Abstract
Alcohol and tobacco are the most commonly used addictive substances, with high comorbidity rates between alcohol use disorder and tobacco use disorder. Risk for alcohol and nicotine addiction is highly heritable, and they share common genetic factors. A GWAS in over 1 million individuals has revealed 566 genetic variants in 406 loci associated with multiple stages of alcohol and tobacco use. Three novel genes—SLC39A8, GRK4 and HGFAC—within loci associated with altered alcoholic drinks per week (ADW) or cigarettes per day (CPD) were selected to further study their role in alcohol and tobacco use disorder. The role of these genes was assessed using the two-bottle choice addiction paradigm in transgenic mice for each of the genes. We found significant decreases in chronic alcohol consumption and preference in female Hgfac knockout (KO) mice, and decreased nicotine preference in male Hgfac KO compared with wild-type (WT) mice. Additionally, male Slc39a8 hypomorph mice showed greater overall nicotine preference compared with WT mice, while no differences were detected for Grk4 KO mice in alcohol or nicotine consumption and preference in either sex. Thus, this study implicates Hgfac and Slc39a8 in alcohol and tobacco use in a sex-specific manner.
Similar content being viewed by others
Introduction
Alcohol and nicotine are the most commonly misused substances, both in the U.S. and worldwide1,2. In 2019, 27.8% of adults 21 and older used alcohol products, and 69.5% of adults 18 and older used tobacco products in the past year in the U.S. In addition, over 14.5 million individuals aged 12 and older met criteria for an alcohol use disorder (AUD), and nearly 26 million individuals reported nicotine dependence in 2019, respectively3. Moreover, alcohol and tobacco use is highly comorbid, and epidemiological studies indicate that a large proportion of individuals who meet criteria for either an AUD or tobacco use disorder (TUD) also meet criteria for both4,5,6. These traits are highly heterogenous, and numerous complex factors contribute to increased AUD and TUD liability, including genetic and environmental influences. Heritability estimates have shown that genetic factors contribute to approximately 40–60% of the population variability in developing an AUD or TUD, a proportion of which is also common to both disorders7,8,9,10. However, many of the unique and shared genetic factors remain unknown and understudied. Identifying novel genetic factors can provide a better understanding of the molecular and neural mechanisms that contribute to AUD and/or TUD, and potentially identify targets for the development of pharmacotherapies.
One approach for identifying novel variants and gene loci associated with substance use behaviors is the application of genome-wide association studies (GWAS), which perform agnostic tests of association between a phenotype and common genetic variation across the genome. In recent years, GWAS sample sizes have become sufficiently well-powered to detect robust and replicable loci for substance use disorders (SUDs) and related traits. These GWAS findings also demonstrate that alcohol and tobacco use are highly polygenic, composed of a large number of implicated variants and loci, each with small effect. Nonetheless, large-scale GWAS of alcohol and tobacco use phenotypes to date have yielded significant associations for variants and loci in genes with clear biological relevance for addiction, such as those involved in substance metabolism, neural targets, and dopaminergic neurotransmission (for recent reviews of notable SUD GWAS findings, see9,11,12). Many genetic loci that are identified are novel targets for AUD and/or TUD related biology, representing untapped potential in further understanding addiction biology and for developing novel therapies.
Our recent work investigating genetic associations in over one million individuals discovered numerous novel genetic loci and genes that are associated with alcohol and tobacco use13. How these novel genes contribute to the neurobiology of AUD and/or TUD has yet to be determined. The objective of this study was to provide pre-clinical behavioral data on novel genes that were associated with alcoholic drinks per week (ADW) or cigarettes smoked per day (CPD) that were identified in the GWAS study13, using a mouse model of alcohol and nicotine consumption. We chose genes closest to the target genetic loci where the single nucleotide polymorphism (SNP) predicted a non-synonymous mutation, and which had not been widely investigated in the context of AUD and/or TUD in humans: HGFAC, SLC39A8, and GRK4. HGFAC and GRK4 had not been previously associated with AUD and/or TUD in humans, whereas SLC39A8 is highly pleiotropic and has previously been associated with AUD14,15. HGFAC encodes hepatocyte growth factor activator, which is a coagulation factor XII-like serine endopeptidase16 and was associated with ADW13. SLC39A8 encodes for Zrt- and Irt-like protein 8 (ZIP8), a zinc transporter17,18 and was also associated with ADW13. Lastly, GRK4 encodes for G-protein coupled receptor kinase 419 and was associated with CPD13. To observe a potential maximal effect of these genes, we used transgenic mouse lines where the protein products of these genes were deleted or function was severely disrupted. We obtained and tested Hgfac global knock-out (KO) mice, Grk4 global KO mice, and Slc39a8 heterozygote (HET) hypomorph mice (homozygote Slc39a8 gene disruption is lethal)20 and assessed voluntary, alcohol and nicotine consumption in 2-bottle choice tests13. The HGFAC and SLC39A8 variants were associated with reduced ADW in humans13, thus we hypothesized that disruption of Hgfac and Slc39a8 would reduce alcohol consumption and preference in mice. The GRK4 variant was associated with decreased CPD in humans13, thus we hypothesized that disruption of Grk4 would reduce nicotine consumption and preference in mice. We found significant decreases in chronic alcohol consumption and preference in female Hgfac KO mice, and differences in nicotine preference in male Hgfac KO mice compared with wild-type littermates. There were no changes in nicotine consumption in Grk4 KO mice. Male Slc39a8 hypomorph mice showed no changes in alcohol consumption or preference, and an increase in nicotine preference.
Methods
Animals and drugs
Adult male and female mice used in all experiments were a minimum of 8 weeks old. The mice began experiments between 8 and 13 weeks old and all cohorts of mice consisted of age-matched littermate pairs. The Grk4 global KO transgenic breeder mice were obtained from The Jackson Laboratory. The Hgfac global KO breeders were a generous gift from Professor Kataoka at Miyazaki University, Japan. The Slc39a8 HET hypomorph breeders were a generous gift from Dr. Daniel Nebert at the University of Cincinnati and Dr. Zijuan Liu at Oakland University, Michigan, USA. Homozygote Slc39a8 hypomorph mice are estimated to have 10–15% of wild-type protein levels and are not viable20; therefore, all experiments were conducted with Slc39a8 HET and WT mice. All three transgenic lines were maintained on an C57BL/6 genetic background and bred in-house at the University of Minnesota. For each transgenic line, we investigated behaviors in male and female transgenic mice (KO mice for the Hgfac and Grk4 lines, HET mice for the Slc39a8 line) and in the WT littermate controls for each of the three lines. All mice were group housed with a maximum of 4 males and maximum of 5 females in a cage under a standard 12-h light/dark cycle until the start of experiments. Mice were individually housed during the voluntary consumption experiments. All animal procedures and experiments were approved by the Institutional Animal Care and Use Committee (IACUC) at the University of Minnesota, and were in accordance with the recommendations of the ARRIVE guidelines21.
Alcohol (Decon Labs, King of Prussia, PA), and nicotine tartrate salt (Acros Organics, Thermo Fisher Scientific, Chicago, IL) were mixed with tap water to the concentrations reported for each experiment. The concentrations of nicotine described herein are reported as free base. Nicotine solutions were not filtered or pH adjusted, and neither the alcohol nor nicotine solutions contained sweetener. Saccharine and quinine (MilliporeSigma, Burlington, MA) were mixed with tap water to the concentrations reported for the taste experiments.
