Abstract
Methods to spatially induce apoptosis are useful for cancer therapy. To control the induction of apoptosis, methods using light, such as photochemical internalization (PCI), have been developed. We hypothesized that photoinduced delivery of microRNAs (miRNAs) that regulate apoptosis could spatially induce apoptosis. In this study, we identified pre-miR-664a as a novel apoptosis-inducing miRNA via mitochondrial apoptotic pathway. Further, we demonstrated the utility of photoinduced cytosolic dispersion of RNA (PCDR), which is an intracellular RNA delivery method based on PCI. Indeed, apoptosis is spatially regulated by pre-miR-664a and PCDR. In addition, we found that apoptosis induced by pre-miR-664a delivered by PCDR was more rapid than that by lipofection. These results suggest that pre-miR-664a is a nucleic acid drug candidate for cancer therapy and PCDR and pre-miR-664a-based strategies have potential therapeutic uses for diseases affecting various cell types.
Similar content being viewed by others
Introduction
Apoptosis is a programmed cell death that elicits no inflammatory responses. Thus, methods to spatially induce apoptosis can be useful for treatments such as cancer therapy1. To control the induction of apoptosis, methods that use light, such as photodynamic therapy (PDT) and photochemical internalization (PCI), have been developed. For instance, PDT, which uses a combination of a photosensitizer and harmless light to generate reactive oxygen species, induces photo-dependent apoptosis2,3. Methods combining PDT and small interfering RNA (siRNA) have also been developed to obtain synergistic effects4,5.
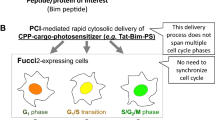
PCI, which uses a photosensitizer and harmless light, delivers targeted molecules such as drugs, peptides, and nucleic acids into the cytosol6,7,8. We developed a PCI-based apoptosis-inducing molecule, TatBim-Alexa. This molecule, which is a conjugate of the Tat cell-penetrating peptide, the BH3 domain from the apoptosis-inducing protein Bim, and the photosensitizer Alexa, can spatially control apoptosis, depending on photo-activation9,10.
MicroRNAs (miRNAs) have been attracting attention as nucleic acid drugs11. miRNAs are small non-coding RNAs that play important roles in the posttranscriptional regulation of gene expression by targeting the 3′-untranslated regions (UTRs) of mRNA12. Primary-miRNA (pri-miRNA), which consists of a stem-loop structure, is transcribed by RNA polymerase II, and cleaved by Drosha-DiGeorge Syndrome Critical Region 8 (Drosha-DGCR8) complex to produce precursor miRNA (pre-miRNA)12,13,14. Then, pre-miRNA is exported from the nucleus and cleaved by Dicer to produce 21–23 base pair mature miRNA12,14. Importantly, miRNAs regulate biological and physiological processes such as apoptosis, proliferation, and cell differentiation11,15,16. For instance, miR-15 and miR-16 induce apoptosis by suppressing the anti-apoptotic B cell lymphoma 2 protein17. In addition, miR-466 induces apoptosis by downregulating the Runt-related transcription factor, which is associated with osteoblast differentiation and inhibits tumor growth in vivo18.
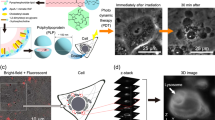
We developed a photoinduced cytosolic dispersion of RNA (PCDR) method, which is an intracellular RNA delivery method8,19,20. The PCDR method is based on the PCI mechanism and can photo-dependently deliver RNA into the cytosol via the endocytic pathway. In other words, the PCDR method can spatially regulate RNA delivery into the cytosol. In this study, we used the PCDR method to deliver apoptosis-inducing miRNA and thereby spatially regulate apoptosis. In addition, we compared the apoptosis induction achieved via the PCDR method and that induced by lipofection.
Methods
Preparation of TatU1A-Alexa546 molecules
TatU1A protein was prepared as described previously19. Purified TatU1A protein and Alexa Fluor 546 C5 (Life Technologies, CA, USA), which contains a thiol-reactive maleimide group, were mixed and then incubated at 25 °C for 1 h. The Alexa Fluor 546-modified TatU1A molecules (TatU1A-Alexa546) were purified in a Centri-Sep spin column (Princeton Separations, NJ, USA) equilibrated with T buffer containing 20 mM Hepes–KOH (pH 7.6), 115 mM NaCl, 5.4 mM KCl, 1.8 mM CaCl2, 0.8 mM MgCl2, and 13.8 mM glucose. Protein concentrations were determined using a Protein Assay Kit (Nacalai Tesque, Kyoto, Japan). The TatU1A labeling efficiency was calculated by measuring Alexa Fluor 546 absorbance. Then, labeling efficiency was adjusted to 35% using unlabeled TatU1A protein.
Preparation of pre-miR-664a, pre-miR-664a-U1A, and pre-miRNA-U1A control
In this study, pre-miR-664a, pre-miR-664a-U1A, and pre-miRNA-U1A control (non-targeting pre-miRNA containing the U1A recognition sequence) were prepared by in vitro transcription. To generate DNA templates for transcription, primer extension was performed using 2 µM of each primer (shown in Table S1) in a reaction mixture containing KOD Dash DNA polymerase (TOYOBO, Osaka, Japan). The resulting template DNA was precipitated with 2-propanol. T7 RNA polymerase was prepared as described previously21. The transcription reaction was carried out at 37 °C for 4 h in a reaction mixture containing 40 mM Tris–HCl (pH 8.0), 24 mM MgCl2, 5 mM dithiothreitol, 10 mM guanosine monophosphate, 2 mM of each NTP, 1.8 U/mL inorganic pyrophosphatase (Sigma, St. Louis, MO, USA), 26.2 µg/mL purified T7 RNA polymerase, and 10 µg/mL DNA template. The pre-miRNA transcripts were purified using an 8% denaturing polyacrylamide gel. All pre-miRNAs were renatured by incubating for 1 min at 85 °C, followed by slow cooling to 4 °C.
