Allele-aware chromosome-level genome assembly and efficient transgene-free genome editing for the autotetraploid cultivated alfalfa. Nat. Commun. 11, 2494. https://doi.org/10.1038/s41467-020-16338-x (2020).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinform. (Oxf., Engl.) 25, 1754–1760. https://doi.org/10.1093/bioinformatics/btp324 (2009).
Garrison, E. & Marth, G. Haplotype-based variant detection from short-read sequencing. arXiv:1207.3907. https://doi.org/10.48550/arXiv.1207.3907 (2012).
Browning, B. L., Zhou, Y. & Browning, S. R. A one-penny imputed genome from next-generation reference panels. Am. J. Hum. Genet. 103, 338–348. https://doi.org/10.1016/j.ajhg.2018.07.015 (2018).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959. https://doi.org/10.1093/genetics/164.4.1567 (2000).
Falush, D., Stephens, M. & Pritchard, J. K. Inference of population structure using multilocus genotype data: Linked loci and correlated allele frequencies. Genetics 164, 1567–1587. https://doi.org/10.1093/genetics/164.4.1567 (2003).
Falush, D., Stephens, M. & Pritchard, J. K. Inference of population structure using multilocus genotype data: Dominant markers and null alleles. Mol. Ecol. Notes 7, 574–578. https://doi.org/10.1111/j.1471-8286.2007.01758.x (2007).
Hubisz, M. J., Falush, D., Stephens, M. & Pritchard, J. K. Inferring weak population structure with the assistance of sample group information. Mol. Ecol. Resour. 9, 1322–1332. https://doi.org/10.1111/j.1755-0998.2009.02591.x (2009).
Jombart, T., Devillard, S. & Balloux, F. Discriminant analysis of principal components: A new method for the analysis of genetically structured populations. BMC Genet. 11, 1–15. https://doi.org/10.1186/1471-2156-11-94 (2010).
Jombart, T. et al. Package ‘adegenet’. Bioinform. Appl. Note 24, 1403–1405. https://doi.org/10.1093/bioinformatics/btn129 (2008).
R Core Team. R: A language and Environment for Statistical Computing https://www.R-project.org (2019).
Evanno, G., Regnaut, S. & Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol. Ecol. 14, 2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x (2005).
Earl, D. A. & von-Holdt, B. M. STRUCTURE HARVESTER: A website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resourc. 4, 359–361. https://doi.org/10.1007/s12686-011-9548-7 (2012).
Jakobsson, M. & Rosenberg, N. A. CLUMPP: A cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23, 1801–1806. https://doi.org/10.1093/bioinformatics/btm233 (2007).
Francis, R. M. pophelper: An R package and web app to analyse and visualize population structure. Mol. Ecol. Resour. 17, 27–32. https://doi.org/10.1111/1755-0998.12509 (2017).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly (Austin) 6, 80–92. https://doi.org/10.4161/fly.19695 (2012).
Covarrubias-Pazaran, G. Genome-assisted prediction of quantitative traits using the R package sommer. PLoS ONE 11, e0156744. https://doi.org/10.1371/journal.pone.0156744 (2016).
Acknowledgements
The author would like to thank Dashnyam Byambatseren for technical assistance, and Digital Research Alliance of Canada for providing the high-performance computing service as well as technical support.
Funding
This project was funded by the Agriculture Development Fund of Saskatchewan and the Western Grain Research Foundation.
Author information
Authors and Affiliations
Contributions
B.B. conceived and designed the experiments. H.W. performed the experiments, analyzed the data and wrote the paper. B.B., B.C., Y.B., and B.T. participated in discussion and revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, H., Coulman, B., Bai, Y. et al. Genetic diversity and local adaption of alfalfa populations (Medicago sativa L.) under long-term grazing. Sci Rep 13, 1632 (2023). https://doi.org/10.1038/s41598-023-28521-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-28521-3
This article is cited by
-
Analysis of forage quality, volatile organic compounds and metabolic pathways in alfalfa (Medicago sativa L.) at different stages based on electronic nose and GC-MS
Chemical and Biological Technologies in Agriculture (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.