Abstract
Calmodulin-like proteins (CML) are important calcium signal transduction proteins in plants. CML genes have been analyzed in several plants. However, little information on CML in Phaseolus vulgare is available. In this study, we identified 111 PvCMLs distributed on eleven chromosomes. Phylogenetic analysis classified them into seven subfamilies. Cis-acting element prediction showed that PvCML contained elements related to growth and development, response to abiotic stress and hormones. Moreover, the majority of PvCMLs showed different expression patterns in most of the nine tissues and developmental stages which indicated the role of PvCML in the growth and development of common bean. Additionally, the common bean was treated with melatonin by seed soaking, and root transcriptome at the 5th day and qRT-PCR of different tissue at several stages were performed to reveal the response of PvCML to the hormone. Interestingly, 9 PvCML genes of subfamily VI were detected responsive to exogenous melatonin, and the expression dynamics of nine melatonin response PvCML genes after seed soaking with melatonin were revealed. Finally, the protein interaction network analysis of nine melatonin responsive PvCMLs was constructed. The systematic analysis of the PvCML gene family provides theoretical support for the further elucidation of their functions, and melatonin response molecular mechanism of the CML family in P. vulgaris.
Similar content being viewed by others
Introduction
Calcium (Ca2+) is a dynamic element and one of the most important second messengers in plant cell signal transduction1. Calcium signaling plays a fundamental role in cellular activities. Different Ca2+ sensing proteins can detect different Ca2+ signals of cytoplasmic free calcium2. One of the most important protein chelators for Ca2+ is the protein with the EF-Hand domain3. Four important protein families contain EF-hand domains, including CAMs (Calmodulins)4, CMLs (calmodulin-like proteins)5, CDPKs (Ca2+-dependent protein kinases)6, and CBLs (Calcineurin B-like proteins)7.
CMLs are novel plant-specific Ca2+ sensors with the plant-specific EF-Hand domain8. The structure and biochemical features of CML proteins indicate that they have similar properties to CAM3. However, there is one Dx3D motif in CML and four calcium-binding DxD motifs in CAM9. It is an important distinction between CML and CAM. By bioinformatic methods, CML genes have been identified and analyzed in many plants, such as 50 CML genes in Arabidopsis thaliana5, 32 CML genes in rice10, 52 CML genes in tomato11, 144 CML genes in soybean12, 46 CML genes in Medicago truncatula13, 62 CML genes in grapevine14, and 79 CML genes in Brassica rapa L.15. In general, closely related members of CML family have high sequence similarity, and motif distribution and gene structure were arranged similarly in the same subfamily. Moreover, CML genes have been shown to play important roles in plant growth and development, cell metabolism, innate immunity, abiotic and biological stress disease resistance5,16,17,18,19. For example, the ShCML44 gene of tomato involved in responses to various stress, and the overexpression of ShCML44 can improve the tolerance of plants to cold, drought, and salt stress20. AtCML8 expression can be induced by salicylic acid or NaCl treatment21. The expression of AtCML9 was induced by abscisic acid and some abiotic stresses, which indicated that AtCML9 plays a role in salt tolerance by modulating ABA-mediated pathways16. AtCML20 inhibited ABA-induced stomatal closure and drought tolerance22, whereas AtCML24 inhibited pollen tube growth, autophagy, abscisic acid response, and ion stress23. In addition, a large number of cis-elements related to abiotic stress and hormones were predicted in the promoter region of the CML gene. Furthermore, previous experiments reveal that these CML genes showed different expression profiles after stress and hormone treatment20.
Melatonin (MT) was discovered in the bovine pineal gland in 195824. Plant melatonin appears to be a multi-regulatory molecule with a variety of hormone activities in plants, including the regulation of seed germination rate, plant growth, and development process25,26. Phaseolus vulgaris L. is an edible leguminous plant27. Due to its economic value, especially in developing countries, many researchers have focused on the yield increase of the common bean28. Exogenous application of melatonin can regulate the growth and development of the common bean and significantly improve root growth of the common bean under salt stress29. Moreover, previous studies revealed that exogenous melatonin was sensed by receptor CAND2/PMTR1, which activated downstream of Ca2+ signal transduction depending on calcium sensors, including CMLs (Cam-like proteins)30. Therefore, exogenous melatonin response in the common bean is closely related to intracellular calcium and CML. However, the features, expression patterns, and the response to melatonin of the CML family in common bean is unclear until now.
In this paper, the CML gene family of common beans was identified, and there were 111 PvCMLs in P. vulgaris. Its gene structure and cis-acting element distribution were analyzed. Subsequently, we analyzed publicly available transcriptome databases and determined the spatial–temporal expression pattern of all PvCML genes in several tissues. Our root transcriptomic data further revealed specific melatonin responsive PvCML, which belonged to a CMLs subfamily VI. Spatial–temporal expression profiling in different tissues of nine specific melatonin responsive CMLs was verified using qRT-PCR. Meanwhile, we also analyzed the expression dynamics of nine melatonin response PvCML genes in the root at different time points after melatonin seed-soaking treatment using qRT-PCR. This work provides a foundation for further functional study of CML genes and the molecular mechanism of melatonin.
Materials and methods
Identification of CML in P. vulgaris
Protein sequences of P. vulgaris (v2.1) were downloaded from Phytozome database (v13) (https://phytozome.jgi.doe.gov)31. The reported CML protein sequences in A. thaliana and rice were downloaded from the TAIR database (https://www.arabidopsis.org)32 and TIGR database (http://rice.plantbiology.msu.edu)33, respectively. Then, 32 OsCML and 50 AtCML proteins were used as query sequences to perform BLASTP searching (E-value < 1e−5)34. To remove false PvCML genes, we screened the candidate CML genes according to the main character of conserved EF-hand domains9. Subsequently, InterPro 86.0 (http://www.ebi.ac.uk/interpro) and SMART 9.0 (http://smart.embl-heidelberg.de/) software was applied to verify the reliability of the EF-hand domain prediction. Additionally, a Dx3D motif was also checked to distinguish the CML family from the CAM family.