The FAM-labeled RNA (RNA-FAM) was purchased from Hokkaido System Science (Hokkaido, Japan). The RNA-FAM sequence was as follows: 5′-GAU UAU GUC CGG UUA UGU ACA UUG CAC UCC GUA CAU AAC CGG ACA UAA UCdT dT-FAM-3′ (the U1A binding sequence is underlined).
Cell culture
HeLa cells were obtained from RIKEN BRC, which participates in the National Bio-Resource Project of the MEXT, Japan. HeLa cells were maintained in RPMI 1640 medium (Nacalai Tesque) with 10% heat-inactivated fetal bovine serum (FBS) (Sigma) and 1% antibiotic–antimycotic solution (Gibco, Gaithersburg, MD, USA) at 37 °C and 5% CO2.
SH-SY5Y cells (DS Pharma Biomedical, Osaka, Japan) were seeded onto collagen-coated dishes (Corning, Cambridge, MA, USA) and maintained in Dulbecco’s modified Eagle’s medium-F12 (1:1) medium (Nacalai Tesque) supplemented with 10% heat-inactivated FBS and 1% penicillin–streptomycin (Gibco) at 37 °C and 5% CO2.
Flp-In-293 cells (Thermo Fisher Scientific, Waltham, MA, USA) were seeded onto collagen-coated dishes and maintained in Dulbecco’s modified Eagle’s medium (Nacalai Tesque) supplemented with 10% heat-inactivated FBS and 1% penicillin–streptomycin at 37 °C and 5% CO2.
Cellular RNA delivery by lipofection
The miRNA-664a-5p mimic and miRNA mimic control were purchased from Dharmacon (Lafayette, CO, USA). The cells were transfected with 10 nM miRNA-664a-5p mimic or 10 nM pre-miR-664a using Lipofectamine RNAiMAX (Invitrogen, Carlsbad, CA, USA) for 24 h at 37 °C and 5% CO2. For control experiments, the cells were also transfected with miRNA mimic control or pre-miRNA-U1A control. The next day, the medium was replaced with fresh growth medium or differentiation medium containing 30 µM retinoic acid (Wako Pure Chemical Industries, Osaka, Japan). The cells were again transfected with miRNA mimics, pre-miR-664a, or pre-miRNA-U1A control in growth medium or differentiation medium after 2 days. After 24 h, the medium was replaced with fresh growth medium or differentiation medium, and the cells were cultured for an additional 2 days.
To induce apoptosis, the cells were transfected with 10 nM pre-miR-664a or pre-miR-664-U1A using Lipofectamine 3000 (Invitrogen) for 24 h at 37 °C and 5% CO2. For control experiments, the cells were also transfected with pre-miRNA-U1A control.
Cellular RNA delivery by PCDR
TatU1A-Alexa546 (2 µM) and pre-miRNA or RNA-FAM (0.2 µM) were mixed in Opti-MEM (Gibco). The cells were grown in 96-well plates and treated for 4 h with TatU1A-Alexa546/RNA complexes prepared as described above. After washing with growth medium, HeLa cells were irradiated with 530–550 nm light at 20 J/cm2 intensity using a mercury arc lamp passed through a MWIG mirror unit and a 40×, 20×, or 4× objective lens using a fluorescence microscope (IX51, Olympus, Tokyo, Japan). SH-SY5Y cells were irradiated with 530–550 nm light at 30 J/cm2. After photo-irradiation, the cells were grown for 24 h at 37 °C and 5% CO2.
Confirming whether the RNA-FAM was delivery efficiently depended on the photo-irradiation energy, the cells were irradiated with 530–550 nm light from 0 to 40 J/cm2.
Detection of apoptotic cells by Nucview488 caspase-3 Assay Kit
Apoptosis was detected using NucView488 caspase-3 Assay Kit (Biotium, Freemont, CA, USA). Apoptotic cells were visualized using an Olympus IX51 fluorescence microscope, and 0.25 µM staurosporine (Wako Pure Chemical Industries) was used as a positive control.
Immunocytochemistry
HeLa cells were transfected with 10 nM pre-miR-664a or pre-miR-664-U1A using Lipofectamine 3000 for 24 h at 37 °C and 5% CO2. Immunocytochemistry was performed as described previously16. Mouse anti-Cytochrome c (dilution 1:300) was purchased from Cell Signaling Technology (Danvers, MA, USA). Alexa 594-conjugated secondary antibody was purchased from Invitrogen, and 4′,6-diamidino-2-phenylindole (DAPI) was obtained from Dojindo. The stained cells were examined using an Olympus IX51 fluorescence microscope.
Mitochondrial permeability transition pore opening
HeLa cells were transfected with 10 nM pre-miR-664a or pre-miR-664-U1A using Lipofectamine 3000 for 20 h at 37 °C and 5% CO2. Transfected HeLa cells were treated with HBSS containing 1 µM calcein-AM (Dojindo, Kumamoto, Japan) and 5 mM cobalt chloride, which quenched the cytosolic and nuclear fluorescence of calcein, for 20 min at 37 °C and 5% CO2. Cells were washed with HBSS, and then were observed using an Olympus IX51 fluorescence microscope.
