Abstract
The emao, a traditional beer starter used in the North–East regions of India produces a high quality of beer from rice substrates; however, its microbial community structure and functional metabolic modules remain unknown. To address this gap, we have used shot-gun whole-metagenome sequencing technology; accordingly, we have detected several enzymes that are known to catalyze saccharification, lignocellulose degradation, and biofuel production indicating the presence of metabolic functionome in the emao. The abundance of eukaryotic microorganisms, specifically the members of Mucoromycota and Ascomycota, dominated over the prokaryotes in the emao compared to previous metagenomic studies on such traditional starters where the relative abundance of prokaryotes occurred higher than the eukaryotes. The family Rhizopodaceae (64.5%) and its genus Rhizopus (64%) were the most dominant ones, followed by Phaffomycetaceae (11.14%) and its genus Wickerhamomyces (10.03%). The family Leuconostocaceae (6.09%) represented by two genera (Leuconostoc and Weissella) was dominant over the other bacteria, and it was the third-highest in overall relative abundance in the emao. The comprehensive microbial species diversity, community structure, and metabolic modules found in the emao are of practical value in the formulation of mixed-microbial cultures for biofuel production from plant-based feedstocks.
Similar content being viewed by others
Introduction
The ethnic beer (or wine) fermentation process with the use of traditional starter cultures1 and without starter cultures (spontaneous)2 continues to be prevalent in many parts of the world that possess sociocultural and scientific values. The microorganisms found in the ethnic beer fermentations are diverse1,2 and are mostly comprised of yeasts, molds, and bacteria. Each type of ethnic beer fermentation produces desirable end-product(s) with unique taste and flavor due to the complex microbial interactions, succession, and metabolite productions during the fermentation process. A proper understanding of the microbial community structure and the role of individual microorganisms associated with an environmental sample is crucial for bioprospection. However, the key microbial ingredients in the emao and the fermentation process in the preparation of several different traditional ethnic beers remain incompletely known.
The analysis of microbial community structure using culture-dependent methods has several limitations, for example, non-cultivability of certain microbes in the laboratory conditions3, competition for nutrients, and dominance of some microbes over the others in culture media, and the different growth kinetics among the species. In contrast, culture-independent methods, such as metagenomics through Next Generation Sequencing (NGS), allow the microbial communities to study without the need for microbial isolation and cultivation3. Although recently PCR-amplicon sequencing of 16S rRNA genes and ITS rDNA regions have been used for the determination of microbial diversity in many environmental samples including traditional starter cultures, this method is vulnerable to flaws4,5 due to the inherent PCR problems including, template competition, primer mismatch, biased amplification due to different template copy number in the analyses of metagenomic DNAs. Alternatively, metagenomic DNAs can be used for whole metagenome sequencing via high throughput NGS platforms, and the data so obtained can be used for protein sequence prediction and subsequent extraction of ribosomal protein (r-protein) sequences for community structure determination of the associated microorganisms. The use of r-protein sequences in taxonomic identification and community structure determination in metagenomics represents a novel methodology in microbial ecology due to the occurrence of a single copy of each ribosomal protein gene in an individual6.
The craft of brewing rice beer (zu or zou) using emao (also written as amao) is an age-old tradition among the Bodo (or Boro) community, one of the earliest settlers of the North-East India7,8. The origin of the Bodo community and their traditional brewing methodology are not clearly documented. The Bodo, Dimasa, and Garo communities of the North East India might have emerged from a common ancestor about 1500 years ago9. There are similarities in their language, culture, and brewing traditions. In the Bodo traditional methodology, the emao is prepared from non-sticky rice powder by adding a combination of specific herbs and a small portion of emao from the existing stock (Fig. 1, Table 1). The traditional reason for using the herbs in emao is to stimulate specific taste and flavor that is unique to Bodo beer, depending on the combination of herbs used. Customarily, 3–5 herbs are used in the preparation of emao, sometimes some of these herbs may not be available at the time of its preparation. Several recent studies have reported the microbial diversity occurring on the other similar traditional starters across the world using both culture-dependent10,11,12,13,14,15 and culture-independent methods16,17,18,19,20,21,22,23,24. Nonetheless, the study of microbial diversity in the emao is limited, as only four fungal species, one mold and three non-Saccharomyces yeasts, have been reported with the use of culture-dependent methods25,26,27. However, bacterial diversity and culture-independent studies as they relate to emao have not been reported in the literature.
