Abstract
Amyloid precursor protein (APP) is a transmembrane protein that plays critical role in the pathogenesis of Alzheimer's disease (AD). It is also involved in many types of cancers. Increasing evidence has shown that the tyrosine phosphorylation site Y682 in the intracellular tail of APP is crucial for APP function. Here, we report that Vav2, a guanine nucleotide exchange factor (GEF) for Rho family GTPase, is a novel interaction partner of APP. We found that Vav2-SH2 domain was able to bind directly to the Y682-phosphorylated intracellular tail of APP through isothermal titration calorimetry and NMR titrating experiments. The crystal structure of Vav2-SH2 in complex with an APP-derived phosphopeptide was determined to understand the structural basis of this recognition specificity. The interaction of APP and Vav2 in a full-length manner was further confirmed in cells by GST pull-down, co-immunoprecipitation and immunofluorescence staining experiments. In addition, we found overexpression of Vav2 could inhibit APP degradation and markedly increase the protein levels of APP and its cleavage productions in 20E2 cells, and this function of Vav2 required a functional SH2 domain.
Similar content being viewed by others
Introduction
Amyloid precursor protein (APP) is a type-I transmembrane protein that is crucial for neuronal development and homeostasis1,2. It is best known for its role in the pathogenesis of Alzheimer’s disease (AD)3. AD is a neurodegenerative disease and is the most often found form of dementia among elderly people. Currently, there is no effective treatment for this disease4. The pathological hallmark of AD is the presence of amyloid plaques in the brain of AD patients5,6. APP is proteolytically cleaved by β- and γ-secretase to generate the amyloid β peptide (Aβ), which is the major component of amyloid plaques7,8,9. In addition, APP has been found to be overexpressed in various cancers, including breast cancer and can promote cancer cell migration and invasion1,10,11.
In human, there are three main isoforms of APP derived by alternative splicing: a 695-residue form expressed primarily in the central nervous system and 751- and 770-residue forms that are ubiquitously expressed12. All three forms share a similar architecture with a large ectodomain, a single transmembrane domain and a short intracellular tail13,14. The short intracellular tail of APP is essential for APP function15,16. Numerous cytosolic proteins are found to bind directly to the intracellular domain of APP17. It is 47 amino acids in length and contains several functional motifs and phosphorylation sites, including Y682, whose phosphorylation level has been shown to be significantly increased in AD brains18,19. Most interactions of APP involve the Y682 phosphorylation site. Y682 modulates the interactions of APP with different proteins through its phosphorylation and dephosphorylation20. Some proteins, such as X1121,22,23, Fe6524,25,26, and JIP-127 interact with APP when Y682 is not phosphorylated, whereas others, such as Grb228,29, Shc30, only when Y682 is phosphorylated. Increasing evidences show that these interactions play key roles in the regulation of APP processing and function, and in AD15,31,32,33,34,35. For example, Mint and Fe65 family modulate cellular trafficking and the processing of APP and affect Aβ production36,37. JIP-1 facilitates APP axonal trafficking and regulates APP-dependent axonal transport of synaptic vesicles38. Notably, a recent study showed that targeting the APP-Mint2 protein–protein interaction with a peptide-based inhibitor reduced amyloid β formation, which may present an alternative strategy in the pursuit of new therapeutic approaches in AD treatment39.
Herein, we report the identification of Vav2 as a novel interaction partner for APP. It is a guanine nucleotide exchange factor (GEF) for Rho GTPases belonging to the Vav family (Vav1, Vav2 and Vav3)40,41. Vav2 is broadly expressed in human tissues and is involved in regulating various biological processes, including cell spreading and migration, neuronal development, angiogenesis, and cancer cell motility42,43,44,45,46,47. Vav2 consists of multiple domains, including a calponin homology (CH) domain, an Acidic (Ac) region, a catalytic Dbl homology (DH), a pleckstrin homology (PH) domain, a zinc finger (ZF) domain, a Src homology 2 (SH2) domain and two Src homology 3 (SH3) domains48. Among all Ras superfamily GEFs, only Vav family proteins possess an SH2 domain, a common protein interaction module that specifically recognizes phosphotyrosine motif49,50. Through its SH2 domain, Vav2 can bind to the tyrosine-phosphorylated cytoplasmic domains of several membrane receptors and then mediate different extracellular signals to intracellular responses51,52,53.
Our research demonstrated that Vav2 can interact with APP through its SH2 domain which binds directly to the Y682-phosphorylated APP intracellular tail. A crystal structure of Vav2-SH2 domain in complex with the APP-derived phosphopeptide APP-pY682 (QNG-pY-ENPT, residues 679–686 of APP695) was determined at 2.45 Å resolution, which revealed a conserved recognition mechanism. The interaction of APP and Vav2 in a full-length manner was further confirmed by GST pull-down experiments, co-immunoprecipitation and immunofluorescence staining. Moreover, we found that overexpression of Vav2 significantly increased APP protein level and promoted Aβ40 generation in 20E2 cells, an AD cell model. We further show that Vav2 overexpression inhibited APP protein degradation. This function of Vav2 requires its SH2 domain. Together, these findings uncover a novel interaction between Vav2 and APP and a regulatory role of Vav2 in APP turnover.
Results
Identification and characterization of a direct interaction between Vav2-SH2 domain and Y682-phosphorylated APP peptide
Our previous work showed that Vav2-SH2 domain prefers mostly to recognize a consensus motif of pY-E-X-P, where X denotes any amino acid54,55. We found that the tyrosine phosphorylation site Y682 in the intracellular domain of APP, within the sequence of YENP, exactly matched this consensus motif. Therefore, we speculated that Y682-phosphorylated APP might be able to bind the SH2 domain of Vav2. To explore the possibility, we performed NMR titration and isothermal titration calorimetry (ITC) experiments using the recombinantly expressed Vav2-SH2 protein and a synthesized phosphotyrosine peptide derived from residue Y682 in APP (termed APP-pY682: QNG-pY-ENPT, residues 679–686 of APP695) (Fig. 1A and Table S1). As a control, the non-phosphorylated form of this peptide (referred to as APP-Y682) was also included. The purified Vav2-SH2 protein was determined by SDS-PAGE with high purity (> 98%) (Fig. S1), and the final protein yields were about 15 mg and 10 mg per L culture for the unlabeled and 15N-labeled protein, respectively.
