Abstract
Interactions between the prokaryotic microbiome and eukaryotic parasites in the vertebrate gut may affect overall host health and disease. While intertropical areas exhibit a high rate of parasites carriers, such interactions are understudied in these populations. Our objectives were to (1) describe the gut microbiome of individuals living in Madagascar, (2) identify potential associations between bacterial taxa and parasites colonizing the digestive tract and (3) highlight main determinants of the gut microbiota composition in this developing country. Metadata (socioeconomic, diet, clinical) and fecal samples were collected from 219 volunteers from North-West Madagascar (Mahajanga). Fecal microbiome was assessed through 16S rRNA gene sequencing and metabolomics, and related to dietary habits and parasites carriage. We highlight important Malagasy gut microbiome peculiarities. Out of three detected enterotypes, only one is similar to that observed in Westernized countries (Ruminococcus-driven). Functions associated with the two others (Clostridium sensu stricto-driven and Escherichia/Shigella-driven) are mostly directed toward amino acids biosynthesis and degradation, respectively. Diet and protozoan carriage were the main drivers of microbiota composition. High protozoan carriage was associated with higher diversity, richness and microbial functionalities. The gut microbiome of Malagasy strongly differs from that of Westernized countries. Asymptomatic protozoan carriage and dietary habits are the external factors with the deepest impact on gut microbiome. Further studies are needed to understand whether gut microbial richness constitute a predilection niche for protozoans colonization, due to their gazing features, or whether the parasites themselves induce a higher bacterial richness.
Similar content being viewed by others
Introduction
The trillions of bacteria colonizing our intestines are known to play key roles in human health, ranging from digestion, to immunity development, yet, complexity of the human gut microbiome is far from being fully decrypted. Most studies are performed in westernized countries and, when focusing on lower income countries, often aim at linking the gut microbiome with nutritional status. Hence, one of the first studied populations from rural Africa highlighted a specific microbiome in 14 children from Burkina Faso, geared toward degradation of polysaccharides1. Another study emphasized starch degradation capabilities of the gut microbiome in a Malawi children cohort (n = 83 infants)2. When analyzing the microbiota of adult population, isolated territories or people are often selected to describe the reciprocal impact of genetic versus environment on the gut ecosystem. Most of these studies describe a higher bacterial diversity and richness in these populations, either originating from the East African tropical savanna (n = 27 Hadza hunters-gathers3) or the southwestern Pacific Ocean (n = 40 individuals originating from two traditional societies of Papua New Guinea4).
Parasitic eukaryotes members of the human microbiome are often underestimated and underexplored, even though their importance in the gut ecosystem, mainly as immunity drivers, is widely accepted. Protozoans’ colonization ranges from 5 to 100% of the population, depending on geographical origin, with many carriers remaining asymptomatic. Some parasitic protozoa, such as Entamoeba histolytica, Giardia intestinalis, and Tritrichomonas suis, can modify intestinal mucus abundance and composition, enabling them to penetrate the mucus barrier during pathogenesis5. Many gut protozoa can also damage the epithelium during pathogenesis (disruption of tight junctions, cell invasion and destruction)6. Both mucosal and epithelial damages will directly alter the host's interaction with their microbiome during infection. However, asymptomatic parasitic colonization also impacts the host microbiome and one of the most studied organisms is Blastocystis sp. (abbreviated as Blastocystis in the present paper). While historically considered as parasite and assumed to have a detrimental effect on the host organism, the vision of a potential commensalism and beneficial role of Blastocystis starts to emerge7. A recurrent finding is the robust link between gut colonization by Blastocystis and higher microbiome diversity and/or richness8,9,10,11,12, together with a negative association with the Bacteroides genus abundance and Bacteroides enterotypes9,12,13.
Still, these results are seen through the prism of westernized countries (France, Sweden, Belgium and Mexico) and compiled data from shotgun metagenomic studies. Considering the higher rate of parasitic carriers in intertropical area, we focused our research on volunteers from Madagascar and previously described a wide panel of intestinal parasites14. Protozoa were the most prevalent with 72.8% of the population colonized whereas helminths and microsporidia were detected in less than 10% of the volunteers. Blastocystis was the most prevalent protozoa followed by various amoebas and flagellates.
The objectives of the present study were to (1) describe the gut microbiota composition of individuals living in Madagascar, (2) identify potential associations between gut bacterial taxa and parasites colonizing the digestive tract and (3) highlight main determinants of the gut microbiota composition in this developing country.
It is, to our knowledge the first paper intending to describe the general gut microbiome profiles of healthy adults from Madagascar, and, together with the Flemish gut project12, linking parasitic status to microbiome in a cohort of more than 200 individuals.