Chromosomal location and physicochemical characterization of PvCML genes
The sequences of PvCML were used to retrieve their chromosomal locations in the P. vulgaris genome databases Phytozome 12. The software TBtools was used to analyze the chromosomal location. Each PvCML gene was named on the basis of its precise position on the chromosome.
The physicochemical properties such as amino acid number, isoelectric point (pI), molecular weight, and coefficient of protein were analyzed by Protparam tool of ExPASy (https://web.expasy.org/protparam). The Plant-mPLoc tool (http://www.csbio.sjtu.edu.cn/bioinf/plant-multi) was used to predict subcellular localization of CMLs in P. vulgaris.
Phylogenetic analysis of CML
Multiple sequences alignment of 111 PvCML proteins were performed using Clustal W, and a phylogenetic tree was constructed by MEGA X with the Maximum-likelihood method, and Jones–Taylor–Thornton (JTT) + G model was set, and 1000 bootstrap replicates was performed.
Analysis of conserved domain, gene structure and cis-acting element of PvCML genes
The exon–intron structure analysis of PvCML genes was conducted using the TBtools program with default parameters. The conserved motifs were analyzed with MEME (https://meme-suite.org). The 2.0 kb upstream sequences of PvCML genes were analyzed using the PlantCARE (http://bioinformatics.psb.ugent.be/webtools/plantcare/html) to identify the cis-acting elements in the promoter region of PvCML.
Duplication events analysis of CML
The PvCML was mapped to the chromosomes according to the chromosomal locations provided by Ensembl plants database (https://plants.ensembl.org/index.html). The gene duplication events were analyzed by Multiple Collinearity Scan toolkit (MCScanX) with default parameters. We used TBtools with default parameters to calculate Ka, Ks, Ka/Ks.
Tissue specific expression profile of PvCML gene family
RNA-seq data from 9 tissues or organs were retrieved from the phytozome database. The transcript data of the PvCML gene family were selected and sorted, then a heatmap was constructed by the TBtools software. Fragments per kilobase of exon per million fragments mapped (FPKM) value was transformed to log2 (value + 1)35.
Gene expression analysis of PvCML gene response to melatonin treatments
Common bean varieties are provided by the National Coarse Cereals Engineering Research Center. In this study, P. vulgaris L. var. Jiyin 1 was analyzed and the formal identification of the plant materials was undertaken by Mr. M. Li. We got the permission to collect the plant samples and all methods were carried out in accordance with relevant guidelines and regulations. Our root transcriptome data of P. vulgaris after melatonin treatment at 5 day was analyzed as above, PvCML genes response to melatonin treatments were identified. The assembled gene dataset, deposited at the National Center for Biotechnology Information under the accession number PRJNA55837615 (http://www.ncbi.nlm.nih.gov/bioproject/).
To further study the melatonin effect on the expression of these PvCML genes, P. vulgaris L. var. Jiyin 1 (JY) was treated with 100 µmol/L melatonin by seeds soaking, which had the most obvious effect on the growth of common bean, and distilled water was used as a control (H2O). Then, JY was cultured in petri dish (ϕ12 × 1.5 cm) for 7 days at the temperature 25 °C in dark, and radicle were sampled at different stages (3 d, 5 d, 7 d). Meanwhile, some JY treated by melatonin was planted in soil and cultured in greenhouse with 18 h/8 h light at 25 °C. The leaf, stem, hypocotyl, and root of JY were sampled at 10 d. All samples were immediately frozen in liquid nitrogen and stored at − 80 °C for further experiments.
Total RNA of all samples (hypocotyls, stems, leaves, and roots) was extracted by TRIzol reagent (BIOMARS, Beijing, China). RNA was subsequently reverse-transcripted to cDNA using a SuperMix (Innovagene, Beijing). For qRT-PCR primers of ninmelatonin responsive genes, all 111 PvCML nucleotide sequences were aligned, and the primers were designed in the region of difference. Moreover, these specific primers were further confirmed by primer-blast analysis. The qRT-PCR primers were listed in Supplementary Table S1. The normal PCR was performed by these primers, and the PCR products were detected on agarose gel electrophoresis to make sure of the purity of amplification. Then, qRT-PCR was conducted using the CFX96 qPCR system (Bio-Rad). The qRT-PCR protocol was as follows: 95 °C for 3 min; 40 cycles of 95 °C for 15 s and 60 °C for 30 s. Three biological replicates and three technical replicates were performed. Actin (KF569629) was used as an internal reference gene, and the relative expression level of PvCML genes were calculated using 2−ΔΔCt36.
Protein interaction analysis and visualization
The protein interaction network of PvCML proteins which responded to exogenous melatonin was analyzed using the STRING website (https://www.string-db.org), and the protein interaction network was visualized with Cytoscape software37. Moreover, annotation of these proteins was performed in the KEGG website (https://www.kegg.jp/)38.
Results
Identification and characterization of CML family members in P. vulgaris
In total, 124 CML protein sequences with EF-hand domains were identified in P. vulgaris using BLASTp, and 111 genes (PvCML1-111) were confirmed and classified in the CML family by InterPro and SMART analysis. These PvCML genes sequences were further used to retrieve the chromosomal locations of PvCML genes. The results showed that PvCML genes were distributed on all chromosomes of P. vulgaris. Specifically, 16 PvCMLs, 15 PvCMLs and 18 PvCMLs were found on chromosomes 1, 2 and 3, respectively. Three PvCMLs were distributed on chromosomes 4 and 10; chromosomes 5 and 9 contained nine PvCMLs; chromosome 6, 7, 8, and 11 contained eight, twelve, thirteen and five PvCMLs, respectively (Fig. 1).
