Abstract
In potentially severe diseases in general and COVID-19 in particular, it is vital to early identify those patients who are going to progress to severe disease. A recent living systematic review dedicated to predictive models in COVID-19, critically appraises 145 models, 8 of them focused on prediction of severe disease and 23 on mortality. Unfortunately, in all 145 models, they found a risk of bias significant enough to finally "not recommend any for clinical use". Authors suggest concentrating on avoiding biases in sampling and prioritising the study of already identified predictive factors, rather than the identification of new ones that are often dependent on the database. Our objective is to develop a model to predict which patients with COVID-19 pneumonia are at high risk of developing severe illness or dying, using basic and validated clinical tools. We studied a prospective cohort of consecutive patients admitted in a teaching hospital during the “first wave” of the COVID-19 pandemic. Follow-up to discharge from hospital. Multiple logistic regression selecting variables according to clinical and statistical criteria. 404 consecutive patients were evaluated, 392 (97%) completed follow-up. Mean age was 61 years; 59% were men. The median burden of comorbidity was 2 points in the Age-adjusted Charlson Comorbidity Index, CRB was abnormal in 18% of patients and basal oxygen saturation on admission lower than 90% in 18%. A model composed of Age-adjusted Charlson Comorbidity Index, CRB score and basal oxygen saturation can predict unfavorable evolution or death with an area under the ROC curve of 0.85 (95% CI 0.80–0.89), and 0.90 (95% CI 0.86 to 0.94), respectively. Prognosis of COVID-19 pneumonia can be predicted without laboratory tests using two classic clinical tools and a pocket pulse oximeter.
Similar content being viewed by others
Introduction
In potentially severe diseases in general, and COVID-19 in particular, it is vital to early identify those patients who are going to progress to severe disease. Physicians often care for patients who present a clearly mild or severe profile and do not need to calculate a predictive index to make decisions. However, in other cases, it is not easy to anticipate the clinical course, and that is when a prediction rule can be helpful, e.g., in a patient with mild to moderate acute symptoms, but with a weak baseline situation; or in a patient who does have striking acute symptoms but is young and healthy. It can also be helpful for healthcare managers when need to quantify the healthcare demand that it is going to be faced and prepare the necessary resources in advance. Finally, a suitable predictive rule would be useful as a quality control tool for both, clinical physicians, and healthcare managers1.
Motivated by the urgent need to characterise COVID-19 there has been a blast of publications (more than 80,000 up to early December 2020); the symptoms and initial characteristics of the disease are well known, but the determinants of its course are less clear. A living systematic review dedicated to predictive models in COVID-191, in its latest version (search updated May 5), has found 145 models, 8 of them focused on prediction of severe disease and 23 on mortality. Unfortunately, in all 145 models, they found a risk of bias significant enough to finally "not recommend any for clinical use". The most frequent bias issues referred to the analysis; however, the most serious ones were those related to sampling. Authors recommend concentrating on avoiding biases in sampling and prioritising the study of already identified predictive factors, rather than the identification of new ones that are often dependent on the database. Our objective is to develop a model to predict which patients with COVID-19 pneumonia are at high risk of developing severe illness, using readily available clinical data in the absence of laboratory or sophisticated computing/artificial intelligence.
Methods
Study design
Prospective cohort study, formed by all patients consecutively admitted to the Hospital Universitario Virgen de la Victoria (HUVV) with COVID-19 pneumonia, during the first wave: March 1 to April 28, 2020. Follow-up lasted until the discharge of the last patient: July 21, 2020. HUVV is a 506-bed hospital, classified as level 2, located in Málaga (southern Spain), which directly serves a population of 470,000 inhabitants.
Participants and source of data
The inclusion criteria were: confirmed, symptomatic SARS-CoV-2 infection; and requiring hospital admission. Exclusion criteria were: age under 14 years. When the patient had consulted several times at the Emergency Department, data was collected from the consultation in which the acute infection by SARS-CoV-2 was diagnosed.
SARS-CoV-2 infection was confirmed by real-time reverse transcription polymerase chain reaction (RT-PCR), or detection of IgM antibodies with enzyme immunoassay techniques (ELISA).
Hospital admission was based on respiratory symptoms plus radiological infiltrates or significant comorbidity. Radiologists examined the plain chest radiographs of patients suspected of having SARS-CoV-2 infection. Admission to Intensive Care Unit (ICU) was based on the development of severe disease and recoverability.