Alcohol two-bottle choice
Drug naïve male and female mice from the three transgenic lines were singly housed in double grommet cages and underwent a continuous access alcohol two-bottle choice procedure that we and other groups have previously used to evaluate alcohol consumption in mice22,23,24,25. The two-bottle choice procedure was performed as described in our past work investigating voluntary alcohol consumption in mice22,23. Briefly, the procedure involves presenting mice with a bottle of water and a bottle containing alcohol in their home cage. The following escalating alcohol concentrations were presented for 4 days each: 3, 6, 10, 14, and 20% v/v in water. Bottle positions were switched every 2 days to account for side preferences. Bottles were weighed every 2 days and mice were weighed each week. Mice had ad libitum access to food and water during the procedure.
Nicotine two-bottle choice
Drug naïve WT, KO or HET male and female mice were singly housed in double grommet cages and underwent a continuous access nicotine two-bottle choice procedure22,26,27. Mice were presented with a bottle of water and a bottle containing nicotine dissolved in water. The nicotine concentration was presented at the following concentrations for 1 week each: 30, 50, 75, and 100 µg/mL. The bottles were weighed every 2–3 days and replaced with fresh solutions every 3–4 days. The positions of the bottles were switched at every weighing to account for side preferences. The mice were weighed each week. Mice had ad libitum access to food and water during the procedure.
Taste preference test
To determine whether the genes of interest influenced taste preference, and thus could confound the interpretation of the drug 2-bottle choice tests, we assessed the preference for sweet and bitter tasting substances. The taste preference tests were performed in a similar manner as our past work examining sweet and bitter tastes in transgenic mice23. In this study, a subset of mice that completed the alcohol or nicotine two-bottle choice tests were given water for a week prior to beginning the taste test. Briefly, the procedure involves a saccharine two-bottle choice test followed by a quinine two-bottle choice with 3 days of water only in between the tests. The mice were presented with one bottle of water and one bottle of saccharine or quinine in their home cage. The saccharine concentrations presented were 1.5 and 15 mM, and the quinine concentrations presented were 0.01 and 0.1 mM. Each concentration was presented for 2 days each and the bottle positions switched after every day to account for side preferences. Mice were weighed each week. Mice had ad libitum access to food and water during the procedure.
Statistical analysis
Drug consumption (either g/kg/day for alcohol or mg/kg/day for nicotine) was calculated using the weights of the fluid consumed, the concentration and density (for alcohol only) of the solutions and the weights of the mouse. The percent preference for the bottle of interest was calculated as the amount of fluid (g) consumed from the bottle of interest divided by overall fluid consumed from all available bottles multiplied by 100. The consumption and preference were calculated using 2-way repeated measures ANOVA followed by appropriate post-hoc multiple comparison tests using Prism 6.0 (GraphPad, La Jolla, CA). If sex differences in drug consumption were observed in the WT mice, male and female mice were analyzed separately (for each transgenic line), as in past studies22,24,28.
Results
Hgfac—alcohol consumption and preference
As HGFAC was associated with ADW in the GWAS, we first assessed alcohol consumption and preference using a chronic 2-bottle choice test. The Hgfac transgenic line is maintained on a C57BL/6 background, and this genetic background has a well-documented sex difference, in which female mice have greater alcohol consumption and preference compared with male mice22,25,29,30,31. We first compared alcohol consumption between male and female Hgfac WT mice and found a significant interaction between sex and alcohol concentration (Finteraction(4,300) = 15.71, P < 0.0001, Fconcentration(4,300) = 90.69, P < 0.0001, Fsex(1,75) = 82.91, P < 0.0001). Sidak’s multiple comparison testing showed that female mice consumed significantly more alcohol compared with male mice at all concentrations except for 3% (P < 0.05 for 6, 10, 14 and 20% between sexes). For alcohol preference, we also found a significant interaction between sex and alcohol concentration (Finteraction(4,292) = 2.445, P = 0.047, Fconcentration(4,292) = 59.90, P < 0.0001, Fsex(1,73) = 22.02, P < 0.0001). Sidak’s multiple comparisons testing showed that female mice also had higher preference for alcohol at all concentrations except for 3% (P < 0.05 for 6, 10, 14 and 20% between sexes), similar to the results observed for alcohol consumption. Given the observed sex difference in alcohol consumption and preference, we examined the effects of Hgfac separately by sex as we have done in prior work22,24,28.
For alcohol consumption in female Hgfac WT and KO mice, we found a significant interaction between alcohol concentration and genotype (Finteraction(4,260) = 4.046, P = 0.003, Fconcentration(4,260) = 82.19, P < 0.0001, Fgenotype(1,65) = 8.473, P = 0.005, Fig. 1A). Sidak’s multiple comparisons testing showed that female WT mice consumed more alcohol compared with KO littermates at the 10% and 14% concentrations. For alcohol preference, we found main effects of genotype and alcohol concentration with no interaction (Finteraction(4, 256) = 1.400, P = 0.23, Fconcentration(4, 256) = 49.43, P < 0.0001, Fgenotype(1, 64) = 9.804, P = 0.003, Fig. 1B). For alcohol consumption in male drug naïve Hgfac WT and KO mice, we found a significant main effect of alcohol concentration with no interaction between concentration and genotype, and no main effect of genotype (Finteraction(4,288) = 0.773, P = 0.55, Fconcentration(4,228) = 70.35, P < 0.0001, Fgenotype(1,72) = 3.505, P = 0.07, Fig. 1C). For alcohol preference in male mice, we found a main effect of concentration, no main effect of genotype and no significant interaction between genotype and concentration (Finteraction(4,288) = 0.644, P = 0.63, Fconcentration(4,228) = 41.57, P < 0.0001, Fgenotype(1,72) = 0.621, P = 0.43, Fig. 1D). Overall, we found that female Hgfac KO mice showed decreased alcohol consumption and preference compared with WT littermates, with no differences observed in male mice.
Alcohol consumption and preference in Hgfac WT and KO mice. (A) Female Hgfac KO mice had significantly reduced alcohol consumption at the 10% and 14% alcohol concentrations. Sidak’s multiple comparisons testing **P < 0.01 and ***P < 0.001 compared with WT mice. (B) There was a reduction in overall alcohol preference in female Hgfac KO mice compared with WT mice. **P = 0.003 for main effect of genotype. There was no difference between (C) alcohol consumption and (D) alcohol preference between the male Hgfac KO and WT littermates. Data are presented as mean ± SEM, n = 28–39 female mice per genotype, n = 36–38 male mice per genotype.
Hgfac—Nicotine consumption and preference
We first examined nicotine consumption and preference in Hgfac WT males and females to determine whether the documented sex difference in nicotine intake in C57BL/6 mice22,32,33 was also present in this C57BL/6-based transgenic line. For nicotine consumption, we found a significant interaction between sex and nicotine concentration (Finteraction(3,102) = 8.623, P < 0.0001, Fconcentration(3,102) = 50.53, P < 0.0001, Fsex(1,34) = 20.86, P < 0.0001). Sidak’s multiple comparisons testing showed that female Hgfac WT mice consumed more nicotine at the 75 and 100 μg/mL concentrations compared with male Hgfac WT mice. For nicotine preference, we found that female Hgfac WT mice showed greater overall nicotine preference compared with male Hgfac WT mice, with significant main effects of sex and nicotine concentration without an interaction between sex and nicotine concentration (Finteraction(3,102) = 0.014, P = 0.99, Fconcentration(3,102) = 3.469, P = 0.02, Fsex(1,34) = 6.514, P = 0.02).