Physicochemical properties analysis showed that the predicted molecular weight of PvCML genes ranged from 9240.24 Da (PvCML 24) to 89,386.22 Da (PvCML 1), and the number of amino acids of the PvCMLs ranged from 80 AA (PvCML 24) to 789 AA (PvCML 1). The predicted pI varied from 3.90 (PvCML 73) to 9.79 (PvCML 97). The subcellular localization results showed that 54 PvCMLs were located on the cell membrane, 29 PvCMLs on the nucleus, 3 PvCMLs on the cell membrane and nucleus, and the remaining PvCMLs on the cytoplasm, vacuole, and chloroplast (Supplementary Table S2).
Evaluation of gene structures and conserved motifs of the CMLs in P. vulgaris
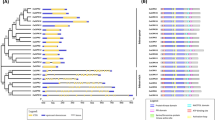
A phylogenetic tree was constructed based on the alignment of PvCMLs full-length amino acid sequences by MEGA software to demonstrate the structural classification of the PvCMLs. PvCMLs could be divided into seven subfamilies (I–VII) (Fig. 2a). Furthermore, conserved motifs analysis of these proteins showed that all 111 PvCMLs contained at least one EF-hands domain with D-x3-D structure. Ten different motifs were identified and their distribution were shown in Fig. 2b. The conserved sequences of ten motifs were listed in Supplementary Fig. S1. Each PvCML subfamily had a unique motif distribution model. Motif 1, 4, 6, 7, 8, and 10 were specifically presented in the subfamily I members. These motifs are protein kinase domains which are usually associated with ATP binding and protein kinase activity. Other family members (II-VII) were different in order or quantity of motif 2, 3, 5, and 9 (Fig. 2b). These four motifs were EF-hand domains associated with calcium-binding. Meanwhile, according to the motif sequence (Fig. S1), motifs 2, 3, 5, and 9 are EF-hand domains containing Dx3D structures. Dx3D was in the first place of EF-hand domain in 111 PvCML, which was consistent with the characteristics of CML proteins.
Phylogenetic relationships, gene structure, and motif compositions of PvCML genes. (a) Phylogenetic tree of PvCMLs. The phylogenetic tree was constructed with the maximum-likelihood method of MEGA X (1000 bootstrap replicates). The size of the blue circle represented bootstrap value. (b) Motif structure of PvCMLs. Different motifs were represented by different colors. Motif lengths were presented proportionally. (c) The gene structure of PvCMLs. The yellow box represented exon, and the green box represented the untranslated region. Intron represents by solid line.
To gain information about the conservation and difference of PvCMLs genes, we analyzed the exon–intron organization of PvCMLs genes. The results showed that the number of exons in PvCMLs ranged from 1 to 12, and forty-four PvCMLs (39.63%) contained only one exon. There were twenty-six PvCMLs (23.42%) with more than 8 exons. Most PvCMLs (53.15%) contained 2 to 7 exons (Fig. 2c). The subfamily I–V members and some members of subfamily VII contained multiple exons. The majority of subfamily VI members contained one exon. Above all, the phylogenetic tree and gene structure analysis demonstrated that the structural difference between PvCMLs may be related to its gene function within this family.
Prediction of cis-elements in the promoter sequences of PvCMLs genes
To explore the transcriptional regulation of PvCMLs gene, cis-elements in PvCMLs promoter regions (the 2000 bp upstream region from transcriptional start site) were predicted using Plant CARE. A total of 99 cis-regulatory elements were identified. Partial cis-acting elements of PvCML were further classified into plant hormone response element, growth and development response element, and abiotic stress response element. Phytohormone (ABA, MeJA, GA, and SA) responsive elements included TATC-box, ABRE, TCA-element, CGTCA-motif, P-box, GARE-motif, TGACG-motif, etc. (Fig. 3a). Cis-elements involved in plant growth and development were widely distributed in all genes (Fig. 3b). Additionally, four types of abiotic stresses (drought, low-temperature, anaerobic, and light) related elements, such as ABRE, ARE, LTR, GT1-motif, MBS, etc. were identified (Fig. 3c). The top ten cis-regulatory elements, with the exception of CAAT and TATA-box, were visualized, including 504 Box 4, 263 ERE, 241 G-box, 211 ARE, 208 ABRE, 142 GT1-motif, 106 TCT motif, 101 CGTCA-motif, 101 WUN-motif (Fig. 3d). These results indicated that PvCMLs genes might play critical roles not only in various plant developmental events, but also in phytohormone and abiotic responses in P. vulgaris.
Cis-elements in the promoter regions of PvCMLs. (a) The number distribution of phytohormones responsive element, (b) plant growth and development related element, (c) abiotic stress response element and (d) distribution of top 10 cis-regulatory elements in each PvCMLs gene. Element is indicated by different colors in the grid.
Phylogenetic analysis of PvCML proteins
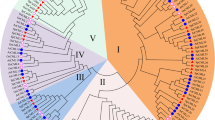
To evaluate the evolutionary relationships of the CMLs among common bean (P. vulgaris), rice (Oryza sativa L.) and A. thaliana, multiple sequence alignment was performed using the CML amino acid sequences. A comprehensive phylogenetic tree was constructed with 193 CML protein sequences, including 111 sequences from common bean (PvCMLs), 32 from rice (OsCMLs), and 50 from Arabidopsis (AtCML). These CML genes were classified into nine subgroups (Group I–Group IX) (Fig. 4). Group I included 13 CML members (2 AtCML, 4 OsCMLs and 7 PvCMLs); Group II included 35 CML members (12 AtCMLs, 3 OsCMLs and 20 PvCMLs); Group III contained 13 CML members (5 AtCMLs and 8 PvCMLs); Group IV included 20 CML members (5 AtCMLs, 4 OsCMLs and 11 PvCMLs); Group V and VIII subgroups only contained PvCML members, PvCML 28/96/29/43/17/106/4/67/85/34/38 and PvCML 64/93/33/1/2/23/27/62/89/14/80/76/79/25/102/71/72/94/12/78/108/21/40/86/22/45/111 formed a single V and VIII subgroups in the phylogenic tree; Group VI contained 13 CML members (6 AtCMLs, 3 OsCMLs and 4 PvCMLs); Group IX included 46 CML members (15 AtCMLs, 13 OsCMLs and 18 PvCMLs).