Results
Malagasy population exhibits a specific gut microbiome
A total of 6428 OTUs was detected in the fecal samples provided by 219 Malagasy individuals. On average, a Malagasy microbiome harboured 439 ± 106 OTUs classified in 12 microbial phyla. Main phyla were affiliated to Firmicutes (58.24% ± 19.96), Bacteroidetes (15.90% ± 13.65), Proteobacteria (14.01% ± 19.80), Actinobacteria (5.62% ± 6.07) and Spirochaetes (1.48% ± 4.61). Microbial diversity and richness indices (Simpson: 0.90 ± 0.10 and number of observed OTUs: 439 ± 106, respectively) were high. The bacterial load estimated by qPCR depicted a dense bacterial ecosystem composed on average of 1.67 × 1010 CFU/g of feces (1.14 × 106–2.05 × 1012 CFU/g).
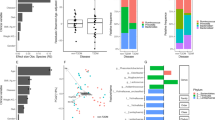
Ten bacterial families were identified as taxa contributing the most to the variance observed between microbiota profiles (96% of the β-diversity variance explained) (Fig. 1A) and were affiliated to Enterobacteriaceae, Peptostreptococcaceae, Clostridiaceae 1, Porphyromonadaceae, Streptococcaceae, Ruminococcaceae, Bacteroidaceae, Prevotellaceae, Bifidobacteriaceae, and Lachnospiraceae. Three distinct clusters or enterotypes (Ent) were statistically detected within the microbiome of the Malagasy population (Fig. 1B). Enterotypes stratification was mainly supported by the relative abundance of Ruminococcus (Ent1—n = 92 individuals), Clostridium sensu stricto (Ent2—n = 100 individuals) and Escherichia/Shigella (Ent3—n = 27 individuals) genera (Fig. 1C). Ent1 was driven by Ruminococcus bromii (100% identity). An uncultured rumen clone (98%—EF436397) and Clostridium disporicum (100%) were associated with Ent2. Escherichia coli (100%) was found more abundant in Ent3. Both bacterial diversity and richness were significantly lower in Ent3 as compared to Ent1 and Ent2 (p = 1e−13 and 1.1e−09 and p = 8e−07 and 8e−06 respectively) (Additional file 1: Fig. S1A). However, the bacterial load estimated by qPCR was not statistically different between the three enterotypes (ANOVA p = 0.19). (Additional file 1: Fig. S1B).
Malagasy gut microbiota general description and enterotypes classification. (A) Bacterial families (n = 10) contributing the most to the dissimilarity observed within the 219 microbiomes. Taxa contribution was determined using canonical correspondence analysis of family abundances and was plotted on the two first PCoA dimensions. Arrows length is proportional to family contribution (%). Dots indicate individual’s location on PCoA. (B) PCA inter-class analysis depicting the enterotype stratification of individuals from Madagascar. Each dot represents a fecal sample. (C) Bacterial genera driving the most the enterotype description. Bacterial genera were selected following a random forest classification based on enterotypes stratification. (D) Factors associated with the inter-individual variation based on the gut microbiota composition. A total of 19 factors (FDR < 0.1) including medical, demographical, dietary and protozoan carriage were associated with inter-individual variation of the gut microbiota composition. The bar plot indicates the contribution (%) of each factor at explaining the inter-individual variation observed within the faecal bacterial composition of Malagasy individuals.
Intestinal colonization by parasites strongly affects the gut microbiome
The contribution of demographical (n = 5), medical (n = 8), dietary (n = 7) and parasitism status (n = 14) factors on inter-individual microbial composition (β-diversity) was assessed by PERMANOVA analysis. At a false discovery rate of 0.1, 19 factors were associated with dissimilarity in microbial composition (Fig. 1D). Altogether, these factors explained 25.79% of the observed variation between individuals. Neither bacterial diversity nor bacterial richness correlated with age, gender or BMI, while variation in the microbiota composition within Malagasy population was slightly but significantly affected by individual’s age (1% of total microbiota variance).
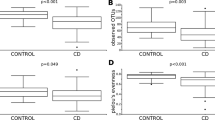
Parasitic status accounted for most of the bacterial microbiome dissimilarity (8.39%) and especially Entamoeba coli (1%), Blastocystis (0.94%), Entamoeba histolytica/dispar (0.73%) and Chilomastix mesnili (0.7%). Within our cohort, 85% of individuals were colonized by one or more parasites. Blastocystis was the most prevalent (77.6% of the cohort) and frequently associated with Dientamoeba fragilis (22.8%). While most parasites were associated with higher bacterial richness, microbiome diversity was also significantly increased in individuals colonized by Blastocystis sp., Entamoeba coli, and Enteromonas (Additional file 2: Fig. S2A). On the other hand, individuals positive for Blastocystis, Ameoba and C. mesnili displayed lower bacterial load. Strikingly, cumulative parasite colonization of the Malagasy gut increased bacterial richness and diversity (Fig. 2A). Relative abundances of 21 taxa shifted according to the presence of parasites with Blastocystis and E. coli having the greatest impact on gut microbiome. Colonized patients had higher relative abundance of commensal bacteria but lower proportions of Streptococcus and Lactobacillus (Fig. 2C). A higher parasitic cumulative diversity was associated with a significantly lower abundance of Lactobacillus (Fig. 2B), and with a low sanitary level (Additional file 2: Fig. S2B).