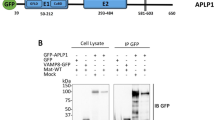
Binding of Vav2-SH2 to phosphotyrosine peptide APP-pY682 derived from APP. (A) Schematic representation of human Vav2 and APP. Sequence of APP intracellular domain and phosphotyrosine peptide APP-pY682 are labeled. (B) Overlay of 1H–15N HSQC spectra of Vav2-SH2 in the absence (black) and in the presence of increasing amounts of the peptide APP-pY682 (QNG-pY-ENPT). The molar ratios of the protein to the peptide are shown in the inset: 1:0 (black), 1:0.25 (blue), 1:0.5 (cyan), 1:1.0 (green), 1:2.0 (yellow) and 1:3.0 (red). Residues R698 and R700 are zoomed in. (C) ITC measurements of the binding affinity of APP-pY682 and APP-Y682 to Vav2-SH2.
For NMR titrating experiments, 2D 1H–15N HSQC spectra56 of 15N-labeled Vav2-SH2 domain at a series of protein to peptide ratios were collected, respectively. As shown in Fig. S2, titration with the non-phosphorylated APP peptide did not induce any chemical shift perturbations in the 1H–15N HSQC spectra of Vav2-SH2 domain. In contrast, titration with the phosphopeptide APP-pY682 caused substantial chemical shift perturbations (CSPs) in the protein, indicating direct binding (Fig. 1B). During the titration, many resonances corresponding to the free state of Vav2-SH2 disappeared, while another set of crosspeaks, corresponding to the bound state appeared. All these residues were mapped on the structure of Vav2-SH2 (Fig. S3). This pattern of CSPs demonstrated the formation of a complex that is in slow to intermediate exchange on the chemical shift NMR time scale. ITC reported a binding Kd of 0.86 μM for Vav2-SH2 in complex with APP-pY682 peptide and no binding of Vav2-SH2 and APP-Y682 (Fig. 1C).
Crystal structure of Vav2-SH2 domain in complex with APP-pY682 peptide
To further understand the structural basis for the specific recognition of Y682-phosphorylated APP by Vav2-SH2 domain, we solved the crystal structure of Vav2-SH2 in complex with the APP-pY682 peptide. The structure was refined to 2.45 Å resolution (Fig. 2 and Table 1) (PDB entry: 7WFY).
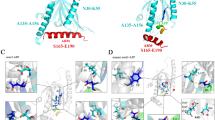
Specific interaction of the Vav2-SH2 domain and the peptide APP-pY682. (A) Overall stereoview of Vav2-SH2 domain in complex with peptide APP-pY682. Vav2-SH2 is colored purple, with the secondary structures are labeled. The APP-pY682 peptide (green) is shown in stick mode, with the residues N680′, G681′, pY682′, E683′, N684′, P685′ and T686′ are labeled. (B) Detailed interactions of the peptide APP-pY682 with SH2 domain. The peptide APP-pY682 and the side chains of crucial residues are shown in stick mode and labeled respectively. Selected hydrogen bonds are colored in black. (C) Electrostatic potential surface representation of Vav2-SH2 domain bound to APP-pY682 peptide (green). The pY and pY + 3 pockets are indicated by magenta and blue circles separately.
In the complex, the general fold of Vav2-SH2 domain is almost the same with that observed in its free state55, which consists of a short N-terminal α-helix (αN), a central β-sheet (βB–βD), two α-helices (αA and αB) and a small β-sheet (βD’ and βE) (Fig. 2A). The peptide lies in an extended backbone conformation roughly perpendicular to the central β-strands of the SH2 domain (Fig. 2A,B). Phosphotyrosine pY682′ inserts into the canonical pY-binding pocket formed by R680, R698, R700, H719 and K721 (Fig. 2B,C). The phosphate moiety of pY682′ forms a hydrogen bond network with the side chains of R680, R698 and R700. The aromatic moiety of this residue is packed against residues H719 and K721. Residue E683′ (pY + 1) makes hydrophobic interactions with the side chains of residues K718, I720, F756 and L759. This residue is further stabilized by forming a hydrogen bond between its backbone amide nitrogen and carbonyl oxygen atom of residue H719. Residue N684′ (pY + 2) does not interact with SH2 domain. Residue P685′ (pY + 3) patches on the surface of the pY + 3 pocket formed by EF and BG loops and makes contacts with T732, S755 and F756 (Fig. 2B,C). Other residues of the APP-pY682 peptide show little or no interaction with the SH2 protein.
Comparison of the recognitions of Y682-phosphorylated APP by Vav2-SH2 and Grb2-SH2
The SH2 domain of Grb2 can also bind to Y682-phosphorylated APP. The crystal structure of Grb2-SH2 in complex with a Y682-phosphorylated APP peptide was reported previously29. Structure comparison reveals a significant difference in the mechanisms of the peptide binding to the SH2 domains of Vav2 and Grb2. As shown above and in Fig. 3A, the APP-pY682 peptide binds to Vav2-SH2 in an extended conformation. However, when bound to Grb2-SH2 domain the peptide adopts a U-shape conformation (Fig. 3B). This is due to the presence of a bulky Tryptophan residue (W121) in the EF loop of Grb2-SH2, which is a Threonine residue (T732) at the corresponding position of Vav2-SH2. The large sidechain of W121 occupies the pY + 3 pocket and sterically hinders the phosphopeptide from assuming an extended conformation and forces it into the U-shape conformation. In the complex of Grb2-SH2 with pY682-phosphorylated APP peptide, both residues N684′ (pY + 2) and T686′ (pY + 4) of APP interact strongly with the protein. N684′ forms a network of hydrogen bonds with K109 and L120, while T686′ interacts with L111 and K109 in Grb2-SH2. In contrast, in the complex of Vav2-SH2 with APP-pY682, neither N684′ nor T686′ contacts with the protein.
Comparison of the phosphopeptide-binding sites in Vav2-SH2 and Grb2-SH2. (A,B) The presentation of the structures of Vav2-SH2 complexed with APP-derived phosphopeptide (682pY-ENPT686) and Grb2-SH2 (PDB code: 3MXC). Vav2-SH2 domain (in purple) and Grb2-SH2 domain (in grey) are shown in ribbon and surface representation, APP-derived phosphopeptide (in green) and the side chains of crucial residues is shown in stick representation. Conformation of APP-pY682 as seen in complex with Vav2-SH2 domain is hindered by W121 (in red) of EF loop and BG loop when bound to the Grb2-SH2 domain. Selected hydrogen bonds are colored in black.
