Abstract
Fruits of Garcinia mangostana L. (mangosteen) are rich in nutrients with xanthones found in the pericarp having great pharmaceutical potential. Mangosteen variety Mesta is only found in Malaysia, which tastes sweeter than the common Manggis variety in Southeast Asia. In this study, we report the complete mitogenome of G. mangostana L. variety Mesta with a total sequence length of 371,235 bp of which 1.7% could be of plastid origin. The overall GC content of the mitogenome is 43.8%, comprising 29 protein-coding genes, 3 rRNA genes, and 21 tRNA genes. Repeat and tandem repeat sequences accounted for 5.8% and 0.15% of the Mesta mitogenome, respectively. There are 333 predicted RNA-editing sites in Mesta mitogenome. These include the RNA-editing events that generated the start codon of nad1 gene and the stop codon of ccmFC gene. Phylogenomic analysis using both maximum likelihood and Bayesian analysis methods showed that the mitogenome of mangosteen variety Mesta was grouped under Malpighiales order. This is the first complete mitogenome from the Garcinia genus for future evolutionary studies.
Similar content being viewed by others
Introduction
Mitochondria are the main organelle of energy production for cell sustainability. The first land plant mitochondrial genome (mitogenome) sequenced was Marchantia polymorpha1. Mitogenome sizes are more variable, which range from 66 kb2 to 11.3 Mb3 compared to plastomes that are more conserved with the length range from 110 to 200 kb4. Plant mitogenomes are complex due to the rearrangement, duplication, recombination, and horizontal gene transfer between nucleus and organelles (plastids and mitochondria)5,6,7. Despite the variation in plant mitogenome size, the functional genes found in plant mitogenomes are conservative8,9 with low point mutation rates10. Hence, mitochondrial genomes (mitogenomes) serve as an important genetic information for the phylogenetic and evolutionary studies11 in addition to chloroplast genomes (plastomes) with distinct genetic inheritance.
Most of the plant phylogenetic studies were focused on plastomes. To date (as of 28 April 2022), there are a total of 7260 and 1104 plant chloroplast and plastid genomes respectively deposited into the GenBank (https://www.ncbi.nlm.nih.gov/genome/browse#!/organelles/) compared to 348 plant mitogenome. There is no mitogenome from the family of Clusiaceae that exists in the GenBank Organelle Genome database. This massive difference in the number of organelle genomes deposited in GenBank is due to the repetitive regions and rearrangement events that are common in mitogenomes, which hinder the assembly using short-read sequencing12. However, with the advancement of third-generation sequencing technology such as Pacific Biosciences (PacBio) and Oxford Nanopore Technologies (ONT), which can sequence over 100 kb per molecule and up to 1 million base pairs (bp), long-read sequencing is becoming more commonly applied in mitogenome assembly13,14,15. Hence, exploring the plant mitogenomes with a slower evolution rate compared to plastomes and nucleus genomes16 can complement evolutionary studies.
Mangosteen (Garcinia mangostana L.) is well-known as the “Queen of fruits” with sweet and juicy fruit pulp. It is a tropical fruit under the Clusiaceae family17 that can be found in Southeast Asia countries18. In Malaysia, there is a unique variety Mesta, which is characterized by an oblong shape, thicker mesocarp, and relatively fewer and smaller seeds compared to the common Manggis variety. Despite the eight published plastomes from Garcinia species19, there is no mitogenome from Garcinia species reported to date.
In this study, we assembled a complete mitogenome of Mesta using PacBio data20 and polished using Illumina short reads21. We also compared its structure and gene contents with five closely related species from the same order of Malpighiales and another two species from the Brassicales order, namely Arabidopsis thaliana and Carica papaya (reference used during assembly). The study reports the first mitogenome from the family of Clusiaceae and provides a reference for future evolutionary studies of the Garcinia genus.