Association between intestinal colonization by parasites and the gut microbiome richness and composition. (A) Microbial richness (OTU number) as function of cumulative number of different protozoans detected in an individual (ranged from 0 to > 4). (B) Relative abundance of Lactobacillus genus as function of the cumulative number of protozoans detected in an individual (ranged from 0 to > 4). (C) Cladogram representation of bacterial groups significantly increased (in red) or decreased (in green) in the presence of the four main parasitic colonizers (Entamoeba coli, Blastocystis sp., Chilomastix mesnili and Entamoeba histolytica/dispar).
The proportion of individuals positive for Blastocystis, D. fragilis, E. nana and E. coli was lower in the Escherichia/Shigella Ent3 as compared to others (Additional file 2: Fig. S2C). On the contrary, most of the individuals colonized by three or more different parasites clustered within the Ruminococcus-driven Ent1 (Additional file 2: Fig. S2D).
We further investigated the relative importance of parasites symptomatology and carriage on the gut bacterial composition. While for most parasites, asymptomatic and symptomatic carriers share a similar microbiome, individuals colonized with Blastocystis and presenting gut symptoms of blastocystosis exhibited lower relative abundance of both Acetanaerobacterium and Flavonifractor (Additional file 2: Fig. S2E) than asymptomatic carriers.
Dietary habits strongly impact the Malagasy gut microbiome
Dietary habits had a significant impact (6.74%) on the inter-individual microbiota composition (Fig. 1D). From all collected dietary information, daily liquid, rice and fruit consumptions had the most significant influence on gut microbiota variance (1.54%, 1.54% and 1.64% respectively). Low liquid consumption (< 1 L/day) was associated with significantly higher proportions of Senegalimassilia, Roseburia, Haemophilus and Actinomyces (Fig. 3A). Microbiota of individuals not eating fruits harboured higher percentages of bacteria from Actinomyces, Bifidobacterium and Acetanaerobacterium (Fig. 3A) whereas moderate fruit eaters (1–2) displayed a higher abundance of Prevotella, Haemophilus and Lactobacillus. Daily consumption of 3 or more fruits increased the abundance of Sporobacter, Oscillibacter, Flavonifactor, Coprococcus, Clostridium IV and Butyricicoccus genera.
Impact of dietary habits on gut microbiome composition in Malagasy volunteers. (A) Heatmap depicting the bacterial genera having an abundance modulated by daily dietary habits. Liquid consumption is expressed in litre (L), rice in gram (g), fruits in number and other dietary information as yes/no question. Red, blue and green boxes indicate whether the abundance is higher, equal or lower in individuals positive for the listed food type. ANOVA (group > 2) and Wilcoxon test (group = 2) were run all over genera count dataset according to diet. Only boxes tagged with a star are considered statistically informative. The heatmap was generated using R software (https://cran.r-project.org/) with in-house code that uses basic functions included in "stats" and "graphics" packages. It uses genera abundance (as listed in rows) and highlights, for daily consumption of liquid, rice and fruit items, the category that are the most significantly different from others. For Meat, Fish, Yogurt-HM and Yogurt-I items, the heatmap indicates genera whose abundances are increased/decreased in individuals consuming these items. (B) Proportion of individuals included in each enterotype according to food type’s consumption. (C) Bacterial load distribution among meat and fish eaters (Y) as compared to non-eaters (N). (D) Proportion of individuals eating either yogurt (Cons+) (home-made yogurt, industrial yogurt or both (I&HM)), as function of cumulative number protozoans detected in an individual.
Malagasy gut bacterial load was significantly lower in individuals eating either meat or fish meals (Fig. 3C). Individuals eating both meat and fish harboured higher proportions of Eubacterium and Olsenella than non-eaters. Microbiota of meat eaters was characterized by less Actinomyces, whereas fish eaters’ microbiota exhibited more Acetanaerobacterium and Senegalimassilia. Low to moderate daily rice consumers (< 400 g/day) had higher relative abundance of bacterial species related to Lactobacillus, Clostridium XIVa and Sporobacter (Fig. 3A).
