Abstract
The sterlet (Acipenser ruthenus) is one of the 27 sturgeon species and is well-known for its wide distribution and small body size in comparison to other sturgeons. For assessing the population genetics and parentage identification of sterlet, ten microsatellites developed for Chinese sturgeon and cross-amplified in sterlet were tested by 40 individuals of sterlet. The ten microsatellites were developed using transcriptome sequencing of Chinese sturgeon. The expected heterozygosity (HE), observed heterozygosity (HO), Shannon-Weiner diversity indices (H′) and polymorphic information content (PIC) of the 10 microsatellites ranged from 0.466 to 0.751, from 0.438 to 0.938, from 0.66 to 1.51 and from 0.368 to 0.716, respectively. Combined exclusion probability based on the genotype of pair parent known (CE-PP), one parent known (CE-2P), and no parent known (CE-1P) of the 10 microsatellites were 99.99%, 99.96%, and 99.49%, respectively. These result showed that the 10 microsatellites should be helpful for assessing the population genetics and parentage identification of sterlet.
Similar content being viewed by others
Sturgeon is famous for its caviar. Due to the influence of human activities, the number of all sturgeon is decreasing. The sterlet (Acipenser ruthenus, Linnaeus, 1758) is one of the 27 sturgeon species and is well-known for its wide distribution and small body size in comparison to other sturgeons1. Genetic breeding of farmed sterlet is an effective means to meet the demand for human consumption of caviar and food, as well as for conservation of wild sturgeon resources. The sexual maturation period of sterlet is usually 2–3 years and 3–4 years for males and females, respectively. Therefore, sterlet is an ideal model to study the sturgeon due to earliest maturation2. The aquaculture development of sterlet has two major bottlenecks, which is the larval rearing and the broodstock reproduction3. Due to many years of artificial breeding, inbreeding has become an urgent problem in the breeding of small sturgeons.
Microsatellite refer to simple sequence repeats (SSRs) or short tandem repeats (STRs) and are considered as tracts of DNA motifs, which is ranging from one to 10 nucleotides in length, and the DNA motifs is repeating of 5–50 times4. Microsatellite play an important role on studying the structure and the genetic diversity of populations for supportive stocking programs. Microsatellite can be used to evaluate animals breeding and develop pedigree animal populations, supporting genetic improvement5. The microsatellite DNA has been widely used for genetic analysis of aquatic animals, such as Acipenser sinensis6, Exopalaemon carinicauda7, O. niloticus8, Acipenser dabryanus9, Acipenser schrenckii10, and Siniperca scherzeri11. Moreover, duplex PCR panels for microsatellite loci co-amplification can reduce time and costs in genetic analysis of aquaculture animals. Microsatellite for genetic studies of the sterlet have been developed12. In that study, six species-specific microsatellites for starlet were developed to examined in 67 sterlet individuals (20 farmed and 47 wild-caught), and the result suggested the wild sterlet population had indications of inbreeding. However, no study has been reports about parentage identification for sterlet. The application of microsatellites using in parentage verification have produced highly effective and accurate results13. Therefore, there is a need for developing microsatellites that are suitable for accurate parentage testing in sterlet. Hence, in the present study, we aimed to develop microsatellites that are should be helpful for accurate parentage testing in sterlet.
Methods
Ethics statement
The present study was approved by the Animal Administration and Ethics Committee of Xi’an University of Technology. All experiments were carried out in accordance with relevant guidelines and regulations. The study was carried out in compliance with the ARRIVE guidelines.
Sample collection
Chinese sturgeon (Acipenser sinensis) using in transcriptome assembly was collected in the Chinese Sturgeon Research Institute, (Yichang, China). The 16 individuals of sterlet were randomly selected from the corresponding cultures in a farm in Xi’an, China. These individuals were used for validating polymorphic microsatellite markers. In addition, A full-sib families of sterlet was obtained by artificial insemination in the farm. 22 progenies and their parents were collected for DNA isolation. A total of 40 individuals were used in this study.