Data were collected from patients or their relatives, the computerised medical record, and the daily handover list of unstable COVID patients in the wards. It was collected within the framework of "International COVID-19 Clinical Evaluation Registry: HOPE-COVID 19" that was evaluated by the Ethics and Research Committee of the Hospital Clínico San Carlos in Madrid. The database records were entered anonymized, with an alphanumeric code and the identifying data were kept in a different file guarded by the local researchers; following data protection laws in force: Ley Orgánica 15/1999, of December 13, de Protección de Datos de Carácter Personal; Ley 41/2002, of November 14, Básica Reguladora de la Autonomía del Paciente y Derechos y Obligaciones en materia de Información y Documentación Clínica. Ley 14/2007, of July 3, de Investigación Biomédica; and Ethical Principles for Medical Research on Human Beings established in the Declaration of Helsinki by the World Medical Association. Written informed consent was waived by the Ethics and Research Committee of the Hospital Clínico San Carlos, due to the nature of the anonymized registry and the severity of the situation.
Model development and reporting followed the TRIPOD (Transparent Reporting of a multivariable prediction model for Individual Prediction Or Diagnosis) guidelines2.
Patients and public involvement
Patients and public have not been involved in the development of the research question, outcome measures, design nor execution of this study.
Outcomes
The primary outcome was the development of severe disease, defined by the presence of one of the following criteria: a respiratory failure that needs an inspiratory fraction of oxygen (FiO2) equal to or greater than 0.6, shock or severe dysfunction of another organ, or death. The secondary outcome was vital status at hospital discharge (alive/dead).
Predictors
In each patient, we collected demographic characteristics (gender, age, provenance), comorbidities, baseline functional situation and usual medication, the situation at admission (symptoms and signs, complementary examinations), evolution during hospitalisation, and status at discharge.
As an indicator of acute physiological injury, we calculated the CRB scale3,4 on admission (Supplementary Material); it is a validated version of the CURB-65 scale5; endorsed by British Thoracic Society6 and NICE7,8. Arterial hypotension and tachypnea were defined as in CRB score: systolic arterial pressure < 90 mmHg or diastolic arterial pressure ≤ 60 mmHg; tachypnea was defined by a respiratory rate ≥ 39 per minute. Fever was defined as temperature ≥ 38 °C. At admission, radiologists reported the chest radiographs of patients with suspected SARS-CoV-2 infection.
As a summary variable of comorbidity, we calculated the Age-Adjusted Charlson Comorbidity Index9,10 (Age-Charlson) (Supplementary Material). We chose to evaluate age as a part of the comorbidity index instead of in CRB-65 for two reasons: first, because of clinical significance, as we consider that increasing age provides information about non-explicit comorbidity, it is a kind of "hidden comorbidity index"; and second, for statistical reasons, to minimise the number of predictor variables while maximising the exploitation of a continuous variable.
Statistical analysis
The sample size was determined by the evolution of the pandemic. No imputation of values has been made in the missing data.
In the descriptive analysis, absolute and relative frequencies were calculated in the categorical variables, mean and standard deviation (SD) in the continuous ones with Normal distribution, and median and interquartile range (IQR) in the continuous ones with non-Normal distribution.
For the bivariate analysis, according to the outcomes of interest, we calculated p values with Chi-squared, Student’s t or Fisher’s exact tests as appropriate. P value less than 0.05 are considered statistically significant, no adjustment for multiple comparations were done. All test were 2-tailed.
Multivariate analysis was carried out with forward conditional stepwise logistic regression. Dependent variables were the primary or secondary outcomes; independent variables were selected by clinical and statistical criteria in several stages. For statistical analysis, we used the IBM SPSS Statistics package, version 25.
Ethics approval and consent to participate
This study has been done within the framework of "International COVID-19 Clinical Evaluation Registry: HOPE-COVID 19" project that was approved by the Ethics and Research Committee of Hospital Clínico San Carlos in Madrid(20/241-E) and Agencia Española de Medicamentos y productos Sanitarios (EPA- = D). The database records were entered anonymized, with an alphanumeric code and the identifying data were kept in a different file guarded by the local researchers; following data protection laws in force: Ley Orgánica 15/1999, of December 13, de Protección de Datos de Carácter Personal; Ley 41/2002, of November 14, Básica Reguladora de la Autonomía del Paciente y Derechos y Obligaciones en materia de Información y Documentación Clínica. Ley 14/2007, of July 3, de Investigación Biomédica; and Ethical Principles for Medical Research on Human Beings established in the Declaration of Helsinki by the World Medical Association. Written informed consent was waived because the characteristics of the anonymized registry and the severity of the situation.