Yogurt consumption had also a deep impact on the microbiota. Both homemade and industrialized yogurt consumption led to significantly higher abundance of Bifidobacterium and Lactobacillus. Thirteen other genera were significantly associated with homemade yogurt consumption, most of them being less represented. Dietary habits were overall similar between Ent1 and Ent2 individuals whereas Ent3 was characterized by higher volume drinkers, lesser fruit and rice eaters and more homemade yogurt consumers (Fig. 3B).
A high proportion of parasites-free volunteers consumed homemade yogurt (61%), while highly parasites-colonized individuals did not (58%) (Fig. 3D).
Metabolism of the gut microbiome in Malagasy individuals
A global metabolomic approach has been applied on a subset of fecal samples (nEnt1 = 5, nEnt2 = 7, nEnt3 = 6) and 256 peaks were detected. Similarly to gut microbiota composition, metabolomic profiles allowed distinguishing three groups of individuals (Fig. 4A). A total of 59 peaks, belonging to six classes (Amino Acids, Peptides, and Analogues—Aromatic Compounds—Aliphatic Compounds—Carboxylic acids and derivatives—Lipids—Organic Acids and Derivatives) were differentially represented between enterotypes (Fig. 4B). Specific metabolites associated with each enterotype are described in Additional file 4 (Table S2). Metabolites relating to (1) phenylalanine, tyrosine and tryptophan biosynthesis, (2) tryptophan metabolism, (3) valine, leucine, isoleucine biosynthesis and (4) lysine biosynthesis were specific of Clostridium-driven Enterotype (Ent2). On the other hand, Pantothenate and CoA biosynthesis as well as beta-Alanine metabolism were distributed in both Ent2 and Escherichia-driven Ent3. Finally, metabolites associated with Ruminococcus-driven Ent1 related to metabolism more than biosynthesis (Glycine, serine, threonine metabolism—Cyanoaminoacid metabolism—Riboflavin metabolism).
Metabolomic specificities of the microbiome in Malagasy volunteers. Eighteen volunteers were selected according to their enterotypes affiliation (nEnt1 = 5, nEnt2 = 7, nEnt3 = 6) and analysed through metabolomic approach. (A) The inter-class analysis computed upon PCA data emphasizes the stratification of these metabolomes in three clusters, similarly to microbiome enterotypes distribution. (B) Metabolic peaks (n = 51) discriminating the Malagasy metabolomes according to enterotypes stratification. (C) Metabolic functions inferred from PICRUSt that pointed out the functional specificities associated with each enterotypes. Functions listed in the heatmap had a significantly different distribution between enterotypes (p ≤ 0.01) and were selected according to Kruskal Walis’s test followed by a post-hoc Dunn’s test assessing the intergroup (enterotypes) comparison. All tests were corrected for false discovery rate. (D) Functions differentially distributed among protozoan carrier types (0 to > 4).
A statistical inference of metabolic functions with 16S rRNA data was also performed with PICRUSt to cover the overall population. The most important shift in terms of microbiome gut functions was observed between Ent3 and the two other Enterotypes (Fig. 4C and Additional files 4 and 6: Tables S2 and S4). Xenobiotics biodegradation and metabolism such as aminobenzoate, fluorobenzoate, toluene degradation were enriched in Ent3. The metabolism of Terpenoids and Polyketides in Ent3 was marked by the biosynthesis of siderophore group non-ribosomal peptides whereas Ent1 and Ent2 harboured more functions classified in the biosynthesis of ansamycins, butirosin and neomycin. Ent3 was characterized by a lower abundance of functions related to pentose phosphate pathway, starch and sucrose metabolism. The amino acid metabolism was totally directed to the catabolism of gluthatione, lysine, tryptophan, valine, leucine and isoleucine. On the opposite, Ent1 and 2 were more prone to the biosynthesis of these amino acid units. The increased amino acid biosynthesis observed in Ent1 and 2 was interconnected with enriched protein export capabilities. Functions related to replication and repair, transcription and translation were less represented in Ent3. Similar results were observed for membrane transport and signal transduction functions.
Metabolic functions deduced from PICRUSt were also screened in Malagasy gut microbiomes in accordance to increasing asymptomatic protozoans carriage (Fig. 4D and Additional files 5 and 7: Tables S3 and S5). Microbial functions such as replication and repair, translation, amino acid and lipid metabolism, protein export and terpenoid backbone biosynthesis were enriched in individuals with higher protozoans carriage (n ≥ 3). On the opposite, xenobiotics biodegradation and metabolism such as aminobenzoate and chlorocyclohexane and chlorobenzene degradation together with unclassified amino acid metabolism related functions were increased within microbiome of low protozoans carriers (n < 3).