Full-length APP and Vav2 interact in mammalian cells
To determine whether the SH2 domain of Vav2 can interact with full-length APP in a cellular environment, we purified wild-type GST-Vav2-SH2 protein and its R680A mutant (as a negative control) and then performed GST pull-down assays55. ITC experiments showed that mutation of R680 to Ala in Vav2-SH2 domain nearly abolished its binding with APP-pY682 (Fig. S4). In the GST pull-down assays, we used lysates from 20E2 cells. Immunoprecipitation of APPsw and immunoblotting with an anti-phosphotyrosine antibody revealed that APPsw is tyrosine phosphorylated in 20E2 cells (Fig. S5). The GST pull-down assays showed that wild-type GST-Vav2-SH2, but not GST alone or the R680A mutant bound selectively to full-length APPsw (Fig. 4A). To investigate whether the tyrosine phosphorylation site Y682 of APP is involved in binding with Vav2-SH2 domain, we generated an Y682A mutant of APPsw (APPswY682A). GST pull-down assays were performed to test the binding of APPswY682A to Vav2-SH2 domain using HEK293 cells lysate expressing APPswY682A. HEK293 cells lysate expressing wild type APPsw was used as a positive control. As expected, the results showed that APPswY682A abolish its binding to Vav2-SH2 (Fig. 4B). To test whether the interaction between full-length APPsw and Vav2 occurs in cells, a co-immunoprecipitation experiment was performed. HEK293 cells were transiently co-transfected with pAPPsw and myc-tagged Vav2 plasmids. As shown in Fig. 4C, Vav2 was efficiently precipitated by an antibody against APPsw (C20), but not by the control IgG. Moreover, immunofluorescence staining experiments show that APPsw were co-localized with Vav2 (Fig. 4D). Taken together, these results indicated that Vav2 can interact with APP through its SH2 domain in mammalian cells.
Interaction of Vav2 with full-length APP in cells. (A) GST pull-down assay of APP from lysate of cells stable expressing APP (20E2 cells) with the purified GST fusion proteins. GST fusion proteins were detected by Coomassie blue staining. FL-APPsw were detected by western blot using C20 antibody and the Tyr-phosphorylated APP were detected by western blot using anti-Phospho-Tyrosine antibody. Original blots and gels are presented in Supplementary Figures. (B) GST pull-down assay of the wild type and mutant of APP with purified GST-SH2 fusion proteins. Original blots and gels are presented in Supplementary Figures. (C) Co-IP was performed using HEK293 cells transfected with Vav2-Myc and pAPPsw. C20 was used as the pull-down antibody and anti-myc antibody was used as detection antibody. Original blots are presented in Supplementary Figures. (D) Vav2 co-localizes with APP in HEK293 cells. HEK293 cells transfected with Vav2-Myc and pAPPsw were stained with anti-myc (red) and C20 (green) antibodies, DAPI (blue) was used to indicate the nucleus. Images were captured by LSM 880 fluorescent confocal microscope. The line with an arrow in the third picture indicates the selected line used in co-localization analysis (right).
Overexpression of Vav2 markedly increases APP protein level and Aβ40 generation
To investigate whether Vav2 affects APP metabolism, 20E2 cells were transfected with Vav2 plasmid or empty vector (as control). 48 h after transfection, we measured the levels of FL-APPsw as well as its cleaved products C99 and C83 by western blot. As shown in Fig. 5, overexpression of Vav2 significantly increased the levels of FL-APPsw, C99 and C83 compared with control. In addition, we measured the levels of Aβ40 in the conditioned media and also observed a significant increase in the level of Aβ40 in Vav2 overexpressing cells (Fig. 5E).
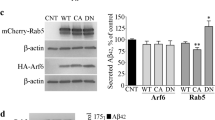
Vav2 upregulates the APP protein level and Aβ40 production. (A) Vav2 regulates the protein level of APP, C99 and C83 in 20E2 cells. 20E2 cells were transfected with Vav2WT, Vav2R680A, Vav2E205A and empty control, respectively. Vav2, FL-APPsw, C99 and C83 protein levels were examined by Western blot, β-actin was used as loading control. Original blots are presented in Supplementary Figures. (B–D) Quantification of (A) using ImageJ software. All quantified results were obtained from three independent experiments. Data were presented as means ± SEM (n = 3); *p < 0.05, p values were calculated by Student’s t test. (E) Elisa was performed to measure Aβ40 in culture media from 20E2 cells transfected with Vav2WT and its mutations, respectively. All quantified results were obtained from three independent experiments. Data were presented as means ± SEM.(n = 3); *p < 0.05, p values were calculated by Student’s t test. (F) HEK293 cells were co-transfected with APPsw or APPswY682A in the presence or absence of Vav2. Western blot was performed to detect the level of FL-APPsw protein level. β-Actin was used as loading control. ImageJ software was used for quantification. Original blots are presented in Supplementary Figures. All quantified results were obtained from three independent experiments. Data were presented as means ± SEM (n = 3); *p < 0.05, p values were calculated by one-way ANOVA with Bonferroni’s multiple comparisons post hoc test.
Given that Vav2 binds to Y682-phosphorylated APP via its SH2 domain, we then analyzed whether a functional SH2 domain is required for Vav2 to upregulate the levels of APP and its productions. We designed a R680A mutant of Vav2 (Vav2R680A). As shown in Fig. 5, when cells were transfected with the mutant Vav2R680A, the increase of FL-APPsw and its productions was significantly diminished, compared with Vav2WT. This observation suggests that the SH2 domain is important for Vav2 to increase the levels of APPsw and its productions. In addition, to test whether the GEF activity of Vav2 is involved in the APPsw levels increased by Vav2 overexpression, we constructed a GEF activity-dead mutant of Vav2 (Vav2E205A)57. Similar to the overexpression of wild type Vav2, overexpression of Vav2E205A mutant in 20E2 cells also led to a significant increase in the levels of APPsw and its productions, suggesting that the GEF activity is not required in this process.
To further explore whether tyrosine phosphorylation site Y682 in APP is required for its protein level elevation caused by Vav2 overexpression, we co-transfected wild type APPsw or APPswY682A mutant with Vav2 into HEK293 cells and the levels of APPsw and APPswY682A were measured, respectively. As shown in Fig. 5F,G, Vav2 sharply increased the protein level of APPsw, but not APPswY682A.
Vav2 overexpression inhibits APP degradation
Next, we investigated the effects of Vav2 on APP protein degradation by cycloheximide (CHX) chase assay. 20E2 cells were transfected with Vav2 and empty control respectively and then chased with cycloheximide. In CHX chase assay, APPsw degradation rate in Vav2 transfected 20E2 cells was markedly disrupted comparing to control cells (Fig. 6A,B). These results suggest that Vav2 overexpression inhibited APP degradation.