Results
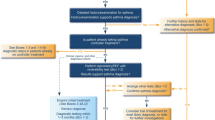
The first COVID-19 patient was admitted to our hospital on March 1, 2020, and the last of this "first wave" on April 28, 2020; during that period 413 records were included in the database. From them, eight records were deleted because of duplication; twelve patients were excluded because they were transferred to another hospital due to administrative reasons, without being admitted at HUVV; and one patient was excluded because he had an asymptomatic SARS-CoV-2 infection and was hospitalised because of non-related condition (atrioventricular block). Therefore, 392 patients with COVID-19 are analysed. Figure 1 shows the participants flowchart. Follow-up lasted until the discharge of the last patient: July 21, 2020. One hundred and four patients developed severe disease (27% of the study group), and fifty-two died (13%). In the Supplementary Material, Fig. S1 shows the daily flow of admissions, discharges and patients hospitalised.
Baseline characteristics are shown in Table 1. The mean age was 61 years; 59% were men. The median burden of comorbidity was 2 points in the Age-Charlson scale, being significantly higher in patients who developed severe disease (median 4.5 versus 2 in the non-severe), and in those who died (median 6.5 points versus 2 in those who survived). Fourteen per cent of the patients had some degree of dependency in activities of daily living. The most prevalent pathological background was arterial hypertension (46%). In the bivariate analysis, the variables most clearly associated with the development of severe disease or death were: age (71 years in the severe vs 57 in the non-severe), cerebrovascular disease, chronic heart disease and arterial hypertension.
The situation on arrival at the Emergency Department is summarised in Tables 2 and 3. The average duration of symptoms was 7 days (median), being significantly shorter in patients with clinical deterioration (6 days in those who developed severe disease, and 5 days in those who eventually died). The CRB score was 0 points in more than 80% of the cases, but any increase was strongly associated with adverse evolution. Baseline pulse oximetry saturation on arrival was the simple complementary examination most strongly associated with a negative outcome in the unadjusted bivariate analysis.
During hospitalisation, most of the patients received hydroxychloroquine (92%) and lopinavir/ritonavir (80%). Drugs aimed at attenuating the inflammatory response (corticosteroids, tocilizumab, interferon) were used less frequently (around 20%) and preferentially in the most severely ill patients. Remdesivir was not administered due to availability issues (Table 4). One hundred and four patients developed severe disease (27% of the sample), at a median of 9 days from the onset of symptoms, forty (10% of the sample) were admitted to the ICU. Fifty-two (13%) died, sixteen of them in the ICU (40% of all admitted to ICU). The median hospital stay of the total sample was 8 days, with two clearly differentiated patterns: shorter stays in patients with moderate disease and in patients who die (median of 7 and 7.5 days respectively), and longer stays in patients who survived despite developing severe disease (median 22 days, IQR: 13–42.2); these differences are clinically, epidemiologically, and statistically significant (Fig. 2).
The final multivariate model for prediction of the primary outcome (development of severe disease), is shown in Table 5. It contains only three variables: Age-Charlson scale, CRB scale, and baseline desaturation by pulse oximetry. There are no missing data, so 392 patients are analysed. Cox and Snell’s R2 is 0.28, and Nagelkerke’s 0.42; Hosmer–Lemeshow test p = 0.22; C statistic: 0.85 (95% CI 0.80–0.89), global sensitivity: 93%, specificity: 55%. Figure 3 displays the receiver operating characteristic curve (ROC curve). Logistic regression requirements are met. Among the rest of the factors that could have independent prognostic value, only CRP, LDH and heart failure had statistically significant coefficients but did not improve the overall performance of the model (Supplementary Material, Table S1 and Fig. S2). Gender, hypertension, previous dependence, days from onset of symptoms to arrival at the hospital, DD, AST, or acute renal failure upon admission were not significant; troponin, ferritin and PCT were not be evaluated in the multivariate analysis because there are few cases with valid data in the first day. We also do not evaluate inpatient drug therapy because their administration has been highly biased by the severity perceived by the physician assisting the patient, and it was not possible to control this confounding factor.
In the multivariate analysis for prediction of the secondary outcome (death), we arrived at a model with the same predictors, and remarkably similar performance (Table 6, Fig. 4). Hosmer Lemeshow test p = 0.85; Cox and Snell’s R2: 0.24 and Nagelkerke’s 0.45. The C statistic: 0.90 (95% CI 0.86–0.94), overall sensitivity 97%, specificity 40%. Among the rest of the variables that could be independent risk factors, only LDH and lymphocyte count reached statistical significance but did not significantly improve the model (Supplementary Material, Table S2 and Fig. S3). Gender, arterial hypertension, dependence in activities of daily living, DD, CRP, PCT, AST, leukocytes, haemoglobin, platelets, sodium, acute renal failure or days from the onset of symptoms to arrival at the hospital were not significant; troponin or ferritin cannot be explored in the multivariate model because there are few cases with valid data.