Maximum likelihood tree of CML proteins from P. Vulgaris, O. sativa and A. thaliana. The CML proteins in the branches from different species were indicated by different color: purple, P. Vulgaris; yellow, O. sativa; blue, A. thaliana. Different colors of branches represented different subfamilies. The size of the blue circle represented the bootstrap value.
Gene duplication and synteny analysis
To clarify the PvCML gene duplication events in common bean, the segmental duplication events in PvCML gene family were investigated. As shown in Fig. 5a, twenty-one gene duplication events formed by genes from different chromosomes. Chromosome 4 contains a duplicated segment, and chromosome 1, 2, 3, 6, and 7 each contain more than five PvCML duplicated segments. Other chromosomes (4, 5, 8, 9, 10, and 11) each contain no more than five duplicated segments. There are two pairs of duplicated segments on the same chromosome. To elucidate the role of selection pressure in evolution, the Ka, Ks, and Ka/Ks values of duplicated genes of the PvCML family were calculated using the TBtools software. Except for no value of two gene pairs (PvCML 39/PvCML 103 and PvCML 56/PvCML 68), the value of Ka/Ks of other duplicated PvCML gene pairs were obtained (Supplementary Table S4). The results showed that the mean Ka/Ks of dispersed duplication (35%) was 0.1834, while the mean Ka/Ks of the segmental duplication (65%) was 0.1534. The Ka/Ks values of all duplication events were < 1 which indicated that the driving force for PvCML family evolution was the purification selection.
In addition, the synteny relationship of CMLs between P. vulgaris and A. Thaliana was analysed. Twenty-five pairs of homologous CMLs were identified between P. vulgaris and A. thaliana (Fig. 5b). Seven genes (PvCML 13/15/19/31/73/74/81) had two homologs in Arabidopsis, while one gene (PvCML 60) had three homologs in Arabidopsis (Supplementary Table S4).
Expression profile analysis of PvCML genes in different tissues and melatonin treatment
Global gene expression data of. P. vulgaris are publicly available, including expression profiles of PvCMLs gene at different developmental stages. To explore the possible functions of the PvCMLs, the public transcript data of the PvCMLs genes in 9 different tissues and organs, including leaves, stem, nodule, root, flowers, flower buds, young pod, green mature pods, young trifoliates, were analyzed. The majority of PvCMLs showed different expression patterns in most of 9 tissues and developmental stages (Fig. 6a). Three PvCML genes (PvCML 19/63/90) were highly expressed in all tissues, whereas five PvCML genes (PvCML 59/14/71/2/28) were expressed at low expression levels in most stages. Some PvCMLs genes showed a tissue specific expression pattern. PvCML 17 and PvCML 24 were expressed only in flower buds. PvCML 53 was expressed at low expression levels in the root. PvCML 10 and PvCML 103 were highly expressed in flowers and flower buds. In addition, some PvCML genes showed similar expression patterns, implying that they may have similar functions in plant growth and development.
Transcriptome analysis of PvCML gene under different tissues and melatonin treatment. (a) Expression pattern of PvCML genes in different tissues based on the transcriptome data. (b) The expression profiles of PvCML genes in root after melatonin treatment. Transcript levels were depicted by different colors on the scale. Blue and red represented low and high expression levels, respectively.
In view of the presence of hormone-related cis elements in PvCML gene promoters, we further analyzed the effects of melatonin on the expression of PvCML genes. According to the previous experiment, we treated seeds of common bean with 100 μM melatonin by seed soaking, and the root transcriptome on the fifth day was sequenced.
Our transcriptome data from melatonin-treated roots revealed that nine PvCML genes (PvCML 6/8/38/41/44/53/54/82/107) in the exogenous melatonin treatment group displayed significantly higher expression levels than that of the control group, and the nine genes all belonged to the subfamily VI (Fig. 6b). This result indicated these genes in group VI plays a more important role in plant response to melatonin. Meanwhile, eight of the nine PvCML transcription patterns were identified from public transcriptome data except PvCML 82. PvCML 6 and PvCML 41 were highly expressed in green mature pods, PvCML 8 in flower tissues and roots, PvCML 38 in flower-related tissues, PvCML 44 in nodules, PvCML 53 and PvCML 54 in green mature pods and roots; Transcript level of PvCML 107 in all tissues were consistent.
qRT-PCR analysis of nine melatonin responsive PvCML genes
In order to verify the reliability of transcriptomic data, we used qRT-PCR to detect the expression levels of nine melatonin responsive PvCML genes at four developmental stages (3 d root, 5 d root, 7 d root, and 10 d root) after melatonin treatment and in different tissues (10 d root, 10 d stem, 10 d hypocotyl, and 10 d leave).
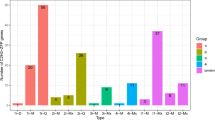
In the control group (H2O), the expression changes dynamics of nine genes in roots at different time points can be divided into three types. PvCML 6/38/107 displayed a consistently increasing trend. PvCML 8/41/53/54 decreased at first, then showed an increasing trend across the subsequent two stages. However, PvCML 44/82 decreased in the initial two stages, then increased obviously at 7 d, and decreased at 10 d again (Fig. 7). In the treatment group, PvCML 6/38 displayed a consistent increasing trend along the four stages in the root, and PvCML 41/107 displayed an increasing trend at most stages except for a decrease at 7 d and 5 d, respectively. The expressions of PvCML 44/53/54/82 were slightly upregulated in the initial three stages and then decreased at 10 d. PvCML 8 kept a stable expression trend along the four stages (Fig. 7). In addition, the expression of nine PvCML genes were detected in different tissues in the melatonin treatment and control group. The control group showed higher content in roots and hypocotyls, for example, six genes (PvCML 6/8/38/41/44/82) were dominantly expressed in hypocotyls, while two genes (PvCML 53/107) were dominantly expressed in roots. PvCML 54 was dominantly expressed in stems. Expression of these nine genes in leaves was at a low transcript level (Fig. 7).
