Abstract
Udder measures have been used to assess milk yield of sheep through classical methods of estimation. Artificial neural networks (ANN) can deal with complex non-linear relationships between input and output variables. In the current study, ANN were applied to udder measures from Pelibuey ewes to estimate their milk yield and this was compared with linear regression. A total of 357 milk yield records with its corresponding udder measures were used. A supervised learning was used to train and teach the network using a two-layer ANN with seven hidden structures. The globally convergent algorithm based on the resilient backpropagation was used to calculate ANN. Goodness of fit was evaluated using the mean square prediction error (MSPE), root MSPE (RMSPE), correlation coefficient (r), Bayesian’s Information Criterion (BIC), Akaike’s Information Criterion (AIC) and accuracy. The 15–15 ANN architecture showed that the best predictive milk yield performance achieved an accuracy of 97.9% and the highest values of r2 (0.93), and the lowest values of MSPE (0.0023), RMSPE (0.04), AIC (− 2088.81) and BIC (− 2069.56). The study revealed that ANN is a powerful tool to estimate milk yield when udder measures are used as input variables and showed better goodness of fit in comparison with classical regression methods.
Similar content being viewed by others
Introduction
Traditionally sheep rearing in tropical regions of Latin America is focused on lamb production. However, recently, farmers have increased their interest on diversifying sheep products. Pelibuey is a breed used mainly for meat production as it has maternal ability and prolificity, and over the last decade its milk production performance has been explored1. Milk production and milk composition from Pelibuey sheep has been analyzed as pure breed2 and as a crossbred with specialized dairy breeds1.The improvement of milk production in this breed requires milk yield recording and in order to favor an adequate productive performance, determination of udder morphological characteristics is needed2.
In sheep as in other dairy animals, the anatomical characteristics of udder are important for milk production and milkability3. The mammary gland morphology is related with milk production and aptitude for mechanical milking, which has already been used in genetic programs for dairy sheep4,5. For instance, Rovai et al.6 suggested that sheep with greater milk yields showed large cisternal udders that were able to secrete high milk volumes. In the same line, McKusick et al.7 reported that increasing one centimeter of udder circumference and udder height results in an increased from 0.06 to 0.11 L of milk in East Friesian ewes. This relationship allows to estimate milk yields from the udder measures.
Several approaches have been used to estimate milk production based on morphological measures of the udder and those have yielded to large differences in their predictive accuracy. The ability to estimate dairy performance depends on the number and type of udder measures, stage of lactation and analytical tools used to assess the udder-milk yield relationship8. Analyzing the relationships between udder morphology and milk yield from dairy ewes started back in the 1950s, and until now, all studies focused on this topic have analyzed this relationship using classical estimation methods9,10, without exploring new computational analyzing tools that increase the ability to explain milk yield performance based on udder measures. In this regard, most of the studies use multiple regressions as an approach to estimate milk production using udder measures as main predictor. However, these traditional approaches have several limitations such as sensibility for missing data, need of normal distribution data, and overparameterization when many predictors are used, which limits their use and fitting performance.
The used artificial neural networks (ANN) are capable to deal with complex non-linear relationships between input and output variables, with no need for rigid a priori models11. The AAN is a nonlinear information processing system inspired by the biological nervous system. The ANN is based on interconnected elementary processing devices called neurons. The networking begins from the input information through one (or) more hidden layers, to the input layers. The ANN application has several advantages as good self-learning ability, nonlinearity, massive parallelism, easy implementation, adaptability, real-time operation, and fault tolerance ability12,13. The ANN has been used in animal science for classification and prediction purposes related to nutritional management13,14, dairy performance15,16, animal health17, genetic improvement11,18 and processing of dairy products19. However, in ruminants there is no evidence of their used to estimate the relationship between milk yield and mammary gland measures in Pelibuey ewes.
The ANN are flexible models in terms of model assumptions and structure with accurate and precise predictions13. However, its use is challenged by defining optimal neuron network architecture, which implies that the determination of hidden units required to mimic the analyzed system using input–output examples is needed. The approach to this issue requires considering the relation between the model complexity and training performance. For example, using Akaikes’s Information Criterion (AIC) and other criteria of goodness of fit allow choosing adequate ANN architecture20. Therefore, the aim of this study is to assess the different ANN architectures applied to udder measures from Pelibuey ewes to estimate milk yield and comparing them with predictions from linear regressions.