DNA isolation
The fin of each individuals was taken and preserved in alcohol. Genomic DNA was extracted from fin of all samples of sterlet. The standard proteinase-K digestion and phenol/chloroform method was used for DNA isolation. The fin of sterlet was put in the tube with 10 µl proteinase-K for 50 min at 56 °C. Subsequently, 600 µl of extraction buffer (0.5 M Tris pH 8.0, 0.5 M EDTA, 5 M NaCl, 10% SDS) was added to this tube, and incubated for 10 min at 95 °C. Then, 500 µl phenol was put in the tube. The mixture was violented shock for 10 s and centrifuged for 15 min at 12000 g at 4 °C. 200 µl supernatant of mixture was transferred to a fresh tube. 250 µl chloroform was put this tube, and the mixture was centrifuged for 15 min at 12,000 g at 4 °C. The supernatant of mixture was transferred to another a fresh tube. 500 µl isopropanol was put this tube, and the mixture was centrifuged for 15 min at 12,000 g at 4 °C. The supernatant of mixture was discarded and 1 ml 75% ethanol was put in this tube to precipitate the pellet. And the mixture was centrifuged for 15 min at 12,000 g at 4 °C, and the supernatant of mixture was discarded. At last, the pellet was dissolved using 200 µl sterile water. 1% agarose gel was used to test DNA integrity.
Transcriptome assembly
The fin of Chinese sturgeon was used to extract RNA using Trizol (Invitrogen, USA) following the manufacturer’s instructions. 1% agarose gel was used to test RNA integrity. Then, the RNA of Chinese sturgeon was used to construct cDNA libraries by using Illumina truSeq stranded mRNA sample preparation protocol. The Illumina HiSeq™ 4000 with 100 bp paired-end sequencing was used to sequence the cDNA libraries with 100 or 2 × 90 bp paired-end sequencing. SOAPnuke (v1.5.6)14 was used to filter the raw reads of libraries according to three criteria: (i) reads with ≥ 20% bases Q ≤ 20 were removed, (ii) reads with a high proportion of unknown sequences were removed (> 5%), (iii) reads with contained adaptors (adaptors with ≤ 20% mismatches were allowed) were discarded15. The Trinity program (version: release-2013-08-14) was used to assemble the clean reads. The TGICL16 was used to cluster the transcripts. The redundant sequences were eliminated and the non-redundant sequences of > 200 bp were retained.
Identification of microsatellites
The Microsatellite Identification tool17 was used to select sequences that is constraining perfect repeat motifs of 4 bp from the libraries data. And the Microsatellite Identification tool was set for search criteria for identification of at least 6 repeat units. The Primer Premier 5.0 software18 was used to design microsatellite primers from all the selected sequences. The primer pairs flanking each microsatellite was identified with a annealing temperature of an optimum at 60 °C, PCR products with expected lengths between 100 and 400 bp.
Marker selection
The DNA of 16 randomly individuals of sterlet were used to screen for polymorphic microsatellites and to optimize amplification conditions. PCR amplification was performed in 25 μl volumes containing 0.25 μM each primer, 1.5 mM MgCl2, 0.25 U of Taq polymerase (Takara, China), 0.25 μM PCR buffer (Takara, China), 0.25 μM dNTPs, about 50–100 ng of template DNA, and ultra-pure water. PCR amplification was conducted under the following conditions: an initial step at 94 °C for 3 min, followed by 35 cycles of denaturing 30 s at 94ºC, 60 °C for 30 s, and 72 °C for 30 s, with a final prolonging at 72 °C for 10 min. The PCR products were resolved with 10% polyacrylamide gel electrophoresis (PAGE). The pBR322 DNA/Mspl marker (Takara) was used as a molecular size standard.
Establishing duplex PCR panels for sterlet and their application in parentage identification
According to the result of marker selection of sterlet, duplex PCR reactions was chosen. The primer dimers and potential hairpin structures must be avoided in choosing of duplex PCR reactions. If primer dimers and potential hairpin structures appear in PCR reaction products, this combination of primers is filtered out. Five duplex PCR reactions were assembled, each containing two microsatellites. The duplex PCR reactions were performed at a total volume of 25 μl, containing 0.25 μM each primer, 1.5 mM MgCl2, 0.5 U of Taq polymerase (Takara, China), 0.5 μM PCR buffer (Takara, China), 0.5 μM dNTPs, about 80–100 ng of template DNA, and ultra-pure water. The duplex PCR reactions were performed under the following profile: an initial step at 94 °C for 3 min, followed by 35 cycles of denaturing 30 s at 94 °C, 60 °C for 30 s, and 72 °C for 30 s, with a final prolonging at 72ºC for 10 min. The PCR products were resolved with 10% polyacrylamide gel electrophoresis (PAGE). The pBR322 DNA/Mspl marker (Takara) was used as a molecular size standard. The selected microsatellites for duplex PCR reactions were used for parentage identification of the full-sib families of sterlet.