Discussion
The gut microbiome of people from the tropical region of Mahajanga, an urban community of Madagascar, strongly differs from that of westernized countries or isolated populations. While westernized countries microbiota can be clustered into three or four enterotypes15 driven by Bacteroides, Prevotella and Ruminococcus, only the latter genus could be described as an enterotype driver in this population. However, the microbiome was still statistically stratified into three enterotypes, the other two being driven by Escherichia/Shigella and Clostridium sensu stricto. Yet while Prevotella still accounts for approximately 7% in average of total bacteria in the overall population (median = 3%), proportions of Bacteroides are lower than 5% (4.4% and median = 0.99%).
Similarly, Pasolli et al. applied shotgun metagenomic gut microbiome profiling of two rural communities with non-Western lifestyles from north-eastern Madagascar and detected that the species-level genome bins profiles of the Madagascar population were profoundly different from that of Western-style populations16. A substantial proportion of genomic species related to Prevotella while Bacteroides was nearly absent as compared to westernized countries. Both our study and Pasolli et al. highlighted a depletion of Alistipes, Parabacteroides and Akkermansia in the Malagasy gut microbiomes.
Diet strongly explains this distribution. High proteins and animal fat intake has previously been associated with higher abundance of the Bacteroides dominant enterotype17. Its absence in Madagascar probably reflects a diet mostly based on rice. Fruits consumption had the highest impact on microbiome variance within the population. Fruits and vegetables are important sources of dietary fibers that will undergo microbial metabolism in the digestive tract. Fibers are known to increase microbiota diversity18,19. Consuming two or more fruits per day in the present population was associated with higher levels of Firmicutes and butyrate-producing bacteria such as Prevotella or Coprococcus. The gut microbiota of individuals ingesting important rice portions (≥ 500 g/day) was enriched in Ruminococcus, Sporobacter and rice consumption was higher in individuals belonging to Ent1. Individuals within this enterotype also ingested less yogurt (both home-made and commercial). Yogurt was associated with significantly higher proportions of Lactobacillus and Bifidobacterium, and lower levels of Prevotella and Enterorhabdus.
At a functional level, Ent1 and Ent2 were mostly enriched in functions dedicated to the biosynthesis of several amino acids as well as central bacterial cell mechanisms such as replication and repair, translation and transcription. On the opposite, numerous functions sustaining degradation of amino acid units and xenobiotics metabolism were enriched in Ent3 as compared to Ent1 and Ent2. This observation is coupled with higher metabolism of nitrogen compounds. The energy metabolism was more directed towards nitrogen metabolism in Ent3 while carbon fixation in photosynthetic organisms and methane metabolism were increased in Ent1 and 2. Previous reports have indicated that aminobenzoate metabolism influences the microbial composition and intermediaries of aminobenzoate degradation are able to enhance a stress response20 and promote the growth and virulence of Enterobacteriaceae21. Similarly, the metabolism of aromatic amino acid such as tryptophan, over-represented in Ent3, has been documented for the noxious potential of derivative compounds issued from tryptophan degradation on the immune system and neuronal functions22,23.
Ent3 harboured less functions classified in the biosynthesis of ansamycins, butirosin and neomycin. Neomycin is a well-characterized antibiotic with excellent activity against Gram-negative bacteria. This could partly explain the Escherichia/Shigella specificity of Ent3.
A main finding of our study is the robust link between the cumulative number of colonizing parasites and the increase gut microbial diversity and richness. An association between Blastocystis colonization and microbiota diversity has previously been described in both westernized and developing countries12,13. In more than 600 healthy European (Flemish adults cohort), Tito et al. reported a higher bacterial diversity associated with the detection of Blastocystis. Moreover, Blastocystis prevalence was unevenly distributed over Westernized country-type enterotypes, with the Bacteroides clusters comprising significantly less Blastocystis-carriers compared with both Ruminococcaceae and Prevotella enterotypes. This is in line with our observation since > 75% of our cohort was colonized by Blastocystis and we would not depict a Bacteroides cluster in this population, underlying the link between this enterotype and the absence of Blastocystis. We here highlight for the first time that gut colonization by a multi-species community of parasites is associated with higher microbiota richness, increasing with the number of different parasites (from 1 to more than 5 different parasites species). Interestingly, while microbial richness increased with the number of parasites colonizing the gut, this was inversely associated with proportions of Lactobacillus, which were significantly increased in yogurt consumers. Hence, one could speculate that yogurt consumption might, in this specific population, be disadvantageous for parasitic colonization and microbiome diversity and richness. An impact of Lactobacillus on fungi and parasites growth was previously described, mostly in the vagina, as a protective therapeutic option against candidiasis24. A key role of hydrogen peroxide and lactic acid production in this antifungal function is well documented. However, lactic acid was not detected in the metabolomic dataset. At the functional level, higher parasites carriage was associated with enrichment of metabolic activities related to replication and repair, translation and transcription. On the opposite, xenobiotics biodegradation and metabolism (amino acid metabolism, aminobenzoate and chlorocyclohexane/chlorobenzene degradation) were enriched in individuals harbouring low parasite diversity.