In female drug naïve Hgfac WT and KO mice, we found a main effect of nicotine concentration with no significant interaction between nicotine concentration and genotype, and no main effect of genotype (Finteraction(3,99) = 0.275, P = 0.84, Fconcentration(3,99) = 68.62, P < 0.0001, Fgenotype(1,33) = 0.069, P = 0.79, Fig. 2A). For nicotine preference in female mice, we found a main effect of concentration with no interaction between nicotine concentration and genotype, and no main effect of genotype (Finteraction(3,99) = 1.212, P = 0.31, Fconcentration(3,99) = 9.707, P < 0.0001, Fgenotype(1,33) = 0.518, P = 0.48, Fig. 2B). For nicotine consumption in male drug naïve Hgfac WT and KO mice, we found a significant interaction between nicotine concentration and genotype (Finteraction(3,99) = 3.267, P = 0.03, Fconcentration(3,99) = 35.25, P < 0.0001, Fgenotype(1,33) = 2.619, P = 0.12, Fig. 2C). Sidak’s multiple comparisons test showed that the male WT mice consumed more nicotine at the 100 μg/mL concentration compared with KO littermates. For nicotine preference, we found a significant main effect of genotype without an interaction between nicotine concentration and genotype or a main effect of genotype (Finteraction(3,99) = 1.857, P = 0.14, Fconcentration(3,99) = 2.459, P = 0.07, Fgenotype(1,33) = 4.451, P = 0.04, Fig. 2D). Overall, we found that male Hgfac KO mice showed reduced nicotine consumption and preference compared with WT littermates, with no differences observed in female mice.
Nicotine consumption and preference in Hgfac WT and KO mice. Female Hgfac WT and KO mice showed no significant differences in (A) nicotine consumption or (B) preference. (C) Male Hgfac KO mice consumed less nicotine at the 100 μg/mL concentration compared with WT littermates. Sidak’s multiple comparisons testing *P < 0.05 between WT and KO mice. (D) Male Hgfac KO mice consumed less nicotine overall compared with WT mice. *P = 0.04 for main effect of genotype. Data are presented as mean ± SEM, n = 17–18 female mice per genotype, n = 16–19 male mice per genotype.
Hgfac—Taste preference
As alcohol and nicotine preference can be influenced by differences in taste preference, we compared saccharine and quinine preference in Hgfac WT and KO mice. In female mice, both saccharine and quinine produced a main effect of concentration only with no interaction between concentration and genotype and no main effect of genotype (saccharine: Finteraction(1,24) = 0.184, P = 0.67, Fconcentration(1,24) = 6.396, P = 0.02, Fgenotype(1,24) = 1.427, P = 0.24, Fig. 3A; quinine: Finteraction(1,23) = 1.995, P = 0.17, Fconcentration(1,23) = 47.39, P < 0.0001, Fgenotype(1,23) = 1.589, P = 0.22, Fig. 3B). Similar results were observed in the male Hgfac WT and KO mice, with a main effect of concentration only for both saccharin and quinine preference (saccharine: Finteraction(1,25) = 0.479, P = 0.50, Fconcentration(1,25) = 6.412, P = 0.02, Fgenotype(1,25) = 0.255, P = 0.62, Fig. 3C; quinine: Finteraction(1,25) = 0.784, P = 0.38, Fconcentration(1,25) = 82.10, P < 0.0001, Fgenotype(1,25) = 0.099, P = 0.76, Fig. 3D). Overall, we observed no changes in sweet or bitter taste preference in either sex of Hgfac KO mice.
Saccharine and quinine consumption and preference in Hgfac WT and KO mice. Female Hgfac WT and KO mice showed no significant differences in (A) saccharine or (B) quinine preference. Male Hgfac WT and KO mice also showed no significant differences in (C) saccharine or (D) quinine preference. Data are presented as mean ± SEM, n = 12–13 female mice per genotype, n = 13–14 male mice per genotype.
Slc39a8—Alcohol consumption and preference
SLC39A8 was associated with ADW in the GWAS, thus we examined 2-bottle choice alcohol consumption. As the Slc39a8 transgenic line is also maintained on a C57BL/6 background, we first examined alcohol consumption and preference across sex in WT mice. For alcohol consumption, we found a significant interaction between sex and alcohol concentration (Finteraction(4,232) = 17.48, P < 0.0001, Fconcentration(4,232) = 47.47, P < 0.0001, Fsex(1,58) = 45.58, P < 0.0001). Sidak’s multiple comparisons testing showed that female Slc39a8 WT mice consumed more alcohol compared with male mice at all concentrations except for 3% (P < 0.05 for 6, 10, 14 and 20% compared with male Slc39a8 WT mice). For alcohol preference, we also found a significant interaction between sex and alcohol concentration (Finteraction(4,236) = 2.691, P = 0.03, Fconcentration(4,236) = 16.53, P < 0.0001, Fsex(1,59) = 6.027, P = 0.02). Sidak’s multiple comparisons testing showed that female Slc39a8 WT mice had a greater preference for alcohol at the 10 and 14% concentrations compared with male Slc39a8 WT mice.
We tested chronic 2-bottle choice alcohol consumption and preference in drug naïve Slc39a8 HET and WT mice. For alcohol consumption in female drug naïve Slc39a8 HET and WT mice, we found a main effect of alcohol concentration with no interaction between concentration and genotype, and no main effect of genotype (Finteraction(4,216) = 0.235, P = 0.92, Fconcentration(4,216) = 56.01, P < 0.0001, Fgenotype(1,54) = 0.998, P = 0.32, Fig. 4A). We observed similar results for alcohol preference in the female mice, with a main effect of alcohol concentration only (Finteraction(4,216) = 0.228, P = 0.92, Fconcentration(4,216) = 14.75, P < 0.0001, Fgenotype(1,54) = 0.144, P = 0.71, Fig. 4B). In the male drug naïve Slc39a8 HET and WT mice, we also found a significant main effect of alcohol concentration only for both alcohol consumption (Fig. 4C) and preference (Fig. 4D) (alcohol consumption: Finteraction(4,252) = 1.649, P = 0.16, Fconcentration(4,252) = 33.79, P < 0.0001, Fgenotype(1,63) = 0.554, P = 0.46; alcohol preference: Finteraction(4,256) = 1.384, P = 0.24, Fconcentration(4,256) = 23.10, P < 0.0001, Fgenotype(1,64) = 0.1159, P = 0.73). Overall, we observed no effect of the Slc39a8 hypomorph on alcohol consumption or preference in either sex.
Alcohol consumption and preference in Slc39a8 WT and HET mice. No differences between Slc39a8 WT and HET mice were observed for (A) female alcohol consumption, (B) female alcohol preference, (C) male alcohol consumption and (D) male alcohol preference. Data are presented as mean ± SEM, n = 28 female mice per genotype, n = 32–33 male mice per genotype.
