Abstract
Only up to 25% of the cases in which there is a familial aggregation of breast and/or ovarian cancer are explained by germline mutations in the well-known BRCA1 and BRCA2 high-risk genes. Recently, the BRCA1-associated ring domain (BARD1), that partners BRCA1 in DNA repair, has been confirmed as a moderate-risk breast cancer susceptibility gene. Taking advantage of next-generation sequencing techniques, and with the purpose of defining the whole spectrum of possible pathogenic variants (PVs) in this gene, here we have performed a comprehensive mutational analysis of BARD1 in a cohort of 1946 Spanish patients who fulfilled criteria to be tested for germline pathogenic mutations in BRCA1 and BRCA2. We identified 22 different rare germline variants, being 5 of them clearly pathogenic or likely pathogenic large deletions, which account for 0.26% of the patients tested. Our results show that the prevalence and spectrum of mutations in the BARD1 gene might vary between different regions of Spain and expose the relevance to test for copy number variations.
Similar content being viewed by others
Introduction
Breast cancer (BC) is the most common malignancy in women and the second cause of death due to cancer in this sex in the world1. BRCA1 and BRCA2 are the main high-risk susceptibility genes responsible for Hereditary Breast and Ovarian Cancer (HBOC), however their germline mutations only explain up to 25% of the cases in which a familial aggregation of these diseases is observed. Other moderate-high susceptibility genes have been identified in recent years, whose association with breast and/or ovarian cancer susceptiblity has been accurately established, thanks to advances in next-generation sequencing (NGS) technologies2,3. Recently, the two largest BC case–control studies published so far, have confirmed those genes that have an actual association with the disease, and therefore should be included in diagnostic testing, in addition to BRCA1/2. These genes are ATM, BARD1, CDH1, CHEK2, PALB2, RAD51C, RAD51D and TP534,5. All of those were already considered clinically relevant6,7,8, except BARD1, whose role in BC susceptibility had not been so clearly established until now.
The BRCA1-associated ring domain (BARD1) gene encodes a 777 amino acid protein which was discovered in 1996 to directly interact with BRCA1, through their homologous N-terminal RING domains, to form a stable complex9. BARD1 also contains two tandem conserved C-terminal BRCT domains shared with BRCA1, and four tandem ankyrin repeats. The tumour suppressor complex BRCA1-BARD1 interacts with DNA and other DNA damage response factors in order to repair double-strand breaks, including damaged replication forks, via homologous recombination. Besides, the heterodimer acts as an efficient E3 ubiquitin ligase, targeting proteins involved in cell-cycle, genome stability and hormone signaling10,11,12,13. BARD1 has several BRCA1-independent functions, including the inhibition of mRNA maturation and the induction of apoptosis by means of p53 stabilization14. Furthermore, BARD1 spliced isoforms lacking regions of functional domains, have been found overexpressed in some breast and ovarian tumors in comparison with the full-length protein, and have shown oncogenic properties15.
The two studies by Dorling et al. and Hu et al. have definitely confirmed the role of BARD1 as a moderate BC susceptibility gene (OR: 2.09; 95% CI, 1.35–3.23; P = 0.00098, for protein-truncation variants4). These studies also report a stronger association of pathogenic variants (PVs) in BARD1 with ER-negative tumors and, more remarkably, with triple-negative breast cancer (TNBC).
Numerous studies have been performed in the last years aiming to define the prevalence of mutations in the BARD1 gene in different cohorts16. However, to our knowledge, none of them has taken in account the whole spectrum of possible pathogenic variants (PV) in the gene. Regarding the Spanish population, only one study has investigated the prevalence of BARD1 PVs in an important number of patients, focusing on the region of Catalonia17. With the purpose of continuing to deepen our knowledge of the gene in breast and ovarian cancer predisposition, here we report a comprehensive mutational analysis of the BARD1 gene in a cohort of 1946 patients who fulfilled criteria to be tested for pathogenic mutations in the BRCA1/2 genes18 from different regions of Spain.