Workflow of the traditional method of emao preparation. Rice is used as the base material. The number of herbs might vary as listed in Table 1. Fresh cakes made from mixing rice and herbs are dusted all over with the mother starter powder, thereafter kept covered with a clean cloth or rice straw or fern leaves for 3–4 days. The surface of new cakes turns white due to the growth of microbes. Newly prepared emao cakes are sun-dried for 2–3 days and preserved in bags or bamboo containers in aeration condition.
The occurrence of amylolytic, alcohol-producing and lactic acid bacteria (LAB) that represent three major groups of microorganisms, in several ethnic beer starters has been reported1. In addition, the occurrence of acetic acid bacteria such as Acetobacter orientalis and A. pasteurianus in traditional starter Banh men28, and Gluconobacter sp. in traditional starter Marcha29 were also reported. Thus, in most traditional beer fermentation processes several microorganisms work together in consortia mode, which produces unique beer. The traditional method entails the preparation of starter culture and maintenance; however, the substrate used for fermentation varies from one locality to the next, and one community to the other1. These variations also add diversity to the microbial species that catalyze the production of beer, therefore the fermented products are expected to taste different from one to the other. Close to two dozens of traditional beer starters are found throughout the North East India30, and only a few of them have been studied using culture-independent methods16,22,24. As biofuel-producing microorganisms are important for sustainable biowaste management and energy production, therefore, the identification of biofuel-producing microorganisms is an ongoing effort. The production and use of biofuels derived from organic materials including biowastes are necessary, thereby reducing the CO2 emissions associated with the fossil fuel combustion. The traditional beer starter cultures of Asian origin are already in use for bio-ethanol production from cereals or fruits. Therefore, these starter cultures could potentially serve as a source of other forms of biofuel-producing microorganisms too. Here, for the first time, we report the microbial community structure in the emao based on the ribosomal protein sequences derived from the whole-metagenome sequences and profile the metabolic potentials of emao in relation to biofuel production.
Results
Whole-metagenome overview
The number of nucleotide sequence reads recorded in the emao sample was 13,060,410, which gave a total of 6,530,205 merged reads with ≥ 400 nucleotides per sequence (Table 2). We obtained a total of 1,285,880 amino acid (AA) sequences having ≥ 50 AA per sequence, which was subsequently used as an input for taxonomic binning, enzyme identification, and metabolic module analyses.
Community structure and species diversity
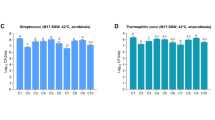
We retrieved a total of 8562 non-redundant ribosomal protein (r-protein) sequences from the module M91000 for all organisms (Table 2) and the taxonomic binning of these r-protein sequences revealed 92% Eukarya and 8% Bacteria with a ratio of 9:2:1 for Mucoromycota (molds), Ascomycota (yeasts), and Firmicutes (bacteria), respectively (Fig. 2a). The relative abundance of molds was the highest (73.44%), followed by yeasts (18.02%) and lactic acid bacteria (LAB) (7.87%) in emao (Fig. 2b). However, the yeast species diversity was four times higher than the bacteria and molds in emao (Supplementary Data Table S1). The relative abundance of the genus Rhizopus and its family Rhizopodaceae were the highest among the groups in emao (Fig. 2c,d). Among the yeasts, the relative abundance of the genus Wickerhamomyces and its family Phaffomycetaceae were the highest.
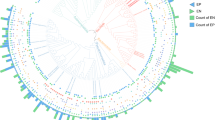
We identified a total of 74 microbial species in emao based on r-protein sequences (Fig. 3, Supplementary Data Table S1). Rhizopus delemar (syn. R. oryzae) was the highest with 56% overall relative abundance (ORA), followed by R. microsporus (7% ORA) and Mucor circinelloides (5% ORA) among the molds. Among the yeasts, Wickerhamomyces anomalus was the most dominant (9% ORA), followed by W. ciferrii (1% ORA), Ascoidea rubescens (0.6% ORA), Cyberlindnera fabiani (0.6%), Pachysolen tannophilus (0.6% ORA), Candida tropicalis (0.5% ORA), Saccharomyces cerevisiae (0.4% ORA), and a few more ethanol-producing species with low ORA (Supplementary Data Table S1). Among LAB, the most dominant species was Leuconostoc mesenteroides (1.9% ORA), followed by Weissella confusa (1.8% ORA) and Lactococcus garvieae (1% ORA).