Slc39a8—nicotine consumption and preference
We first examined nicotine consumption in Slc39a8 WT male and female mice and found an interaction between nicotine concentration and sex (Finteraction(3,153) = 3.910, P = 0.01, Fconcentration(3,153) = 45.77, P < 0.0001, Fsex(1,51) = 14.23, P < 0.001). Sidak’s multiple comparisons testing showed that female Slc39a8 WT mice consumed more nicotine compared with male mice at the 50, 75 and 100 μg/mL concentrations (all P < 0.05 between sex). For nicotine preference, we found a main effect of sex such that female Slc39a8 WT mice had greater overall nicotine preference compared with male mice. There was no interaction between sex and concentration and no main effect of nicotine concentration itself (Finteraction(3,153) = 0.640, P = 0.59, Fconcentration(3,153) = 0.177, P = 0.91, Fsex(1,51) = 6.565, P = 0.01).
We tested nicotine consumption in female drug naïve Slc39a8 WT and HET mice, and found a main effect of nicotine concentration with no interaction between nicotine concentration and genotype, and no main effect of genotype (Finteraction(3,123) = 0.05, P = 0.99, Fconcentration(3,123) = 46.16, P < 0.0001, Fgenotype(1,41) = 0.09, P = 0.77, Fig. 5A). For nicotine preference in female drug naïve Slc39a8 WT and HET mice, we found no significant interaction between nicotine concentration and genotype, and no main effects of genotype or nicotine concentration (Finteraction(3,123) = 0.541, P = 0.66, Fconcentration(3,123) = 0.213, P = 0.89, Fgenotype(1,41) = 0.039, P = 0.85, Fig. 5B). For nicotine consumption in male drug naïve Slc39a8 WT and HET mice, we found a main effect of nicotine concentration only and no main effect of genotype or an interaction between genotype and concentration (Finteraction(3,204) = 0.9384, P = 0.42, Fconcentration(3,204) = 37.87, P < 0.0001, Fgenotype(1,68) = 1.422, P = 0.24, Fig. 5C). However, for nicotine preference in male drug naïve Slc39a8 mice, we found a main effect of genotype such that the Slc39a8 HET mice showed greater overall nicotine preference compared with WT littermates (Fig. 5D). We also observed a trend for a significant interaction between nicotine concentration and genotype and no main effect of nicotine concentration itself (Finteraction(3,204) = 2.395, P = 0.07, Fconcentration(3,204) = 0.1849, P = 0.91, Fgenotype(1,68) = 4.034, P = 0.049, Fig. 5D). To further examine drinking behavior, we analyzed the total fluid consumed over the 4-week experiment normalized by body weight (g/kg) in the male Slc39a8 HET and WT mice. We found a main effect of week and a main effect of genotype, with no interaction between week and genotype (Finteraction(3,204) = 0.549, P = 0.65; Fweek(3,204) = 9.442, P < 0.0001; Fgenotype(1,68) = 4.983, P = 0.03). Male Slc39a8 HET mice drank an average of 11.6% less total fluid by body weight compared with male WT littermates throughout the entire experiment. There was also no difference in the average body weight between male Slc39a8 HET and WT mice (HET: 26.6 ± 0.5 g, WT: 26.3 ± 0.6 g, t = 0.78, P = 0.63). Overall, we observed an increase in nicotine preference in male Slc39a8 HET mice with no effect on nicotine consumption. This increased preference was present despite a small reduction in overall fluid intake in the male Slc39a8 HET mice. No genotype effects were observed for female mice.
Nicotine consumption and preference in Slc39a8 WT and HET mice. No differences between female Slc39a8 WT and HET mice were observed for (A) nicotine consumption or (B) nicotine preference. (C) No genotype differences were observed in nicotine consumption between male Slc39a8 WT and HET mice. (D) Male Slc39a8 HET mice showed greater overall nicotine preference compared with WT littermates. *P = 0.049 for main effect of genotype. Data are presented as mean ± SEM, n = 21–22 female mice per genotype, n = 32–38 male mice per genotype.
Slc39a8—taste preference
In female Slc39a8 WT and HET mice, there was no main effect of saccharin concentration or an interaction between saccharin concentration and genotype (Finteraction(1,11) = 0.043, P = 0.84, Fconcentration(1,11) = 1.440, P = 0.26, Fgenotype(1,11) = 0.5012, P = 0.49, Fig. 6A). Similarly, we found a main effect of quinine concentration without an interaction between quinine concentration and genotype (Finteraction(1,11) = 0.028, P = 0.87, Fconcentration(1,11) = 23.71, P < 0.001, Fgenotype(1,11) = 0.432, P = 0.52, Fig. 6B). In male Slc39a8 WT and HET mice, we observed main effects of saccharin and quinine concentration with no main effect of genotype or an interaction between concentration and genotype (saccharine: Finteraction(1,15) = 1.707, P = 0.21, Fconcentration(1,15) = 8.768, P = 0.01, Fgenotype(1,15) = 0.9721, P = 0.34, Fig. 6C; quinine: Finteraction(1,15) = 0.3869, P = 0.54, Fconcentration(1,15) = 20.74, P < 0.001, Fgenotype(1,15) = 0.032, P = 0.86, Fig. 6D).
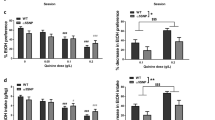
Saccharine and quinine consumption and preference in Slc39a8 WT and HET mice. Female Slc39a8 WT and HET mice showed no significant differences in (A) saccharine or (B) quinine preference. Male Slc39a8 WT and HET mice also showed no significant differences in (C) saccharine or (D) quinine preference. Data are presented as mean ± SEM, n = 5–8 female mice per genotype, n = 7–10 male mice per genotype.
Grk4—nicotine consumption and preference
GRK4 was associated with CPD in the GWAS, thus we first assessed nicotine 2-bottle choice. As this transgenic line is also maintained on a C57BL/6 background, we examined sex differences in WT mice. For nicotine consumption, we found a significant interaction between nicotine concentration and sex (Finteraction(3,93) = 9.959, P < 0.0001, Fconcentration(3,93) = 58.20, P < 0.0001, Fsex(1,31) = 18.24, P < 0.0001). Sidak’s multiple comparisons testing showed that female Grk4 WT mice had greater nicotine consumption compared with male mice at the 50, 75 and 100 μg/mL concentrations. For nicotine preference, we found a main effect of sex such that female Grk4 WT mice had higher overall preference compared with male mice. There was a main effect of nicotine concentration, and no interaction between nicotine concentration and sex (Finteraction(3,93) = 2.483, P = 0.07, Fconcentration(3,93) = 5.707, P = 0.0001, Fsex(1,31) = 4.277, P = 0.04).