Genetic analysis
The ATetra1.2 software19 was used to calculate observed heterozygosity (HO), the mean expected heterozygosity (HE), and Shannon-Weiner diversity indices (H′). The polymorphic information content (PIC) was calculated using the formula \(PIC=1-\sum_{i=1}^{n}{P}_{i}^{2}-\sum_{i=1}^{n-1}{\sum }_{j=i+1}^{n}2{P}_{i}^{2}{P}_{j}^{2}\), Pi and Pj are the frequencies of I and J allele in each microsatellite loci. The Gene Marker software was used to calculate the size of allele. Microsoft Office Excel 2007 was used to process the genotypic data. Based on the allele phenotypes, a dendrogram of ten random individuals of sterlet was constructed using MEGA software20. The FaMoz software21 was used to calculate the exclusion and combined exclusion probability based on the allele frequencies. During the analysis the genetic data of the experiment, the dropout and false allele should be avoid.
Results
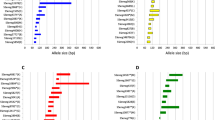
In this study, a total of 160 microsatellites primers were designed to develop duplex PCR reactions. 16 sterlet individuals were used to test the 160 microsatellites, and 10 microsatellites showed clearly polymorphism (Table 1). The observed allele number of the 10 microsatellites ranged from 2 to 6. The expected heterozygosity (HE), observed heterozygosity (HO), Shannon-Weiner diversity indices (H′) and polymorphic information content (PIC) of the 10 microsatellites ranged from 0.466 to 0.751, from 0.438 to 0.938, from 0.66 to 1.51 and from 0.368 to 0.716, respectively (Table 2). According to the results for the 10 microsatellites tested in the 16 sterlet individuals, we build five duplex PCR reactions (Table 1). The 16 sterlet individuals were accurately reconstructed in the dendrogram based in the 10 microsatellites by Mega software (Fig. 1). The five duplex PCR reactions were also used in the parentage analysis of the full-sib families of sterlet. Combined exclusion probability based on the genotype of pair parent known (CE-PP), one parent known (CE-2P), and no parent known (CE-1P) increased with the number of microsatellites (Fig. 2). Combined exclusion probabilities based on the genotype of CE-PP, CE-2P, and CE-1P of the 10 microsatellites were 99.99%, 99.96%, and 99.49%, respectively. Combined exclusion probability of CE-PP, CE-2P, and CE-1P can be reach 99% when used 10, 8, 5 microsatellites, respectively. These result showed that the 10 microsatellites should be helpful for parentage identification of sterlet.
Combined exclusion probabilities based on the genotype of pair parent known (CE-PP), one parent known (CE-2P), and no parent known (CE-1P) of 10 microsatellites in a full-sib families of sterlet. Note: CE-1P: Combined exclusion probability (no parent known); CE-2P: Combined exclusion probability (one parent known); CE-PP: Combined exclusion probability (pair parent).
Discussion
Application of microsatellite markers play important role in genetic conservation and management of fish, and many studies have demonstrated that microsatellite is usefulness for the broodstock management and assessment of population structures22,23,24,25,26. In the present study, five duplex PCR reactions were built to be used in the population genetics and parentage identification for sterlet. The microsatellite for sterlet have been developed in previous studies12,27. However, only seven tetranucleotide microsatellites were reported12. The risk of obtaining false alleles can be reduced by using tetranucleotide microsatellites instead of dinucleotide microsatellites28. The dinucleotide microsatellites are more likely to occur as shadow bands than tetranucleotide microsatellites during the PCR process29. Furthermore, tetranucleotide microsatellites are more stable and accurate than dinucleotide microsatellites30. Therefore, the 10 tetranucleotide microsatellites should be helpful for genetic study of sterlet.
The observed heterozygosity and expected heterozygosity of the 10 microsatellites in this study is higher than the previous microsatellites12,27. In this study, 9 of 10 microsatellites with PIC > 0.5, which is higher than the microsatellites that be reported in beluga sturgeon31, Amur sturgeon10, however, is lower than the microsatellites that be reported in Chinese sturgeon6, Dabry’s Sturgeon9. Although the method of alleles scoring (PAGE) used in this study is cheap and quick, it has many drawbacks, such as result interpretation error, misreading false allele and so on. In this study, there is no evidence of stutter bands, null alleles, or large allele dropout. The reason for this result may be due to the small sample size. These result suggested that the 10 microsatellites were helpful in studying of the population genetics of sterlet.