The qRT-PCR expression patterns of CML genes in P. vulgaris. The reactions were normalized using the actin reference gene. The standard deviations were represented by the error bars from three independent technical replicates. 3 Day-Root (3D-R), 5 Day-Root (5D-R), 7 Day-Root (7D-R), 10 Day-Root (10D-R), 10 Day-Hypocotyl (1D-H), 10 Day-Stems (1D-S) and 10 Day-Leaf (1D-L). The expression levels of each gene were expressed as a ratio relative to that of 3D-R (H2O). t-test was used to analyze the mean expression levels of three replicates in melatonin treatment (MT) with control treatment (H2O). Star(s) above the bars indicated significant differences among the treatments. A star represented significant level (p < 0.05), two stars represented significant level (p < 0.01), and three stars represented significant level (p < 0.001).
After melatonin treatment, five genes (PvCML 6/8/38/44/54/82) of treatment group showed a downregulation trend compared with the control group on the third day of root development. Three genes (PvCML 41/53/107) in treatment group were up-regulated. On the fifth day of root development, melatonin treatment significantly upregulated eight genes except PvCML 107. The results were primarily consistent with the transcriptome data. On the seventh day of root development, seven genes (PvCML 6/38/44/53/54/82/107) were up-regulated compared with the control group. On the tenth day of root development, all genes except PvCML 53 were down-regulated by melatonin. On the tenth day of hypocotyl development, genes except PvCML 44/82 were down-regulated by melatonin treatment. On the tenth day of stem tissue, genes except PvCML 41/44 were down-regulated by melatonin treatment. On the tenth day of leaf tissue, genes except PvCML 8/38/41 were up-regulated by melatonin treatment (Fig. 7).
Interaction network of PvCML proteins
STRING database was used to construct an interaction network of nine melatonin responsive PvCML proteins. KEGG results indicated that PvCML 6/8/38/41/44/53/54/82/107 were mainly involved in plant–pathogen interaction pathways. Only three genes were found to interact with PvCML 6/8/38/41/44/53/54/82/107 genes (Fig. 8). Among them, XP _007153935.1 contains motif CH (actin binding domain), ARM, and IQ (calmodulin binding motif). XP_007156352.1 and XP_007158053.1 both contain motif MyTH4 (plant driver protein), B41 (plasma membrane binding domain), and KISc (driver protein). By GO analysis, it was found that XP_007153935.1, XP_007158053.1, and XP_007156352.1 genes are jointly involved in ATP binding, ATP-dependent microtubule motor activity, microtubule binding, oxidoreductase activity, and calmodulin binding pathway. XP_007137801 and XP_007163765 interacted with PvCML 6 and PvCML 44, respectively (Fig. 8). So far, XP_007137801 and XP_007163765 were uncharacterized proteins.
Discussion
Under biotic or abiotic stress, plant cells will produce a specific pattern of intracellular calcium flux, which will trigger the signal cascade and eventually affect the calcium concentration in cell39,40. As a kind of primary calcium sensors, plant CML proteins play key roles in cellular signaling networks by regulating various targets41,42. With the accumulation of various plant genome sequences, CML gene family has become a typical feature of plant genome43. However, the characteristics of CML gene in common bean have not been systematically studied. In this study, 111 PvCML genes were identified from the common bean genome by bioinformatic methods. The amount of PvCML in common bean was equivalent to that in other leguminous crops, such as soybeans12. Compared with A. thaliana44, O. sativa10 and Chinese cabbage15, common beans have more CML genes. The number of CML gene family members varies significantly among plant genomes, and it is independent of genome size. The variation of CML gene number among different species may be caused by the natural variation of different species and their adaptation to the complex growth environment.
Gene length, amino acid number, and pI of each PvCML were different. However, compared with CML gene families in other dicotyledons, the average number of amino acids of PvCML (282) was similar to that of MdCML (231). The mean pI of PvCML (5.17) and MdCML (5.22) were essentially the same (Supplementary Table S5). Some PvCML genes that shared close phylogenetic relationships displayed similar EF-hand distributions and intron–exon structures, which indicate that these genes may have similar biological functions and expression characteristics. The function of CML members depends on the number of EF-Hand motifs involved in Ca2+ binding properties45. The number of EF-Hand motifs varies among plant species. For example, AtCML usually contains 2–6 EF-Hand motifs5. Whereas MtCML contains 1–4 EF-Hand motifs13. In common beans, 2–4 conserved EF-Hand motifs were found in PvCML. Some motifs, such as motifs 2, 3, 5, and 9, calcium-binding EF-Hand domains, were conserved in all subfamilies of PvCML, and these motifs were related to the basic function of PvCML. Motifs 1, 4, 6, 7, 8, and 10, protein kinases, were unique in the subfamily I, which may affect the ATP binding and protein kinase activity of the subfamily I PvCML. And each PvCML contains a Dx3D motif. In addition, intron–exon structure analysis in M. truncatula and Brassica rapa showed that 78% of MtCML genes and 76.92% of BraCML genes did not contain introns15. Our Intron–exon structure analysis showed that 36.04% of PvCML genes did not contain introns. Compared with MtCML and BraCML, PvCML contains fewer genes without introns. It is thought that the fewer introns in a gene, the faster a plant responds to environmental changes46,47. In order to further understand PvCML, a phylogenetic tree was constructed to explore the evolutionary relationship of PvCML. In fact, 23 PvCML duplication pairs (20.72%) in 111 PvCMLs were identified in the common bean genome which suggested that some CML genes may be generated by segmental duplication events. Meanwhile, 25 PvCML were homologous with AtCMLs. Similarly, 20 pairs of homologous CML genes were found in M. truncatula and Arabidopsis13. At the same time, previous study suggested that whole-genome duplication (WGD) of legumes plays an important role in shaping the genome46. Therefore, WGD and segmental duplication may be involved in the evolution of PvCML gene.