For nicotine consumption in drug naïve female Grk4 WT and KO mice, we found a main effect of nicotine concentration, no significant interaction between nicotine concentration and genotype, and no main effect of genotype (Finteraction(3,117) = 0.436, P = 0.73, Fconcentration(3,117) = 84.20, P < 0.0001, Fgenotype(1,39) = 0.059, P = 0.81, Fig. 7A). For nicotine preference in female mice, we found similar results with a main effect of nicotine concentration, no interaction between nicotine concentration and genotype, and no main effect of genotype (Finteraction(3,117) = 0.751, P = 0.52, Fconcentration(3,117) = 18.82, P < 0.0001, Fgenotype(1,39) = 0.004, P = 0.95, Fig. 7B). For nicotine consumption in drug naïve male Grk4 WT and KO mice, we found a main effect of nicotine concentration with no interaction between nicotine concentration and genotype, and no main effect of genotype (Finteraction(3,93) = 1.720, P = 0.17, Fconcentration(3,93) = 34.90, P < 0.0001, Fgenotype(1,31) = 0.971, P = 0.33, Fig. 7C). For nicotine preference in male mice, we observed no interaction between nicotine concentration and genotype, and no main effects of nicotine concentration or genotype (Finteraction(3,93) = 0.392, P = 0.76, Fconcentration(3,93) = 1.286, P = 0.28, Fgenotype(1,31) = 0.5394, P = 0.47, Fig. 7D). Overall, we observed no effect of Grk4 deletion in nicotine consumption or preference in either sex.
Nicotine consumption and preference in Grk4 WT and KO mice. No differences between female Grk4 WT and KO mice were observed for (A) nicotine consumption or (B) nicotine preference. No genotype differences were observed in (C) nicotine consumption or (D) nicotine preference between male Grk4 WT and KO mice. Data are presented as mean ± SEM, n = 16–25 female mice per genotype, n = 16–17 male mice per genotype.
Grk4—alcohol consumption and preference
For alcohol consumption in the Grk4 WT mice, we found an interaction between alcohol concentration and sex (Finteraction(4,68) = 5.526, P < 0.001, Fconcentration(4,68) = 16.78, P < 0.0001, Fsex(1,17) = 23.98, P < 0.001). Sidak’s multiple comparisons testing showed that female Grk4 WT mice had higher alcohol consumption at the 10, 14 and 20% concentrations compared with male mice. For alcohol preference, we found a main effect of alcohol concentration. There was a non-significant trend for a main effect of sex, and no interaction between sex and alcohol concentration (Finteraction(4,80) = 0.1416, P = 0.97, Fconcentration(4,80) = 42.07, P < 0.0001, Fsex(1,20) = 4.021, P = 0.06).
For alcohol consumption in drug naïve female Grk4 WT and KO mice, we found a main effect of alcohol concentration with no interaction between alcohol concentration and genotype, and no main effect of genotype (Finteraction(4,76) = 0.2627, P = 0.90, Fconcentration(4,76) = 41.21, P < 0.0001, Fgenotype(1,19) = 0.1614, P = 0.69, Fig. 8A). For alcohol preference in female mice, we observed similar results with only a main effect of alcohol concentration (Finteraction(4,76) = 1.644, P = 0.17, Fconcentration(4,76) = 69.30, P < 0.0001, Fgenotype(1,19) = 0.7833, P = 0.39, Fig. 8B). For alcohol consumption in drug naïve male Grk4 WT and KO mice, we observed a main effect of alcohol concentration with no interaction between alcohol concentration and genotype, and no main effect of genotype (Finteraction(4,64) = 2.061, P = 0.10, Fconcentration(4,64) = 21.05, P < 0.0001, Fgenotype(1,16) = 3.234, P = 0.09, Fig. 8C). Similarly, for alcohol preference in male mice, we observed a main effect of alcohol concentration only (Finteraction(4,64) = 0.9941, P = 0.42, Fconcentration(4,64) = 26.99, P < 0.0001, Fgenotype(1,16) = 0.011, P = 0.92, Fig. 8D). As there were no genotype effects for either alcohol or nicotine 2-bottle choice tests, we did not evaluate the effect of Grk4 deletion on taste preference.
Alcohol consumption and preference in Grk4 WT and KO mice. No differences between female Grk4 WT and KO mice were observed for (A) alcohol consumption or (B) alcohol preference. No genotype differences were observed in (C) alcohol consumption or (D) alcohol preference between male Grk4 WT and KO mice. Data are presented as mean ± SEM, n = 10–11 female mice per genotype, n = 7–11 male mice per genotype.
Discussion
Despite moderate success in the identification of replicable SUD-related genes, numerous challenges preclude discovery and validation of additional genetic variants and loci, including methodological limitations of genotyping microarrays and genotype imputation, diagnostic and phenotypic heterogeneity, genetic signal divergence across related but distinct behaviors, traits, and measures, and the relative lack of diverse genetic ancestry representation. In addition, the majority of genome-wide significant variants and loci are located in non-coding or regulatory regions of the genome with unclear functional consequence, and the extent of linkage disequilibrium complicates efforts to ascertain the true causal variant(s) and gene(s) within a locus. As such, methods and study designs that can isolate the effects of a specific gene mutation or change in gene product in a tightly controlled experimental environment represent ideal conditions under which to validate the effects of novel candidate genes on a phenotype and clarify the biological mechanisms underlying these associations. One such promising approach uses model organisms to employ back-translation of human GWAS findings for orthologous genes and comparable behavioral phenotypes in non-human species. Identifying the role of novel genes in addiction-relevant behaviors, such as drug consumption and withdrawal, requires genetic manipulation in an animal model that can exhibit the relevant behaviors. Transgenic mice are advantageous as they exhibit a wide variety of complex addiction-related behaviors and are frequently used as pre-clinical models. One example of back-translation using mouse models was a GWAS meta-analysis that identified an intronic variant in KLB to be significantly associated with alcohol consumption, and subsequently validated the effects of this gene on alcohol consumption in brain-expressed β-Klotho knockout mice34. These findings provide preliminary support for the application of back-translation of human GWAS findings to validate novel gene loci and biological relevance, and test for cross-species convergence for SUD-related traits (for additional discussions on the use of model organisms in the 'post-GWAS' era, see35,36).
Based on the results of our large-scale GWAS study13, we selected three genes, HGFAC, SLC38A9 and GRK4, that were associated with altered ADW or CPD to directly test in pre-clinical mouse models of alcohol and nicotine consumption. ADW and CPD are more amenable to modeling in pre-clinical studies since they are consummatory behaviors that can be measured in mice. We used 2-bottle choice tests in which the mice voluntarily consume alcohol or nicotine in their home cage over several weeks. These tests are widely used in the pre-clinical setting to assess voluntary alcohol and nicotine consumption in mice, particularly in the high drug-consuming C57BL/6 mouse strain24,28,32,33,37,38,39,40,41,42. Mice voluntarily consume physiologically relevant levels of drugs as they can exhibit withdrawal symptoms after the drug bottles are withheld22,29,43,44. Additionally, these tests do not require animal training, nor food or fluid restriction at any time. All three of the transgenic lines were maintained on the C57BL/6 background, which is the most frequently used for genetic deletion studies and behavioral assessments40,42,45,46. Within the C57BL/6 strain, female mice are known to consume more alcohol and nicotine compared with male mice22,25,29,30,31,32,33. All three transgenic lines also showed this sex difference in alcohol and nicotine intake, with WT female mice consuming more alcohol and nicotine compared with WT littermate male mice. Thus, we examined the sexes separately within each transgenic line as in our prior work22,24,28,31.