Vav2 stabilizes APP protein turnover. (A) Vav2 influences the degradation rate of APP protein. 20E2 cells were transfected with Vav2WT or empty control. Forty-eight hours after transfection, cells were treated with 150 μg/mL of CHX for indicated times. Cell lysates were detected for FL-APPsw and Vav2 by Western blot, β-actin was used as loading control. Original blots are presented in Supplementary Figures. (B) Quantification of (A). All quantified results were obtained from three independent experiments. Data were presented as means ± SEM (n = 3); *p < 0.05, p values were calculated by Student’s t test at the same time point.
Discussion
In the present work, we have identified that Vav2 is a novel APP-interacting protein. It is a ubiquitous guanine nucleotide exchange factor (GEF) for Rho family GTPases. Vav2 is reported to interact with several tyrosine-phosphorylated cell surface receptors through its SH2 domain and is involved in regulating a wide range of biological processes58,59. The interaction between Vav2 and APP is mediated by the SH2 domain of Vav2 and the Y682 phosphorylation site in the intracellular domain of APP. Y682 has been shown to play a crucial role in modulating the binding and unbinding of APP to specific cytosolic proteins through its phosphorylation state.
Our ITC and NMR data showed that Vav2-SH2 domain can interact directly with the Y682-phosphorylated APP peptide APP-pY682. The complex structure revealed that this phosphopeptide bound to the Vav2-SH2 domain adopted a typical extended conformation. Previous study has shown that Grb2 can also bind to Y682-phosphoryled APP through its SH2 domain and the crystal structure of Grb2-SH2 in complex with Y682-phosphorylted peptide is available29. The reported binding Kd of Grb2-SH2 to Y682-phosphorylated peptide is 0.29 μM which is comparable to the Kd of Vav2-SH2 binding to APP-pY682, suggesting that Vav2-SH2 and Grb2-SH2 bind to Y682-phosphorylated APP with similar affinities. However, there are obvious differences in the recognition mechanism of the phosphopeptide by Vav2-SH2 and Grb2-SH2. In Grb2-SH2 complex, the phosphopeptide does not adopt an extended conformation as that seen in the Vav2-SH2 complex, but presents a folded “U” shaped structure. Both the residues Y + 2 and Y + 4 are found to contact directly with Grb-SH2. However, in the Vav2-SH2 complex, no interaction of these two residues to Vav2-SH2 domain was observed. It should be noted that although our biochemical and structural data using APP phosphopeptide has established that the Y682 phosphorylation site of APP can act as a docking site for Vav2-SH2 domain, it is unclear whether other regions of APP as well as the membrane environment would affect this interaction. Further study should be carried out using full-length APP protein under a proper model membrane system.
In this study, we have also showed that Vav2 overexpression can inhibit APP degradation and thus lead to a significantly enhancement of the levels of APP and its productions in both 20E2 cells and HEK293 cells. Using the SH2 domain mutant (R680A) and GEF-dead mutant (E205A), we found that the SH2 domain but not the GEF activity is required for Vav2 to elevate APP level. Moreover, Vav2 overexpression has no effect on the level of APPswY682A mutant. These results may suggest a potential role of Vav2-APP interaction in the regulation of APP level.
Notably, the phosphorylation level of Y682 is significantly elevated in AD patient18,19. The high levels of APP Y682 phosphorylation may enhance the interaction between Vav2 and APP in AD patient. The potential role of APP-Vav2 interaction in AD need to be further studied. In addition, APP is found to be overexpressed in multiple cancers, such as breast cancer60,61,62. It has been shown recently to promote cancer cell migration and invasion11. However, the underline mechanism is not clear. It is well known that Vav2 is also overexpressed in most human cancers and promotes cancer cell migration and invasion in several types of human cancer43,63,64,65. Therefore, the identification of the interaction between Vav2 and APP may open up a novel avenue for further research on the role of APP in cancer.
Methods
Plasmid construction
For NMR and ITC experiments, the DNA fragment encoding the SH2 domain of Vav2 (residues 659–771) was cloned into a pET28a (+) (Novagen) plasmid as described previously54, generating a fusion protein with an N-terminus 6 × His tag. For GST pull-down experiments, the Vav2-SH2 was subcloned into pGEX4T-1 vector. The APP expression plasmid pAPPsw and the Vav2 expression plasmid pCMV5-myc-Vav2 were constructed as previously described55,66. Mutants were generated by PCR mediated site-directed mutagenesis. All the constructs were verified by DNA sequencing.
Recombinant protein expression, purification and peptide synthesis
The recombinant plasmids harboring His-tagged or GST-tagged Vav2-SH2 were transformed into Escherichia coli BL21 (DE3) (Novagen) strain. Cells were grown at 37 °C up to an A600 nm of 0.8, and then were induced with 0.5 mM IPTG at 25 °C for 8 h. For the production of uniformly 15N-labeled samples, cells were grown in minimal medium using 15NH4CL (0.5 g/L) as the sole nitrogen source. 15N-NH4Cl was purchased from Cambridge Isotope Laboratories, Inc. The His-tagged proteins were purified by a chelated-nickel column followed by Thrombin protease treatment to remove the tag as previous described54. The GST and GST fusion proteins were purified using immobilized glutathione. All proteins were further purified on a Superdex75 gel-filtration column (GE Healthcare, Piscataway, NJ, USA). The purity of proteins was confirmed by SDS–PAGE. Protein concentrations were estimated with absorbance spectroscopy using the molar absorption coefficient.
The phosphotyrosine peptide APP-pY682 (QNG-pY-ENPT) corresponding to residues 679–686 of APP695 with Y682 phosphorylated and unphosphorylated peptide APP-Y682 (QNGYENPT) were synthesized by GL Biochem Ltd. (Shanghai).
NMR sample preparation and NMR experiments
The NMR sample of the apo Vav2-SH2 contained 0.1 mM of protein was dissolved in NMR buffer: 20 mM Tris/HCl with 100 mM NaCl, 3 mM DTT in 90% H2O, 10% D2O (pH 7.0). D2O was purchased from Cambridge Isotope Laboratories, Inc. For the SH2/peptide complex samples, the 15N-labeled Vav2-SH2 (0.1 mM) mixed with phosphotyrosine peptide APP-pY682 (8 mM) or APP-Y682 (8 mM) which were dissolved in the same NMR buffer and the solutions were adjusted to pH 7.0 using NaOH. A series of 1H–15N HSQC spectra were recorded as the titrant gradually titrated into the protein solutions using the Bruker hsqcfpf3gpphwg pulse program. NMR experiments were carried out at 293 K on a Bruker Avance 600 MHz NMR spectrometer equipped with cryoprobes. NMR data were processed with NMRPipe67 and analyzed using Sparky3 (Goddard and Kneller, University of California, San Francisco). The assignment of Vav2-SH2 was extracted from a previous study55.