In this study, we found that at least one stress response cis-element, growth and development related element, and hormone response element were detected in each PvCML gene. The results indicated that PvCML plays an important role in plant growth and development, stress response, and hormone response. For example, AtCML 44 is a homologous gene of PvCML 6. AtCML 44 is up-regulated under drought and salt stress. OsCML 31 is a homologous gene of PvCML 41. OsCML 31 is strongly induced under drought stress, and its overexpression can enhance plant drought tolerance48. AtCML 41 is a homologous gene of PvCML 44. AtCML 41 has been proved to be a stress response gene and plays an inhibitory role in the basic defense response of plants49,50. Cis-acting element analysis also showed that PvCML 6, PvCML 41, and PvCML 44 contain MBS (MYB binding site drought response element). AtCML 42 is a homologous of PvCML 84 and PvCML 98. AtCML 42 has been shown to affect plant response by changing plant hormone signals. Cis-acting element analysis showed that PvCML 84 and PvCML 98 contained TGACG cis-acting elements in response to hormones.
In addition, the tissue specific expression pattern of PvCML gene is thought to be related to its potential biological function. Transcriptome data analysis showed that the expression of the same subfamily members in the same tissue was not consistent. PvCML gene was highly expressed in different tissues, which was basically consistent with previous reports of M. sativa and Arabidopsis. For example, previous experiments showed that AtCML 3 was highly expressed in flower organs51. AtCML 3 homologous PvCML 107 also showed a higher expression in floral apparatus in transcriptome data.
Transcriptomic analysis of common bean treated with melatonin showed that nine PvCML genes were differentially expressed in the root of common bean. Subsequently, these nine genes were verified by qRT-PCR, and the results were consistent with the transcriptome data. In normal treatment or melatonin treatment, the expression of nine PvCMLs in roots was up-regulated or fluctuated with time, and reached the highest on the 7th or 10th day. In previous studies, CMLs in other plants were also significantly affected by hormone treatment, for example, MeJA induced high expression of AtCML 39 with more than 100 times52. CMLs expressions were different at different development stages in common bean. qRT-PCR results showed that the expression levels of nine PvCML genes in leaves were lower than that in other tissues. In addition, exogenous supplementation of low concentration melatonin can promote the growth of hypocotyls, stems, and roots of plants53,54,55. Our study proved that exogenous melatonin affected the expression of PvCML 6, PvCML 8, and PvCML 44 in hypocotyl, but increased the expression of PvCML 44/53/82 in leaves. The expression of PvCML 6/8/54 were downregulated in stems by melatonin. Taken together, differential expression of the PvCML gene at various time points and different tissues of the common bean suggested its response to melatonin treatment. More experiments need to be performed to demonstrate how melatonin regulates PvCML.
Interaction network analysis of genes contributes to the understanding of their functions. Hence, protein interaction net of nine melatonin responsive CML was analyzed. Three proteins were identified (XP_007153935.1, XP_007158053.1, and XP_007156352.1), which were involved in ATP binding, ATP-dependent microtubule movement, and microtubule binding and calmodulin binding pathways, respectively. The interaction between CMLs and these genes requires further study.
Data availability
All data generated or analysed during this study are included in this published article and its supplementary information files.
References
Berridge, M., Lipp, P. & Bootman, M. The versatility and universality of calcium signaling. Nat. Rev. Mol. Cell Biol. 1, 11–21. https://doi.org/10.1038/35036035 (2000).
Sanders, D., Pelloux, J., Brownlee, C. & Harper, J. F. Calcium at the crossroads of signaling. Plant Cell 14, S401–S417. https://doi.org/10.1105/tpc.002899 (2002).
Mohanta, T. K., Kumar, P. & Bae, H. Genomics and evolutionary aspect of calcium signaling event in calmodulin and calmodulin-like proteins in plants. BMC Plant Biol. 17(1), 38. https://doi.org/10.1186/s12870-017-0989-3 (2017).
Cheng, S. H., Willmann, M. R., Chen, H. C. & Sheen, J. Calcium signaling through protein kinases. The Arabidopsis calcium-dependent protein kinase gene family. Plant Physiol. 129, 469–485. https://doi.org/10.1104/pp.005645 (2002).
McCormack, E., Tsai, Y. C. & Braam, J. Handling calcium signaling: Arabidopsis CaMs and CMLs. Trends Plant Sci. 10, 383–389. https://doi.org/10.1016/j.tplants.2005.07.001 (2005).
Shi, S. et al. The Arabidopsis calcium-dependent protein kinases (CDPKs) and their roles in plant growth regulation and abiotic stress responses. Int. J. Mol. Sci. 19, 1900. https://doi.org/10.3390/ijms19071900 (2018).
Mohanta, T. K., Mohanta, N., Mohanta, Y. K., Parida, P. & Bae, H. Genome-wide identification of Calcineurin B-Like (CBL) gene family of plants reveals novel conserved motifs and evolutionary aspects in calcium signaling events. BMC Plant Biol. 15, 189. https://doi.org/10.1186/s12870-015-0543-0 (2015).
Gifford, J. L., Walsh, M. P. & Vogel, H. J. Structures and metal-ion-binding properties of the Ca2+-binding helix-loop-helix EF-hand motifs. Biochem. J. 405, 199–221. https://doi.org/10.1042/BJ20070255 (2007).
Mohanta, T. K. et al. Molecular players of EF-hand containing calcium signaling event in plants. Int. J. Mol. Sci. 20, 6. https://doi.org/10.3390/ijms20061476 (2019).
Boonburapong, B. & Buaboocha, T. Genome-wide identification and analyses of the rice calmodulin and related potential calcium sensor proteins. BMC Plant Biol. 7, 4. https://doi.org/10.1186/1471-2229-7-4 (2007).
Munir, S. et al. Genome-wide identification, characterization and expression analysis of calmodulin-like (CML) proteins in tomato (Solanum lycopersicum). Plant Physiol. Biochem. 102, 167–179. https://doi.org/10.1016/j.plaphy.2016.02.020 (2016).