Discussion
The main conclusion of this study is that the prognosis of a patient with COVID-19 pneumonia can probably be predicted by combining a widely validated comorbidity scale and an acute disease scale; the only "complementary examination" that we include is arterial saturation by pulse oximetry, a measurement that can be done at patient's home as easily as taking blood pressure. We have chosen the most popular comorbidity scale: the Charlson Comorbidity Index11 (age-adjusted version9); and as a pneumonia severity scale, one of the CURB-65 family: the CRB scale3; but surely there will be other options. The main point is to check the validity of an idea with such clinical coherence: the prognosis of a patient essentially depends on the balance between the resistance capacity and the aggressiveness of the acute problem.
The sample we study meets the requirements to be considered representative: confirmed cases, consecutively included, in the same phase of the disease (on admission to hospital), with homogeneous admission criteria, in a naturally delimited time frame, with prospective data collection and complete follow-up (minimal percentage of losses: 12/404, 3%). Furthermore, looking at the proportion of hospital beds occupied by COVID-19 patients (maximum 38%, Supplementary Material, Fig. S1), we get at the impression that the confounding effect that a possible work-overload could have on patient outcomes has been lower in our hospital than in other cases12,13.
Baseline characteristics also support the idea of representativeness; they are very similar to other series12,14,15, predominantly male, with a mean age of 60 years, similar to the USA16 and intermediate between that of China17 (around 55 years), and United Kingdom18 (70 years old). The comorbidity burden was low (Age-Charlson median 2 points), similar to that observed by Casas-Rojo15 in Spain, and in other series that have evaluated age and the Charlson index separately: Italy19, USA20, Denmark21, or China13. In all of them, with such different socio-geographic contexts, both characteristics were independent risk factors, which reinforces the idea of the suitability of combining them in Age-Charlson.
In the first clinical evaluation, CRB and SpO2 were abnormal in only 18% of the patients, but with a strong association with severity. SpO2 could be especially useful in COVID-19 patients, helping to detect what has been called "silent hypoxemia"22,23.
The most widespread model in which data on comorbidity and acute disease are combined in patients with pneumonia is the PSI scale24. However, it has a substantial disadvantage: it cannot be used outside a health centre since 7 of its 19 variables require laboratory or radiology/ultrasound. There are very few studies with predictive models applicable in primary care that, at the same time, implement such intuitive idea as that assessing the prognosis of potentially seriously ill patients requires considering not only the aggressiveness of the acute disease but also the burden of chronic disease that weakens them25. Generally, both components have been studied as alternatives26, and rarely as complementary27,28. In patients with COVID-19, Petrilli29 and ISARIC18,30 are two groups that more closely resemble this study’s objective. Petrilli does not explicitly include a comorbidity scale but empirically reaches the same conclusions: age, comorbidity, oxygenation and inflammation parameters determine the need for hospitalisation and the development of severe disease; the relative weight of each possibly varies depending on the outcome and the population of interest. ISARIC-4C is based on the components of the Charlson Index and CURB-65, along with gender, obesity and CRP to build a model with 8 predictor variables, including 2 biochemical which limits its application outside the hospital context; unexpectedly, hypotension has not reached the final model. In polypathological COVID-19 patients, the usefulness of combining acute damage and comorbidity scales has also been partially reported, in this case not with the Charlson index but with a specific scale for polypathological patients (PROFUND)31.
Regarding other variables that could be important, we have explored the baseline functional situation in terms of dependency for activities of daily living, and though it was significant in the unadjusted bivariate analysis (Table 1), it ceased to be so in the multivariate after incorporating the Age-Charlson scale; however, we think it deserves to be further explored. Casas-Rojo15 have similar results: 16% of dependency for activities of daily living, and association with worse evolution in the bivariate analysis; the multivariate analysis has yet to be published. Bernabeu-Wittel31 in a study focused on multiple pathological patients with COVID-19 incorporates functional status (Barthel index) into the assessment of comorbidity.
The rate of severe disease in this series is 27%; in other studies, it ranges between 15 and 37%29,30,32,33. This variability may be due to differences in the selected sample and in the definition of severe disease:
-
Major differences in sampling: due to differences in the age of the patients (which we will address next); or due to exclusively including patients diagnosed by chest CT34 (which is more sensitive than plain radiography); or excluding patients who already present in a severe condition35,36,37,38 (because their objective is to study the progression from non-severe to severe); or limiting follow-up to a short period which does not allow that a significant proportion of included patients reach the outcome of interest39,40, and therefore rising a significant risk of selection bias that will be later discussed.