Phylogenetic tree of species as recorded in the emao. A total of 74 microbial species and 1 rice species were identified based on a ribosomal protein sequence homology search against the NCBI-nr database. The values within parentheses indicate the relative abundance in percent. Species names in bold (total 19) indicate the lignocellulose/pentose metabolizer as listed in Supplementary Data Table S3). B bacteria, M molds, Y yeasts.
The microbial diversity compared to other traditional beer starters revealed that the emao exhibits some bacterial species, Pediococcus pentosaceus, Leuconostoc citreum, L. mesenteroides, L. pseudomesenteroides, Weissella ciberia, W. confusa, W. paramesenteroides, Lactococcus lactis, L. gravieae; and yeast species, Candida tropicalis, Debariomyces hansenii, Hypopichia burtonii, Pichia kudriavzevii, Meyerozyma guilliermondii, Millerozyma farinosa, Clavispora lusitaniae, Wickerhamomyces anomalus, W. ciferrii, Ogataea parapolymorpha, Kluveromyces dobzhanskii, K. lactis, Torulaspora delbrueckii, Saccharomyces cerevisiae; and mold species, Lichthieimia corymbifera, Mucor circinelloides, M. racemosus, Rhizopus delemar, R. microsporus, R. stolonifer share commonality to other traditional starter cultures, Nuruk from Korea21,31,32, Marcha29 and Xaj-pitha22 from India, Daku from China11,19, and Koji from Japan22 (Supplementary Data Table S1). Some of the dominant species recorded in emao were also common in several other traditional beer starters considered for comparison. Of 24 other traditional starters (OTS)10,11,12,13,14,15,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47 compared to emao, the dominant microbial species, W. anomalus was found common in 20 OTS, S. cerevisiae in 15 OTS, R. delemar in 13 OTS, M. circinelloides in 12 OTS, Pediococcus pentosaceus in 9 OTS, R. microsporus in 7 OTS, and Torulaspora delbrueckii, L. mesenteroides and Candida tropicalis in 6 OTS (Supplementary Data Table S1). However, the other dominant species, R. stolonifer, Choanephora cucurbitarum, Parasitella parasitica, W. ciferrii, C. fabianii, A. rubescens, L. citreum, W. confusa, and L. garvieae recorded in emao, were found only in a few OTS. The rest of the microbial species which accounted for 51% of the total species detected in emao were not reported previously in the 24 OTS considered for comparison (Supplementary Data Table S1).
Identification of carbohydrate-active enzymes (CAZymes) and biofuel producing enzymes (BPZymes) in emao
A total of 19,702 CAZymes48 and 34,493 BPZymes49 were recorded in emao (Fig. 4a,b, Supplementary Data Fig. S1 & S2). Hotpep50, a CAZyme assigning program successfully assigned only 45% of the total CAZymes detected in emao into six CAZyme families and 123 CAZyme sub-families. Among the CAZyme families, glycoside hydrolases (GH) were recorded as the most prevalent (21%), followed by glycosyltransferases (GT, 18%), whereas the rest four types of CAZymes were comparatively less prevalent (≤ 3%). We also emphasized identifying the lignocellulolytic enzymes occurring in emao as essential for mobilizing lignocellulosic substrates into useful products such as biofuel. As such, there is no database for lignocellulolytic enzymes; therefore, we compared the CAZymes and BPZymes of emao to the previous reports on lignocellulolytic enzymes51,52 and identified a total of 1929 lignocellulolytic CAZymes and 5576 lignocellulolytic BPZymes in emao (Fig. 4c,d). A comparison of lignocellulolytic enzymes of emao to pill bug (Armadillidium vulgare) gut microbiome52 revealed seven times higher lignin modifying enzymes, four times higher hemicellulases, and four times higher hemicellulases and/or cellulases in emao, in contrast to four times higher lignocellulose-binding modules in pill bug. In BPZyme analysis, enzymes associated with alcohol production were found two times higher than the enzymes associated with diesel production and fuel cells. We could identify 44% ethanol-producing, 26% fuel-cell producing, 20% diesel producing, and 10% alternate-biofuel producing enzymes (Fig. 4b).