Results
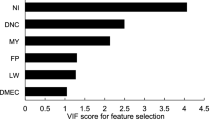
Scatterplots, distributions and correlation coefficients of input and output variables are shown in Fig. 1. Must variables had a normal distribution, but the VDF and MY showed right skewness. The scatterplots showed significant linear relationship between the inputs CI (r = 0.66; p < 0.001) and VDF (r = 0.741; p < 0.001) with the output MY. Also, a significant correlation between CI and IW (r = 0.615; p < 0.001) and VDF (r = 0.661; p < 0.001) was found, which could determine the presence of multicollinearity in the MLR implementation.
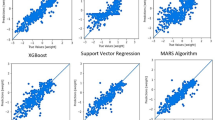
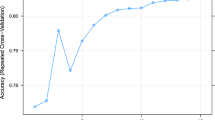
Table 1 shows the goodness of fit of MRL and the seven hidden configurations of the ANN. The proportion of the variance of MY explained by MRL was of 62.0%, with an accuracy of 90.6% and a value of RMSPE of 0.11 kg. The ANN with an architecture of two layers with five neurons (5–5) each one showed similar values of goodness of fit criteria and MLR accuracy. However, with the increase of neurons in both layers improved the goodness of fit. This finding indicated that modifying the number of hidden nodes influenced the model performance and its estimations. Figure 2 shows the best ANN architecture, which was structured by two-layers and 15 neurons each one (15–15). The 15–15 ANN architecture achieved an accuracy of 97.9% and depicted the highest values of r (0.97) and r2 (0.93), and the lowest values of RMSPE (0.04 kg), AIC (− 2088.81) and BIC (− 2069.56).
Figures 3 and 4 shows the residual plots of the MRL and 15–15 ANN models used to estimate milk yield. According to their size, and shown as red colored of circles, in Fig. 3, the MRL model showed the worst estimations of milk yields that were around 0.75 and 1.0 kg. In contrast, the ANN 15–15 showed a lower goodness of fit when estimated milk yields were around 0.25 and 0.5 kg. Finally, when the magnitude of residuals was compared with MRL and ANN 15–15, the artificial intelligence approaches showed lower values along with all milk yield estimations.
Discussion
Our study assessed the usefulness of udder measures to estimate milk production on Pelibuey sheep using machine-learning techniques. The accurate prediction of milk yield is a fundamental step for implementing strategies to improve dairy performance in small ruminants. Using milk yield records, the ANN have been applied to estimate milk yield in dairy cows15,21,22, sheep23, goat24,25 and buffalo26. The udder measures have been widely used to predict milk yield2; however, these predictions have been carried out using traditional statistical tools such as multiple linear regression. The current study is the first reporting the use of ANN to estimate milk yield based on udder measures from sheep.
Several studies have compared the performance of traditional predictive tools (i.e., multiple linear regression and random forest) versus ANN to estimate milk yield15,21, predict mastitis27, forecasting cow locomotion score28, prediction of body weight29,30, modelling rumen fill31 and prediction of carcass tissue composition32. Compared with MLR, results from the current study revealed a better performance of ANN to estimate milk yield based on udder measures. These findings agree with Bhosale and Singh21 who reported that the performance of ANN (r2 = 83.5) was better than the MLR (r2 = 76.21) model for milk yield prediction. In addition, other authors have reported a better performance of ANN versus MLR when estimating milk yields from ruminants15,33,34.
In the current study, an increase of goodness of fit and accuracy was observed as number of nodes in the two hidden layers were increased. Which is explained by the fact that the number of nodes in the hidden layers defines the complexity and power of the neural network model to delineate underlaying relationships and structures inherent to the database. The number of nodes in the hidden layers should be large enough for the correct representation of the studied phenomena, but at same time low enough to have adequate generalizations. Several methods have been proposed to define the optimum number of hidden layer nodes. For instance, the approach proposed by Garson35 is often used to choose the number of hidden neurons and is calculated as follow: Np/[r(Ni + No)]; where Ni and No are input and output layer nodes, respectively, and the number of training samples is represented by Np. The Garson35 approach determines an optimal number of layers to our database in a range of 6 to 12 nodes, which is markedly lower than the best architecture tested in the current study (n = 30).