Zeng, H., Zhang, Y., Zhang, X., Pi, E. & Zhu, Y. Analysis of EF-hand proteins in soybean genome suggests their potential roles in environmental and nutritional stress signaling. Front. Plant Sci. 8, 877. https://doi.org/10.3389/fpls.2017.00877 (2017).
Sun, Q., Yu, S. & Guo, Z. Calmodulin-like (CML) gene family in Medicago truncatula: Genome-wide identification, characterization and expression analysis. Int. J. Mol. Sci. 21, 7142. https://doi.org/10.3390/ijms21197142 (2020).
Dubrovina, A. S. et al. The effect of abiotic stress conditions on expression of Calmodulin (CaM) and Calmodulin-Like (CML) genes in wild-growing grapevine Vitis amurensis. Plants (Basel) 8, 602. https://doi.org/10.3390/plants8120602 (2019).
Nie, S., Zhang, M. & Zhang, L. Genome-wide identification and expression analysis of calmodulin-like (CML) genes in Chinese cabbage (Brassica rapa L. ssp. pekinensis). BMC Genom. 18, 842. https://doi.org/10.1186/s12864-017-4240-2 (2017).
Magnan, F. et al. Mutations in AtCML9, a calmodulin-like protein from Arabidopsis thaliana, alter plant responses to abiotic stress and abscisic acid. Plant J. 56, 575–589. https://doi.org/10.1111/j.1365-313X.2008.03622.x (2008).
Delk, N. A., Johnson, K. A., Chowdhury, N. I. & Braam, J. CML24, regulated in expression by diverse stimuli, encodes a potential Ca2+ sensor that functions in responses to abscisic acid, daylength, and ion stress. Plant Physiol. 139, 240–253. https://doi.org/10.1104/pp.105.062612 (2005).
Cheval, C., Aldon, D., Galaud, J. P. & Ranty, B. Calcium/calmodulin-mediated regulation of plant immunity. Biochim. Biophys. Acta 1833, 1766–1771. https://doi.org/10.1016/j.bbamcr.2013.01.031 (2013).
Chen, C. et al. GsCML27, a gene encoding a calcium-binding EF-hand protein from Glycine soja, plays differential roles in plant responses to bicarbonate, salt and osmotic stresses. PLoS One 10, e0141888. https://doi.org/10.1371/journal.pone.0141888 (2015).
Munir, S. et al. Overexpression of calmodulin-like (ShCML44) stress-responsive gene from Solanum habrochaites enhances tolerance to multiple abiotic stresses. Sci. Rep. 6, 31772. https://doi.org/10.1038/srep31772 (2016).
Park, H. C. et al. AtCML8, a calmodulin-like protein, differentially activating CaM-dependent enzymes in Arabidopsis thaliana. Plant Cell Rep. 29, 1297–1304. https://doi.org/10.1007/s00299-010-0916-7 (2010).
Wu, X. et al. CML20, an Arabidopsis calmodulin-like protein, negatively regulates guard cell ABA signaling and drought stress tolerance. Front. Plant Sci. 8, 824. https://doi.org/10.3389/fpls.2017.00824 (2017).
Yang, X. et al. Arabidopsis thaliana calmodulin-like protein CML24 regulates pollen tube growth by modulating the actin cytoskeleton and controlling the cytosolic Ca (2+) concentration. Plant Mol. Biol. 86, 225–236. https://doi.org/10.1007/s11103-014-0220-y (2014).
Lerner, A. B., Case, J. D., Takahashi, Y., Lee, T. H. & Mori, W. Isolation of melatonin, the pineal gland factor that lightens melanocytes. J. Am. Chem. Soc. 80(10), 2587–2587. https://doi.org/10.1021/ja01543a060 (1958).
Murch, S. J., Campbell, S. S. B. & Saxena, P. K. The role of serotonin and melatonin in plant morphogenesis: Regulation of auxin-induced root organogenesis in in vitro-cultured explants of St. John’s Wort (Hypericum perforatum L.). In Vitro Cell. Dev. Biol. Plant 37, 786–793. https://doi.org/10.1007/s11627-001-0130-y (2001).
Hernández-Ruiz, J., Cano, A. & Arnao, M. B. Melatonin: A growth-stimulating compound present in lupin tissues. Planta 220(1), 140–144. https://doi.org/10.1007/s00425-004-1317-3 (2004).
Padmavathi, B., Babu, A. N., Naveen, R., Kiranmai, K. & Prameel, V. A phytopharmacological review on Phaseolus vulgaris. Int. J. Res. Ayurveda Pharm. 12, 118–123. https://doi.org/10.7897/2277-4343.120386 (2021).
Broughton, W. J. et al. Beans (Phaseolus spp.)—Model food legumes. Plant Soil 252, 55–128. https://doi.org/10.1023/A:1024146710611 (2003).
Han, Y. et al. The molecular events underpinning cultivar differences in melatonin counteracting salt damage in Phaseolus vulgaris. Funct. Plant Biol. 49, 201–217. https://doi.org/10.1071/FP21126 (2022).
Wei, J. et al. Phytomelatonin receptor PMTR1-mediated signaling regulates stomatal closure in Arabidopsis thaliana. J. Pineal Res. 65, e12500. https://doi.org/10.1111/jpi.12500 (2018).
Goodstein, D. M. et al. Phytozome: A comparative platform for green plant genomics. Nucleic Acids Res. 40, D1178–D1186. https://doi.org/10.1093/nar/gkr944 (2012).
Rhee, S. Y. et al. The Arabidopsis Information Resource (TAIR): A model organism database providing a centralized, curated gateway to Arabidopsis biology, research materials and community. Nucleic Acids Res. 31, 224–228. https://doi.org/10.1093/nar/gkg076 (2003).
Kawahara, Y. et al. Improvement of the Oryza sativa Nipponbare reference genome using next generation sequence and optical map data. Rice (N Y) 6, 4. https://doi.org/10.1186/1939-8433-6-4 (2013).
Wheeler, T. J. & Eddy, S. R. Nhmmer: DNA homology search with profile HMMs. Bioinformatics 29, 2487–2489 (2013).