Results
General features of Garcinia mangostana var. Mesta mitogenome
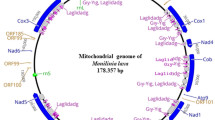
De novo assembly using Organelle_PBA generated two mitogenome contigs with the length of 389,277 bp (scf7180000000010) and 20,340 bp (scf7180000000011), respectively. The smaller contig was the subset of the larger (master) contig (Supplementary Figure S1). For the larger contig that was circular, manual curation was done by removing one of the identical ends (~ 18 kb) and a total of 63 bases were added based on the detected variants when short reads were aligned to the trimmed mitogenome (Supplementary Figure S2). The final complete mitogenome of Garcinia mangostana var. Mesta was 371,235 bp which was slightly larger than Arabidopsis thaliana mitogenome (367,808 bp) but smaller than Carica papaya mitogenome (476,890 bp) (Table 1). The average Mesta mitogenome coverage was 129 × using PacBio subreads (Supplementary Figure S3). The mitogenome comprising 29 protein-coding genes, 3 rRNA genes (rrn5, rrn18, and rrn26), and 21 tRNA genes (Fig. 1 & Table 2). The total length of protein-coding genes was 28,113 bp, which accounted for 7.6% of the mitogenome. There were only five ribosomal proteins (rpl5, rpl10, rpl16, rps3, and rps4) found in the Mesta mitogenome.
The circular mitochondrial genome of G. mangostana variety Mesta. Genes inside the circle are transcribed clockwise while genes outside the circle are transcribed anti-clockwise. The gray bars inside the circle represent the GC content of the sequence. Asterisks (*) indicate genes containing intron(s).
Comparison of mitogenome gene content of different species
Comparison of the mitogenomes gene content of different species (Fig. 2 and Supplementary Table S1) showed that all the mitogenomes encoded the basic core set of 24 protein-coding genes (complex I, III, IV, V, and cytoplasmic membrane proteins). However, the mitogenome of Passiflora edulis encoded two copies of the genes ccmB, nad4L, nad6, and nad7 and four copies of the gene cox2. Mitogenome of Arabidopsis thaliana also encoded two copies of atp6 gene with the length of 1158 bp and 1050 bp (Fig. 2 and Supplementary Table S1), respectively. Two copies of ccmFN gene were also found in the mitogenome of Carica papaya. The nad1 gene in Mesta was found only consisted of 3 exons instead of 5 as observed in other species despite the total length of its CDS sequence being almost like the other species. Both genes, sdh3 and sdh4, which were mitochondrial complex II were not found in Mesta mitogenome. Most of the ribosomal proteins were found missing in both Salicaceae and Clusiaceae families from the order Malpighiales. Noticeably, gene rps12 was missing from Mesta but found in other species in the comparison.
Protein-coding genes in plant mitogenomes. White boxes indicate that the gene is not present in the mitogenome. Yellow, green, and blue boxes indicate that one, two, and four copies exist in the mitogenome, respectively. Purple boxes indicate trans-splicing. Mangosteen (Garcinia mangostana) is shown in bold.
Distribution of tRNAs
The 21 tRNAs identified in Mesta mitogenome only code for 14 amino acids (Ser, Phe, Asn, Met, Pro, Gly, Lys, Gln, Tyr, His, Trp, Asp, Glu, Cys). Two out of 21 tRNAs had a chloroplast-origin while the rest were mitochondrial-origin. However, the tRNA genes code for the other six amino acids (Leu, Ile, Thr, Ala, Val, Arg) were not detected. Among the 21 tRNAs, one of them was predicted to have one intron (Supplementary Table S2).
Plastome-derived sequences
There were five plastome-derived sequences with an identity of more than 80% and a sequence length of at least 100 bp (Table 3) found in the mitogenome. This accounted for a total length of 6214 bp which was 1.7% of the mitogenome. The plastome genes contained were rpl2 (partial), rpl23, trnl-CAU, ndhA (partial), ndhH, rps15 (partial), atpE (partial), atpB, rps3 (partial), and trnD-GUC. Three of the fragments were found at the plastome large single-copy (LSC) region and one from the single-copy (SSC) region and inverted repeat (IR) regions, respectively.
Introns and RNA Editing
There were 20 introns distributed among 8 protein-coding genes (ccmFC, cox1, nad1, nad2, nad4, nad5, nad7, and rps3) in Mesta mitogenome. Among them, nad1, nad2, and nad5 genes were trans-spliced. The maximum number of introns found in a gene were four which can be found at certain nad gene families such as nad2, nad5, and nad7.