Detection of TatU1A-Alexa546 in endosomes using Lysotracker green
TatU1A-Alexa546 (2 µM) and pre-miRNA-U1A control (0.2 µM) were mixed in Opti-MEM. The cells were grown in 96-well plates and treated for 4 h with TatU1A-Alexa546/RNA complexes prepared as described above. Opti-MEM containing those complexes was replaced with fresh Opti-MEM containing Lysotracker green (Thermo Fisher Scientific) for 1 h at 37 °C. Cells were washed twice with Opti-MEM, and then observed using an Olympus IX51 fluorescence microscope.
Detection of polarization of mitochondrial membranes
HeLa and SH-SY5Y cells were transfected with 10 nM pre-miRNA-U1A control and pre-miR-664-U1A using Lipofectamine 3000 for 24 h at 37 °C and 5% CO2. To observe the polarization of the mitochondrial membrane, the JC-1 mitochondrial membrane potential assay kit (Cayman Chemical, Michigan, USA) was used according to the manufacturer’s instructions. The fluorescence images of JC-1 aggregates [red] and monomers [green] were examined using a fluorescence microscope (IX-51). The fluorescence intensities were calculated using CellSens software (Olympus).
RNA-FAM incorporation
The cells were transfected with RNA-FAM using Lipofectamine 3000 or PCDR, as described above. RNA-FAM mean fluorescence intensity within transfected cells was determined by flow cytometry using a Guava easyCyte flow cytometer (Merck Millipore, Burlington, MA, USA). For all experiments, data were acquired from 15,000 cells.
To observe endosomal escape in SH-SY5Y and HeLa cells, RNA-FAM taken up by cells was visualized using an Olympus IX51 fluorescence microscope.
Cell viability in cells transfected with miRNA or pre-miRNA
Cell viability was evaluated using the Cell Counting Kit-8 (Dojindo). After transfection, the medium was replaced with fresh medium containing CCK-8 solution. The absorbance at 450 nm was measured to determine cell viability.
Time-lapse imaging of lipofection- and PCDR-induced apoptosis
Time-lapse imaging was used to observe apoptotic cells. To perform the experiment under the same conditions as the PCDR method, HeLa cells were pre-treated for 4 h with Opti-MEM and transfected with 10 nM pre-miRNA using Lipofectamine 3000. Apoptotic cells were observed at 12, 14, 16, 18, 20, 22, and 24 h after the transfection reagent was added.
TatU1A-Alexa546 (2 µM) and pre-miRNAs (0.2 µM) were mixed in Opti-MEM. HeLa cells were treated with TatU1A-Alexa546/RNA complexes for 4 h. After washing, the cells were irradiated with 530–550-nm light at 20 J/cm2 using a mercury arc lamp passed through the MWIG mirror unit and 40 × objective lens of a fluorescence microscope. Apoptotic cells were observed at 4, 6, 8, 10, 12, 14, 16, 18, and 24 h after photo-irradiation.
Statistical analysis
The number of apoptotic cells was counted using CellSens software. All results are expressed as means ± SEM. R22 and EZR23 software were used for the statistical analyses. The number of the experimental replication is shown in the respective figure legends.
Results and discussion
Pre-miR-664a induces apoptosis
We previously showed that miR-664a-3p promotes neuronal differentiation of SH-SY5Y cells16. When we analyzed the neuronal differentiation mechanism induced by miR-664a-5p, interesting results were obtained. Retinoic acid and miR-664a-5p mimic induced neuronal differentiation of SH-SY5Y cells (Fig. 1A). In contrast, cell death was observed in cells transfected with pre-miR-664a, which is the precursor of miR-664a-5p. In addition, pre-miR-664a inhibited cell growth of the cells (Fig. 1B).
Pre-miR-664a inhibits growth and induces cell death in SH-SY5Y cells. (A) Phase contrast images of SH-SY5Y cells transfected with miR-664a-5p, pre-miR-664a, miRNA mimic, or pre-miRNA-U1A control using Lipofectamine RNAiMAX. Retinoic acid induced SH-SY5Y cell differentiation. Scale bars indicate 100 µm. (B) The viability of SH-SY5Y cells transfected with miR-664a-5p, pre-miR-664a, miRNA mimic, or pre-miRNA-U1A control using Lipofectamine RNAiMAX. Cell viability was normalized to the viability of cells treated with miRNA mimic control. Data represent the means ± SEM of three independent experiments.
To confirm whether pre-miR-664a induced apoptosis, apoptotic cells were evaluated using a NucView 488 caspase-3 assay kit, which can detect apoptosis. As shown in Fig. 2A,B, the ratio of apoptotic cells was significantly increased in SH-SY5Y cells transfected with pre-miR-664a using Lipofectamine 3000 compared to those transfected with pre-miRNA-U1A control (0.23 vs. 0.04, respectively). The ratio of apoptotic cells was also significantly increased in HeLa cells transfected with pre-miR-664a compared to the pre-miRNA-U1A control (0.53 vs. 0.06, respectively). In addition, SH-SY5Y and HeLa cell growth was inhibited by pre-miR-664a treatment, compared to pre-miRNA-U1A control (Fig. 2C).
pre-miR-664a delivered by lipofection induces apoptosis. (A) SH-SY5Y and HeLa cells were transfected with pre-miRNA-U1A control or pre-miR-664a using Lipofectamine 3000. Apoptotic cells were detected using NucView 488 caspase assay kit. Staurosporine was used as a positive control. PC phase contrast images. Scale bars indicate 100 µm. (B) Apoptosis efficiency of SH-SY5Y (black bar) and HeLa cells (white bar) are shown. Data represent the means ± SEM of five independent experiments. *P < 0.01; two-way ANOVA and Tukey’s multiple comparisons test. (C) The growth of SH-SY5Y (black bar) and HeLa cells (white bar) transfected with pre-miRNA-U1A control or pre-miR-664a using Lipofectamine 3000. Data represent the means ± SEM of five independent experiments. *P < 0.05; P-values were calculated using two-way ANOVA and Tukey’s multiple comparisons test.