Isothermal titration calorimetry (ITC)
ITC experiments were performed at 298 K on a PEAQ-ITC titration calorimeter (MicroCal, Northampton, MA). Vav2-SH2 domain wild type and mutants were dialyzed extensively against the titration buffer containing 10 mM Tris/HCl, 100 mM NaCl (pH 7.4). Peptides were dissolved in the same buffer and the solutions were adjusted to pH 7.4 using NaOH. 0.75 mM peptides were titrated into Vav2-SH2 domain wild type or its mutant (30 μM). Control experiments were performed under the same condition by titrating same peptide into buffer alone. ITC data were analyzed with a single-site binding model using Microcal PEAQ-ITC software.
Protein crystallization
For crystallization experiments, purified Vav2-SH2 protein was concentrated to 15 mg/mL in a buffer containing 10 mM Tris/HCl, pH 7.4, 100 mM NaCl and 5 mM β-mercaptoethanol. Purified protein was mixed with pY682 peptide at a 1:1.2 molar ratio and crystallized using the sitting-drop vapour-diffusion method at 16 °C after mixing 0.5 μL of the protein solution with 0.5 μL of the reservoir solution. Crystals were obtained with a reservoir solution containing 25% PEG 3350, 0.2 M MgCl2·6H2O and 0.1 M Tris/HCl, pH 8.4. Before flash-freezing crystals in liquid nitrogen, crystals were soaked in a cryoprotectant consisting of 85% reservoir solution and 15% glycerol.
Data collection and structure determination
X-ray diffraction data were collected at the Shanghai Synchrotron Radiation Facility (SSRF) beam line BL18U. The data were processed with HKL200068 and programs in the CCP4 suite69. The structure of the Vav2-SH2 domain complexed with pY682 was solved by molecular replacement with co-ordinates from PDB entry 3MXC29 and the program PHASER70. All the structural models were subsequently refined by programs REFMAC571, PHENIX72, and COOT73. Crystallographic parameters are listed in Table 1. All structure figures were prepared with PyMOL. The transition state complex interface was calculated in PDBePISA.
Cell cultures, transfection
HEK293 cells were maintained in Dulbecco’s modified Eagle’s medium supplemented with 10% (V/V) fetal calf serum and penicillin/streptomycin. The 20E2 cell line, a Swedish mutant APP stable HEK293 cell line was maintained in complete DMEM supplemented with 100 μg/mL Zeocin. Cells were transfected using Lipofectamine™ 3000 (Invitrogen) reagent according to the manufacturers’ protocols.
Western blotting and antibodies
For immunoblotting analyses, 20E2 cells were lysed in RIPA lysis buffer supplemented with protease and phosphatase inhibitors (Roche Applied Science). Immunoblotting was performed as described previously74. Primary antibodies used are: anti-myc tag mAb (Cell Signaling, Danvers, MA), C20 antibody, anti-β actin mAb (Sigma-Aldrich), anti-Phospho-Tyrosine mAb (Cell Signaling, Danvers, MA). Detection was performed with the Li-Cor Odyssey imaging system and quantitated with ImageJ software.
GST pull-down assays
One hundred microliter glutathione-agarose beads were incubated with 1 mg purified GST or indicated GST-SH2 fusions and the mutant GST-SH2R680A for 2 h at 4 °C in GST binding buffer (20 mM Tris/HCl, pH 7.4, 150 mM NaCl and 1 mM EDTA, 0.5% TritonX-100). The beads were then washed 4 times with GST binding buffer and incubated for another 2 h with 0.2 mg indicated lysates of 20E2 cells stable expressing the indicated APPsw constructs. After washing 4 times with GST binding buffer, beads were boiled in SDS sample buffer, run on 12% SDS–PAGE gel and analyzed by Coomassie or immunoblotting.
Co-immunoprecipitation (co-IP)
HEK293 cells were co-transfected with pAPPsw and pCMV5-myc-Vav2. For co-IP, cells were harvested after 48 h and lysed in 1 mL of 1% NP-40 lysis buffer supplemented with protease and phosphatase inhibitors (Roche Applied Science). Cell lysates were then incubated with primary antibody and protein A/G-agarose beads (Santa Cruz Biotechnology, Santa Cruz, CA) at 4 °C overnight. Mouse or rabbit IgG (Beyotime Institute of Biotechnology, Haimen, China) was performed with protein A/G-agarose beads as negative controls. After washing 3 times with PBS, beads were boiled in SDS sample buffer, run on 8% SDS–PAGE gel and analyzed by immunoblotting with indicated antibodies.
Immunofluorescence
Immunofluorescence was performed as previously described75. Primary antibodies used were C20 antibody and anti-myc tag mAb (Cell Signaling, Danvers, MA). Secondary antibodies were CoraLite488-conjugated Affinipure Goat Anti-Rabbit IgG and CoraLite594-conjugated Affinipure Goat Anti-Mouse IgG (proteintech, Wuhan, China). Images were captured by LSM 880 fluorescent microscope (Carl Zeiss, Jena, Germany) and analyzed with ZEN software.
Elisa
20E2 cells were transfected with Vav2WT, Vav2R680A, Vav2E205A and empty control, respectively. Forty-eight hours after transfection, the level of Aβ40 in the supernatant was measured using the Aβ1–40 Elisa kit (Cloud-Clone Corp, Wuhan, China) according to the manufacturer’s protocol.
Cycloheximide (CHX) pulse-chase assay
The 20E2 cells were transfected with Vav2WT and empty control, respectively. Forty-eight hours after transfection, cells were treated with 150 μg/mL of CHX (MCE, Shanghai, China) and harvested after 0, 3, 6 and 9 h, respectively. The protein level of FL-APPsw was detected by western blotting and quantitated with ImageJ software.
Data analysis
Data are presented as means ± SEM from three to five independent experiments. Student’s t test was performed for differences between two groups. One-way or two-way ANOVA with Bonferroni’s multiple comparisons post hoc test was applied for multigroup comparisons. All analyses were performed with GraphPad Prism 9 software (GraphPad). Differences were defined to be statistically significant at p < 0.05.
Data availability
Raw data is available from the corresponding authors upon reasonable request. Structure data are deposited in the Protein Data Bank with the accession code 7WFY.
References
Lee, H. N., Jeong, M. S. & Jang, S. B. Molecular characteristics of amyloid precursor protein (APP) and its effects in cancer. Int. J. Mol. Sci. 22, 4999. https://doi.org/10.3390/ijms22094999 (2021).
Guo, Y., Wang, Q., Chen, S. & Xu, C. Functions of amyloid precursor protein in metabolic diseases. Metabolism 115, 154454. https://doi.org/10.1016/j.metabol.2020.154454 (2021).