Finally, our results strengthen the observation made by Audebert et al.8 on a potentially positive association between parasitic colonization and microbial diversity and richness. The features observed in Ent3 individuals, at both compositional and functional levels, emphasize potential health complication notably gut inflammation.
Eventhough one cannot presently decipher the causal link between these two types of microorganisms, presence of gut parasites may partly be responsible or associated with the specific microbiome described in this population. While Pasolli et al. suspected a role of the nutrition to explain the very low proportion of Bacteroides, toward increased proportions of Proteobacteria, mostly Enterobacteriaceae and Succinivibrionaceae (previously undescribed bacterial genus Succinatimonas)16, we demonstrate that parasitism is a very strong driver of the human gut microbiome composition, diversity and functionality, which overtake the impact of diet.
Conclusion
On the overall, our study highlights the existence of three specific enterotypes in the Malagasy population. Whereas diet, as previously described in other reports, is one of the most important drivers of the gut microbiota composition, individual protozoans carriage (understudied in westernized countries) accounts for most of the dissimilarity observed in the microbiome of people living in Madagascar. Further studies are needed to understand whether gut microbial richness constitute a predilection niche for protozoans colonization, probably due to their gazing features, or whether the parasites themselves induce a higher bacterial richness. Animal studies may help answering this question.
Materials and methods
Study population and design
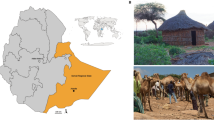
The study was conducted prospectively from February to June 2015 in Mahajanga, North-West Madagascar, and its rural neighbourhood14. The region is characterized by tropical climate. Fecal samples from 219 Malagasy individuals, either healthy volunteers or patients hospitalized in the gastroenterology department of the university hospital PZaGa of Mahajanga, were collected. Two grams of feces were aliquoted and stored at − 20 °C before being routed to France in dry ice. Physicians concomitantly documented individuals’ socioeconomic, demographic and medical information, as well as dietary habits from each individual (Table 1). Exclusion criteria included the use of antibiotic or probiotic 2 months prior to inclusion and/or an age under 18 year-old.
Data are summarized according to protozoan carriage. Individuals' age and BMI are expressed as the mean value computed for each groups. Sex is expressed as the male to female ratio. Protozoan detection, dietary habits, sanitary levels and geographic areas are expressed as the percentage of individuals encountered in each group of parasites load. Information related to the geographic areas can be found in Greigert et al.14.
Bacterial diversity and composition assessment by high-throughput sequencing
After a bead-beating step, total DNA was extracted from 200 mg of feces using the DNA Stool minikit (Qiagen) according to manufacturer recommendations. Bacterial load was assessed by quantitative PCR of the Bacteria domain as previously described25. The V3–V4 region of the 16S rRNA gene was amplified with the following primers: V3F «TACGGRAGGCAGCAG» (bac339F modified from26) and V4R «GGACTACCAGGGTATCTAAT» bac806R. 16S rDNA amplicon libraries were sequenced at GENOTOUL (https://www.genotoul.fr) on a MiSeq device using the 2 × 250 bp V3 kit. Remaining adapter/primer sequences were trimmed and reads were checked for quality (≥ 20) and length (≥ 200 bp) using cutadapt27. Reads were further corrected for known sequencing error using SPAdes28 and then merged using PEAR29. OTUs were identified using a Vsearch pipeline30 set up to dereplicate (–derep_prefix –minuquesize 2), cluster (–unoise3), chimera check (uchime3_denovo) the merged reads. OTU taxonomical classification was performed using both classifier and seqmatch from RDPTools suit31. The training sequence data set required for OTU classification at species level was downloaded from EzBioCloud 16S database for QIIME32. Metagenomic profile inference from 16S dataset was determined using PICRUSt33.
Parasitological examination
All fecal samples were processed in the 30 min following stool emission using both culture (Dobell and Laidlaw biphasic medium) and culture-independent approaches as previously described14. Briefly, culture-independent assays relied on microscopy examinations of both non- and concentrated faeces (MIF and Faust methods). Molecular diagnosis was carried out for Blastocystis, D. fragilis, Cryptosporidium, E. bieneusi and E. intestinalis. We discriminated between E. dispar and E. histolytica by using conventional PCR. Detailed information, including primer sequences, can be found in Greigert et al.14. Stool samples were considered positive if helminth eggs, larvae, cysts and/or trophozoites of protozoans were detected by at least one of the four conventional methods and/or a positive molecular diagnosis.