Nucview488 detects only the final caspase3/7 activity. Therefore, apoptosis of the cells treated with pre-miR-664a was detected using JC-1 dye reagent, which detects depolarization of the mitochondrial membrane. As shown in Fig. S1, the JC-1 fluorescence ratio in SH-SY5Y cells treated with pre-miR-664a was lower compared to that in the cells treated with pre-miRNA-U1A control. In addition, the JC-1 fluorescence ratio in HeLa cells treated with pre-miR-664a was also lower compared to that in the cells treated with pre-miRNA-U1A control (Fig. S1), suggesting that pre-miR-664a induced apoptosis.
Furthermore, we observed cytochrome c (cyt-c) localization and measured mitochondrial permeability transition pore (MPTP) opening, to detect apoptosis induced by pre-miR-664a. Cyt-c localizes into the intermembrane space of mitochondrial membrane under the standard condition, and cyt-c releases from the mitochondria to the cytosol due to MTPT opening during apoptosis24,25. To observe cyt-c localization, we performed immunostaining of cyt-c. Cyt-c in the cells treated with pre-miRNA-U1A control was observed as dots in the cytoplasm (Fig. S2). In contrast, cyt-c in the cells treated with pre-miR-664a was diffused in the cytoplasm, and the nuclear fragmentation and condensation were observed. To determine whether pre-miR-664a induced MPTP opening, MPTP opening was measured by calcein/Co2+-quenching method. The fluorescence intensity of calcein-AM in the cells treated with pre-miR-664a was lower than that in the cells treated with pre-miRNA-U1A control, suggesting that pre-miR-664a induces MPTP opening (Fig. S3). These results suggest that pre-miR-664a induced MPTP opening to promote the release of mitochondrial cyt-c. In summary, these results suggest that pre-miR-664a is a novel miRNA that induces apoptosis via mitochondrial apoptotic pathway.
Pre-miR-664a generates miR-664a-5p and miR-664a-3p. Previous reports showed that miR-664a-5p promotes neuronal differentiation of SH-SY5Y cells and promotes osteogenic differentiation of human bone marrow-derived mesenchymal stem cells16,26. miR-664a-3p promotes cell proliferation in gastric cancer cells27. In addition, apoptosis induction by miR-664a-3p or miR-664a-5p alone has not been reported. Therefore, both miR-664a-3p and miR-664a-5p are likely to be important for the induction of apoptosis.
Regarding potential pre-miR-664a targets relating to apoptotic pathway, siRNA-mediated knockdown of high mobility group AT-hook 2 (HMGA2), which is a target of miR-664a-5p, induces apoptosis in PC3 and DU145 cells26,28. Therefore, apoptosis may be induced by miR-664a-5p-mediated down-regulation of HMGA2. In addition, caspase 3/7 activity of HMGA2- and checkpoint kinase 1 (CHK1)-knockdowned cells is higher than that of HMGA2-knockdowned cells29. CHK1 regulates cell cycle progression and cell cycle arrest induced by DNA damage29. The targetscan7.2 software30 predicts that CHK1 is not a target of miR-664a-3p, however multiple proteins regulated cell cycle progression and cell cycle checkpoint such as cyclin A are target. Hence, pre-miR-664a may induce apoptosis via down-regulations of HMGA2 by miR-664a-5p and cell cycle progression regulated by miR-664a-3p.
Pre-miR-664a-U1A induces apoptosis
We attempted to control region-specific apoptosis induction by combining pre-miR-664a and the PCDR method. However, the PCDR method using the RNA carrier U1A protein can only deliver RNA containing a U1A recognition sequence19. Therefore, we prepared pre-miR-664-U1A, where the pre-miR-664a hairpin structure was replaced with the U1A recognition sequence (Fig. 3A). To confirm whether pre-miR-664-U1A induced apoptosis, we elevated apoptosis of HeLa cells and SH-SY5Y cells transfected with pre-miR-664-U1A using Lipofectamine 3000. As shown in Fig. 3B,C, pre-miR-664-U1A induced apoptosis in HeLa cells (0.77) and SH-SY5Y cells (0.33). In addition, apoptosis efficiency by pre-miR-664a was similar to that by pre-miR-664a-U1A in HeLa and SH-SY5Y cells (Fig. S4). These results indicate that substituting U1A recognition sequence in pre-miR-664a has hardly any effect on apoptosis induction.
Pre-miR-664a-U1A delivered by lipofection induces apoptosis. (A) The pre-miR-664a and pre-miR-664a-U1A sequences are shown. The underlined sequence shown in pre-miR-664a-U1A is replaced with the U1A binding sequence. The red font sequence indicates miR-664a-5p. The light blue font sequence indicates miR-664a-3p. (B) SH-SY5Y and HeLa cells were transfected with pre-miRNA-U1A control or pre-miR-664a-U1A using Lipofectamine 3000. Apoptotic cells were detected using NucView 488 caspase assay kit. Staurosporine was used as a positive control. PC phase contrast images. Scale bars indicate 100 µm. (C) Apoptosis efficiency of SH-SY5Y (black bar) and HeLa cells (white bar). The data represent the means ± SEM of five independent experiments. *P < 0.01; P-values were calculated using two-way ANOVA and Tukey’s multiple comparisons test.