Caudano, F. et al. cGMP favors the interaction between APP and BACE1 by inhibiting Rab5 GTPase activity. Sci. Rep. 10, 1358. https://doi.org/10.1038/s41598-020-58476-8 (2020).
Ugbaja, S. C., Sanusi, Z. K., Appiah-Kubi, P., Lawal, M. M. & Kumalo, H. M. Computational modelling of potent beta-secretase (BACE1) inhibitors towards Alzheimer’s disease treatment. Biophys. Chem. 270, 106536. https://doi.org/10.1016/j.bpc.2020.106536 (2021).
Gouras, G. K., Olsson, T. T. & Hansson, O. beta-Amyloid peptides and amyloid plaques in Alzheimer’s disease. Neurotherapeutics 12, 3–11. https://doi.org/10.1007/s13311-014-0313-y (2015).
Kotler, S. A., Walsh, P., Brender, J. R. & Ramamoorthy, A. Differences between amyloid-beta aggregation in solution and on the membrane: Insights into elucidation of the mechanistic details of Alzheimer’s disease. Chem. Soc. Rev. 43, 6692–6700. https://doi.org/10.1039/c3cs60431d (2014).
Guo, T. et al. Molecular and cellular mechanisms underlying the pathogenesis of Alzheimer’s disease. Mol. Neurodegener. 15, 40. https://doi.org/10.1186/s13024-020-00391-7 (2020).
Sahoo, B. R. et al. Nanodisc-forming scaffold protein promoted retardation of amyloid-beta aggregation. J. Mol. Biol. 430, 4230–4244. https://doi.org/10.1016/j.jmb.2018.08.018 (2018).
Nguyen, P. H. et al. Amyloid oligomers: A joint experimental/computational perspective on Alzheimer’s disease, Parkinson’s disease, type II diabetes, and amyotrophic lateral sclerosis. Chem. Rev. 121, 2545–2647. https://doi.org/10.1021/acs.chemrev.0c01122 (2021).
Galvao, F. Jr., Grokoski, K. C., da Silva, B. B., Lamers, M. L. & Siqueira, I. R. The amyloid precursor protein (APP) processing as a biological link between Alzheimer’s disease and cancer. Ageing Res. Rev. 49, 83–91. https://doi.org/10.1016/j.arr.2018.11.007 (2019).
Wu, X., Chen, S. & Lu, C. Amyloid precursor protein promotes the migration and invasion of breast cancer cells by regulating the MAPK signaling pathway. Int. J. Mol. Med. 45, 162–174. https://doi.org/10.3892/ijmm.2019.4404 (2020).
Wolfe, M. S. & Guenette, S. Y. APP at a glance. J. Cell. Sci. 120, 3157–3161. https://doi.org/10.1242/jcs.03481 (2007).
Thinakaran, G. & Koo, E. H. Amyloid precursor protein trafficking, processing, and function. J. Biol. Chem. 283, 29615–29619. https://doi.org/10.1074/jbc.R800019200 (2008).
Pantelopulos, G. A., Straub, J. E., Thirumalai, D. & Sugita, Y. Structure of APP-C991-99 and implications for role of extra-membrane domains in function and oligomerization. Biochim. Biophys. Acta Biomembr. https://doi.org/10.1016/j.bbamem.2018.04.002 (2018).
Schettini, G., Govoni, S., Racchi, M. & Rodriguez, G. Phosphorylation of APP-CTF-AICD domains and interaction with adaptor proteins: Signal transduction and/or transcriptional role—Relevance for Alzheimer pathology. J. Neurochem. 115, 1299–1308. https://doi.org/10.1111/j.1471-4159.2010.07044.x (2010).
Cha, H. J., Shen, J. & Kang, J. Regulation of gene expression by the APP family in the adult cerebral cortex. Sci. Rep. 12, 66. https://doi.org/10.1038/s41598-021-04027-8 (2022).
Russo, C. et al. The amyloid precursor protein and its network of interacting proteins: Physiological and pathological implications. Brain Res. Rev. 48, 257–264. https://doi.org/10.1016/j.brainresrev.2004.12.016 (2005).
Zhang, T., Chen, D. & Lee, T. H. Phosphorylation signaling in APP processing in Alzheimer’s disease. Int. J. Mol. Sci. 21, 209. https://doi.org/10.3390/ijms21010209 (2019).
Oliveira, J., Costa, M., de Almeida, M. S. C., da Cruz, E. S. O. A. B. & Henriques, A. G. Protein phosphorylation is a key mechanism in Alzheimer’s disease. J. Alzheimers Dis. 58, 953–978. https://doi.org/10.3233/JAD-170176 (2017).
Tamayev, R., Zhou, D. & D’Adamio, L. The interactome of the amyloid beta precursor protein family members is shaped by phosphorylation of their intracellular domains. Mol. Neurodegener. 4, 28. https://doi.org/10.1186/1750-1326-4-28 (2009).
Lau, K. F., McLoughlin, D. M., Standen, C. L., Irving, N. G. & Miller, C. C. Fe65 and X11beta co-localize with and compete for binding to the amyloid precursor protein. NeuroReport 11, 3607–3610. https://doi.org/10.1097/00001756-200011090-00041 (2000).
Sastre, M., Turner, R. S. & Levy, E. X11 interaction with beta-amyloid precursor protein modulates its cellular stabilization and reduces amyloid beta-protein secretion. J. Biol. Chem. 273, 22351–22357. https://doi.org/10.1074/jbc.273.35.22351 (1998).
Borg, J. P., Yang, Y., De Taddeo-Borg, M., Margolis, B. & Turner, R. S. The X11alpha protein slows cellular amyloid precursor protein processing and reduces Abeta40 and Abeta42 secretion. J. Biol. Chem. 273, 14761–14766. https://doi.org/10.1074/jbc.273.24.14761 (1998).
Ando, K., Iijima, K. I., Elliott, J. I., Kirino, Y. & Suzuki, T. Phosphorylation-dependent regulation of the interaction of amyloid precursor protein with Fe65 affects the production of beta-amyloid. J. Biol. Chem. 276, 40353–40361. https://doi.org/10.1074/jbc.M104059200 (2001).
Zambrano, N. et al. Interaction of the phosphotyrosine interaction/phosphotyrosine binding-related domains of Fe65 with wild-type and mutant Alzheimer’s beta-amyloid precursor proteins. J. Biol. Chem. 272, 6399–6405. https://doi.org/10.1074/jbc.272.10.6399 (1997).