Results
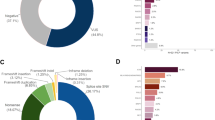
Among the 1946 patients screened using targeted NGS, we identified 22 different BARD1 rare germline variants, 5 Copy Number Variants (CNV), 11 missenses and 6 synonymous, distributed in 49 patients (5 carried a CNV, 26 a missense and 18 a synonymous variant). Strickingly, we did not find any frameshift, nonsense nor splicing variants. The position across the protein of the detected variants is represented in Fig. 1. Each patient carried one single rare variant, except for one case that presented the variants c.1339C > G; p.Leu447Val and c.1059C > G; p.Pro353 simultaneously. c.1977A > G; p.Arg659 = was found in 11 patients, representing the most recurrent variant in our population of study; and c.33G > T; p.Gln11His was the second most common appearing in 10 cases. When applying a Fisher exact test comparing the number of individuals harbouring each of these two variants in our series of cases vs. local controls, no statistically significant differences were found (p = 0.318 and 0.492 respectively), suggesting that these variants are not potentially associated with BC risk.
Map of the BARD1 protein domains showing the location of rare germline variants identified in our cohort. The graphic was generated with MutationMapper tool at cBioPortal for Cancer Genomics (https://cbioportal.org/mutation_mapper).
All variants found were classified according to their pathogenicity based on ACMG guidelines and taking into consideration ClinVar database (https://ncbi.nlm.nih.gov/clinvar/), literature search and Franklin Genoox platform (https://franklin.genoox.com/clinical-db/home) variant interpretation.
0.26% of our cohort of Spanish patients fulfilling criteria for BRCA1/2 testing present pathogenic/likely pathogenic variants in the BARD1 gene
Only the five CNVs detected in our series were considered as pathogenic or likely pathogenic following ACMG guidelines. CNVs were detected from NGS data, based on the depth of coverage which is correlated with the copy number of the region19. All of them were predicted to cause an out of frame RNA, predicted to derive in an incorrect translation of aminoacids that ends up in the appearance of a premature stop codon, involving the loss of an important functional domain of the protein (Fig. 1 and Supplementary Fig. S1). Each one of the five large deletions reported was present just once in our cohort, giving a prevalence of 0.00257 (5/1946), and were not present in control samples. For establishing PM2 (Pathogenic Moderate 2) criteria, (absent from controls in Exome Sequencing Project, 1000 Genomes or ExAC) we took in account structural variants (SVs) detected in BARD1 over more than 21,000 alleles from gnomAD SVs v2.1 dataset. We also took in account data from Fabulous Ladies Over Seventy (FLOSSIES, https://whi.color.com/) database, from which along 9884 noncancer controls, the only deletion reported (once) comprised exons 4–11 with an overall frequency of 0.0001012 in contrast to our 0.00257 prevalence of large deletions in the cases studied. As local controls, we included 1119 DNA samples from Spanish individuals provided by Spanish National Biobank (BNADN), that had been genotyped using Illumina Global Screening Array v1.0 + MD containing 700.078 SNPs. We made use of these data to analyse CNVs in the BARD1 gene, and no CNVs were detected.
Of note, out of the five CNVs detected, only deletion of exon 1 and deletion of the entire gene have been previously described20,21,22. In this study, we report for the first time the deletion of exons 10–11, deletion of exon 9 and deletion of exons 2–11.
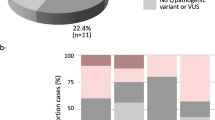
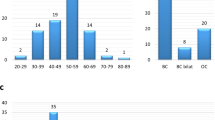
Carriers of these PVs were women diagnosed with BC, whose genotype and phenotype data are summarized in Table 1. A breakdown of the patients based on the risk criterion fulfilled, and percentage of BARD1 PVs found in each group are shown in Table 2.
Reclassification of rare BARD1 synonymous variants
Most of the synonymous variants detected in our population were previously described with conflicting interpretations of pathogenicity in ClinVar (Supplementary Table 1). In an effort to add more evidence to the current classification, we carried out a cDNA assay to elucidate their individual effect in splicing. With RNA derived from blood samples of carriers we performed a RT-PCR, followed by an amplification with specific primer pairs, ending up with Sanger sequencing. All the analyzed variants showed biallelic expression at the cDNA level (Supplementary Fig. S2), confirming that none of them caused an alteration in the splicing process, which agrees with benign strong 3 (BS3) criteria from ACMG guidelines. By applying the BS3 evidence we provide an updated classification to benign for all of them.