Pre-miR-664a-U1A delivered by PCDR method induces apoptosis
We next attempted to induce photo-dependent apoptosis by combining pre-miR-664a-U1A and the PCDR method in HeLa cells and SH-SY5Y cells. The cells transfected with pre-miR-664a-U1A using PCDR showed photo-dependent apoptosis (Fig. 4A,B). In contrast, without photo-irradiation, pre-miR-664a-U1A did not induce apoptosis in these cells. In addition, the cells transfected with pre-miRNA-U1A control by PCDR did not show apoptosis. These results indicate that pre-miR-664a-U1A delivered by the PCDR method induces photo-dependent apoptosis.
pre-miR-664a-U1A delivered by photoinduced cytosolic dispersion of RNA (PCDR) induces apoptosis. (A) pre-miRNA-U1A control or pre-miR-664a-U1A were delivered into SH-SY5Y and HeLa cells using PCDR. Then, the cells were photo-irradiated. Apoptotic cells were detected using NucView 488 caspase assay kit. Staurosporine was used as a positive control. PC: Phase contrast images. Scale bars indicate 100 µm. (B) Apoptosis efficiency of SH-SY5Y (black bar) and HeLa cells (white bar). Data represent the means ± SEM of five independent experiments. *P < 0.01; P-values were calculated using two-way ANOVA and Tukey’s multiple comparisons test.
Cell type-dependency of transfection efficiency of PCDR and lipofection methods
After determining that pre-miR-664a-U1A delivered by PCDR induces apoptosis, we evaluated whether the PCDR delivery method was more efficient than transfection using lipofectamine. Using the lipofection method, apoptosis in HeLa cells was higher than that in SH-SY5Y cells (Fig. 2B). However, apoptosis in HeLa cells was almost the same as in SH-SY5Y cells using the PCDR method (Fig. 4B). We speculate that apoptosis efficiency may depend on the RNA concentration delivered into the cells, which is a consequence of differences in the RNA delivery methods. In other words, the RNA concentration delivered into HeLa cells using the lipofection method may be higher than the RNA concentration delivered into SH-SY5Y cells, while the RNA concentration delivered via PCDR is similar in HeLa and SH-SY5Y cells. Therefore, we used flow cytometry to measure the incorporation of FAM-labeled RNA (RNA-FAM) into HeLa and SH-SY5Y cells by lipofection. The RNA-FAM fluorescence intensity in HeLa cells after lipofection was similar to the fluorescence intensity in SH-SY5Y cells (Fig. 5A). This suggests that the concentration of RNA-FAM delivered into HeLa and SH-SY5Y cells was similar, which cannot explain the difference in pre-miR-664a-U1A/lipofectamine-induced apoptosis between HeLa and SH-SY5Y cells.
RNA-FAM incorporation in the cytosol using Lipofectamine 3000 and photoinduced cytosolic dispersion of RNA (PCDR). (A) The fluorescence intensity of RNA-FAM delivered into SH-SH5Y and HeLa cells using Lipofectamine 3000 was measured using flow cytometry. Data represent the means ± SEM of three independent experiments. The fluorescence intensity of HeLa cells transfected with RNA-FAM was defined as 1.0. (B) RNA-FAM was transfected into SH-SY5Y and HeLa cells using Lipofectamine 3000. RNA-FAM images were obtained after transfection. PC phase contrast images. Scale bars indicate 100 µm. (C) The endosomal escape efficiencies of RNA-FAM were calculated by counting the number of the cells in which FAM fluorescence was dispersed within the cytosol (NF) and the number of the cells (NT). The endosomal escape efficiency was defined as NF/NT. Data represent the means ± SEM of four independent experiments. *P < 0.05; P-values were calculated using one-way ANOVA and Dunnett’s test. (D) The fluorescence intensity of RNA-FAM delivered by PCDR into SH-SH5Y and HeLa cells was measured using flow cytometry. The fluorescence intensity of HeLa cells irradiated at 0 J/cm2 was defined as 1.0. Data represent the means ± SEM of five independent experiments. *P < 0.05; P-values were calculated using one-way ANOVA and Tukey’s test. (E) RNA-FAM was delivered by PCDR into the cytosol of SH-SY5Y and HeLa cells. RNA-FAM images were obtained before and after photoirradiation. SH-SY5Y and HeLa cells were irradiated at 30 J/cm2 and 20 J/cm2, respectively. PC Phase contrast images. Scale bars indicate 50 µm. (F) The endosomal escape efficiencies of TatU1A-Alexa546/RNA complexes were calculated by counting the number of the cells in which FAM fluorescence was dispersed within the cytosol after photoirradiation (NF) and the number of the cells (NT). The endosomal escape efficiency was defined as NF/NT. Data represent the means ± SEM of four independent experiments. *P < 0.05, not significant (n.s); P-values were calculated using one-way ANOVA and Dunnett’s test.
Transfection efficiency by cationic lipids depends on the cell type31. Cationic lipids, including lipofectamine, form polyplexes and deliver RNA into the cells via the endocytic pathway. Therefore, the lipofectamine/RNA-FAM complex might be trapped in endosomes in a cell type-dependent manner. As shown in Fig. 5B,C, cytosolic RNA-FAM dispersion in HeLa cells was more efficient than in SH-SY5Y cells. These results suggest that the difference in cell type-dependent apoptosis efficiency after lipofection depends on the efficiency of RNA delivery into the cytosol rather than the whole cell, which includes endosomes.
The TatU1A/RNA complex enters cells via the endocytic pathway19,32. We confirmed that TatU1A-Alexa546/pre-miRNA-U1A control complexes accumulated into endosomes in SH-SY5Y and HeLa cells before photo-irradiation using Lysotracker green. Lysotracker green detects acidic membranous structures such as endosomes and lysosomes. These complexes were colocalized with Lysotracker green in SH-SY5Y and HeLa cells (Fig. S5), suggesting that TatU1A-Alexa546/pre-miRNA-U1A control complexes accumulated into endosomes via the endocytic pathway.