Microbial metabolomics
Eighteen fecal samples were selected for metabolomics analysis by Profilomic (http://www.profilomic.com/fr). Aliquots (50 mg) were suspended in 50 µL of water and 400 µL of methanol, centrifuged at 4 °C and 10.000g for 15 min and precipitated for 1 h 30 at 4 °C. Following second centrifugation and cleaning, supernatant was evaporated under N2 atmosphere. The pellet was suspended in 250 µL mix of water and AcN (40/60) containing internal standards. The solution was centrifuged at 20,000g and 4 °C for 15 min and injected in the mass spectrometer (Q-Exactive—ThermoFisher Scientific) coupled to liquid chromatography (aSequant ZIC-pHILIC—Merck) machine (LC-HRMS). The analysis was run at high resolution (70,000 FWHM) with alternating negative and positive ionisation mode. Signals were annotated using mass and retention time from each spectrum and a metabolite database issued from Profilomic.
Statistics
Statistical analyses were run using R programming language and software together with gplots, gdata, vegan (http://cran.r-project.org/package=vegan), ade4 and phangorn34 packages. OTU counts were normalized via simple division to their sample size and then multiplication by the size of the smaller sample. α-Diversity and richness were estimated using diversity and estimateR. Distance matrix for β-diversity analysis was computed using vegdist and Bray–Curtis method. Principle Component Analysis and Principle Coordinate Analysis were computed either on compositional data or on distance matrix data using dudi.pca and dudi.pco respectively. Effects of demography, diet, presence of parasite on β-diversity were assessed using adonis function together with Bray–Curtis distance matrix. Wilcoxon test, Kruskal Willis test and anova were used as required to detect differences between groups of variables. P-values were corrected as necessary using False Discovery Rate correction. Enterotypes were defined as previously published15.
Ethics approval and consent to participate
This study was approved by the Malagasy Ministry of Public Health (reference number 33-MSANP/CE). Written informed consents were obtained from all participants. This study was conducted in accordance with the Code of Ethics of the World Medical Association (Declaration of Helsinki).
Data availability
Data are available from the corresponding author. 16S rRNA gene reads are publicly available from NCBI SRA under Bioproject accession number PRJNA600229.
References
De Filippo, C. et al. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Proc. Natl. Acad. Sci. USA 107(33), 14691–14696 (2010).
Yatsunenko, T. et al. Human gut microbiome viewed across age and geography. Nature 486(7402), 222–227 (2012).
Schnorr, S. L. et al. Gut microbiome of the Hadza hunter-gatherers. Nat. Commun. 5, 3654 (2014).
Martinez, I. et al. The gut microbiota of rural papua new guineans: Composition, diversity patterns, and ecological processes. Cell Rep. 11(4), 527–538 (2015).
Kim, J. J. & Khan, W. I. Goblet cells and mucins: Role in innate defense in enteric infections. Pathogens 2(1), 55–70 (2013).
Certad, G., Viscogliosi, E., Chabe, M. & Caccio, S. M. Pathogenic mechanisms of cryptosporidium and giardia. Trends Parasitol. 33(7), 561–576 (2017).
Chabe, M., Lokmer, A. & Segurel, L. Gut protozoa: Friends or foes of the human gut microbiota?. Trends Parasitol. 33(12), 925–934 (2017).
Audebert, C. et al. Colonization with the enteric protozoa Blastocystis is associated with increased diversity of human gut bacterial microbiota. Sci. Rep. 6, 25255 (2016).
Beghini, F. et al. Large-scale comparative metagenomics of Blastocystis, a common member of the human gut microbiome. ISME J. 11(12), 2848–2863 (2017).
Forsell, J. et al. The relation between Blastocystis and the intestinal microbiota in Swedish travellers. BMC Microbiol. 17(1), 231 (2017).
Nieves-Ramírez ME, Partida-Rodríguez O, Laforest-Lapointe I, Reynolds LA, Brown EM, Valdez-Salazar A, Morán-Silva P, Rojas-Velázquez L, Morien E, Parfrey LW et al Asymptomatic Intestinal Colonization with Protist Blastocystis Is Strongly Associated with Distinct Microbiome Ecological Patterns. mSystems 3, Jun 26;3(3):e00007–18 (2018)
Tito RY, Chaffron S, Caenepeel C, Lima-Mendez G, Wang J, Vieira-Silva S, Falony G, Hildebrand F, Darzi Y, Rymenans L, et al. Population-level analysis of Blastocystis subtype prevalence and variation in the human gut microbiota. Gut. Jul;68(7):1180–1189 (2019).