It is worth mentioning that the c.1977A > G silent change had been previously described to alter several exonic splicing enhancer (ESE) motifs causing the deletion of exons 2 to 9 and resulting in a frame-shift which created a premature stop codon (p.Cys53_Trp635delinsfsX12)23. On the contrary, for the only RNA that we had available, out of all the patients carrying this variant, we did not see this alteration (Supplementary Fig. S2). Moreover, this synonymous change resulted to be the rare variant appearing more times in our population of study, which in addition to the global allele frequency shown in GnomAD, support its neutrality (Supplementary Table 1).
Analysis of rare BARD1 missense variants
To our knowledge, this is the first study in which a comprehensive reporting of BARD1 missense variants is performed. Although we did not conduct functional assays, we performed an exhaustive review of the literature to try to add any piece of information that could contribute to their classification (Table 3).
p.Thr716Ala is classified as benign following ACMG guidelines, relying on its relatively high frequency in control populations extracted from independent databases. In our cohort, p.Thr716Ala was only present in one patient. Moreover, several published studies have designated p.Thr716Ala as deleterious, arguing that it abolishes binding to OLA1, a protein interacting in BRCA1-BARD1 complex, leading to the alteration of BARD1 tumor suppressor activity24,25,26. Taking this in account, we consider that it should be regarded as VUS.
A total of 105 BARD1 missense and truncation variants have been previously tested in a functional assay for homology-directed repair (HDR) alteration27,28. Of our detected missense variants, p.Asp612Val, p.Thr343Ile and p.Leu239Gln were described as functional, although for these two last variants a significantly increased loss of heterozygosity (LOH) has been reported. In addition, they recognized p.Thr716Ala as functional, while other studies had described this variant as deleterious as mentioned before. This suggests that HDR assay, although informative, might not be sufficient for classifying BARD1 variants.
Discussion
Familial BC is a polygenic disease that can be partially explained, by the effect of PVs distributed over several susceptibility genes with different levels of risk29. In this context, BARD1 has been recently confirmed as a moderate-risk BC gene by two large case–control studies, sorting out the uncertainity about its association with the disease4,5. With the aim of establishing the mutational spectrum of BARD1 in the Spanish population in order to improve genetic diagnosis, we screened the gene for PVs by a multigene panel analysis in a series of 1946 Spanish patients who fulfilled criteria to be tested for germline mutations in the BRCA1/2 genes. To our knowledge, this is the first study that comprehensively reports all types of rare variants in the BARD1 gene, subsequently discovering 5 large deletions, 11 missense and 6 synonymous from which only the large deletions were finally regarded as pathogenic or likely pathogenic. Surprisingly, no frameshift, nonsense nor splicing variants were found, which could be due to the limited size of our cohort or to populations or even regional differences in the spectrum of BARD1 mutations.
Previously, another study investigated the presence of germline PVs within the BARD1 gene in the Spanish population17. Among 4015 patients with clinical suspicion of HBOC, they identified 19 PVs: 8 truncating, 1 splicing and 2 large deletion, ending up with a PV prevalence of 0.47% which approximately doubles the 0.22% reported by Couch et al. and the 0.25% described by Suszynska and Kozlowski16,30, and triples the 0.13% of protein-truncating variants reported by Dorling et al. and the 0.15% identified by Hu et al.4,5. In our cohort, we found a prevalence of 0.26%, which is similar to previous studies, but lower than that reported in the other Spanish cohort. This could be explained by the lower number of patients studied here, or differences in the ascertainment of the cases. These differences in the frequency of mutations, could also be explained by the possibility of variation in the mutational spectrum of BARD1 between different geographical areas of Spain, as most of our patients proceeded from Madrid and Castilla la Mancha regions, while those reported in Rofes et al. proceeded mainly from Catalonia. These regional differences have been already observed for mutations in the BRCA1/2 genes, and detecting them is very relevant to improve genetic diagnosis31, however, larger studies performed in Spanish population would be necessary to confirm this hypothesis.
Germline CNVs are recognized as the genetic cause of diverse hereditary pathologies, even though their detection via NGS is still challenging32. Only a few studies have characterized the CNV landscape within the BARD1 gene, having reported a total of 8 different large deletions so far in BC or OC patients17,20,21,22,33,34,35. In our study, we have detected the deletion of exon 1 and the deletion of the entire gene previously reported20,21 and, what is more, we have identified for the first time exons 2–11, exon 9 and exons 10–11 deletions present in three BC cases. It is very important to highlight that any deletion can not be assumed to be pathogenic, and special caution should be taken with the ones that affect one exon and/or do not produce a frameshift. In our case, all the large deletions were predicted to cause an out of frame RNA, and classified as pathogenic or likely pathogenic by following the ACMG guidelines adaptation for single-gene copy number variants36.