A total of 333 putative RNA-editing sites had been predicted using PREP-MT (Supplementary Table S4). Among the CDS, gene nad4 contained the highest number of predicted editing sites (35) while there was no editing site found at the atp9 gene. A total of three annotated genes did not start with the start codon ATG (atp6, nad1, and rpl16) (Supplementary Table S3). Among them, the ACG site at the beginning sequence of nad1 was putatively predicted to be one of the editing sites that converted it into the ATG start codon. Similarly, early termination was predicted at the gene ccmFC sequence which converted the CGA into the stop codon TGA.
Repeat sequences of Mesta mitochondrial DNA
A total of 64 repeat sequences comprising the forward and palindromic repeats were detected. The repeat sequences ranged from 51 to 4172 bp with a total size of 21,356 bp which accounted for 5.8% of the total Mesta mitogenome (Supplementary Table S5). There were three forward repeat sequences with lengths of 4172 bp, 2031 bp, and 1641 bp. As for the tandem repeats, it constituted 0.15% of the mitogenome (Supplementary Table S6).
Phylogenomic analysis
A total of 22 (excluding ccmFn and mttB) out of 24 basic core sets of protein-coding genes (Table S7) were used for phylogenomic analysis. Two methods were used for phylogenomic analysis of mitogenome: Maximum likelihood (ML) and Bayesian phylogenetic analysis. ML analysis separated the 15 species into different groups based on the order (Fig. 3). In comparison, the tree topology of both methods was almost identical except for R. communis (Figs. 3 and 4). Nevertheless, both methods grouped Mesta under Malpighiales order together with P. edulis, S. suchowensis, S. purpurea, P. alba, P. tremula, and P. davidiana.
Discussion
Due to the high complexity of plant mitogenome with large repetitive regions, long-read sequencing is superior in mitogenome assembly13,22. In this study, a total of two Mesta mitogenome contigs were obtained using PacBio data. The shorter contig was a subset of the longer one with the size of 371,235 bp (after manual curation) and was considered as the complete Mesta mitogenome. It is not uncommon to have multiple mitogenome contigs in plants to exist in both circular and linear structures due to intramolecular recombination events12,23,24. For instance, there were 10 contigs in Fagopyrum esculentum25 and 13 contigs in Picea sitchensis22.
Mesta mitogenome encoded the basic core set of 24 protein-coding genes26 commonly found in plant mitogenome27. However, Mesta mitogenome size was relatively small compared to other plant mitogenomes28 from the same order, Malpighiales (Table 1), due to the reduced number of ribosomal proteins and missing genes encoding respiratory chain complex II, sdh3 and sdh4 (Fig. 2 and Supplementary Table S3). These protein-coding genes could be lost during evolution and might be transferred to the nuclear genome as observed in other angiosperm mitogenomes9,29,30,31 such as S. latifolia32, S. noctiflora33, P. dactylifera, and A. indica27,30. For instance, rps12 was not found in Mesta mitogenome as well as Oenothera and Zostera marina34,35.
A complete set of tRNAs coding for 20 amino acids is required for protein translation in plant mitogenomes. However, currently, there was no complete set of tRNA genes found in the mitogenome of angiosperm36. For the Mesta mitogenome, a total of six amino acids encoded by tRNAs (Leu, Ile, Thr, Ala, Val, Arg) were not detected and these tRNAs types were generally reported missing in angiosperm mitogenomes36. In the mitogenome of S. latifolia, the majority of the tRNAs were reported lost with only nine types of amino acids32. The loss of tRNAs can be either replaced by tRNAs from the chloroplast or nuclear genome32,37.
There are several factors attributed to the differences of plant mitogenome lengths, including the integration of nuclear and plastid genomes as well as the number and length of non-coding regions26. Chloroplast sequences can be found in plant mitogenomes as there were integration events during evolution. The integration of plastome sequences into mitogenomes can range from 1 to 12%38. For the Mesta mitogenome, the integration rate was 1.7% which was smaller compared to other plants such as watermelon (7.6%)39, P. dactylifera (10.3%)27, and C. pepo (11.6%)40.
The repetitive regions found in the intergenic regions of mitogenome were in variable types such as short repeats, tandem repeats, and long complex repeats27,28,41,42. The large repeats (> 1 kb) might cause homologous recombination and eventually lead to the different configuration of the mitogenomes43. Apart from large repeats, both direct and inverse repeats also contribute to the subgenomic molecules6. Repeats detected in Mesta mitogenome using web-based REPuter was low (5.8%) compared to C. melo (42.7%)44, V. vinifera (6.8%)45, and N. colorata (48.89%)15 but higher than P. dactylifera (2.3%)27. Similarly, the tandem repeats detected in Mesta mitogenome were also low which was 0.15% compared to 0.33% in P. dactylifera27.