The previous study have showed that the CE-1P and CE-2P for parentage identification of Nile tilapia were 0.967 and 0.9999 by using 7 microsatellites32. Using four groups of fluorescent-labeled multiple capillary for parentage assignment, the assignment success rate of O. niloticus, O. niloticus × O. aureus, O. aureus, and their mixed population reached 100% when 7, 8, 9, and 12 microsatellites, respectively33. The parentage identification success rate was highly correlated with the allelic polymorphism of microsatellite34. The 10 microsatellites in this study have showed high allelic polymorphism, suggesting that they are suitable for parentage identification of sterlet. They can be useful tools to characterize the structure and genetic diversity of sterlet, to appropriate breeding schemes and establish broodstocks for supportive stocking programs, to aid the development of conservation measures as well as to monitor genetic changes in farmed strains. Although the 10 microsatellites presented here was meaning for sterlet parentage identification, this methodology still has some limitations: (a) The PIC values of the 10 microsatellites in this study are not sufficiently high to reduce the number of microsatellites for parentage assignment; (b) More stable microsatellites need to be further screened.
In conclusion, these five duplex PCR reactions, which consist of 10 microsatellites, should be helpful for assessing the population genetics and parentage identification of sterlet.
Data availability
Data are available from the corresponding author upon reasonable request.
References
Müller, T. et al. Attempts on artificial induction of sexual maturation of sterlet (Acipenser ruthenus) and identification of late spermatogenesis stage in hermaphroditic fish. Int. Aquatic Res. 10, 293–297 (2018).
Akhavan, S. R., Salati, A. P., Falahatkar, B. & Jalali, S. A. H. Changes of vitellogenin and Lipase in captive Sterlet sturgeon Acipenser ruthenus females during previtellogenesis to early atresia. Fish Physiol. Biochem. 42, 967–978 (2016).
Abdollahpour, H., Falahatkar, B., Efatpanah, I., Meknatkhah, B. & Van Der Kraak, G. Influence of thyroxine on spawning performance and larval development of Sterlet sturgeon Acipenser ruthenus. Aquaculture 497, 134–139 (2018).
Vieira, M. L. C., Santini, L., Diniz, A. L. & Munhoz, Cd. F. Microsatellite markers: what they mean and why they are so useful. Genet. Mole. Biol. 39, 312–328 (2016).
Weising, K., Winter, P., Hüttel, B. & Kahl, G. Microsatellite markers for molecular breeding. J. Crop. Prod. 1, 113–143 (1997).
Hu, Y. et al. Development and characterization of a duplex PCR assay in Chinese sturgeon (Acipenser sinensis) for genetic analysis. Sci. Rep. 10, 1–7 (2020).
Zhang, Q., Zhang, C., Yu, Y., Wang, Q. & Li, F. Characteristic analysis of simple sequence repeats in the ridgetail white prawn Exopalaemon carinicauda genome and its application in parentage assignment. J. World Aquac. Soc. 51, 690–701 (2020).
Zhao, Y. et al. An optimized group mating design and determination of the admixture rate in Nile tilapia families. Aquac. Rep. 3, 45–50 (2016).
Hu, Y. et al. Development of 17 novel cross-species microsatellite markers for Dabry’s Sturgeon (Acipenser dabryanus) from Chinese Sturgeon (Acipenser sinensis) via next-generation sequencing. Pak. J. Zool. 51, 2381–2384 (2019).
Hu, Y. et al. Development of twenty-two novel cross-species microsatellite markers for amur sturgeon (Acipenser schrenckii) from Chinese Sturgeon (Acipenser sinensis) via next-Generation Sequencing. Turkish J. Fish. Aquatic Sci. 19, 175–178 (2018).
Yang, M. et al. Parentage determination in golden mandarin fish (Siniperca scherzeri) based on microsatellite DNA markers. Aquacult. Int. 23, 499–507 (2015).
Kohlmann, K., Kersten, P., Gener, J., Onr, D. & Suciu, R. New microsatellite multiplex PCR sets for genetic studies of the sterlet sturgeon, Acipenser ruthenus. Environ. Biotechnol. 13, 11–17 (2017).
Linacre, A. et al. ISFG: recommendations regarding the use of non-human (animal) DNA in forensic genetic investigations. Forensic Sci. Int. Genet. 5, 501–505 (2011).
Chen, Y. et al. SOAPnuke: A MapReduce acceleration-supported software for integrated quality control and preprocessing of high-throughput sequencing data. Gigascience 7, gix120 (2018).
Du, H. et al. Hypothalamus-pituitary-gonad axis transcriptome profiling for sex differentiation in Acipenser sinensis. Sci. Data 6, 1–8 (2019).
Pertea, G. et al. TIGR Gene Indices clustering tools (TGICL): A software system for fast clustering of large EST datasets. Bioinformatics 19, 651–652 (2003).