Recently, Madhiarasan and Deppa12 tested 151 different criteria of selection of the number of hidden neurons in back propagation networks using the RMSE, Mean Absolute Error (MAE), Mean Relative Error (MRE) and Mean Square Error (MSE) as error criteria to choose the optimal neuron network architecture. These authors also proposed a new approach when selecting the number of hidden neurons: = (8n − 4)/(n − 4) where n is the number of input parameters. The use of this new approach allows selecting the model of ANN with the lowest error value of forecasting and easy implementation. For the current database, the computed number of hidden neurons using the Madhiarasan and Deppa12 approach was 36, which is similar to the best architecture tested in the current study.
Therefore, we considered that an ANN architecture with 2 layers and 15 neurons in each one is the most appropriate for our database structure as it provides the best estimations without affecting its generalization capacity. Our best architecture had the highest number of hidden neurons, which is in accordance with Ince and Sofu23, who referred that a bigger network size would contribute to lower error levels in the training set. However, several authors referred that a more complex ANN architecture not necessarily represents an optimal solution for a given problem15. For this reason, the definition of the most appropriate architecture is the main step of the ANN implementation, which must considerer the context of research, database structure, type of variables, learning methods and objective of implementation (prediction or classification)15,23.
Overall, five udder measures were used to estimate the milk yield from Pelibuey sheep implementing two analytic approaches: multiple linear regression (MLR) and artificial neural networks (ANN). Both approaches were able to predict milk yield. However, the ANN showed better goodness compared with MLR. The accuracy of ANN to estimate milk yield depended on its neuron network architecture with increasing accuracy as the complexity of the ANN was increased. Therefore, the result of the current study reveals that ANN is a powerful tool to estimate milk yield when udder measures are used as an input variable and our data could be used or extrapolated for studies related to dairy ruminants. Further studies could use larger datasets and use them to perform different experiments including hyperparameter optimization by applying different approaches such as grid search, meta-heuristics, evolutionary methods and those based on nature-inspired algorithms.
Methods
Animal care, welfare and procedures were carried out according to the guidelines of the Animal Care Committee of the División Académica de Ciencias Agropecuarias, Universidad Juárez Autónoma de Tabasco under the approved ID project PFI: UJAT-DACA-2015-IA-02.
Database
A total of 357 milk yield records with its corresponding gland udder measures was used in the present study. The records of a total of 38 multiparous Pelibuey ewes were analyzed in the current study. Ewes had an average body weight of 36.3 ± 4.9 kg with single (n = 33) and twin lambing (n = 5). Details about the nutritional, health and lamb management are specified by Arcos-Álvares et al.2. Ewes were milking manually once a day and udder measures were recorded twice a week. Udder measures were taken before and after milking and those were udder circumference (circumference at the udder medium area), udder width (distance between the widest lateral points of the udder), udder height (distance from the udder insertion to the lower extremity) and udder volume were calculated as follows:
where V is the udder volume (cm3), R is radius (cm), CP is circumference perimeter (cm), \(\pi\) is 3.14159265358979 and Ud is the udder depth (cm).
Model training
Arcos-Álvares et al.2 using the same database of the current study to estimate milk yield through fit several multiple linear regression (MLR) models. To estimate daily milk yield (MY), these authors used a stepwise process to choose the best model using the following goodness of fit criteria: mean square prediction error (MSPE), root of MSPE (RMSPE) and coefficient of determination (r2). For that, the inputs were initial udder circumference (CI), final udder heigh (FH), initial udder width (IW), final udder width (FW), and the difference between the initial and final udder volumes (VDF) as input variables. Arcos-Álvares et al.2 used five models with the best goodness of fit following this equation:
where MY is the daily milk yield and the input variables (CI, FH, IW, FW and VDF) as were described above. The asterisks mean the level of significance of the regression coefficients: *p < 0.05; ***p < 0.000.
In order to contrast MLR versus ANN the same inputs were used to carried out machine learning essay. Normalization is very important to build ANN since the database has different range and different units. The proposed approaches use the min–max normalization technique using the formula proposed by Zeng and Yeung36:
where Xi is the real input data, X′min is the least input data, X′max is the greatest input data, X′min is the least target value, X′max is the greatest target values.