RNA-editing events are essential in plant development and stress response46. The most common RNA editing events in plant organelles (mitochondria and plastids) were the conversion of C-to-U47. RNA editing can lead to the start codon/stop codon generation, eliminate premature stop codon, change the splicing site, affect the RNA structure, and cause instability of RNAs46. It is predicted that RNA-editing events generated the start codon in nad1 and the stop codon in ccmFC genes of Mesta mitogenome. The start codon of nad1 gene in several species such as A. alpina48, B. stricta49, and C. rubella50 was also formed by RNA-editing. On the other hand, stop codon prediction in ccmFC gene sequence had also been reported in A. thaliana and C. bursa-pastoris1. The GTG in rpl16 might be a translation start codon as similar observations were found in maize, Marchantia, and Petunia mitogenomes51.
For phylogenomic analysis, 22 protein-coding genes found in all mitogenomes were used to study the evolutionary relationships among the different species under Malpighiales, Brassicales, Cucurbitales, Rosales, Fabales, and Vitales. Despite one taxon (R. communis) that was not clustered into the Malpighiales order in the Bayesian analysis, both ML and Bayesian analyses grouped the Mesta under Malpighiales order. This observation was in concordance with the mangosteen plastome evolutionary study19. However, plastome could not resolve the G. mangostana species into different cultivars (Manggis and Mesta) due to the same protein-coding genes sequences19. Hence, to verify whether mitogenomes provide a better resolution than plastome in the phylogenetic study of Clusiaceae family, especially mangosteen species of different cultivars, extensive mitogenome assembly are needed for future comparison among different relatives and cultivars.
Conclusion
The complete mitogenome of Mesta was successfully assembled. The Mesta mitogenome length was relatively smaller than other species in the same order due to the loss of most ribosomal proteins and both sdh genes. Phylogenomic analysis based on the 22 protein-coding genes among the 15 selected species showed that Mesta was clustered within the Malpighiales order. The mitogenome can serve as a good reference to study the regulation of the mitogenome genes.
Materials and methods
Mitochondrial genome assembly
Genome sequences of Mesta variety were obtained from the NCBI SRA database with the accession numbers SRX2718652 to SRX2718659 for PacBio long-read data (9.5 Gb)20 and SRX270978 for Illumina short reads (50.2 Gb)21. CANU v2.052 was used to perform PacBio raw data correction and trimming using default parameters. Next, non-mitogenome reads were removed by sequence alignment of each read against a Carica papaya mitogenome (accession no. NC_012116.1) which was used as the reference genome. Then, de novo assembly was performed using Organelle_PBA software53. Manual curation was performed to obtain the complete Mesta mitogenome.
Genome annotation
Gene annotation was conducted using both GeSeq54 and MITOFY web server40. Web-based tRNA-scan v2.0 server (http://lowelab.ucsc.edu/tRNAscan-SE/index.html)55 was used to annotate tRNA genes. The physical mitogenome map was generated using Organellar Genome DRAW (OGDRAW v1.3.1) program with default parameters56.
Identification of plastome derived sequences
Plastome-derived sequences were identified by aligning plastome (accession number: MZ823408) and mitogenome of G. mangostana var. Mesta (accession number: OM759996) using NCBI-Nucleotide BLAST (BLASTN) webserver (https://blast.ncbi.nlm.nih.gov/Blast.cgi) with at least 80% sequence identity and alignment length greater than 100 bp.
Analysis of RNA-editing and substitution rate
Putative RNA-editing sites in protein-coding genes of Mesta mitogenome were predicted using PREP-mt web-based program (http://prep.unl.edu/)57. The cut-off value was set at 0.6 to obtain an accurate prediction.
Analysis of repetitive sequences
The repeat sequences were identified using web-based REPuter (https://bibiserv.cebitec.uni-bielefeld.de/reputer/)58 with a minimum length of repeat size of 50 bp. Web-based tool Tandem Repeats Finder version 4.0959 (https://tandem.bu.edu/trf/trf.basic.submit.html) was used to identify tandem repeats in Mesta mt using default parameter.