Lignocellulolytic enzymes predicted in emao based on carbohydrate-active enzymes (CAZymes) and biofuel producing enzymes (BPZymes) analyses. A complete list of CAZymes and BPZymes as recorded in emao is given in Supplementary Data Fig. S1 & S2, respectively. (a) Pie chart indicating the percentage of CAZyme families assigned by Hotpep. The value within parentheses is the total count. (b) Relative percentage of BPZyme categories as classified in BioFuelDB. (c) Lignocellulolytic CAZyme sub-families segregated activity-wise. (d) Lignocellulolytic BPZymes with Enzyme Commission (EC) number extracted from (b) and correlated to (c) (dotted lines).
Metabolic and physiological potentials in emao
An analysis of amino acid sequences in Genomaple53,54 revealed the most feasible functional modules with significant module completion ratio (MCR) and Q-values at an individual taxonomic rank (ITR) or whole microbial count (WC) level. Any module having 100% MCR and/or less than 0.5 Q-value is considered significant and feasible53,55. A total of 489 metabolic KEGG (Kyoto Encyclopedia of Genes and Genomes) modules with > 0% MCR (WC) scores were recorded in emao, out of which 46% (i.e., 28% out of total 804 existing modules in KEGG56) had 100% MCR (WC), and that could be the crucial modules in determining the functionality and uniqueness of emao in beer fermentations (Supplementary Data Table S2). The carbohydrate and lipid metabolisms are involved directly or indirectly in plant biomass degradation and biofuel production. Therefore, we focused on carbohydrate and lipid metabolic modules having significant MCR and Q-values (Fig. 5, Table 3).
Carbohydrate and lipid metabolic maps in the emao. Metabolic modules with a Q-value less than 0.5 for whole microbial communities (WC) were shown in the map (Q-values are provided in Supplementary Data Table S2). The probability of occurrence of a metabolic module increases with decreasing Q-value as the latter is zero if all the genes necessary for a module is complete55. This coarse-grained map was created using the KEGG Atlas map56,87. 1, Galactose degradation; 2, D-galacturonate degradation (fungi); 3, D-galacturonate degradation (bacteria); 4, D-glucuronate degradation; 5, Pectin degradation; 6, Embden-Meyerhof Pathway (glycolysis); 7, Pentose phosphate pathway; 8, Glycolysis core module; 9, Pyruvate oxidation; 10, Citrate cycle; 11, Glyoxylate cycle; 12, Melanoate semialdehyde pathway; 13, Gluconeogenesis; 14, PRPP biosynthesis; 15, Nucleotide sugar biosynthesis; 16, N-glycan metabolism; 17, Acyl glycerol degradation; 18, Phosphatidylcholine biosynthesis; 19, Phosphatidylethanolamine biosynthesis; 20, GPI-anchor biosynthesis, core oligosaccharide; 21, Inositol phosphate metabolism; 22, Ceramide biosynthesis; 23, Sphingosine biosynthesis; 24, Sphingosine degradation; 25, Fatty acid biosynthesis, initiation; 26, Fatty acid biosynthesis, elongation; 27, Fatty acid biosynthesis, elongation (ER); 28, Fatty acid biosynthesis, elongation (mitochondria); 29, Beta-oxidation; 30, Beta-oxidation acyl-CoA synthesis; 31, C5 isoprenoid biosynthesis (Mevalonate); 32, C10-20 isoprenoid biosynthesis; 33, Ergocalciferol biosynthesis.