-
Important differences in the definition of severe disease: most of predictive models developed in China37,40,41 use the definition recommended by the National Health Commission of China, that is broader than ours. In this Chinese definition, a ratio of arterial oxygen pressure to inspiratory oxygen fraction (Pa/Fi) less than 300 is a sufficient criterion to diagnose severe pneumonia. So, for example, a patient that with a FiO2 of 0.3 and a PaO2 of 80 mmHg (Pa/Fi: 80/0.3 = 267), should be considered severe with the Chinese definition, but not with ours. Our definition adopts criteria routinely recommended to consider the admission of a patient with pneumonia to an area of high dependency or an Intensive Care Unit42,43,44, regarding FiO2 it requires to need 0.6 or more. Other studies limit the definition to “admission in ICU or Intermediate Unit”18; overlooking that in order to admit a patient in these units, in addition to severity, patient recoverability and availability of beds are also assessed; this explains the variability in the use of ICUs and why a high percentage of severely ill patients are not treated in ICU45, approximately 60% in our series.
Crude mortality rate in our series is 13% (of hospitalised patients). Again, direct comparison with other series is difficult, even being mortality a more robust outcome than disease severity. In Spain, mortality in multicentre studies of hospitalised patients has been 21–28% 15,32, in the United Kingdom 30%30, Italy 20%46. Age distribution and incomplete follow-up are two factors that could explain not only differences in raw mortality but also in the performance of predictive models.
Mortality varies according to age in all series; in our study, it ranges from 0% under 40 years to almost 40% at ages above 80 years, Fig. 5. A partial solution to improving comparability could be the age-standardised mortality rate, that is the mortality that a population would have if it had the age distribution of a reference population (e.g., the WHO World Standard Population)47,48, although it is not without criticism49. In this series, the age-standardised mortality rate with this reference population is 2.9 deaths per 100 COVID-19 patients admitted.
Mortality rate and model performance can be biased in studies with incomplete follow-up, and cannot be controlled in the analysis phase. In mortality studies published in the first months of the pandemic, it has been frequent to limit the follow-up to 2 weeks of hospital stay; so that only those patients who have died or been discharged during that time were analysed and those who remained hospitalised were excluded16,18,50. The lack of follow-up information on these, most likely biases the estimation of crude mortality and the performance of predictive models of mortality1,33. Figure 2 shows the distribution of hospital stay in our series depending on the outcome, the group of severe but surviving patients had the longest stay, well above 14 days, so would be largely censored for the analysis if the follow-up be limited to two weeks: as it is a "selective" loss of survivors, it leads to overestimate the mortality rate; and in addition, as it is also a "selective" loss of patients with a difficult prognosis (they were severely ill but survived), it leads to overestimate the performance of the prediction model. Our series only has a 3% loss of included patients, and not related to the length of stay nor outcome, but due to transfer from the Emergency Department to another hospital because of their place of residence.
We have aimed to keep the predictive model as applicable and straightforward as possible without compromising performance; that is why we have not included in the final models some laboratory variables that could be in, from a statistical point of view. With the same perspective, our models are advantageous compared with others that require tools with little availability today, such as artificial intelligence or computer applications with copyright34,36,37,46,47. This does not mean that all these issues could not be necessary for other objectives and contexts; v.g., comorbidity variables probably be more decisive in countries with an older population; while variables of acute inflammatory damage do so in countries with a young population29,51,52.
What use can these models have? An essential requirement to apply them with confidence is their validation in independent but representative samples. Once validated, it can have multiple applications, both in the clinical and management area:
-
Support to make clinical decisions when, after a routine initial assessment, the course of action is unclear. In many situations it is necessary to filter patients according to severity, for example, to keep as an outpatient or where to ubicate in a hospital. Usually, the most convenient approach is to start with a screening tool to identify those at high risk to get the most from scarce resources. Screening tools are characterized by high sensitivity, and that is the main feature of our model: the global sensitivity for predicting severe disease is 93%, and specificity 55%. The corresponding nomogram in the Supplementary material would allow to calculate the risk for a particular patient, and together with the clinical judgment get to a conclusion. Those patients considered at high risk could be managed with a short observation period to check the tendency and carry out more specific tests that are often more complex and resource-consuming.
-
Support in decision-making for the management of the infrastructure necessary for the assistance to function as efficiently and effectively as possible.
-
Quality control, through the relationship between observed and expected mortality according to the model53.
Study limitations
The sample came from a single centre during the first months of the pandemic when the standard treatment was not the same as nowadays. Its size is modest, and though sufficient for the study's objective, it still produces wide confidence intervals in relevant variables. There is a potential risk of overfitting in the development of all predictive models, so it is necessary to validate it in an independent sample of patients before any clinical use.
Conclusion
In patients with COVID-19 pneumonia, the prognosis can likely be significantly narrowed by combining a comorbidity scale and a current severity scale of pneumonia, both based in clinical data readily available. This study proposes a predictive model based on Age-Adjusted Charlson index, the CRB scale, and baseline arterial saturation. It can be completed at the first medical contact through standard anamnesis, physical examination, and a pocket pulse oximeter.