Andersen, L. O. & Stensvold, C. R. Blastocystis in health and disease: Are we moving from a clinical to a public health perspective?. J. Clin. Microbiol. 54(3), 524–528 (2016).
Greigert, V. et al. Human intestinal parasites in Mahajanga, Madagascar: The kingdom of the protozoa. PLoS ONE 13(10), e0204576 (2018).
Arumugam, M. et al. Enterotypes of the human gut microbiome. Nature 473(7346), 174–180 (2011).
Pasolli, E. et al. Extensive unexplored human microbiome diversity revealed by over 150,000 genomes from metagenomes spanning age, geography, and lifestyle. Cell 176(3), 649–662.e620 (2019).
Wu, G. D. et al. Linking long-term dietary patterns with gut microbial enterotypes. Science 334(6052), 105–108 (2011).
Sonnenburg, E. D. & Sonnenburg, J. L. Starving our microbial self: The deleterious consequences of a diet deficient in microbiota-accessible carbohydrates. Cell Metab. 20(5), 779–786 (2014).
Tap, J. et al. Towards the human intestinal microbiota phylogenetic core. Environ. Microbiol. 11(10), 2574–2584 (2009).
Eloe-Fadrosh, E. A. & Rasko, D. A. The human microbiome: From symbiosis to pathogenesis. Annu. Rev. Med. 64, 145–163 (2013).
Freestone, P. P., Walton, N. J., Haigh, R. D. & Lyte, M. Influence of dietary catechols on the growth of enteropathogenic bacteria. Int. J. Food Microbiol. 119(3), 159–169 (2007).
Jaglin, M. et al. Indole, a signaling molecule produced by the gut microbiota, negatively impacts emotional behaviors in rats. Front. Neurosci. 12, 216 (2018).
Natividad, J. M. et al. Impaired aryl hydrocarbon receptor ligand production by the gut microbiota is a key factor in metabolic syndrome. Cell Metab. 28(5), 737-749.e734 (2018).
Verdenelli, M. C. et al. Evaluation of antipathogenic activity and adherence properties of human Lactobacillus strains for vaginal formulations. J. Appl. Microbiol. 116(5), 1297–1307 (2014).
Furet, J. P. et al. Comparative assessment of human and farm animal faecal microbiota using real-time quantitative PCR. FEMS Microbiol. Ecol. 68(3), 351–362 (2009).
Wilson, K. H., Blitchington, R. B. & Greene, R. C. Amplification of bacterial 16S ribosomal DNA with polymerase chain reaction. J. Clin. Microbiol. 28(9), 1942–1946 (1990).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. 17(1), 10-12 (2011).
Bankevich, A. et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. J. Comput. Mol. Cell Biol. 19(5), 455–477 (2012).
Zhang, J., Kobert, K., Flouri, T. & Stamatakis, A. PEAR: A fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics 30(5), 614–620 (2014).
Rognes, T., Flouri, T., Nichols, B., Quince, C. & Mahe, F. VSEARCH: A versatile open source tool for metagenomics. PeerJ 4, e2584 (2016).
Cole, J. R. et al. The Ribosomal Database Project: Improved alignments and new tools for rRNA analysis. Nucleic Acids Res. 37(1), D141–D145 (2009).
Yoon, S. H. et al. Introducing EzBioCloud: A taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int. J. Syst. Evol. Microbiol. 67(5), 1613–1617 (2017).
Langille, M. G. et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat. Biotechnol. 31(9), 814–821 (2013).
Schliep, K. P. phangorn: Phylogenetic analysis in R. Bioinformatics 27(4), 592–593 (2011).
Acknowledgements
We thank the volunteers for participating to our study and providing all necessary samples and information. We are grateful to the INRAE MIGALE bioinformatics facility (MIGALE, INRAE, 2020. Migale bioinformatics Facility, https://doi.org/10.15454/1.5572390655343293E12) for providing computing and storage resources. We also thank the Genotoul Get-PlaGe sequencing platform.
Author information
Authors and Affiliations
Contributions
P.P., A.A.B., P.L. and E.C. were involved in the study concept and design, data interpretation, drafting of the manuscript, and supervision of study. S.M. and V.G. were involved in acquisition, analysis and interpretation of data and drafting of the manuscript. J.B., C.N., J.L.R., E.M., R.R., K.L.R., C.M. and M.L. were involved in the acquisition of clinical and microbiota data. M.R. was involved in samples management prior to shipment to France. S.M., P.P., V.G. and P.L. performed statistical analysis. All authors agree with the content of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mondot, S., Poirier, P., Abou-Bacar, A. et al. Parasites and diet as main drivers of the Malagasy gut microbiome richness and function. Sci Rep 11, 17630 (2021). https://doi.org/10.1038/s41598-021-96967-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-96967-4
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.