Singlet oxygen is generated from Alexa Fluor 546 during photo-irradiation, which allows the complex to escape from endosomes and disperse throughout the cytosol32. Therefore, the RNA concentration delivered into the cytosol via the PCDR method may be dependent on the applied light energy. We first examined whether the RNA concentration delivered into the cytoplasm was dependent on light energy. As shown in Fig. 5D, RNA-FAM fluorescence in HeLa cells was increased by photo-irradiation, and peaked at 20 J/cm2. RNA-FAM fluorescence in SH-SY5Y cells was slightly increased by photo-irradiation (Fig. 5D). In addition, the maximum RNA-FAM fluorescence intensity in HeLa cells was similar to that in SH-SY5Y cells, suggesting that the RNA concentration delivered into HeLa cells was approximately the same as that in SH-SY5Y cells. RNA-FAM fluorescence intensity increased after the TatU1A/RNA complex escaped from the endosome32. The increased RNA-FAM fluorescence intensity after photo-irradiation suggests that RNA-FAM was dispersed into the cytosol by photo-irradiation (Fig. 5D).
To observe the escape of RNA-FAM from endosomes, RNA-FAM was delivered into the cell by the PCDR method and visualized using a fluorescence microscope (Fig. 5E). RNA-FAM endosomal escape was observed in HeLa and SH-SY5Y cells (Fig. 5E,F). These results indicate that cell type-independent apoptosis by the PCDR method occurs because endosomal escape occurs in both HeLa and SH-SY5Y cells.
The TatU1A-Alexa546/pre-miRNA complex delivered by the PCDR method enters cells via the endocytic pathway. Singlet oxygen photogenerated from Alexa Fluor 546 facilitates the escape of the complex from endosomes32. Therefore, the endosomal escape efficiency induced by the PCDR method might be cell type-independent, unlike the lipofection method.
In summary, apoptosis induced by lipofection is cell type-dependent because RNA delivery into the cytosol depended on the cell type. In contrast, apoptosis induced by PCDR is cell type-independent because RNA delivery into the cytosol did not depend on the cell type.
Apoptosis is more rapidly induced by PCDR than by the lipofection method
Our previous study showed that the concentration of cytoplasmic RNA delivered by PCDR rapidly increased compared to RNA delivery by lipofection (Lipofectamine 3000)20. Therefore, to examine whether apoptosis induced by PCDR occurred faster than that by lipofection, we observed apoptotic cells using time-lapse imaging. Apoptosis of cells transfected with pre-miR-664a-U1A via lipofection increased in a time-dependent manner, and was significantly induced at 18 h compared to pre-miRNA-U1A control (Fig. 6A). In contrast, apoptosis caused by pre-miR-664a-U1A and PCDR was significantly induced at 6 h compared to pre-miRNA-U1A control (Fig. 6B). Further, apoptosis was observed at 12 h in cells transfected with pre-miR-664a-U1A by PCDR, but not lipofectamine (Fig. 6A,B). Thus, apoptosis induced by PCDR occurred more rapidly than lipofection-mediated apoptosis.
Ratio of time-dependent apoptosis in cells transfected by photoinduced cytosolic dispersion of RNA (PCDR) or lipofection. HeLa cells were transfected with pre-miRNA-U1A control or pre-miR-664a-U1A using Lipofectamine 3000 (A) or PCDR (B). Apoptotic cells were detected at 12, 14, 16, 18, 20, 22, and 24 h (lipofection) or 4, 6, 8, 10, 12, 14, 16, 18, and 24 h (PCDR). The ratio of apoptotic cells treated with pre-miRNA-U1A control by lipofection (gray circles), pre-miR-664a-U1A by lipofection method (gray squares), pre-miRNA control by PCDR (black circles), and pre-miR-664a-U1A by PCDR (black squares). Data represent the means ± SE of four (PCDR) or five (lipofection) independent experiments. *P < 0.05, **P < 0.01; P-values were calculated using two-way ANOVA and Tukey’s multiple comparisons test comparing pre-miR-664a-U1A with pre-miRNA-U1A control at each time point.
Region-specific apoptosis induction by the PCDR method
We attempted to demonstrate region-specific apoptosis induction by combining pre-miR-664a-U1A and PCDR. After SH-SY5Y cells were treated with TatU1A-Alexa546/pre-miRNA complexes, the cells in a specific region were irradiated (Fig. 7, Day 1). TatU1A-Alexa546 entered the cytoplasm only within the photo-irradiated region. Twenty-four hours later, apoptosis was detected using NucView 488 (Fig. 7, Day 2). Apoptotic cells were observed only within the photo-irradiated region. Similar experiments were performed using HeLa cells. Apoptosis was induced only in photo-irradiated cells (Fig. 7). These results indicate that apoptosis can be spatially controlled by pre-miR-664a-U1A delivered by PCDR, regardless of cell type.
Region specific apoptosis of cells transfected with pre-miR-664a-U1A. The cells were treated with TatU1A-Alexa546 and pre-miRNA complexes. Then, the cells to the right of the dotted line were irradiated (Day 1). Phase contrast and TatU1A-Alexa546 fluorescence images were immediately obtained. Apoptotic cells were detected using NucView 488 caspase assay kits (Day 2). PC phase-contrast images. Scale bars indicate 100 µm.