In addition to clearly PVs, to our knowledge, this is the first study in which a comprehensive analysis of all missense and synonymous rare variants is performed. There are already several functional studies reporting that some of these rare variants could be pathogenic24,25,27,28. Unfortunately, we were not able to classify most of the BARD1 missense variants we identified, but we think it is relevant to include them in further screenings and perfom functional assays, as otherwise, the percentage of PVs could be underestimated. Regarding synonymous variants, it is important to carefully analyze the effect of apparently silent changes, taking into consideration that this type of variants are usually classified as (likely) benign, obviating their possible effect in splicing, which could turn out in a missclassification. For all the rare synonymous variants in our population, damaging effect in splicing was experimentally ruled out, which corresponds with BS3 criteria of ACMG guidelines that allowed us to classify them all as definitively benign. These includes the c.1977A > G variant, that had been previously suggested to be deleterious by altering splicing and causing the partial deletion of exons 2–9 of the gene23. In our hands, no alteration in splicing was seen for this variant (Supplementary Fig. S2). Due to this discrepancy, these findings should be confirmed. However, the fact that c.1977A > G was present in 11 patients from our cohort, together with a 0.002 global allele frequency described in GnomAD (including one homozygote individual) and an even higher frequency of 0.005 in local controls (Table 3), support our finding in favour of its neutrality. Actually, frequency in controls, exceed the expected for disorder, that is 0.001 for BARD1 (according to the calculations made by the Franklin Genoox platform). Taking this in account, the variant would meet the BS1 and BP7 criteria and, even in the absence of BS3, could be classified as likely benign.
Finally, many independent studies have associated germline PVs in BARD1 with high risk of TNBC compared to other BC subtypes4,5,17,37,38,39. This assumption turns out to be of great importance, given that these patients often show a more aggressive disease course, with higher rates of metastases, earlier age at diagnosis and recurrence, and worse survival rates40. In our study, only one out of the 5 BC patients who carried a PV, was diagnosed with TNBC, representing 0.4% of the 243 patients that were diagnosed with TNBC in our series, regardless of fulfilling any other criteria. While doubling the overall rate, the reduced size of the TNBC subset does not allow us to drive any conclusion in this regard.
In summary, here we report the second largest study on BARD1 performed in Spanish population, and the first one in which all possible rare variants are comprehensively reported. Our results show that the prevalence and spectrum of mutations might vary between different regions in Spain, and highlights the large deletions in BARD1 as important contributors to BC susceptibility in our population. Larger studies and incorporation of independent functional assays are needed in order to properly classify each variant by its clinical impact, which is essential to determine the degree of susceptibility to the disease for each carrier.
Methods
Patients
A total of 1946 patients who fulfilled criteria to be tested for germline mutations in the BRCA1/2 genes were recruited from the Spanish National Cancer Center (CNIO), Hospital General La Mancha Centro, Complejo Hospitalario Universitario de Albacete and Hospital General Universitario de Ciudad Real. Families fulfilled one or more of the following criteria: (i) one BC diagnosed at < 40 years; (ii) ≥ 2 first-degree relatives diagnosed with BC at least one of them at < 50 years; (iii) one TNBC diagnosed at < 60 years; (iv) ≥ 3 first-degree BC relatives; (v) ≥ 1 male BC case; (vi) ≥ 1 ovarian cancer case; (vii) BC not fulfilling i-vi criteria but being bilateral or having any antecedent of pancreatic or prostate cancer (Table 2). Patients included in the study signed an appropriate informed constent and the proposal was approved by the ethics committee at the Fuenlabrada University Hospital, Madrid, Spain. All methods were carried out in accordance with relevant guidelines and regulations.
Controls
For frequency comparison of single nucleotide change variants detected in our patients, we used data from 125,748 exome sequences and 15,708 whole-genome sequences from unrelated individuals from gnomAD (https://gnomad.broadinstitute.org/) and 2094 Spanish unrelated individuals from the Collaborative Spanish Variant Server (http://csvs.clinbioinfosspa.es/).