A supervised learning was used to train and teach the network using a two-layer ANN with seven hidden structures: 5–5, 5–10, 10–5, 10–10, 10–15, 15–10 and 15–15 neurons. Inputs vales are transferers to the hidden layer that multiplies weight W using tangent sigmoid activation function; the outputs of hidden layers get transferred to the output layers that multiplies with weight V using tangent sigmoid activation function12. A random 70% of the data was used to training and 30% for testing, the process was repeated 100 times. The globally convergent algorithm based on the resilient backpropagation was used to calculate ANN with a maximum step for training ANN of 1 × 107 using the neuralnet package in R software37.
Performance assessment
The fit of ANN was compared with the best multiple regression model (MRM) fitted previously by Arcos-Alvarez et al.2 using same input variables in both models. Goodness of fit was evaluated using:
Mean square of prediction error (MSPE), calculated as:
where et is the residual for the milk yield of t sheep, n is the number of observations in the training dataset and p is the number of parameters of models.
Root-mean-square prediction error (RMSPE):
Akaike’s Information Criterion (AIC) calculated as follows38:
where n is the number of examples in the training dataset, LL is the log-likelihood for the model using the natural logarithm (e.g., the log of the MSPE), and k is the number of parameters in the model.
Bayesian’s Information Criterion (BIC) calculated as follows39:
where n is the number of observations in the training dataset, LL is the log-likelihood for the model using the natural logarithm (e.g. log of the mean squared error), and k is the number of parameters in the model, and log() is the natural logarithm.
Accuracy of estimations was calculated as follow:
Ethics declaration
Animal care, welfare and procedures were carried out according to the guidelines of the Animal Care Committee of the División Académica de Ciencias Agropecuarias, Universidad Juárez Autónoma de Tabasco under the approved ID project PFI: UJAT-DACA-2015-IA-02.
Data availability
Data is available as supplementary material.
References
Angeles-Hernandez, J. C., Ortega, O. A. C., Perez, A. H. R. & Ronquillo, M. G. Effects of crossbreeding on milk production and composition in dairy sheep under organic management. Anim. Prod. Sci. 54(10), 1641–1645. https://doi.org/10.1071/AN14214 (2014).
Arcos-Álvarez, D. et al. Udder measurements and their relationship with milk yield in Pelibuey ewes. Animals 10(3), 518. https://doi.org/10.3390/ani10030518 (2020).
Pourlis, A. Ovine mammary morphology and associations with milk production, milkability and animal selection. Small Rumin. Res. 184, 106009. https://doi.org/10.1016/j.smallrumres.2019.10.010 (2020).
Ayadi, M. et al. Relationship between mammary morphology traits and milk yield of Sicilo-Sarde dairy sheep in Tunisia. Small Rumin. Res. 96(1), 41–45. https://doi.org/10.1016/j.smallrumres.2010.10.013 (2011).
Iñiguez, L., Hilali, M., Thomas, D. L. & Jesry, G. Udder measurements and milk production in two Awassi sheep genotypes and their crosses. J. Dairy Sci. 92(9), 4613–4620. https://doi.org/10.3168/jds.2008-1950 (2009).
Rovai, M., Caja, G. & Such, X. Evaluation of udder cisterns and effects on milk yield of dairy ewes. J. Dairy Sci. 91(12), 4622–4629. https://doi.org/10.3168/jds.2008-1298 (2008).
McKusick, B. C., Marnet, P. G., Berger, Y. M. & Thomas, D. L. Preliminary results: Effects of udder morphology on commercial milk production of East Friesian crossbreed ewes. Proceedings of the 5th Great Lakes Dairy Sheep Symposium. November 4–6, 1999, Brattleboro, Vermont, USA (1999).
Van der Linden, D. S. et al. Comparison of four techniques to estimate milk production in singleton-rearing non-dairy ewes. Small Rumin. Res. 90(1–3), 18–26. https://doi.org/10.1016/j.smallrumres.2009.12.007 (2010).
Emediato, R. M. S., Siqueira, E. R. D., Stradiotto, M. M., Maestá, S. A. & Fernandes, S. Relationship between udder measurements and milk yield in Bergamasca ewes in Brazil. Small Rumin. Res. 75(2–3), 232–235. https://doi.org/10.1016/j.smallrumres.2007.11.006 (2008).