The presence of both 100% MCR for the Embden-Meyerhof pathway with zero Q-value and 75–80% MCR (0.5–0.85 Q-value) for the Entner-Doudoroff pathway (which occurs mainly in some bacteria), revealed the presence of feasible alternative glycolytic pathways in emao (Table 3). Besides, 100% MCR with zero Q-value for pectin, galactose, d-galacturonate, and d-glucuronate degradation pathways are noteworthy, which signifies the possibility of metabolizing those substrates by the microorganisms associated with emao (Fig. 3). Acyl-CoA is necessary for the synthesis of fatty acids precursor, acetyl-CoA57,58. Thus, 100% MCR for both the beta-oxidation module and acyl-CoA synthesis module suffices the fatty acids and isoprenoid biosynthesis potentiality in emao. The presence of 100% MCR for mitochondrial and endoplasmic fatty acid biosynthesis modules and 100% MCR for lipid biosynthesis modules corroborate the involvement of eukaryotes (mainly fungi) in biodiesel production.
Discussion
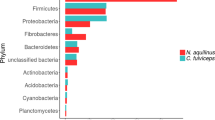
The analyses of yeasts, molds and LAB found in the emao showed the presence of major groups of microbial compositions similar to the Asian ethnic beer starters including Nuruk, Marcha, Thiat, Hamei, Humao, and Xaj-pitha (Supplementary Data Table S1). However, a certain degree of variations is evident among the ethnic starters in their microbial compositions at the species or genus level. One could surmise that variations could be due to the ingredient used and preparation methods followed by different communities. The emao preparation entails rice as a base material and the variation often occurs in the use of quantity and number of herbal species. Of 7–8 different types of herbs, the use of at least 2–3 is the traditional norms of the Bodo people in emao preparation, although the herbs are often used alternatively depending on the availability of the herbs at the time of emao preparation [Source: Local informants]. Emao starters usually produce rice beer containing a maximum of about 17% (w/v) ethanol59, however, the ethanol concentrations can vary depending on the duration of the fermentation period. We found a typical community structure in emao using ribosomal protein-based taxonomic binning where the yeasts, molds, and LAB were seen at a ratio of 9:2:1, respectively. We consider this community structure of emao might be appropriate for producing traditional zou or joubishi containing 5–17% (w/v) ethanol59,60,61 that might have had garnered thousand years ago and is still being maintained in its pristine form by the Bodo peoples through their traditional practices. The findings of higher relative abundance of Firmicutes than Proteobacteria, and higher fungal species counts than bacteria in emao (Fig. 5) differ from the community structures as reported previously in the traditional beer starters Xaj-pitha22, Marcha and Thiat29 from the region. Such a deviation is likely due to differences in methods and approaches followed.
The yeasts, molds, and LAB play unique roles in alcoholic beverage fermentation, especially when starch is the feedstock. Molds are mostly aerobic, do the saccharification, and sensitive to ethanol except for a few species that can produce a low level of ethanol62. The presence of a high number of saccharifying and/or lignocellulose degrading microorganisms is always advantageous and desirable at the beginning of fermentation while using plant-based substrates. Some non-Saccharomyces yeast does both saccharification and ethanol production simultaneously but is mostly non-tolerant to high ethanol concentrations63,64,65,66. Ethanol-sensitive microbes are subsequently killed or arrested by increasing ethanol concentrations at the later stages of ethanol fermentation67. Ethanol production and tolerance level of Saccharomyces yeasts also vary from strain to strain, although they are generally high ethanol tolerant68,69,70. Wild S. cerevisiae cannot utilize starch or other complex carbohydrates directly due to the lack of degrading enzymes for those substrates71 for which they are dependent on other saccharifying microbes.
A low proportion of LAB is desirable in beer fermentation as they can perform some necessary functions despite being known as spoilage agents72,73. They mainly produce lactic acid besides producing bacteriocins against some human pathogens, and they are responsible for maintaining low pH, keeping quality, and taste enhancement in beer74,75,76,77. We did not find any acetic acid-producing bacteria (Acetobacter spp.) in emao, which is a good indicator for this traditional starter of the Bodo community. Acetic acid bacteria are responsible for beer defects78,79, which often happens due to contamination of such spoilage agents if hygienic conditions are compromised during the preparation of starter culture or beer fermentation.