Beier, S., Thiel, T., Münch, T., Scholz, U. & Mascher, M. MISA-web: A web server for microsatellite prediction. Bioinformatics 33, 2583–2585 (2017).
Lalitha, S. Primer premier 5. Biotech Softw. Int. Rep. Comput. Softw. J. Sci. 1, 270–272 (2000).
Van Puyvelde, K., Van Geert, A. & Triest, L. ATETRA, a new software program to analyse tetraploid microsatellite data: Comparison with TETRA and TETRASAT. Mol. Ecol. Resour. 10, 331–334 (2010).
Kumar, S., Tamura, K. & Nei, M. MEGA3: Integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief. Bioinform. 5, 150–163 (2004).
Gerber, S., Chabrier, P. & Kremer, A. FAMOZ: A software for parentage analysis using dominant, codominant and uniparentally inherited markers. Mol. Ecol. Notes 3, 479–481 (2003).
Moghim, M. et al. Application of microsatellite markers for genetic conservation and management of P ersian sturgeon (A cipenser persicus, Borodin, 1897) in the C aspian S ea. J. Appl. Ichthyol. 29, 696–703 (2013).
Welsh, A., Hill, T., Quinlan, H., Robinson, C. & May, B. Genetic assessment of lake sturgeon population structure in the Laurentian Great Lakes. North Am. J. Fish. Manag. 28, 572–591 (2008).
Wozney, K. M., Haxton, T. J., Kjartanson, S. & Wilson, C. C. Genetic assessment of lake sturgeon (Acipenser fulvescens) population structure in the Ottawa River. Environ. Biol. Fishes 90, 183–195 (2011).
Henderson-Arzapalo, A. & King, T. L. Novel microsatellite markers for Atlantic sturgeon (Acipenser oxyrinchus) population delineation and broodstock management. Mol. Ecol. Notes 2, 437–439 (2010).
Boscari, E., Pujolar, J. M., Dupanloup, I., Corradin, R. & Congiu, L. Captive breeding programs based on family groups in polyploid sturgeons. PLoS ONE 9, e110951 (2014).
Dudu, A., Georgescu, S. E., Burcea, A., Florescu, I. & Costache, M. Microsatellites variation in sterlet sturgeon, Acipenser ruthenus from the lower danube. Sci. Papers Animal Sci. Biotechnol./Lucrari Stiintifice Zootehnie si Biotehnologii 46, 90–94 (2013).
Taberlet, P., Waits, L. P. & Luikart, G. Noninvasive genetic sampling: Look before you leap. Trends Ecol. Evol. 14, 323–327 (2008).
Archie, E. A., Moss, C. J. & Alberts, S. C. Characterization of tetranucleotide microsatellite loci in the African Savannah Elephant (Loxodonta africana africana). Mol. Ecol. Notes 3, 244–246 (2003).
Sean, W. P., Fildes, N. J. & Rebecca, R. Sequence analysis and characterization of stutter products at the tetranucleotide repeat locus vWA. Nucleic Acids Res. 24, 2807–2812 (1996).
Toi, K., Ciorpac, M., Taflan, E., Holostenco, D. & Gener, J. Development and assessment of a duplex PCR assay for genetic analyses of microsatellite loci in beluga sturgeon, Huso huso. Genet. Aquatic Org. 2 (2018).
Zhao, Y. et al. An optimized group mating design and determination of the admixture rate in Nile tilapia families. Aquac. Rep. 3, 45–50 (2016).
Liu, Z. et al. Establishment and application of a microsatellite-based parentage identification method for Oreochromis niloticus, O. aureus, and O. niloticus × O. aureus. J Appl Ichthyol 37, 83–88 (2021).
Macavoy, E. S., Wood, A. R. & Gardner, J. Development and evaluation of microsatellite markers for identification of individual Greenshell mussels (Perna canaliculus) in a selective breeding programme. Aquaculture 274, 41–48 (2008).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (No. 51709225).
Author information
Authors and Affiliations
Contributions
J.W. and Y.H. conceived and designed the research. J.W. wrote the manuscript with contributions from Z.S., L.J. and Y.H. Y.H. analysed the data. J.W. and Y.H. carried out the experiment.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, J., Sun, Z., Jiang, L. et al. Developing microsatellite duplex PCR reactions for sterlet (Acipenser ruthenus) and their application in parentage identification. Sci Rep 12, 12036 (2022). https://doi.org/10.1038/s41598-022-16194-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-16194-3
This article is cited by
-
Isolation of tetrameric microsatellite markers and its application on parentage identification in Procambarus clarkii
Aquaculture International (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.