Radzimanowski, J. et al. Structure of the intracellular domain of the amyloid precursor protein in complex with Fe65-PTB2. EMBO Rep. 9, 1134–1140. https://doi.org/10.1038/embor.2008.188 (2008).
Taru, H. et al. Interaction of Alzheimer’s beta -amyloid precursor family proteins with scaffold proteins of the JNK signaling cascade. J. Biol. Chem. 277, 20070–20078. https://doi.org/10.1074/jbc.M108372200 (2002).
Zhou, D., Noviello, C., D’Ambrosio, C., Scaloni, A. & D’Adamio, L. Growth factor receptor-bound protein 2 interaction with the tyrosine-phosphorylated tail of amyloid beta precursor protein is mediated by its Src homology 2 domain. J. Biol. Chem. 279, 25374–25380. https://doi.org/10.1074/jbc.M400488200 (2004).
Das, S., Raychaudhuri, M., Sen, U. & Mukhopadhyay, D. Functional implications of the conformational switch in AICD peptide upon binding to Grb2-SH2 domain. J. Mol. Biol. 414, 217–230. https://doi.org/10.1016/j.jmb.2011.09.046 (2011).
Tarr, P. E., Roncarati, R., Pelicci, G., Pelicci, P. G. & D’Adamio, L. Tyrosine phosphorylation of the beta-amyloid precursor protein cytoplasmic tail promotes interaction with Shc. J. Biol. Chem. 277, 16798–16804. https://doi.org/10.1074/jbc.M110286200 (2002).
Matrone, C., Iannuzzi, F. & Annunziato, L. The Y682ENPTY687 motif of APP: Progress and insights toward a targeted therapy for Alzheimer’s disease patients. Ageing Res. Rev. 52, 120–128. https://doi.org/10.1016/j.arr.2019.04.003 (2019).
Matrone, C. A new molecular explanation for age-related neurodegeneration: The Tyr682 residue of amyloid precursor protein. BioEssays 35, 847–852. https://doi.org/10.1002/bies.201300041 (2013).
Raychaudhuri, M. & Mukhopadhyay, D. Grb2-mediated alteration in the trafficking of AβPP: Insights from Grb2-AICD interaction. J. Alzheimers Dis. 20, 275–292. https://doi.org/10.3233/jad-2010-1371 (2010).
Taru, H. & Suzuki, T. Regulation of the physiological function and metabolism of AbetaPP by AbetaPP binding proteins. J. Alzheimers Dis. 18, 253–265. https://doi.org/10.3233/JAD-2009-1148 (2009).
Lee, Y. S., Chow, W. N. V. & Lau, K. F. Phosphorylation of FE65 at threonine 579 by GSK3beta stimulates amyloid precursor protein processing. Sci. Rep. 7, 12456. https://doi.org/10.1038/s41598-017-12334-2 (2017).
Caster, A. H. & Kahn, R. A. Recruitment of the Mint3 adaptor is necessary for export of the amyloid precursor protein (APP) from the Golgi complex. J. Biol. Chem. 288, 28567–28580. https://doi.org/10.1074/jbc.M113.481101 (2013).
McLoughlin, D. M. & Miller, C. C. The FE65 proteins and Alzheimer’s disease. J. Neurosci. Res. 86, 744–754. https://doi.org/10.1002/jnr.21532 (2008).
Koushika, S. P. “JIP”ing along the axon: The complex roles of JIPs in axonal transport. BioEssays 30, 10–14. https://doi.org/10.1002/bies.20695 (2008).
Wang, L., Zhou, Y., Chen, D. & Lee, T. H. Peptidyl-prolyl cis/trans isomerase Pin1 and Alzheimer’s disease. Front. Cell Dev. Biol. 8, 355. https://doi.org/10.3389/fcell.2020.00355 (2020).
Crespo, P., Schuebel, K. E., Ostrom, A. A., Gutkind, J. S. & Bustelo, X. R. Phosphotyrosine-dependent activation of Rac-1 GDP/GTP exchange by the vav proto-oncogene product. Nature 385, 169–172. https://doi.org/10.1038/385169a0 (1997).
Han, J. et al. Lck regulates Vav activation of members of the Rho family of GTPases. Mol. Cell. Biol. 17, 1346–1353. https://doi.org/10.1128/MCB.17.3.1346 (1997).
Havel, L. S., Kline, E. R., Salgueiro, A. M. & Marcus, A. I. Vimentin regulates lung cancer cell adhesion through a VAV2-Rac1 pathway to control focal adhesion kinase activity. Oncogene 34, 1979–1990. https://doi.org/10.1038/onc.2014.123 (2015).
Ling, Z. et al. Targeting CCL2-CCR4 axis suppress cell migration of head and neck squamous cell carcinoma. Cell Death Dis. 13, 158. https://doi.org/10.1038/s41419-022-04610-5 (2022).
Arora, P. D., Marignani, P. A. & McCulloch, C. A. Collagen phagocytosis is regulated by the guanine nucleotide exchange factor Vav2. Am. J. Physiol. Cell Physiol. 295, C130–C137. https://doi.org/10.1152/ajpcell.00168.2008 (2008).
Cowan, C. W. et al. Vav family GEFs link activated Ephs to endocytosis and axon guidance. Neuron 46, 205–217. https://doi.org/10.1016/j.neuron.2005.03.019 (2005).
Hunter, S. G. et al. Essential role of Vav family guanine nucleotide exchange factors in EphA receptor-mediated angiogenesis. Mol. Cell. Biol. 26, 4830–4842. https://doi.org/10.1128/MCB.02215-05 (2006).
Marignani, P. A. & Carpenter, C. L. Vav2 is required for cell spreading. J. Cell Biol. 154, 177–186. https://doi.org/10.1083/jcb.200103134 (2001).
Schuebel, K. E. et al. Isolation and characterization of murine vav2, a member of the vav family of proto-oncogenes. Oncogene 13, 363–371 (1996).
Bustelo, X. R. Regulatory and signaling properties of the Vav family. Mol. Cell. Biol. 20, 1461–1477. https://doi.org/10.1128/MCB.20.5.1461-1477.2000 (2000).
Rossman, K. L., Der, C. J. & Sondek, J. GEF means go: Turning on RHO GTPases with guanine nucleotide-exchange factors. Nat. Rev. Mol. Cell Biol. 6, 167–180. https://doi.org/10.1038/nrm1587 (2005).
Fang, W. B., Brantley-Sieders, D. M., Hwang, Y., Ham, A. J. & Chen, J. Identification and functional analysis of phosphorylated tyrosine residues within EphA2 receptor tyrosine kinase. J. Biol. Chem. 283, 16017–16026. https://doi.org/10.1074/jbc.M709934200 (2008).