For CNV classification, we used data from more than 21,000 alleles from gnomAD SVs v2.1 dataset, 9884 noncancer controls from Fabulous Ladies Over Seventy (FLOSSIES, https://whi.color.com/). We also included a total of 1119 DNA samples from Spanish individuals provided by Spanish National Biobank (BNADN). DNA samples were genotyped using Illumina Global Screening Array v1.0 + MD containing 700.078 SNPs according to the manufacturer's protocols and scanned using a iScan system. GenomeStudio software v2.0.4 (Illumina) was used for genotype calling. Before CNV analysis, BARD1 region was carefully inspected to ensure enough probe coverage of the gene, resulting in 59 probes thorugh all the gene. CNVs were detected and called using PennCNV v1.0.5 software from genotype, Log R Ratio (LRR) and B Allele Frequency (BAF) data values41. BARD1 gene locus (Chr 2: 215,590,370–215,674,428 GRCh37) and an additional 100 KB up and downstream, were inspected to identify CNVs in this region.
DNA isolation
Genomic DNA was extracted from peripheral blood samples using Maxwell RSC automated instrument (Promega) according to the manufacturer’s protocol. Purified DNA was quantitated making use of Quant-iT PicoGreen dsDNA reagent (Invitrogen).
Next-generation sequencing (NGS)
BARD1 was analyzed within a larger NGS panel, Onco-GeneSGKit (Sistemas Genómicos). Libraries were prepared employing capture-based target enrichment covering 111 genes involved in familial cancer (ACD, AIP, APC, ATM, ATR, AXIN2, BAP1, BARD1, BLM, BMPR1A, BRCA1, BRCA2, BRIP1, BUB1, CDH1, CDK4, CDKN1B, CDKN2A, CHEK2, DDB2, DICER1, DIS3L2, DKC1, ELANE, EPAS1, EPCAM, ERCC1, ERCC2, ERCC3, ERCC4, ERCC5, ERCC6, FANCA, FANCB, FANCC, FANCD2, FANCE, FANCF, FANCG, FANCI, FANCL, FANCM, FH, FLCN, G6PC3, GFI1, GREM1, HOXB13, JAGN1, KIF1B, KIT, MAX, MDH2, MEN1, MET, MITF, MLH1, MNX1, MSH2, MSH6, MSR1, MUTYH, NBN, NF2, NFIX, NHP2, NOP10, NSD1, NTHL1, PALB2, PARN, PDGFRA, PMS1, POLD1, POLE, POLH, POT1, PRKAR1A, PTCH1, PTEN, RAD50, RAD51, RAD51C, RAD51D, RB1, RECQL, RECQL4, RET, RTEL1, SCG5, SLX4, SMAD4, SMARCA4, STK11, SUFU, TERC, TERT, TINF2, TMEM127, TP53, TSC1, TSC2, UBE2T, VHL, VPS45, WAS, WRN, WT1, XPA, XPC, XRCC2). Briefly, 50 ng of total gDNA from cases was tagmented and amplified. Quality and quantity assessment of the resulting libraries was performed on a 2100 Bioanalyzer system (Agilent). Libraries were hybridized with capture probes to targeted regions, followed by hybrid capture with streptavidin-coated magnetic beads. Finally, libraries were indexed by PCR and combined in a single run before sequencing on a MiSeq platform (Illumina). The average read depth per sample was 200X, and, on average, 99.9% of the tardeted regions had a coverage of at least 20X.
Variant calling and classification
FastQ files generated were uploaded in GeneSystems platform (Sistemas Genómicos) to perform variant annotation. GenBank reference sequence NM_000465.3, with numbering starting at the A of the first ATG, was used for BARD1 variant nomenclature following the Human Genome Variation Society (HGVS) guidelines (http://varnomen.hgvs.org/). All coding exons and at least + /− 25 intronic nucleotides flanking each exon were analysed. Variants showing a minor allele frequency (MAF) < 0.01 in every subpopulation of the Genome Aggregation Database v2.1.1 (gnomAD, https://gnomad.broadinstitute.org/) were considered as “rare” and selected for further analysis.
Sequence variants identified in this work were grouped by their clinical significance according to the American College of Medical Genetics and Genomics (ACMG) standards and guidelines as pathogenic, likely pathogenic, uncertain significance (VUS), benign, or likely benign42. In the case of large deletions, the adaptation of this classification to single-gene copy number variants was applied36.