Phylogenomic analysis
The amino acids sequences of the 22 protein-coding genes (Table S7) were concatenated and aligned using MAFFT version 7 online tool (https://mafft.cbrc.jp/alignment/server/)60. To examine the evolutionary relationship of G. mangostana var. Mesta, two methods were used for phylogenomic trees construction: (1) The maximum likelihood (ML) analysis was performed using RAxML-NG v1.0.2 tool61 based on the selected model STMTREV + I + G4 + F in the ModelTest-NG v0.1.662; (2) Bayesian phylogenetic analysis was performed using BEAST v.2.6.763. The right substitution model (JTT + F + I + G4) was determined by using the ModelFinder64 in IQ-TREE web server65 (http://iqtree.cibiv.univie.ac.at/) according to the Corrected Akaike Information Criterion (AICc). Next, BEAUti v2.6.7 (part of the BEAST v.2.6.7 package) was used to generate the BEAST XML input file with the following parameters: strict clock, Yule process, and Markov chain Monte Carlo (MCMC) chain length of 10,000,000 with sampling frequency of every 1000 generations. Tracer v1.7.266 was used to examine the Effective Sample Size (ESS) to ensure the value was more than 200. Then, the first 10% of the sample was discarded as burn-in using TreeAnnotator (part of the BEAST v.2.6.7 package) and phylogenetic tree was visualized using FigTreev1.4.4 (http://tree.bio.ed.ac.uk/software/figtree/).
Data availability
The complete mitogenome sequence of Garcinia mangostana var. Mesta has been submitted to GenBank with the accession number OM759996.
References
Oda, K. et al. Gene organization deduced from the@complete sequence of liverwort Marchantia polymorpha mitochondrial DNA: A primitive form of plant mitochondrial genome. J. Mol. Biol. 223, 1–7 (1992).
Skippington, E., Barkman, T. J., Rice, D. W. & Palmer, J. D. Miniaturized mitogenome of the parasitic plant Viscum scurruloideum is extremely divergent and dynamic and has lost all nad genes. Proc. Natl. Acad. Sci. 112, E3515–E3524 (2015).
Sloan, D. B. et al. Rapid evolution of enormous, multichromosomal genomes in flowering plant mitochondria with exceptionally high mutation rates. PLoS Biol. 10, e1001241 (2012).
Sugiura, M. The chloroplast genome. Essays Biochem. 30, 49–57 (1995).
Bergthorsson, U., Richardson, A. O., Young, G. J., Goertzen, L. R. & Palmer, J. D. Massive horizontal transfer of mitochondrial genes from diverse land plant donors to the basal angiosperm Amborella. Proc. Natl. Acad. Sci. 101, 17747–17752 (2004).
Møller, I. M., Rasmusson, A. G. & Van Aken, O. Plant mitochondria—Past, present and future. Plant J. https://doi.org/10.1111/tpj.15495 (2021).
Sanchez-Puerta, M. V. et al. Genome-scale transfer of mitochondrial DNA from legume hosts to the holoparasite Lophophytum mirabile (Balanophoraceae). Mol. Phylogenet. Evol. 132, 243–250 (2019).
Liu, Y., Wang, B., Li, L., Qiu, Y.-L. & Xue, J. Genomics of Chloroplasts and Mitochondria 159–174 (Springer, 2012).
Adams, K. L., Qiu, Y.-L., Stoutemyer, M. & Palmer, J. D. Punctuated evolution of mitochondrial gene content: High and variable rates of mitochondrial gene loss and transfer to the nucleus during angiosperm evolution. Proc. Natl. Acad. Sci. 99, 9905–9912 (2002).
Palmer, J. D. & Herbon, L. A. Plant mitochondrial DNA evolves rapidly in structure, but slowly in sequence. J. Mol. Evol. 28, 87–97. https://doi.org/10.1007/bf02143500 (1988).
O’Conner, S. & Li, L. Mitochondrial fostering: The mitochondrial genome may play a role in plant orphan gene evolution. Front. Plant Sci. https://doi.org/10.3389/fpls.2020.600117 (2020).
Sloan, D. B. One ring to rule them all? Genome sequencing provides new insights into the ‘master circle’model of plant mitochondrial DNA structure. New Phytol. 200, 978–985 (2013).