Pre-miR-664a induces apoptosis in benign cells, Flp-In-293 cells
SH-SY5Y and HeLa cells, which are malignant cells, were used in the present study. We confirmed that pre-miR-664a induced apoptosis in Flp-In-293 cells, which were created from benign HEK293 cells. As shown in Fig. S6, pre-miR-664a induced apoptosis in Flp-In-293 cells. This result suggests that pre-miR-664a probably induces apoptosis in human cells. Therefore, a method for region-specific delivery of pre-miR-664a, as occurs in the PCDR method, is important for the application of pre-miR-664a in cancer therapy.
Potential therapeutic uses of strategies based on PCDR and pre-miR-664a
The PCI method can effectively deliver targeted molecules to the targeted cells. For instance, pre-clinical studies suggest that PCI can effectively deliver chemotherapeutic agents into tumor cells and improve treatment efficacy6. Thus, a PCI strategy will likely develop as an effective cancer therapy.
The strategy of pre-miR-664a and PCDR based on PCI has potential therapeutic uses for cancer therapy. However, for this strategy to be successful, its problems must be addressed. Naked pre-miR-664a and TatU1A are unstable in blood, and TatU1A-Alexa/pre-miR-664a complexes may be distributed around the whole body because TatU1A has no target specificity. In order to use this method in vivo, subcutaneous or local injections are required. In addition, it will be important to improve the stabilities of pre-miR-664a and TatU1A in blood and the tumor specificity of TatU1A-Alexa546/pre-miRNA complex. For instance, the development of pre-miR-664a with nucleotide modifications33 and TatU1A with modifications34 are important for improving their stability in blood. In order to accumulate TatU1A-Alexa/pre-miR-664a complexes in a tumor, it is necessary to fuse TatU1A with a peptide which accumulates into specific cells, such as RGD peptide35.
We previously developed a nanocarrier named “lactosome”. Lactosome accumulated in mouse tumors through enhanced permeability and retention effects36. In addition, lactosome could photo-dependently deliver siRNA into the cells in vitro37. Therefore, the combination of pre-miR-664a and lactosome may induce apoptosis in a light-dependent manner in vivo.
Conclusions
In this study, we identified pre-miR-664a as a novel apoptosis-inducing miRNA. Furthermore, apoptosis could be spatially regulated by pre-miR-664a-U1A delivered by PCDR. Therefore, pre-miR-664a is a potential nucleic acid drug candidate for cancer therapy. To understand apoptosis induction mechanisms by pre-miR-664a, further analyses of the target genes of pre-miR-664a will be needed.
We demonstrated that while apoptosis efficiency after lipofection depends on the cell type, apoptosis efficiency after PCDR is cell type-independent. Furthermore, apoptosis induced by PCDR occurs more rapidly than that induced by lipofection. The increased apoptosis rate using the PCDR method may be attributed to the rapid dispersion of RNA into the cytosol after photo-stimulation. Therefore, PCDR and pre-miR-664a-based strategies have potential therapeutic uses for diseases affecting various cell types.
References
Xu, X., Lai, Y. & Hua, Z. Apoptosis and apoptotic body: disease message and therapeutic target potentials. Biosci. Rep. 39, BSR20180992 (2019).
Dougherty, T. J. et al. Photoradiation therapy for the treatment of malignant tumors. Cancer Res. 38, 2628–2635 (1978).
Sook, H., Heejun, H., Jieun, S. & Kun, H. Combination of photodynamic therapy (PDT) and anti-tumor immunity in cancer therapy. J. Pharm. Investig. 48, 143–151 (2018).
Lecaros, R. L. G., Huang, L., Lee, T. & Hsu, Y. Nanoparticle delivered VEGF-A siRNA enhances photodynamic therapy for head and neck cancer treatment. Mol. Ther. 24, 106–116 (2016).
Chen, W. et al. Nanoparticle delivery of HIF1α siRNA combined with photodynamic therapy as a potential treatment strategy for head-and-neck cancer. Cancer Lett. 359, 65–74 (2015).
Šoši, L. et al. Photochemical Internalization: Light paves way for new cancer chemotherapies and vaccines. Cancers (Basel) 12, 165 (2020).
Lai, P., Lou, P., Peng, C., Pai, C. & Yen, W. Doxorubicin delivery by polyamidoamine dendrimer conjugation and photochemical internalization for cancer therapy. J. Control. Release 122, 39–46 (2007).
Miyoshi, Y. et al. Endosomal escape of peptide-photosensitizer conjugates is affected by amino acid sequences near the photosensitizer. Bioconjug. Chem. 31, 916–922 (2020).
Watanabe, K., Fujiwara, H., Kitamatsu, M. & Ohtsuki, T. Photoinduced apoptosis using a peptide carrying a photosensitizer. Bioorg. Med. Chem. Lett. 26, 3115–3118 (2016).
Kim, H., Watanabe, S., Kitamatsu, M. & Watanabe, K. Cell cycle dependence of apoptosis photo-triggered using peptide-photosensitizer conjugate. Sci. Rep. 10, 19087 (2020).
Shirjang, S. et al. MicroRNAs in cancer cell death pathways: Apoptosis and necroptosis. Free Radic. Biol. Med. 139, 1–15 (2019).
Brien, J. O., Hayder, H., Zayed, Y. & Peng, C. Overview of microRNA biogenesis, mechanisms of actions, and circulation. Front. Endocrinol. (Lausanne). 9, 402 (2018).
Landthaler, M., Yalcin, A. & Tuschl, T. The human DiGeorge syndrome critical region gene 8 and its D. melanogaster homolog are required for miRNA biogenesis. Curr. Biol. 14, 2162–2167 (2004).