Copy number variation (CNV) analysis
Gain/loss of large DNA segments in the BARD1 gene were predicted by the CNid method developed by Sistemas Genómicos based on a read depth (RD) approach. Briefly, this approach is based on the comparison of the tested sample with a pool of normal samples, under the hypothesis that depth of coverage in a genomic region is correlated with the copy number of the region19. Identified CNVs were confirmed by Multiplex Ligation-dependent Probe Amplification (MLPA). The assay was performed using the MLPA Probemix P489 BARD1 (MRC-Holland) according to manufacturer’s instructions. MLPA results were analyzed with Peak Scanner Software (Thermo Fisher Scientific). For MLPA analysis, each sample harbouring a candidate CNVs in the BARD1 gene, was compared with five reference samples. Data were analysed and visualized using the Coffalyser.net software (MRC-Holland) (Supplementary Fig. 1). For the sample harbouring the single exon 9 deletion, sanger sequencing of the genomic region close to the ligation site of the half-probes (underlined in red) specific for exon 9, was performed to confirm the absence of single nucleotide polymorphisms that could have caused a decrease of the fluorescence signal (Supplemetary Fig. 1).
Splicing studies
Synonymous variants were tested for its possible effect in splicing. Starting with RNA extracted from blood samples of the patients, a RT-PCR was performed and the cDNA obtained was amplified with specific primer pairs targeting the exons of interest (available upon request). PCR products were sequenced with the Sanger method using the Big Dye Terminator Cycle sequencing kit on an ABI 3730 Sequencer (Applied Biosystems). Results were visualized in FinchTV software.
Data availability
All variants reported in this study have been submitted to https://databases.lovd.nl/shared/genes/BARD1 with accession numbers 0000841043, 0000841044, 0000841045, 0000841046, 0000841047, 0000841048, 0000841050, 0000841051, 0000841052, 0000841053, 0000841054, 0000841055, 0000841058, 0000841059, 0000841060, 0000841061, 0000842164, 0000842165, 0000842166, 0000842167, 0000842168, 0000842169. All primer sequences used for Sanger validation of the variants are available upon request.
References
Fahad Ullah, M. Breast cancer: Current perspectives on the disease status. Adv. Exp. Med. Biol. 1152, 51–64. https://doi.org/10.1007/978-3-030-20301-6_4 (2019).
Nielsen, F. C., Van Overeem Hansen, T. & Sørensen, C. S. Hereditary breast and ovarian cancer: New genes in confined pathways. Nat. Rev. Cancer 16, 599–612. https://doi.org/10.1038/nrc.2016.72 (2016).
Yoshida, R. Hereditary breast and ovarian cancer (HBOC): Review of its molecular characteristics, screening, treatment, and prognosis. Breast Cancer https://doi.org/10.1007/s12282-020-01148-2 (2020).
Dorling, L. et al. Breast cancer risk genes—Association analysis in more than 113,000 women. N. Engl. J. Med. 384, 428–439. https://doi.org/10.1056/NEJMoa1913948 (2021).
Hu, C. et al. A population-based study of genes previously implicated in breast cancer. N. Engl. J. Med. 384, 440–451. https://doi.org/10.1056/NEJMoa2005936 (2021).
Easton, D. F. et al. Gene-panel sequencing and the prediction of breast-cancer risk. N. Engl. J. Med. 372, 2243–2257. https://doi.org/10.1056/NEJMsr1501341 (2015).
Tung, N. et al. Counselling framework for moderate-penetrance cancer-susceptibility mutations. Nat. Rev. Clin. Oncol. 13, 581–588. https://doi.org/10.1038/nrclinonc.2016.90 (2016).
Yang, X. et al. Ovarian and breast cancer risks associated with pathogenic variants in RAD51C and RAD51D. J. Natl. Cancer Inst. 112, 1242–1250. https://doi.org/10.1093/jnci/djaa030 (2020).
Wu, L. C. et al. Identification of a RING protein that can interact in vivo with the BRCA1 gene product. Nat. Genet. 14, 430–440. https://doi.org/10.1038/ng1296-430 (1996).
Cimmino, F., Formicola, D. & Capasso, M. Dualistic role of BARD1 in cancer. Genes (Basel) https://doi.org/10.3390/genes8120375 (2017).
Stewart, M. D. et al. BARD1 is necessary for ubiquitylation of nucleosomal histone H2A and for transcriptional regulation of estrogen metabolism genes. Proc. Natl. Acad. Sci. U. S. A. 115, 1316–1321. https://doi.org/10.1073/pnas.1715467115 (2018).