Omelchenko, D. O. et al. Assembly and analysis of the complete mitochondrial genome of Capsella bursa-pastoris. Plants 9, 469 (2020).
Liao, X. et al. Complete sequence of kenaf (Hibiscus cannabinus) mitochondrial genome and comparative analysis with the mitochondrial genomes of other plants. Sci. Rep. 8, 1–13 (2018).
Dong, S. et al. The complete mitochondrial genome of the early flowering plant Nymphaea colorata is highly repetitive with low recombination. BMC Genomics 19, 614. https://doi.org/10.1186/s12864-018-4991-4 (2018).
Li, S. et al. Progress on sequencing and alignment analysis of higher plant mitochondrial genomes. J. China Agric. Univ. 16, 22–27 (2011).
Nazre, M., Newman, M., Pennington, R. & Middleton, D. Taxonomic revision of Garcinia section Garcinia (Clusiaceae). Phytotaxa 373, 1–52 (2018).
Aizat, W. M., Jamil, I. N., Ahmad-Hashim, F. H. & Noor, N. M. Recent updates on metabolite composition and medicinal benefits of mangosteen plant. PeerJ 7, e6324. https://doi.org/10.7717/peerj.6324 (2019).
Wee, C. C. et al. Plastomes of Garcinia mangostana L. and comparative analysis with other Garcinia species. bioRxiv https://doi.org/10.1101/2022.02.22.481552 (2022).
Midin, M. R. et al. SMRT sequencing data for Garcinia mangostana L. variety Mesta. Genom. Data 12, 134–135. https://doi.org/10.1016/j.gdata.2017.04.003 (2017).
Abu Bakar, S., Kumar, S., Loke, K. K., Goh, H. H. & Mohd Noor, N. DNA shotgun sequencing analysis of Garcinia mangostana L. variety Mesta. Genom. Data 12, 118–119. https://doi.org/10.1016/j.gdata.2017.05.001 (2017).
Jackman, S. D. et al. Complete mitochondrial genome of a gymnosperm, Sitka spruce (Picea sitchensis), indicates a complex physical structure. Genome Biol. Evol. 12, 1174–1179 (2020).
Kozik, A. et al. The alternative reality of plant mitochondrial DNA: One ring does not rule them all. PLoS Genet. 15, e1008373 (2019).
Mower, J. P., Case, A. L., Floro, E. R. & Willis, J. H. Evidence against equimolarity of large repeat arrangements and a predominant master circle structure of the mitochondrial genome from a monkeyflower (Mimulus guttatus) lineage with cryptic CMS. Genome Biol. Evol. 4, 670–686. https://doi.org/10.1093/gbe/evs042 (2012).
Logacheva, M. D., Schelkunov, M. I., Fesenko, A. N., Kasianov, A. S. & Penin, A. A. Mitochondrial genome of Fagopyrum esculentum and the genetic diversity of extranuclear genomes in buckwheat. Plants 9, 618 (2020).
Zardoya, R. Recent advances in understanding mitochondrial genome diversity. F1000Research 9, 270 (2020).
Fang, Y. et al. A complete sequence and transcriptomic analyses of date palm (Phoenix dactylifera L.) mitochondrial genome. PLoS ONE 7, e37164 (2012).
Wynn, E. L. & Christensen, A. C. Repeats of unusual size in plant mitochondrial genomes: identification, incidence and evolution. G3 Genes Genomes Genet. 9, 549–559 (2019).
Asaf, S. et al. Mitochondrial genome analysis of wild rice (Oryza minuta) and its comparison with other related species. PLoS ONE 11, e0152937 (2016).
Choi, K.-S. & Park, S. Complete plastid and mitochondrial genomes of Aeginetia indica reveal intracellular gene transfer (IGT), horizontal gene transfer (HGT), and cytoplasmic male sterility (CMS). Int. J. Mol. Sci. 22, 6143 (2021).
Palmer, J. D. et al. Dynamic evolution of plant mitochondrial genomes: Mobile genes and introns and highly variable mutation rates. Proc. Natl. Acad. Sci. 97, 6960–6966 (2000).
Sloan, D. B., Alverson, A. J., Štorchová, H., Palmer, J. D. & Taylor, D. R. Extensive loss of translational genes in the structurally dynamic mitochondrial genome of the angiosperm Silene latifolia. BMC Evol. Biol. 10, 1–15 (2010).