We examined transgenic mouse lines with constitutive, global gene deletions (Hgfac and Grk4 KO) or genes with severely impacted function (Slc39a8 hypomorph HET). Assessing genetic KO mice evaluates the role of the target transcript and protein in these behaviors, which may not be the biological mechanism underlying the association observed in the GWAS. However, a gene deletion may maximize the probability of detecting a genotype effect on alcohol and nicotine consumption in pre-clinical models. Generating a mouse transgenic line that harbors the target variant is a more translational approach and allows for direct testing of the deleteriousness of the variant in addition to behavioral phenotyping; however, this is offset by the time and cost involved in generating the transgenic mice and may not be amenable to testing large numbers of target genes. In addition, transgenic lines with target gene deletions may be readily available if the gene is of interest in research fields other than SUD. Additional limitations of using constitutive, global genetic KO mice are the regulatory processes impacting gene expression are not directly tested and the role of these genes in specific tissues cannot be assessed. Nevertheless, pre-clinical back-translation approaches provide a valuable starting point for investigating the role of novel genes and proteins in AUD and/or TUD biology.
The SNP in the HGFAC gene (rs3748034) was predicted to generate a nonsynonymous missense mutation, which was associated with lower levels of ADW in the GWAS13. HGFAC encodes for hepatocyte growth factor activator and its primary function is as a liver-secreted serum proteinase that converts hepatocyte growth factor inactive precursor (proHGF) to active hepatocyte growth factor (HGF). HGFAC dysfunction has been implicated in impaired tissue injury and repair, impaired gastrointestinal function, inflammation, fibrosis and cancer16. Low levels of HGFAC mRNA have been found in extrahepatic organs including the gastrointestinal tract, kidneys, lungs and central nervous system16,47. Hgfac KO mice are viable and do not show overt abnormalities or tumorigenesis after 1 year of age48. We found reduced alcohol consumption at 10% and 14% alcohol concentrations, and an overall reduction in alcohol preference in the female Hgfac KO mice compared with WT littermates. A change in both the consumption and preference increases the likelihood that Hgfac deletion produces a meaningful change in alcohol intake in the female mice. No changes in alcohol consumption or preference were observed in the male Hgfac KO mice. As the GWAS data on HGFAC was not stratified by sex13, these data suggest an intriguing potential sex difference in the role of Hgfac in alcohol consumption that should be further explored.
Although the mutation in HGFAC was not associated with CPD, we tested nicotine consumption and preference in drug naïve Hgfac WT and KO mice since alcohol and nicotine addiction mechanisms are known to share genetic factors7,8,9,10. Interestingly, we found a reduction in nicotine consumption in the male Hgfac KO mice at the 100 μg/mL concentration, and an overall reduction in preference for nicotine compared with WT littermates. This reduction in nicotine consumption and preference was not observed for the female Hgfac KO mice, again illustrating an intriguing sex difference in alcohol and nicotine responses.
As alcohol consumption is influenced by sweet taste responses49,50 and nicotine has a bitter taste40,51, we measured preference for saccharin (a non-caloric sweetener) and quinine (a bitter substance) in male and female Hgfac KO and WT mice and saw no genotype differences in preference for either sweet or bitter solutions. Thus, the change in alcohol consumption and preference in the female Hgfac KO mice, and change in nicotine consumption and preference in the male Hgfac KO mice are not confounded by altered sweet and bitter taste perception.
Based on the GWAS findings13, these 2-bottle choice tests supported our hypothesis that Hgfac was involved in alcohol intake and also provided unexpected data implicating Hgfac in nicotine intake. Thus, Hgfac may be another common molecular mechanism that is involved in both alcohol and nicotine addiction, and its involvement appears to differ by sex. A recent study examining SUD-related gene interactions found sex differences in the pattern of genetic interactions between PPP1R12B, a member of the DARPP-32 signaling family important for neurotransmission, and HGFAC52. The meta-analysis identified PPP1R12B as an interacting gene with HGFAC in human females but not in males52. In addition to genetic studies, studies using different pre-clinical models will be required to better understand what other aspects of addiction-related behavior are altered in Hgfac KO mice. As the changes in alcohol and nicotine consumption each occurred in one sex only, further investigation into the molecular and biological mechanisms will be needed to understand how altered Hgfac expression influences alcohol and nicotine addiction biology with a particular emphasis on sex differences in the underlying biological mechanisms.
The SNP in SLC39A8 (rs13107325) was predicted to generate a nonsynonymous missense mutation, which was associated with lower levels of ADW in the GWAS13. This SNP is highly pleiotropic and has been associated with schizophrenia, Crohn’s disease, serum manganese and body mass index and others53. Slc39a8 has not been studied in the context of alcohol or nicotine consumption phenotypes in pre-clinical models. Slc39a8 encodes for a Zrt- and Irt-like protein 8 (ZIP8) which functions as a zinc transporter17,18. ZIP8 is expressed widely expressed in the body, with higher levels in the lung, testis, kidney, liver and lower levels in the brain, heart, intestines and pancreas in adult mice54,55,56. The Slc39a8 hypomorph allele consists of a retained neomycin cassette that results in ~ 5–8% of normal mRNA expression levels20. Homozygous Slc39a8neo/neo mice are not viable, thus we used Slc39a8+/neo HET mice in our experiments. Adult Slc39a8 HET mice exhibit increased incidence of spontaneous liver neoplastic nodules between 13 and 21 months of age57—we performed all our experiments in mice well under the age of 13 months. Although the SNP in SLC39A8 was associated with a decrease in ADW, we did not find genotype differences in alcohol consumption or preference in male or female Slc39a8 WT and HET mice. Furthermore, we found no differences in nicotine consumption or preference in between the female Slc39a8 WT and HET mice. Male Slc39a8 HET mice showed slightly greater overall nicotine preference, with no changes in nicotine consumption, compared with WT mice. The increased nicotine preference in male Slc39a8 HET mice cannot be explained by differences in total fluid consumption, as Slc39a8 HET mice consumed less fluid per body weight compared with WT littermates throughout the experiment, and male Slc39a8 HET mice did not show differences in body weight compared with WT littermates. Further experiments examining nicotine conditioned place preference or operant nicotine responding in male mice will be useful in understanding the role of Slc39a8 in nicotine-related phenotypes.
The SNP in GRK4 (rs1024323) was predicted to generate a nonsynonymous missense mutation, which was associated with lower levels of CPD in the GWAS13. GRK4 encodes for G-protein coupled receptor kinase 4, which contributes to the regulation of G protein coupled receptor (GPCR) activity. After activation of a GPCR, GRK proteins phosphorylate the receptor blocking further receptor activation and recruiting cellular processes that internalize the GPCR. Grk4 transcript is expressed in the kidney, bone, heart, testes, intestine and brain19,58,59. Grk4 has primarily been implicated in hypertension19,60. Although the SNP in Grk4 was associated with a decrease in CPD, we found no difference in nicotine consumption and preference in Grk4 KO mice compared with WT littermates in either sex. In addition, there was no effect of Grk4 deletion in alcohol consumption or preference in either sex.