Tarsounas, M. & Sung, P. The antitumorigenic roles of BRCA1-BARD1 in DNA repair and replication. Nat. Rev. Mol. Cell Biol. 21, 284–299. https://doi.org/10.1038/s41580-020-0218-z (2020).
Li, W. et al. A synergetic effect of BARD1 mutations on tumorigenesis. Nat. Commun. 12, 1243. https://doi.org/10.1038/s41467-021-21519-3 (2021).
Irminger-Finger, I., Ratajska, M. & Pilyugin, M. New concepts on BARD1: Regulator of BRCA pathways and beyond. Int. J. Biochem. Cell Biol. 72, 1–17. https://doi.org/10.1016/j.biocel.2015.12.008 (2016).
Li, L. et al. Oncogenic BARD1 isoforms expressed in gynecological cancers. Cancer Res. 67, 11876–11885. https://doi.org/10.1158/0008-5472.Can-07-2370 (2007).
Suszynska, M. & Kozlowski, P. Summary of BARD1 mutations and precise estimation of breast and ovarian cancer risks associated with the mutations. Genes (Basel) https://doi.org/10.3390/genes11070798 (2020).
Rofes, P. et al. BARD1 pathogenic variants are associated with triple-negative breast cancer in a Spanish hereditary breast and ovarian cancer cohort. Genes (Basel) https://doi.org/10.3390/genes12020150 (2021).
Gonzalez-Santiago, S. et al. SEOM clinical guidelines in hereditary breast and ovarian cancer (2019). Clin. Transl. Oncol. 22, 193–200. https://doi.org/10.1007/s12094-019-02262-0 (2020).
Teo, S. M., Pawitan, Y., Ku, C. S., Chia, K. S. & Salim, A. Statistical challenges associated with detecting copy number variations with next-generation sequencing. Bioinformatics 28, 2711–2718. https://doi.org/10.1093/bioinformatics/bts535 (2012).
Tung, N. et al. Frequency of mutations in individuals with breast cancer referred for BRCA1 and BRCA2 testing using next-generation sequencing with a 25-gene panel. Cancer 121, 25–33. https://doi.org/10.1002/cncr.29010 (2015).
Kwong, A. et al. Germline mutation in 1338 BRCA-negative Chinese hereditary breast and/or ovarian cancer patients: Clinical testing with a multigene test panel. J. Mol. Diagn. 22, 544–554. https://doi.org/10.1016/j.jmoldx.2020.01.013 (2020).
Carter, N. J. et al. Germline pathogenic variants identified in women with ovarian tumors. Gynecol. Oncol. 151, 481–488. https://doi.org/10.1016/j.ygyno.2018.09.030 (2018).
Ratajska, M. et al. Cancer predisposing BARD1 mutations in breast-ovarian cancer families. Breast Cancer Res. Treat. 131, 89–97. https://doi.org/10.1007/s10549-011-1403-8 (2012).
Sauer, M. K. & Andrulis, I. L. Identification and characterization of missense alterations in the BRCA1 associated RING domain (BARD1) gene in breast and ovarian cancer. J. Med. Genet. 42, 633–638. https://doi.org/10.1136/jmg.2004.030049 (2005).
Laufer, M. et al. Structural requirements for the BARD1 tumor suppressor in chromosomal stability and homology-directed DNA repair. J. Biol. Chem. 282, 34325–34333. https://doi.org/10.1074/jbc.M705198200 (2007).
Otsuka, K., Yoshino, Y., Qi, H. & Chiba, N. The function of BARD1 in centrosome regulation in cooperation with BRCA1/OLA1/RACK1. Genes (Basel) https://doi.org/10.3390/genes11080842 (2020).
Lee, C. et al. Functional analysis of BARD1 missense variants in homology-directed repair of DNA double strand breaks. Hum. Mutat. 36, 1205–1214. https://doi.org/10.1002/humu.22902 (2015).
Adamovich, A. I. et al. Functional analysis of BARD1 missense variants in homology-directed repair and damage sensitivity. PLoS Genet. 15, e1008049. https://doi.org/10.1371/journal.pgen.1008049 (2019).
Wendt, C. & Margolin, S. Identifying breast cancer susceptibility genes—a review of the genetic background in familial breast cancer. Acta Oncol. 58, 135–146. https://doi.org/10.1080/0284186x.2018.1529428 (2019).