Zhang, B. et al. Genome-wide analysis of GRAS transcription factor gene family in Gossypium hirsutum L. BMC Genom. 19, 348. https://doi.org/10.1186/s12864-018-4722-x (2018).
Borges, A., Tsai, S. M. & Caldas, D. G. Validation of reference genes for RT-qPCR normalization in common bean during biotic and abiotic stresses. Plant Cell Rep. 31, 827–838. https://doi.org/10.1007/s00299-011-1204-x (2012).
Ragueneau, E. et al. IntAct App: A Cytoscape application for molecular interaction network visualisation and analysis. Bioinformatics 37, 3684–3685. https://doi.org/10.1093/bioinformatics/btab319 (2021).
Kanehisa, M. & Goto, S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 28, 27–30. https://doi.org/10.1093/nar/28.1.27 (2000).
Zebelo, S. A. & Maffei, M. E. Role of early signaling events in plant–insect interactions. J. Exp. Bot. 66, 435–448. https://doi.org/10.1093/jxb/eru480 (2015).
Ranty, B., Aldon, D. & Galaud, J. P. Plant calmodulins and calmodulin-related proteins: Multifaceted relays to decode calcium signals. Plant Signal. Behav. 1, 96–104. https://doi.org/10.4161/psb.1.3.2998 (2006).
Zhang, X., Wang, T., Liu, M., Sun, W. & Zhang, W. H. Calmodulin-like gene MtCML40 is involved in salt tolerance by regulating MtHKTs transporters in Medicago truncatula. Environ. Exp. Bot. 157, 79–90. https://doi.org/10.1016/j.envexpbot.2018.09.022 (2018).
Perochon, A., Aldon, D., Galaud, J. P. & Ranty, B. Calmodulin and calmodulin-like proteins in plant calcium signaling. Biochimie 93, 2048–2053. https://doi.org/10.1016/j.biochi.2011.07.012 (2011).
Shi, J. & Du, X. Identification, characterization and expression analysis of calmodulin and calmodulin-like proteins in Solanum pennellii. Sci. Rep. 10, 7474. https://doi.org/10.1038/s41598-020-64178-y (2020).
McCormack, E. & Braam, J. Calmodulins and related potential calcium sensors of Arabidopsis. New Phytol. 159, 585–598. https://doi.org/10.1046/j.1469-8137.2003.00845.x (2003).
Scholz, S. S., Reichelt, M., Vadassery, J. & Mithöfer, A. Calmodulin-like protein CML37 is a positive regulator of ABA during drought stress in Arabidopsis. Plant Signal. Behav. 10, e1011951. https://doi.org/10.1080/15592324.2015.1011951 (2015).
Pfeil, B. E., Schlueter, J. A., Shoemaker, R. C. & Doyle, J. J. Placing paleopolyploidy in relation to taxon divergence: A phylogenetic analysis in legumes using 39 gene families. Syst. Biol. 54, 441–454. https://doi.org/10.1080/10635150590945359 (2005).
Jeffares, D. C., Penkett, C. J. & Bähler, J. Rapidly regulated genes are intron poor. Trends Genet. 24(8), 375–378. https://doi.org/10.1016/j.tig.2008.05.006 (2008).
Xu, G. Y. et al. A novel rice calmodulin-like gene, OsMSR2, enhances drought and salt tolerance and increases ABA sensitivity in Arabidopsis. Planta 234, 47–59. https://doi.org/10.1007/s00425-011-1386-z (2011).
Denoux, C. et al. Activation of defense response pathways by OGs and Flg22 elicitors in Arabidopsis seedlings. Mol. Plant 1, 423–445. https://doi.org/10.1093/mp/ssn019 (2008).
Baev, V. et al. Identification of RNA-dependent DNA-methylation regulated promoters in Arabidopsis. Plant Physiol. Biochem. 48, 393–400. https://doi.org/10.1016/j.plaphy.2010.03.013 (2010).
Chigri, F. et al. The Arabidopsis calmodulin-like proteins AtCML30 and AtCML3 are targeted to mitochondria and peroxisomes, respectively. Plant Mol. Biol. 78, 211–222. https://doi.org/10.1007/s11103-011-9856-z (2012).
Vanderbeld, B. & Snedden, W. A. Developmental and stimulus-induced expression patterns of Arabidopsis calmodulin-like genes CML37, CML38 and CML39. Plant Mol. Biol. 64, 683–697. https://doi.org/10.1007/s11103-007-9189-0 (2007).
Xiong, F. et al. Hypocotyl elongation inhibition of melatonin is involved in repressing brassinosteroid biosynthesis in Arabidopsis. Front. Plant Sci. 10, 1082. https://doi.org/10.3389/fpls.2019.01082 (2019).
Chen, L. et al. Melatonin promotes seed germination under salt stress by regulating ABA and GA3 in cotton (Gossypium hirsutum L.). Plant Physiol. Biochem. 162, 506–516. https://doi.org/10.1016/j.plaphy.2021.03.029 (2021).
Chen, Q. et al. Exogenously applied melatonin stimulates root growth and raises endogenous indoleacetic acid in roots of etiolated seedlings of Brassica juncea. J. Plant Physiol. 166, 324–328. https://doi.org/10.1016/j.jplph.2008.06.002 (2008).
Acknowledgements
This research was funded by the National Natural Science Foundation of China: 3187102245; Heilongjiang Bayi Agricultural University "Three Longitudinal" Foundation Cultivation Project: ZRCPY202111; Natural Science Foundation of Heilongjiang Province (LH2021C065).
Author information
Authors and Affiliations
Contributions
H.Z. performed the experiments and wrote the manuscript; Y.G. prepared the materials and edit the manuscript; J.D. and Y.D. provided technical assistance; Y.H. designed the experiments and analyzed data. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhao, H., Gao, Y., Du, Y. et al. Genome-wide analysis of the CML gene family and its response to melatonin in common bean (Phaseolus vulgaris L.). Sci Rep 13, 1196 (2023). https://doi.org/10.1038/s41598-023-28445-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-28445-y
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.