Sloan, D. B., MacQueen, A. H., Alverson, A. J., Palmer, J. D. & Taylor, D. R. Extensive loss of RNA editing sites in rapidly evolving silene mitochondrial genomes: Selection vs. retroprocessing as the driving force. Genetics 185, 1369–1380. https://doi.org/10.1534/genetics.110.118000 (2010).
Grohmann, L., Brennicke, A. & Schuster, W. The mitochondrial gene encoding ribosomal protein S12 has been translocated to the nuclear genome in Oenothera. Nucleic Acids Res. 20, 5641–5646 (1992).
Petersen, G. et al. Mitochondrial genome evolution in Alismatales: Size reduction and extensive loss of ribosomal protein genes. PLoS ONE 12, e0177606 (2017).
Michaud, M., Cognat, V., Duchêne, A. M. & Maréchal-Drouard, L. A global picture of tRNA genes in plant genomes. Plant J. 66, 80–93 (2011).
Dietrich, A., Small, I., Cosset, A., Weil, J. & Maréchal-Drouard, L. Editing and import: Strategies for providing plant mitochondria with a complete set of functional transfer RNAs. Biochimie 78, 518–529 (1996).
Mower, J. P., Sloan, D. B. & Alverson, A. J. Plant mitochondrial genome diversity: The genomics revolution. Plant Genome Divers. 1, 123–144 (2012).
Cui, H. et al. Comparative analysis of nuclear, chloroplast, and mitochondrial genomes of watermelon and melon provides evidence of gene transfer. Sci. Rep. 11, 1–9 (2021).
Alverson, A. J. et al. Insights into the evolution of mitochondrial genome size from complete sequences of Citrullus lanatus and Cucurbita pepo (Cucurbitaceae). Mol. Biol. Evol. 27, 1436–1448. https://doi.org/10.1093/molbev/msq029 (2010).
Alverson, A. J., Zhuo, S., Rice, D. W., Sloan, D. B. & Palmer, J. D. The mitochondrial genome of the legume Vigna radiata and the analysis of recombination across short mitochondrial repeats. PLoS ONE 6, e16404 (2011).
Figueroa-Martinez, F., Nedelcu, A. M., Smith, D. R. & Reyes-Prieto, A. The plastid genome of Polytoma uvella is the largest known among colorless algae and plants and reflects contrasting evolutionary paths to nonphotosynthetic lifestyles. Plant Physiol. 173, 932–943 (2017).
Gualberto, J. M. et al. The plant mitochondrial genome: Dynamics and maintenance. Biochimie 100, 107–120 (2014).
Rodríguez-Moreno, L. et al. Determination of the melon chloroplast and mitochondrial genome sequences reveals that the largest reported mitochondrial genome in plants contains a significant amount of DNA having a nuclear origin. BMC Genomics 12, 1–14 (2011).
Goremykin, V. V., Salamini, F., Velasco, R. & Viola, R. Mitochondrial DNA of Vitis vinifera and the issue of rampant horizontal gene transfer. Mol. Biol. Evol. 26, 99–110 (2009).
Tang, W. & Luo, C. Molecular and functional diversity of RNA editing in plant mitochondria. Mol. Biotechnol. 60, 935–945 (2018).
Hao, W. et al. RNA editing and its roles in plant organelles. Front. Genet. 12, 1747 (2021).
Xu, Y. & Bi, C. The complete mitochondrial genome sequence of an alpine plant Arabis alpina. Mitochondrial DNA Part B 3, 725–727 (2018).
Li, J., Bi, C., Tu, J. & Lu, Z. The complete mitochondrial genome sequence of Boechera stricta. Mitochondrial DNA Part B 3, 896–897 (2018).
Lin, H. & Bai, D. The complete mitochondrial genome of a highly selfing species Capsella rubella. Mitochondrial DNA Part B 4, 1907–1908 (2019).
Bock, H., Brennicke, A. & Schuster, W. Rps3 and rpl16 genes do not overlap in Oenothera mitochondria: GTG as a potential translation initiation codon in plant mitochondria?. Plant Mol Biol 24, 811–818. https://doi.org/10.1007/bf00029863 (1994).
Walker, B. J. et al. Pilon: An integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 9, e112963. https://doi.org/10.1371/journal.pone.0112963 (2014).