Data availability
The datasets supporting the conclusions of this article are included within the article and its additional file. Further details are available from the corresponding author, PNO, upon reasonable request.
References
Wynants, L. et al. Prediction models for diagnosis and prognosis of covid-19 infection: Systematic review and critical appraisal. BMJ 369, m1328. https://doi.org/10.1136/bmj.m1328 (2020).
Collins, G. S., Reitsma, J. B., Altman, D. G. & Moons, K. G. M. Transparent Reporting of a multivariable prediction model for Individual Prognosis Or Diagnosis (TRIPOD): The TRIPOD Statement. Br. J. Surg. 102, 148–158. https://doi.org/10.1002/bjs.9736 (2015).
Bauer, T. T., Ewig, S., Marre, R. & Suttorp, N. CRB-65 predicts death from community-acquired pneumonia. J. Intern. Med. https://doi.org/10.1111/j.1365-2796.2006.01657.x (2021).
Fan, G. et al. Comparison of severity scores for COVID-19 patients with pneumonia: A retrospective study. Eur. Respir. J. https://doi.org/10.1183/13993003.02113-2020 (2020).
Lim, W. S. et al. Defining community acquired pneumonia severity on presentation to hospital: An international derivation and validation study. Thorax 58, 377–382. https://doi.org/10.1136/thorax.58.5.377 (2003).
Lim, W. S. et al. BTS guidelines for the management of community acquired pneumonia in adults: Update 2009. Thorax 64(Suppl 3), iii1-55. https://doi.org/10.1136/thx.2009.121434 (2009).
National Institute for Health and Care Excellence (UK). Pneumonia in adults: diagnosis and management. 2020. https://www.ncbi.nlm.nih.gov/pubmed/31940163 (National Institute for Health and Care Excellence, 2021)
Overview | COVID-19 rapid guideline: managing suspected or confirmed pneumonia in adults in the community | Guidance | NICE. https://www.nice.org.uk/guidance/ng165 (2021), (Accessed 30 Aug 2020).
Charlson, M., Szatrowski, T. P., Peterson, J. & Gold, J. Validation of a combined comorbidity index. J. Clin. Epidemiol. 47, 1245–1251. https://doi.org/10.1016/0895-4356(94)90129-5 (1994).
de Groot, V., Beckerman, H., Lankhorst, G. J. & Bouter, L. M. How to measure comorbidity. A critical review of available methods. J. Clin. Epidemiol. 56, 221–229. https://doi.org/10.1016/s0895-4356(02)00585-1 (2003).
Charlson, M. E., Pompei, P., Ales, K. L. & MacKenzie, C. R. A new method of classifying prognostic comorbidity in longitudinal studies: Development and validation. J. Chronic Dis. 40, 373–383. https://doi.org/10.1016/0021-9681(87)90171-8 (1987).
Borobia, A. M. et al. A cohort of patients with COVID-19 in a major teaching hospital in Europe. J. Clin. Med. Res. 9, 1733. https://doi.org/10.3390/jcm9061733 (2020).
Liang, W. et al. Development and validation of a clinical risk score to predict the occurrence of critical illness in hospitalized patients with COVID-19. JAMA Intern. Med. https://doi.org/10.1001/jamainternmed.2020.2033 (2020).
Martos Pérez, F. et al. Comorbilidad y factores pronósticos al ingreso en una cohorte COVID-19 de un hospital general. Rev. Ordem. Med. https://doi.org/10.1016/j.rce.2020.05.017 (2020).
Casas-Rojo, J. M. et al. Características clínicas de los pacientes hospitalizados con COVID-19 en España: resultados del Registro SEMI-COVID-19. Rev. Clin. Esp. https://doi.org/10.1016/j.rce.2020.07.003 (2020).
Richardson, S. et al. Presenting characteristics, comorbidities, and outcomes among 5700 patients hospitalized with COVID-19 in the New York City area. JAMA https://doi.org/10.1001/jama.2020.6775 (2020).
CDC Weekly C, The Novel Coronavirus Pneumonia Emergency Response Epidemiology Team. The epidemiological characteristics of an outbreak of 2019 novel coronavirus diseases (COVID-19)—China, 2020. China CDC Weekly. 2, 113–122. https://doi.org/10.46234/ccdcw2020.032 (2020).
Docherty, A. B. et al. Features of 20 133 UK patients in hospital with covid-19 using the ISARIC WHO Clinical Characterisation Protocol: Prospective observational cohort study. BMJ 369, m1985. https://doi.org/10.1136/bmj.m1985 (2020).
Iaccarino, G. et al. Age and multimorbidity predict death among COVID-19 patients: Results of the SARS-RAS study of the Italian Society of Hypertension. Hypertension 76, 366–372. https://doi.org/10.1161/HYPERTENSIONAHA.120.15324 (2020).