Pandey, A. et al. Analysis of receptor signaling pathways by mass spectrometry: Identification of vav-2 as a substrate of the epidermal and platelet-derived growth factor receptors. Proc. Natl. Acad. Sci. U.S.A. 97, 179–184. https://doi.org/10.1073/pnas.97.1.179 (2000).
Tamas, P. et al. Mechanism of epidermal growth factor regulation of Vav2, a guanine nucleotide exchange factor for Rac. J. Biol. Chem. 278, 5163–5171. https://doi.org/10.1074/jbc.M207555200 (2003).
Ge, L. et al. Biochemical and NMR characterization of the interactions of Vav2-SH2 domain with lipids and the EphA2 juxtamembrane region on membrane. Biochem. J. 477, 3791–3801. https://doi.org/10.1042/BCJ20200300 (2020).
Wu, B. et al. Identification and structural basis for a novel interaction between Vav2 and Arap3. J. Struct. Biol. 180, 84–95. https://doi.org/10.1016/j.jsb.2012.06.011 (2012).
Bodenhausen, G. & Ruben, D. J. Natual abundance nitrogen-15 NMR by enhanced heteronuclear spectroscopy. Chem. Phys. Lett. 69, 185–189. https://doi.org/10.1016/0009-2614(80)80041-8 (1980).
Chrencik, J. E. et al. Structural basis of guanine nucleotide exchange mediated by the T-cell essential Vav1. J. Mol. Biol. 380, 828–843. https://doi.org/10.1016/j.jmb.2008.05.024 (2008).
Rodriguez-Fdez, S. et al. Vav2 catalysis-dependent pathways contribute to skeletal muscle growth and metabolic homeostasis. Nat. Commun. 11, 5808. https://doi.org/10.1038/s41467-020-19489-z (2020).
Lorenzo-Martin, L. F. et al. VAV2 signaling promotes regenerative proliferation in both cutaneous and head and neck squamous cell carcinoma. Nat. Commun. 11, 4788. https://doi.org/10.1038/s41467-020-18524-3 (2020).
Lim, S. et al. Amyloid-beta precursor protein promotes cell proliferation and motility of advanced breast cancer. BMC Cancer 14, 928. https://doi.org/10.1186/1471-2407-14-928 (2014).
Takagi, K. et al. Amyloid precursor protein in human breast cancer: An androgen-induced gene associated with cell proliferation. Cancer Sci. 104, 1532–1538. https://doi.org/10.1111/cas.12239 (2013).
Tsang, J. Y. S. et al. Proteolytic cleavage of amyloid precursor protein by ADAM10 mediates proliferation and migration in breast cancer. EBioMedicine 38, 89–99. https://doi.org/10.1016/j.ebiom.2018.11.012 (2018).
Ojala, V. K. et al. The guanine nucleotide exchange factor VAV3 participates in ERBB4-mediated cancer cell migration. J. Biol. Chem. 295, 11559–11571. https://doi.org/10.1074/jbc.RA119.010925 (2020).
Hu, J., Meng, Y., Zeng, J., Zeng, B. & Jiang, X. Ubiquitin E3 Ligase MARCH7 promotes proliferation and invasion of cervical cancer cells through VAV2-RAC1-CDC42 pathway. Oncol. Lett. 16, 2312–2318. https://doi.org/10.3892/ol.2018.8908 (2018).
Ruggiero, C. & Lalli, E. VAV2: A novel prognostic marker and a druggable target for adrenocortical carcinoma. Oncotarget 8, 88257–88258. https://doi.org/10.18632/oncotarget.21448 (2017).
Sun, X., He, G. & Song, W. BACE2, as a novel APP theta-secretase, is not responsible for the pathogenesis of Alzheimer’s disease in Down syndrome. FASEB J. 20, 1369–1376. https://doi.org/10.1096/fj.05-5632com (2006).
Delaglio, F. et al. NMRPipe—A multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293. https://doi.org/10.1007/Bf00197809 (1995).
Otwinowski, Z. & Minor, W. [20] Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326. https://doi.org/10.1016/S0076-6879(97)76066-X (1997).
Collaborative Computational Project Network. The CCP4 suite: Programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763. https://doi.org/10.1107/S0907444994003112 (1994).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674. https://doi.org/10.1107/S0021889807021206 (2007).
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255. https://doi.org/10.1107/S0907444996012255 (1997).
Adams, P. D. et al. PHENIX: A comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221. https://doi.org/10.1107/S0907444909052925 (2010).
Emsley, P. & Cowtan, K. Coot: Model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132. https://doi.org/10.1107/S0907444904019158 (2004).
Tian, S., Jia, W., Lu, M., Zhao, J. & Sun, X. Dual-specificity tyrosine phosphorylation-regulated kinase 1A ameliorates insulin resistance in neurons by up-regulating IRS-1 expression. J. Biol. Chem. 294, 20164–20176. https://doi.org/10.1074/jbc.RA119.010809 (2019).
Liu, H., Wang, P., Song, W. & Sun, X. Degradation of regulator of calcineurin 1 (RCAN1) is mediated by both chaperone-mediated autophagy and ubiquitin proteasome pathways. FASEB J. 23, 3383–3392. https://doi.org/10.1096/fj.09-134296 (2009).
Acknowledgements
All the NMR experiments were performed at High Magnetic Field Laboratory, Chinese Academy of Sciences.
Funding
This work was supported by grants from the National Natural Science Foundation of China (21673244 and 91849130), the Foundation of Anhui Technologies Major Program (201903a07020026), and the Collaborative Innovation Program of Hefei Science Center, CAS (2020HSC-CIP008). A portion of this work was supported by the High Magnetic Field Laboratory of Anhui Province.
Author information
Authors and Affiliations
Contributions
Y.Z. and X.Y.: Methodology, Investigation, Writing—original draft preparation. Y.L.: Investigation. L.G.: Methodology, Validation, Formal analysis. J.W. (Jiarong Wang): Formal analysis. X.S.: Conceptualization, Resources, Supervision, Funding acquisition. B.W.: Conceptualization, Validation, Writing—review and editing, Supervision, Funding acquisition. J.W. (Junfeng Wang): Conceptualization, Resources, Writing—review and editing, Supervision, Funding acquisition. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, Y., Yang, X., Liu, Y. et al. Vav2 is a novel APP-interacting protein that regulates APP protein level. Sci Rep 12, 12752 (2022). https://doi.org/10.1038/s41598-022-16883-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-16883-z
This article is cited by
-
Exploring the interaction between extracellular matrix components in a 3D organoid disease model to replicate the pathophysiology of breast cancer
Journal of Experimental & Clinical Cancer Research (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.