Zeng, Y., Yi, R. & Cullen, B. R. Recognition and cleavage of primary microRNA precursors by the nuclear processing enzyme Drosha. EMBO J. 24, 138–148 (2005).
Hon, K. W., Abidin, S. A. Z., Othman, I. & Naidu, R. Insights into the role of microRNAs in colorectal cancer (CRC) metabolism. Cancers (Basel) 12, 2462 (2020).
Watanabe, K., Yamaji, R. & Ohtsuki, T. MicroRNA-664a-5p promotes neuronal differentiation of SH-SY5Y cells. Genes Cells 23, 225–233 (2018).
Cimmino, A. et al. miR-15 and miR-16 induce apoptosis by targeting BCL2. Proc. Natl. Acad. Sci. USA 102, 13944–13949 (2005).
Colden, M. et al. MicroRNA-466 inhibits tumor growth and bone metastasis in prostate cancer by direct regulation of osteogenic transcription factor RUNX2. Cell Death Dis. 8, e2572 (2017).
Endoh, T., Sisido, M. & Ohtsuki, T. Cellular siRNA delivery mediated by a cell-permeant RNA-binding protein and photoinduced RNA interference. Bioconjug. Chem. 19, 1017–1024 (2008).
Shiraga, K., Soe, T. H., Matsumoto, S., Watanabe, K. & Ohtsuki, T. Red and near-infrared light-directed cytosolic delivery of two different RNAs using photosensitive RNA carriers. Bioconjug. Chem. 29, 3174–3179 (2018).
Miyoshi, Y., Ohtsuki, T., Kashida, H., Asanuma, H. & Watanabe, K. In-stem molecular beacon targeted to a 5′-region of tRNA inclusive of the D arm that detects mature tRNA with high sensitivity. PLoS ONE 14, e0211505 (2019).
R Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/ (2020).
Kanda, Y. Investigation of the freely available easy-to-use software‘EZR’ for medical statistics. Bone Marrow Transpl. 48, 452–458 (2013).
Bossy-Wetzel, E., Newmeyer, D. D. & Green, D. R. Mitochondrial cytochrome c release in apoptosis occurs upstream of DEVD-specific caspase activation and independently of mitochondrial transmembrane depolarization. EMBO J. 17, 37–49 (1998).
Liu, G. et al. Mitochondrial permeability transition and its regulatory components are implicated in apoptosis of primary cultures of rat proximal tubular cells exposed to lead. Arch. Toxicol. 90, 1193–1209 (2016).
Zhang, Y. et al. MicroRNA-664a-5p promotes osteogenic differentiation of human bone marrow-derived mesenchymal stem cells by directly downregulating HMGA2. Biochem. Biophys. Res. Commun. 521, 9–14 (2020).
Wang, L. et al. miR-664a-3p functions as an oncogene by targeting Hippo pathway in the development of gastric cancer. Cell Prolif. 52, 1–14 (2019).
Shi, Z. et al. Silencing of HMGA2 promotes apoptosis and inhibits migration and invasion of prostate cancer cells. J. Biosci. 41, 229–236 (2016).
Natarajan, S., Hombach-Klonisch, S., Dröge, P. & Klonisch, T. HMGA2 inhibits apoptosis through interaction with ATR-CHK1 signaling complex in human cancer cells. Neoplasia 15, 263–280 (2013).
Agarwal, V., Bell, G. W., Nam, J. W. & Bartel, D. P. Predicting effective microRNA target sites in mammalian mRNAs. Elife 4, e05005 (2015).
Wang, T., Larcher, L. M., Ma, L. & Veedu, R. N. Systematic screening of commonly used commercial transfection reagents towards efficient transfection of single-stranded oligonucleotides. Molecules 23, 2564 (2018).
Ohtsuki, T. et al. The molecular mechanism of photochemical internalization of cell penetrating peptide-cargo-photosensitizer conjugates. Sci. Rep. 5, 18577 (2015).
Hu, B. et al. Therapeutic siRNA: state of the art. Signal Transduct. Target. Ther. 5, 101 (2020).
Schumacher, D., Hackenberger, C. P. R., Leonhardt, H. & Helma, J. Current status: Site-specific antibody drug conjugates. J. Clin. Immunol. 36, 100–107 (2016).
Roveri, M., Bernasconi, M., Leroux, J. C. & Luciani, P. Peptides for tumor-specific drug targeting: State of the art and beyond. J. Mater. Chem. B 5, 4348–4364 (2017).
Akahoshi, A. et al. Enhanced cellular uptake of lactosomes using cell-penetrating peptides. Sci. Technol. Adv. Mater. 17, 245–252 (2016).
Lim, M. S. H. et al. Lactosome-conjugated siRNA nanoparticles for photo-enhanced gene silencing in cancer cells. J. Pharm. Sci. 110, 1788–1798 (2021).
Acknowledgements
This work was supported in part by the Okayama Foundation for Science and Technology, Wesco Scientific Promotion Foundation, the NOVARTIS Foundation (Japan) for the Promotion of Science, and the Terumo Life Science Foundation (to KW). JSPS KAKENHI Grant Number JP18H02090 (to T.O.).
Author information
Authors and Affiliations
Contributions
K.W. designed the experiments. K.W., T.N., and R.O. performed experiments. K.W. and T.O. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Watanabe, K., Nawachi, T., Okutani, R. et al. Photocontrolled apoptosis induction using precursor miR-664a and an RNA carrier-conjugated with photosensitizer. Sci Rep 11, 14936 (2021). https://doi.org/10.1038/s41598-021-94249-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-94249-7
This article is cited by
-
Photo-dependent cytosolic delivery of shRNA into a single blastomere in a mouse embryo
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.