Soorni, A., Haak, D., Zaitlin, D. & Bombarely, A. Organelle_PBA, a pipeline for assembling chloroplast and mitochondrial genomes from PacBio DNA sequencing data. BMC Genomics 18, 49–49. https://doi.org/10.1186/s12864-016-3412-9 (2017).
Tillich, M. et al. GeSeq - versatile and accurate annotation of organelle genomes. Nucleic Acids Res 45, W6-w11. https://doi.org/10.1093/nar/gkx391 (2017).
Chan, P. P. & Lowe, T. M. Gene Prediction 1–14 (Springer, 2019).
Greiner, S., Lehwark, P. & Bock, R. OrganellarGenomeDRAW (OGDRAW) version 1.3.1: Expanded toolkit for the graphical visualization of organellar genomes. Nucleic Acids Res. 47, W59–W64. https://doi.org/10.1093/nar/gkz238 (2019).
Mower, J. P. The PREP suite: Predictive RNA editors for plant mitochondrial genes, chloroplast genes and user-defined alignments. Nucleic Acids Res. 37, W253–W259. https://doi.org/10.1093/nar/gkp337 (2009).
Kurtz, S. et al. REPuter: The manifold applications of repeat analysis on a genomic scale. Nucleic Acids Res. 29, 4633–4642. https://doi.org/10.1093/nar/29.22.4633 (2001).
Benson, G. Tandem repeats finder: A program to analyze DNA sequences. Nucleic Acids Res 27, 573–580. https://doi.org/10.1093/nar/27.2.573 (1999).
Katoh, K., Rozewicki, J. & Yamada, K. D. MAFFT online service: Multiple sequence alignment, interactive sequence choice and visualization. Brief. Bioinform. 20, 1160–1166 (2019).
Kozlov, A. M., Darriba, D., Flouri, T., Morel, B. & Stamatakis, A. RAxML-NG: A fast, scalable and user-friendly tool for maximum likelihood phylogenetic inference. Bioinformatics 35, 4453–4455. https://doi.org/10.1093/bioinformatics/btz305 (2019).
Darriba, D. et al. ModelTest-NG: A new and scalable tool for the selection of DNA and protein evolutionary models. Mol. Biol. Evol. 37, 291–294. https://doi.org/10.1093/molbev/msz189 (2020).
Bouckaert, R. et al. BEAST 2.5: An advanced software platform for Bayesian evolutionary analysis. PLOS Comput. Biol. 15, e1006650. https://doi.org/10.1371/journal.pcbi.1006650 (2019).
Kalyaanamoorthy, S., Minh, B. Q., Wong, T. K., Von Haeseler, A. & Jermiin, L. S. ModelFinder: Fast model selection for accurate phylogenetic estimates. Nat. Methods 14, 587–589 (2017).
Trifinopoulos, J., Nguyen, L.-T., von Haeseler, A. & Minh, B. Q. W-IQ-TREE: A fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Res. 44, W232–W235. https://doi.org/10.1093/nar/gkw256 (2016).
Rambaut, A., Drummond, A. J., Xie, D., Baele, G. & Suchard, M. A. Posterior summarization in Bayesian phylogenetics using Tracer 1.7. Syst Biol 67, 901–904. https://doi.org/10.1093/sysbio/syy032 (2018).
Acknowledgements
We would like to acknowledge the support of this research by Universiti Kebangsaan Malaysia (UKM) Research University grants DIP-2020-005 and AP-2012-018. The group is also supported by NIG-JOINT grant 2021 (2A2021) (YN and H-HG), Japan.
Author information
Authors and Affiliations
Contributions
C.C.W. and H.H.G. conceived and planned the experiment. C.C.W. wrote the paper and analyzed the data. Y.N. contributed to the data analysis. H.H.G., N.A.N.M, V.K.S., M.A., and Y.N. reviewed and edited the manuscript. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wee, CC., Nor Muhammad, N.A., Subbiah, V.K. et al. Mitochondrial genome of Garcinia mangostana L. variety Mesta. Sci Rep 12, 9480 (2022). https://doi.org/10.1038/s41598-022-13706-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-13706-z
This article is cited by
-
The first mitochondrial genome of Calophyllum soulattri Burm.f.
Scientific Reports (2024)
-
Characterization and phylogenetic analysis of the complete mitochondrial genome sequence of Photinia serratifolia
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.