Espinosa-Mendoza, R. I. et al. Predicting milk yield in Pelibuey ewes from the udder volume measurement with a simple method. J. Dairy Res. 87(3), 341–343. https://doi.org/10.1017/S002202992000076X (2020).
Bakoev, S. et al. PigLeg: Prediction of swine phenotype using machine learning. PeerJ 8, e8764. https://doi.org/10.7717/peerj.8764 (2020).
Madhiarasan, M. & Deepa, S. N. A novel criterion to select hidden neuron numbers in improved back propagation networks for wind speed forecasting. Appl. Intel. 44(4), 878–893. https://doi.org/10.1007/s10489-015-0737-z (2016).
Li, M. M., Sengupta, S. & Hanigan, M. D. Using artificial neural networks to predict pH, ammonia, and volatile fatty acid concentrations in the rumen. J. Dairy Sci. 102(10), 8850–8861. https://doi.org/10.3168/jds.2018-15964 (2019).
Cravener, T. L. & Roush, W. B. Prediction of amino acid profiles in feed ingredients: Genetic algorithm calibration of artificial neural networks. Anim. Feed Sci. Technol. 90(3–4), 131–141. https://doi.org/10.1016/S0377-8401(01)00219-X (2001).
Grzesiak, W., Lacroix, R., Wójcik, J. & Blaszczyk, P. A comparison of neural network and multiple regression predictions for 305-day lactation yield using partial lactation records. Can. J. Anim. Sci. 83(2), 307–310. https://doi.org/10.4141/A02-002 (2003).
Usman, S. M., Singh, N. P., Dutt, T., Tiwari, R. & Kumar, A. Comparative study of artificial neural network algorithms performance for prediction of FL305DMY in crossbred cattle. J. Entomol. Zool. Stud. 8, 516–520. https://doi.org/10.22271/j.ento (2020).
Montout, A. X., Bhamber, R. S., Lange, D. S., Ndlovu, D. Z., Morgan, E. R., Ioannou, C. C. & Dowsey, A. W. Accurate and interpretable prediction of poor health in small ruminants with accelerometers and machine learning. bioRxiv. https://doi.org/10.1101/2020.08.03.234203 (2020).
Ehret, A., Hochstuhl, D., Gianola, D. & Thaller, G. Application of neural networks with back-propagation to genome-enabled prediction of complex traits in Holstein-Friesian and German Fleckvieh cattle. Genet. Sel. Evol. 47(1), 1–9. https://doi.org/10.1186/s12711-015-0097-5 (2015).
Hernández-Ramos, P. A., Vivar-Quintana, A. M. & Revilla, I. Estimation of somatic cell count levels of hard cheeses using physicochemical composition and artificial neural networks. J. Dairy Sci. 102(2), 1014–1024. https://doi.org/10.3168/jds.2018-14787 (2019).
Murata, N., Yoshizawa, S. & Amari, S. I. Network information criterion-determining the number of hidden units for an artificial neural network model. IEEE Trans. Neural Netw. 5(6), 865–872. https://doi.org/10.1109/72.329683 (1994).
Bhosale, M. D. & Singh, T. P. Comparative study of feed-forward neuro-computing with multiple linear regression model for milk yield prediction in dairy cattle. Curr. Sci. 108, 2257–2261 (2015).
Liseune, A., Salamone, M., Van den Poel, D., Van Ranst, B. & Hostens, M. Predicting the milk yield curve of dairy cows in the subsequent lactation period using deep learning. Comput. Electron. Agric. 180, 105904. https://doi.org/10.1016/j.compag.2020.105904 (2021).
Ince, D. & Sofu, A. Estimation of lactation milk yield of Awassi sheep with artificial neural network modeling. Small Rumin. Res. 113(1), 15–19. https://doi.org/10.1016/j.smallrumres.2013.01.013 (2013).
Fernández, C. et al. Weekly milk prediction on dairy goats using neural networks. Neural. Comput. Appl. 16(4), 373–381. https://doi.org/10.1007/s00521-006-0061-y (2007).
Kaygisiz, F. & Sezgin, F. H. Forecasting goat milk production in Turkey using Artificial Neural Networks and Box-Jenkins models. Anim. Rev. 4(3), 45–52. https://doi.org/10.18488/journal.ar.2017.43.45.52 (2017).