Couch, F. J. et al. Associations between cancer predisposition testing panel genes and breast cancer. JAMA Oncol. 3, 1190–1196. https://doi.org/10.1001/jamaoncol.2017.0424 (2017).
Diez, O., Gutiérrez-Enríquez, S. & Balmaña, J. Heterogeneous prevalence of recurrent BRCA1 and BRCA2 mutations in Spain according to the geographical area: Implications for genetic testing. Fam. Cancer 9, 187–191. https://doi.org/10.1007/s10689-009-9301-5 (2010).
Moreno-Cabrera, J. M. et al. Evaluation of CNV detection tools for NGS panel data in genetic diagnostics. Eur. J. Hum. Genet. 28, 1645–1655. https://doi.org/10.1038/s41431-020-0675-z (2020).
Adedokun, B. et al. Prevalence of inherited mutations in breast cancer predisposition genes among women in Uganda and Cameroon. Cancer Epidemiol. Biomarkers Prev. 29, 359–367. https://doi.org/10.1158/1055-9965.Epi-19-0506 (2020).
Kaneyasu, T. et al. Prevalence of disease-causing genes in Japanese patients with BRCA1/2-wildtype hereditary breast and ovarian cancer syndrome. NPJ Breast Cancer 6, 25. https://doi.org/10.1038/s41523-020-0163-1 (2020).
Zeng, C. et al. Evaluation of pathogenetic mutations in breast cancer predisposition genes in population-based studies conducted among Chinese women. Breast Cancer Res. Treat. 181, 465–473. https://doi.org/10.1007/s10549-020-05643-0 (2020).
Brandt, T. et al. Adapting ACMG/AMP sequence variant classification guidelines for single-gene copy number variants. Genet. Med. 22, 336–344. https://doi.org/10.1038/s41436-019-0655-2 (2020).
De Brakeleer, S. et al. Frequent incidence of BARD1-truncating mutations in germline DNA from triple-negative breast cancer patients. Clin. Genet. 89, 336–340. https://doi.org/10.1111/cge.12620 (2016).
Buys, S. S. et al. A study of over 35,000 women with breast cancer tested with a 25-gene panel of hereditary cancer genes. Cancer 123, 1721–1730. https://doi.org/10.1002/cncr.30498 (2017).
Shimelis, H. et al. Triple-negative breast cancer risk genes identified by multigene hereditary cancer panel testing. J. Natl. Cancer Inst. 110, 855–862. https://doi.org/10.1093/jnci/djy106 (2018).
Afghahi, A., Telli, M. L. & Kurian, A. W. Genetics of triple-negative breast cancer: Implications for patient care. Curr. Probl. Cancer 40, 130–140. https://doi.org/10.1016/j.currproblcancer.2016.09.007 (2016).
Wang, K. et al. PennCNV: An integrated hidden Markov model designed for high-resolution copy number variation detection in whole-genome SNP genotyping data. Genome Res. 17, 1665–1674. https://doi.org/10.1101/gr.6861907 (2007).
Richards, S. et al. Standards and guidelines for the interpretation of sequence variants: A joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 17, 405–424. https://doi.org/10.1038/gim.2015.30 (2015).
Acknowledgements
We thank all patients and their families that participated in this study. We thank all members of the Familial Cancer Unit and former members of the Human Cancer Genetics Program of the Spanish National Cancer Research Centre for all their support. The genotyping was performed at the Spanish National Cancer Research Centre, in the Human Genotyping lab, a member of CeGen, PRB3 and is supported by grant PT17/0019, of the PE I+D+i 2013-2016, funded by ISCIII and ERDF.
Funding
A.O. is partially funded by FIS PI19/00640 supported by FEDER funds and the Spanish Network on Rare Diseases (CIBERER).
Author information
Authors and Affiliations
Contributions
Study conception and design: A.O. Acquisition of data: B.B-S., A.B., F.V., F.M, R.N-T., G.P., L.P. R.M-Ch., JM.C-C., M.U., A.O. Analysis and interpretation of data: B.B-S., A.B., V.F., F.M, R.N-T., G.P., A.G-N., A.O. Drafting of the manuscript: B.B-S., A.O.. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Benito-Sánchez, B., Barroso, A., Fernández, V. et al. Apparent regional differences in the spectrum of BARD1 pathogenic variants in Spanish population and importance of copy number variants. Sci Rep 12, 8547 (2022). https://doi.org/10.1038/s41598-022-12480-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-12480-2
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.