Of the three genes tested in this study, we found altered drug consumption and preference in Hgfac that supported our hypotheses from the GWAS data. Although we did not find a genotype effect in the Slc39a8 and Grk4 transgenic lines in these chronic 2-bottle choice tests, it is possible that Slc39a8 and Grk4 influence alcohol and nicotine addiction behaviors and mechanisms that are not captured by this procedure. As with all pre-clinical models, chronic 2-bottle choice tests assess only a small aspect of alcohol and nicotine addiction-related behaviors. Indeed, variations of the alcohol 2-bottle choice test have been implemented to capture different aspects of alcohol intake compared with continuous chronic access, such as the binge drinking-in-the-dark procedure61 and models that incorporate repeated cycles of abstinence29. In addition, examining the impact of these genes on alcohol and nicotine metabolism can provide further insight into potential mechanisms of action. Identifying additional behavioral models and tests will be important in future studies of the impact of these genes in AUD and/or TUD biology.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on request.
References
Creamer, M. R. et al. Tobacco product use and cessation indicators among adults—United States, 2018. MMWR Morb. Mortal Wkly Rep. 68, 1013–1019 (2019).
US Department of Health and Human Services (2014) The Health Consequences of Smoking- 50 Years of Progress: A Report of the Surgeon General. Atlanta: U.S. Department of Health and Human Services, Centers for Disease Control and Prevention, National Center for Chronic Disease Prevention and Health Promotion, Office on Smoking and Health.
Center for Behavioral Health Statistics and Quality (2020) Results from the 2019 National Survey on Drug Use and Health: Detailed tables. Rockville, MD: Substance Abuse and Mental Health Services Administration. Retrieved from https://www.samhsa.gov/data/.
Batel, P., Pessione, F., Maitre, C. & Rueff, B. Relationship between alcohol and tobacco dependencies among alcoholics who smoke. Addiction 90, 977–980 (1995).
Littleton, J., Barron, S., Prendergast, M. & Nixon, S. J. Smoking kills (alcoholics)! shouldn’t we do something about it. Alcohol Alcohol. 42, 167–173 (2007).
Van Skike, C. E. et al. Critical needs in drug discovery for cessation of alcohol and nicotine polysubstance abuse. Prog. Neuropsychopharmacol. Biol. Psychiatry. 65, 269–287 (2016).
Funk, D., Marinelli, P. W. & Le, A. D. Biological processes underlying co-use of alcohol and nicotine: Neuronal mechanisms, cross-tolerance, and genetic factors. Alcohol Res. Health. 29, 186–192 (2006).
True, W. R. et al. Common genetic vulnerability for nicotine and alcohol dependence in men. Arch. Gen. Psychiatry. 56, 655–661 (1999).
Hancock, D. B., Markunas, C. A., Bierut, L. J. & Johnson, E. O. Human genetics of addiction: New insights and future directions. Curr. Psychiatry Rep. 20, 8 (2018).
Schlaepfer, I. R., Hoft, N. R. & Ehringer, M. A. The genetic components of alcohol and nicotine co-addiction: From genes to behavior. Curr. Drug Abuse Rev. 1, 124–134 (2008).
Deak, J. D. & Johnson, E. C. Genetics of substance use disorders: A review. Psychol. Med. 51, 2189–2200 (2021).
Gelernter, J. & Polimanti, R. Genetics of substance use disorders in the era of big data. Nat. Rev. Genet. 22, 712–729 (2021).
Liu M., Jiang Y., Wedow R., Li Y., Brazel D.M., Chen F. et al. Association studies of up to 1.2 million individuals yield new insights into the genetic etiology of tobacco and alcohol use. Nat. Genet. 51, 237–244 (2019).
Sanchez-Roige S., Palmer A.A., Fontanillas P., Elson S.L., 23andMe Research Team T.S.U.D.W.G.O.T.P.G.C., Adams M.J. et al. Genome-wide association study meta-analysis of the alcohol use disorders identification test (AUDIT) in two population-based cohorts. Am. J. Psychiatry. 176, 107–118 (2019).
Pickrell, J. K. et al. Detection and interpretation of shared genetic influences on 42 human traits. Nat. Genet. 48, 709–717 (2016).
Fukushima, T., Uchiyama, S., Tanaka, H. & Kataoka, H. Hepatocyte growth factor activator: a proteinase linking tissue injury with repair. Int. J. Mol. Sci. 19, E3435 (2018).
He, L. et al. ZIP8, member of the solute-carrier-39 (SLC39) metal-transporter family: Characterization of transporter properties. Mol. Pharmacol. 70, 171–180 (2006).
Wang, C. Y. et al. ZIP8 is an iron and zinc transporter whose cell-surface expression is up-regulated by cellular iron loading. J. Biol. Chem. 287, 34032–34043 (2012).
Yang, J., Villar, V. A., Jones, J. E., Jose, P. A. & Zeng, C. G protein-coupled receptor kinase 4: Role in hypertension. Hypertension 65, 1148–1155 (2015).
Chen, J. et al. In utero gene expression in the Slc39a8(neo/neo) knockdown mouse. Sci. Rep. 8, 10703 (2018).
Percie du Sert N., Ahluwalia A., Alam S., Avey M.T., Baker M., Browne W.J. et al. Reporting animal research: Explanation and elaboration for the ARRIVE guidelines 2.0. PLoS Biol. 18, e3000411 (2020).
O’Rourke, K. Y., Touchette, J. C., Hartell, E. C., Bade, E. J. & Lee, A. M. Voluntary co-consumption of alcohol and nicotine: Effects of abstinence, intermittency, and withdrawal in mice. Neuropharmacology 109, 236–246 (2016).
Lee, A. M. et al. Deletion of Prkcz increases intermittent ethanol consumption in mice. Alcohol Clin. Exp. Res. 38, 170–178 (2014).
Moen, J. K., DeBaker, M. C., Myjak, J. E., Wickman, K. & Lee, A. M. Bidirectional sex-dependent regulation of α6 and β3 nicotinic acetylcholine receptors by protein kinase Cε. Addict. Biol. 26, e12954 (2021).
Kamens, H. M., Andersen, J. & Picciotto, M. R. Modulation of ethanol consumption by genetic and pharmacological manipulation of nicotinic acetylcholine receptors in mice. Psychopharmacology 4, 253–260 (2010).
Wong A.L., McElroy S.M., Robinson J.M., Mulloy S.M., El Banna F.K., Harris A.C. et al. Flavor-specific enhancement of electronic cigarette liquid consumption and preference in mice. Drug Alcohol Dependence. 211, 107995 (2020).
Meliska, C. J., Bartke, A., McGlacken, G. & Jensen, R. A. Ethanol, nicotine, amphetamine, and aspartame consumption and preferences in C57BL/6 and DBA/2 mice. Pharmacol. Biochem. Behav. 50, 619–626 (1995).
DeBaker, M. C., Moen, J. K., Robinson, J. M., Wickman, K. & Lee, A. M. Unequal interactions between alcohol and nicotine co-consumption: suppression and enhancement of concurrent drug intake. Psychopharmacology 237, 967–978 (2020).
Hwa, L. S. et al. Persistent escalation of alcohol drinking in C57BL/6J mice with intermittent access to 20% ethanol. Alcohol. Clin. Exp. Res. 35, 1938–1947 (2011).
Sneddon, E. A., White, R. D. & Radke, A. K. Sex differences in binge-Like and aversion-resistant alcohol drinking in C57BL/6J mice. Alcohol. Clin. Exp. Res. 43, 243–249 (2019).
Touchette, J. C., Maertens, J. J., Mason, M. M., O’Rourke, K. Y. & Lee, A. M. The nicotinic receptor drug sazetidine-A reduces alcohol consumption in mice without affecting concurrent nicotine consumption. Neuropharmacology 133, 63–74 (2018).