Singh, N. P. et al. Comparative analysis of artificial neural network algorithms for prediction of FL305DMY in Murrah Buffalo. Int. J. Livest. Res. 10(9), 205–209. https://doi.org/10.5455/ijlr.20200704062936 (2020).
Ankinakatte, S., Norberg, E., Løvendahl, P., Edwards, D. & Højsgaard, S. Predicting mastitis in dairy cows using neural networks and generalized additive models: A comparison. Comput. Electron. Agric. 99, 1–6. https://doi.org/10.1016/j.compag.2013.08.024 (2013).
Norouzian, M. A., Bayatani, H. & Alavijeh, M. V. Comparison of artificial neural networks and multiple linear regression for prediction of dairy cow locomotion score. Vet. Res. Forum 12, 33. https://doi.org/10.30466/vrf.2019.98275.2346 (2021).
Khorshidi-Jalali, M., Mohammadabadi, M., Koshkooieh, A. E., Barazandeh, A. & Babenko, O. Comparison of artificial neural network and regression models for prediction of body weight in Raini Cashmere goat. Iran. J. Appl. Anim. Sci. 9, 453–461 (2019).
Akkol, S., Akilli, A. & Cemal, I. Comparison of artificial neural network and multiple linear regression for prediction of live weight in hair goats. Yyu J. Agric. Sci 27, 21–29 (2017).
Adebayo, R. A., Moyo, M., Gueguim-Kana, E. B. & Nsahlai, I. V. The use of artificial neural networks for modelling rumen fill. Can. J. Anim. Sci. 101, 427–437. https://doi.org/10.1139/cjas-2019-0101 (2020).
Ekiz, B., Baygul, O., Yalcintan, H. & Ozcan, M. Comparison of the decision tree, artificial neural network and multiple regression methods for prediction of carcass tissues composition of goat kids. Meat Sci. 161, 108011. https://doi.org/10.1016/j.meatsci.2019.108011 (2020).
Dallago, G. M. et al. Predicting first test day milk yield of dairy heifers. Comput. Electron. Agric. 166, 105032. https://doi.org/10.1016/j.compag.2019.105032 (2019).
Nobari, K., Baneh, H., Esmaeilkhanian, S., Yussefi, K. & Samiei, R. Comparison of linear model and artificial neural network to prediction of milk yield using first recorded parity. J. Rumin. Res. 6(4), 89–100 (2019).
Garson, G. D. Neural Networks: An Introductory Guide for Social Scientists (Sage, 1998).
Zeng, X. & Yeung, D. S. Hidden neuron pruning of multilayer perceptrons using a quantified sensitivity measure. Neurocomputing 69, 825–837. https://doi.org/10.1016/j.neucom.2005.04.010 (2006).
Fritsch S., Guenther F. & Wright M. N. (2019). neuralnet: Training of Neural Networks. R package version 1.44.2. https://CRAN.R-project.org/package=neuralnet
Akaike, H. A new look at the statistical model identification. EEE Trans. Autom. Control 19, 716–723. https://doi.org/10.1109/TAC.1974.1100705 (1974).
Bhat, H. S. & Kumar, N. On the derivation of the bayesian information criterion. School Nat. Sci. Univ. California 99, 1–4 (2010).
Acknowledgements
J.C. Angeles-Hernandez grateful to Mexican Ministry of Education and the Autonomous University of Hidalgo for the grant “Apoyo de Fomento a la Permanencia Institucional”.
Funding
The current work was funding for the Project UAEH-PTC-823 of the Mexican Ministry of Education.
Author information
Authors and Affiliations
Contributions
J.C.A.H., A.J.C.C. and E.V.B.P. conceived of the presented idea and planned the experiments. J.C.A.H. and F.A.C.E. performed the computational procedures. J.C.A.H. and E.V.B.P. wrote the manuscript with support from A.J.C.C., F.A.C.E., A.P.A. and J.A.S.M. J.C.A.H. and E.V.B.P. supervised the project. All authors approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Angeles-Hernandez, J.C., Castro-Espinoza, F.A., Peláez-Acero, A. et al. Estimation of milk yield based on udder measures of Pelibuey sheep using artificial neural networks. Sci Rep 12, 9009 (2022). https://doi.org/10.1038/s41598-022-12868-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-12868-0
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.