Imam, Z. et al. Older age and comorbidity are independent mortality predictors in a large cohort of 1305 COVID-19 patients in Michigan, United States. J Intern Med. https://doi.org/10.1111/joim.13119 (2020).
Christensen, D. M. et al. Charlson Comorbidity Index score and risk of severe outcome and death in Danish COVID-19 patients. J. Gen. Intern. Med. https://doi.org/10.1007/s11606-020-05991-z (2020).
Xie, J. et al. Critical care crisis and some recommendations during the COVID-19 epidemic in China. Intensive Care Med. 46, 837–840. https://doi.org/10.1007/s00134-020-05979-7 (2020).
Shah, S. et al. Novel use of home pulse oximetry monitoring in COVID-19 patients discharged from the emergency department identifies need for hospitalization. Acad. Emerg. Med. https://doi.org/10.1111/acem.14053 (2020).
Fine, M. J. et al. A prediction rule to identify low-risk patients with community-acquired pneumonia. N. Engl. J. Med. 336, 243–250. https://doi.org/10.1056/NEJM199701233360402 (1997).
Kolditz, M., Braeken, D., Ewig, S. & Rohde, G. Severity assessment and the immediate and long-term prognosis in community-acquired pneumonia. Semin. Respir. Crit. Care Med. 37, 886–896. https://doi.org/10.1055/s-0036-1592127 (2016).
Wesemann, T. et al. Pneumonia severity, comorbidity and 1-year mortality in predominantly older adults with community-acquired pneumonia: A cohort study. BMC Infect. Dis. 15, 2. https://doi.org/10.1186/s12879-014-0730-x (2015).
Bahlis, L. F., Diogo, L. P., Kuchenbecker, R. D. S. & Fuchs, S. C. Clinical, epidemiological, and etiological profile of inpatients with community-acquired pneumonia in a public hospital in the interior of Brazil. J. Bras. Pneumol. 44, 261–266. https://doi.org/10.1590/S1806-37562017000000434 (2018).
Watanabe Tejada, L. C. et al. Effect of comorbidities on clinical outcomes in low-risk curb-65 patients. J. Gen. Intern. Med. 28, S67 (2013).
Petrilli, C. M. et al. Factors associated with hospital admission and critical illness among 5279 people with coronavirus disease 2019 in New York City: Prospective cohort study. BMJ 369, m1966. https://doi.org/10.1136/bmj.m1966 (2020).
Knight, S. R. et al. Risk stratification of patients admitted to hospital with covid-19 using the ISARIC WHO Clinical Characterisation Protocol: Development and validation of the 4C Mortality Score. BMJ 370, m3339. https://doi.org/10.1136/bmj.m3339 (2020).
Bernabeu-Wittel, M. et al. Death risk stratification in elderly patients with covid-19 A comparative cohort study in nursing homes outbreaks. Arch. Gerontol. Geriatr. 91, 104240. https://doi.org/10.1016/j.archger.2020.104240 (2020).
Berenguer, J. et al. Characteristics and predictors of death among 4,035 consecutively hospitalized patients with COVID-19 in Spain. Clin. Microbiol. Infect. https://doi.org/10.1016/j.cmi.2020.07.024 (2020).
Wu, Z. & McGoogan, J. M. Characteristics of and important lessons from the coronavirus disease 2019 (COVID-19) outbreak in China: Summary of a report of 72 314 cases from the Chinese Center for Disease Control and Prevention. JAMA https://doi.org/10.1001/jama.2020.2648 (2020).
Colombi, D. et al. Well-aerated lung on admitting chest CT to predict adverse outcome in COVID-19 pneumonia. Radiology 296, E86–E96. https://doi.org/10.1148/radiol.2020201433 (2020).
Ji, D. et al. Prediction for progression risk in patients with COVID-19 pneumonia: The CALL score. Clin. Infect. Dis. https://doi.org/10.1093/cid/ciaa414 (2020).
Singh, K. et al. Validating a widely implemented deterioration index model among hospitalized COVID-19 patients. medRxiv. https://doi.org/10.1101/2020.04.24.20079012 (2020).
Bai, X. et al. Predicting COVID-19 malignant progression with AI techniques. medRxiv. https://doi.org/10.1101/2020.03.20.20037325 (2020).
Huang, H. et al. Prognostic factors for COVID-19 pneumonia progression to severe symptom based on the earlier clinical features: A retrospective analysis. medRxiv. https://doi.org/10.1101/2020.03.28.20045989 (2020).
Carr, E., Bendayan, R, O’Gallagher, K., Bean, D. & Dobson, R. Supplementing the National Early Warning Score (NEWS2) for anticipating early deterioration among patients with COVID-19 infection. https://doi.org/10.1101/2020.04.24.20078006 (Accessed 27 Aug 2020) (2020).