Bagdas, D. et al. Assessing nicotine dependence using an oral nicotine free-choice paradigm in mice. Neuropharmacology 157, 107669 (2019).
Klein, L. C., Stine, M. M., Vandenbergh, D. J., Whetzel, C. A. & Kamens, H. M. Sex differences in voluntary oral nicotine consumption by adolescent mice: A dose-response experiment. Pharmacol. Biochem. Behav. 78, 13–25 (2004).
Schumann, G. et al. KLB is associated with alcohol drinking, and its gene product β-Klotho is necessary for FGF21 regulation of alcohol preference. Proc. Natl. Acad. Sci. U S A. 113, 14372–14377 (2016).
Palmer R.H.C., Johnson E.C., Won H., Polimanti R., Kapoor M., Chitre A. et al. Integration of evidence across human and model organism studies: A meeting report. Genes Brain Behav. e12738 (2021).
Reynolds, T. et al. Interpretation of psychiatric genome-wide association studies with multispecies heterogeneous functional genomic data integration. Neuropsychopharmacology 46, 86–97 (2021).
Belknap, J. K., Richards, S. P., O’Toole, L. A., Helms, M. L. & Phillips, T. J. Short-term selective breeding as a tool for QTL mapping: ethanol preference drinking in mice. Behav. Genet. 27, 55–66 (1997).
Cao, J., Gautier, N. M. & Li, M. D. CD-1 mice show individual differences in nicotine preference in a modified two-bottle oral self-administration model. Front. Psychiatry. 3, 28 (2012).
DeBaker, M. C., Robinson, J. M., Moen, J. K., Wickman, K. & Lee, A. M. Differential patterns of alcohol and nicotine intake: Combined alcohol and nicotine binge consumption behaviors in mice. Alcohol 85, 57–64 (2020).
Glatt, A. R., Denton, K. & Boughter, J. D. J. Variation in nicotine consumption in inbred mice is not linked to orosensory ability. Chem. Senses. 34, 27–35 (2009).
Lee, A. M. & Messing, R. O. Protein kinase C epsilon modulates nicotine consumption and dopamine reward signals in the nucleus accumbens. Proc. Natl. Acad. Sci. U S A. 108, 16080–16085 (2011).
Yoneyama, N., Crabbe, J. C., Ford, M. M., Murillo, A. & Finn, D. A. Voluntary ethanol consumption in 22 inbred mouse strains. Alcohol 42, 149–160 (2008).
Locklear, L. L., McDonald, C. G., Smith, R. F. & Fryxell, K. J. Adult mice voluntarily progress to nicotine dependence in an oral self-selection assay. Neuropharmacology 63, 582–592 (2012).
Renda, A., Penty, N., Komal, P. & Nashmi, R. Vulnerability to nicotine self-administration in adolescent mice correlates with age-specific expression of α4* nicotinic receptors. Neuropharmacology 108, 49–59 (2016).
Belknap, J. K., Crabbe, J. C., Riggan, J. & O’Toole, L. A. Voluntary consumption of morphine in 15 inbred mouse strains. Psychopharmacology 112, 352–358 (1993).
Kuzmin, A. & Johansson, B. Reinforcing and neurochemical effects of cocaine: Differences among C57, DBA, and 129 mice. Pharmacol. Biochem. Behav. 65, 399–406 (2000).
Kataoka, H. & Kawaguchi, M. Hepatocyte growth factor activator (HGFA): pathophysiological functions in vivo. FEBS J. 277, 2230–2237 (2010).
Itoh, H. et al. Regeneration of injured intestinal mucosa is impaired in hepatocyte growth factor activator-deficient mice. Gastroenterology 127, 1423–1435 (2004).
Blednov, Y. A. et al. Perception of sweet taste is important for voluntary alcohol consumption in mice. Genes Brain Behav. 7, 1–13 (2008).
Brasser, S. M., Norman, M. B. & Lemon, C. H. T1r3 taste receptor involvement in gustatory neural responses to ethanol and oral ethanol preference. Physiol. Genomics. 41, 232–243 (2010).
Gyekis, J. P. et al. Gustatory, trigeminal, and olfactory aspects of nicotine intake in three mouse strains. Behav. Genet. 42, 820–829 (2012).
Liu, K. et al. Epistatic evidence for gender-dependant slow neurotransmission signalling in substance use disorders: PPP1R12B versus PPP1R1B. EBioMedicine 61, 103066 (2020).
Costas, J. The highly pleiotropic gene SLC39A8 as an opportunity to gain insight into the molecular pathogenesis of schizophrenia. Am. J. Med. Genet. B Neuropsychiatr. Genet. 177, 274–283 (2018).
Wang, B. et al. Enhanced cadmium-induced testicular necrosis and renal proximal tubule damage caused by gene-dose increase in a Slc39a8-transgenic mouse line. Am. J. Physiol. Cell Physiol. 292, C1523–C1535 (2007).
Girijashanker, K. et al. Slc39a14 gene encodes ZIP14, a metal/bicarbonate symporter: similarities to the ZIP8 transporter. Mol. Pharmacol. 73, 1413–1423 (2008).
Fujishiro, H. et al. The role of ZIP8 down-regulation in cadmium-resistant metallothionein-null cells. J. Appl. Toxicol. 29, 367–373 (2009).
Liu, L. et al. Hepatic ZIP8 deficiency is associated with disrupted selenium homeostasis, liver pathology, and tumor formation. Am. J. Physiol. Gastrointest. Liver Physiol. 315, G569–G579 (2018).
Sallese, M., Lombardi, M. S. & De Blasi, A. Two isoforms of G protein-coupled receptor kinase 4 identified by molecular cloning. Biochem. Biophys. Res. Commun. 199, 848–854 (1994).
Premont R.T., Macrae A.D., Aparicio S.A., Kendall H.E., Welch J.E.& Lefkowitz R.J. et al. The GRK4 subfamily of G protein-coupled receptor kinases. Alternative splicing, gene organization, and sequence conservation. J. Biol. Chem. 274, 29381–29389 (1999).
Zhang, H., Sun, Z. Q., Liu, S. S. & Yang, L. N. Association between GRK4 and DRD1 gene polymorphisms and hypertension: A meta-analysis. Clin. Interv. Aging. 11, 17–27 (2016).
Rhodes, J. S., Best, K., Belknap, J. K., Finn, D. A. & Crabbe, J. C. Evaluation of a simple model of ethanol drinking to intoxication in C57BL/6J mice. Physiol. Behav. 84, 53–63 (2005).
Acknowledgements
This work was supported by R01 AA026598 (AML), R01 DA044283 (SIV).
Author information
Authors and Affiliations
Contributions
F.K.E.B., S.M.M., W.T., S.M.M., A.L.W., G.C. collected and analyzed the data. F.K.E.B., J.M.O., S.I.V. and A.M.L. wrote and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Banna, F.K.E., Otto, J.M., Mulloy, S.M. et al. Back-translating GWAS findings to animal models reveals a role for Hgfac and Slc39a8 in alcohol and nicotine consumption. Sci Rep 12, 9336 (2022). https://doi.org/10.1038/s41598-022-13283-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-13283-1
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.