Gong, J. et al. A tool to early predict severe 2019-novel coronavirus pneumonia (COVID-19): A multicenter study using the risk Nomogram in Wuhan and Guangdong, China. MedRxiv. https://doi.org/10.1101/2020.03.17.20037515 (2020).
Guo, Y. et al. Development and validation of an early warning score (EWAS) for predicting clinical deterioration in patients with coronavirus disease 2019. Infectious diseases (except HIV/AIDS). medRxiv https://doi.org/10.1101/2020.04.17.20064691 (2019).
Smith, G. & Nielsen, M. ABC of intensive care. Criteria for admission. BMJ 318, 1544–1547. https://doi.org/10.1136/bmj.318.7197.1544 (1999).
Metlay, J. P. et al. Diagnosis and treatment of adults with community-acquired pneumonia. An official clinical practice guideline of the American Thoracic Society and Infectious Diseases Society of America. Am. J. Respir. Crit. Care Med. 200, e45–e67. https://doi.org/10.1164/rccm.201908-1581ST (2019).
Christian, M. D. et al. Chapter 7. Critical care triage. Recommendations and standard operating procedures for intensive care unit and hospital preparations for an influenza epidemic or mass disaster. Intensive Care Med. 36(Suppl 1), S55-64. https://doi.org/10.1007/s00134-010-1765-0 (2010).
Ewig, S. et al. New perspectives on community-acquired pneumonia in 388 406 patients. Results from a nationwide mandatory performance measurement programme in healthcare quality. Thorax 64, 1062–1069. https://doi.org/10.1136/thx.2008.109785 (2009).
Das, A., Mishra, S. & Gopalan, S. S. Predicting community mortality risk due to CoVID-19 using machine learning and development of a prediction tool. medRxiv https://doi.org/10.1101/2020.04.27.20081794 (2020).
Zeng, L. et al. Risk assessment of progression to severe conditions for patients with COVID-19 pneumonia: A single-center retrospective study. medRxiv. https://doi.org/10.1101/2020.03.25.20043166 (2020).
Giacomelli, A. et al. 30-day mortality in patients hospitalized with COVID-19 during the first wave of the Italian epidemic: A prospective cohort study. Pharmacol Res. 158, 104931. https://doi.org/10.1016/j.phrs.2020.104931 (2020).
Ahmad, O. B. et al. Age standardization of rates: A new WHO standard. 2001; 9. https://www.who.int/healthinfo/paper31.pdf (World Health Organization, 2021)
Rao, C., Doi, S. & Williams, G. Enhanced comparisons of COVID-19 mortality across populations. MedRxiv https://doi.org/10.1101/2020.06.14.20131318 (2020).
Spuntarelli, V. et al. COVID-19: is it just a lung disease? A case-based review. SN Compr. Clin. Med. 2,1–6 (2020).
Arcari, L. et al. Incidence and determinants of high-sensitivity troponin and natriuretic peptides elevation at admission in hospitalized COVID-19 pneumonia patients. Intern Emerg. Med. 15, 1467–1476 (2020).
Bottle, A., Jarman, B. & Aylin, P. Strengths and weaknesses of hospital standardised mortality ratios. BMJ 342, c7116. https://doi.org/10.1136/bmj.c7116 (2010).
Acknowledgements
We want to state our acknowledgement to all the hospital staff, who have made a special effort to assist these patients. Last but not least, we want to dedicate a mention to all patients and their families; they entrusted us with what they loved most.
Author information
Authors and Affiliations
Consortia
Contributions
P.N.O.: conceptualization, design, coordination, result interpretation, draft writing and critical review. C.R.A.: patient care and data acquisition. F.D.M.: patient care and data acquisition. V.M.B.M.: design and coordination. L.R.F.: analysis and result interpretation, M.A.E.F.: conceptualization. HOPE-HUVV group: patient care.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nuevo-Ortega, P., Reina-Artacho, C., Dominguez-Moreno, F. et al. Prognosis of COVID-19 pneumonia can be early predicted combining Age-adjusted Charlson Comorbidity Index, CRB score and baseline oxygen saturation. Sci Rep 12, 2367 (2022). https://doi.org/10.1038/s41598-022-06199-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-06199-3
This article is cited by
-
Development of a proteomic signature associated with severe disease for patients with COVID-19 using data from 5 multicenter, randomized, controlled, and prospective studies
Scientific Reports (2023)
-
Prognostic models in COVID-19 infection that predict severity: a systematic review
European Journal of Epidemiology (2023)
-
Potential and limitations of machine meta-learning (ensemble) methods for predicting COVID-19 mortality in a large inhospital Brazilian dataset
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.