Our findings of the saccharifying, lignocellulolytic (Supplementary Data Table S3), and different biofuels producing microorganisms in corroboration with probable metabolic functionomes (Fig. 5, Table 3) in emao is noteworthy, and the current community structure as found in emao could help develop an effective lignocellulolytic bio-consortia necessary for second-generation biofuel production. Many simple carbohydrate degrading microorganisms do not easily break down the pentose sugars such as D-xylose and L-arabinose that comprise up to 20% of lignocellulosic biomass80. The breaking of lignocellulose into its subunits and subsequently mobilizing them as energy sources for biofuel production is complex. It requires various enzymes to catalyze the metabolic processes, and an organism bearing all the essential enzymes together is rare. However, further experimentation is required to validate the utility of emao as such or in combination with other lignocellulolytic microbes. As the species diversity and the metabolic potentials of emao as reflected from this study are diverse, a pyramiding of target microorganisms from such a natural bio-consortium towards achieving a target product from specific substrates by necessary functional potentials could be a new avenue in tapping natural bioresources for bioprospection. Such an alternative approach can pave the way for bio-consortia formulation to produce biofuels from lignocellulosic materials.
The Genomaple system provides an effective platform to visualize the module completion ratio (MCR) along with the taxonomic information at Phylum or Class level that reflects their functional activity in completing different metabolic modules in any environmental sample considered for an investigation53,54,55,81. However, it is dependent on the KEGG database, which includes only the species with complete genome sequence information. Draft genome information of mold species belonging to Zygomycota is available in the public domain, but not the complete genome information due to which none of these are considered yet in the KEGG database56. In absence of reference taxonomic information, it becomes difficult to specify and segregate the predicted metabolic modules among the taxonomic groups below Phylum or Class level. For the same reason, we were unable to specify and segregate the metabolic modules among the molds and yeasts in this functional metagenome study on emao. Molds are ubiquitous in distribution and play critical ecological roles similar to other fungal groups82. Complete genome information on molds is needed in the public domain to understand better their roles in natural environments and brewing.
Conclusions
The present study is the first to characterize the comprehensive community structure and probable metabolic potentials of the microorganisms associated with the traditional starter culture emao. The presence of diverse groups of microorganisms in corroboration with amylolytic, lignocellulolytic, biofuels producing enzymes as recorded in emao is noteworthy. It could be a pathfinder in the field of microbial consortia bioformulation for biofuel production from otherwise recalcitrant plant biomasses. We found the ribosomal protein-based community structure enumeration a suitable approach in metagenome study. Complete genome information on molds is equally essential in the line of other fungal groups to better understand their roles in traditional brewing and other natural environments. Some dominant microbial species found in emao are similar to some other traditional starters reported previously from the region. Therefore, a comparative study of traditional beer starters that carry microbial genetic information could be of paramount significance in the elucidation and understanding of the history of human population migration and civilization through the ages, like archaeology and philology.
Materials and methods
Sample collection
Six representative emao samples traditionally prepared by the Bodo people were collected from different Bodo-dominated villages of Assam (Table 1, Fig. 6). Information on the method of preparation and herbs used in starter culture were also recorded. We ensured that no beer defect had been experienced during sample collection in using emao from the same stock that we brought for scientific investigation. Traditionally, more than one-year-old emao samples are generally not used for rice beer fermentation; instead, the starter culture is revived in the fresh rice-based medium before completing one year. Therefore, only the active samples, i.e., less than one year from the date of preparation, were considered for this study.

taken from d-maps.com (https://d-maps.com/pays.php?num_pay=84&lang=en & ) and the sampling sites were marked upon it using Adobe Photoshop CS (Version 8.0, 2003). The photographs of the emao samples were taken by D. Narzary and Nitesh Boro.
Map showing the emao sampling sites. (a) Map of India with Assam state highlighted in dark, (b) map of Assam marked with the emao sampling sites (red). Sample numbers (S. No.) are as given in Table 1. The map templates were
Total DNA isolation
Two grams of each of six emao samples were taken and mixed by grinding in pre-sterilized mortar by a pestle, from which a 10 g sample was taken to isolate the total DNA. To recover quality metagenomic DNA, we modified the method of Zhou et al.83 where the extraction buffer was supplemented with 1% activated charcoal and 10 mM MgCl2 as recommended by Sharma et al.84, and the DNA obtained from the modified method of Zhou et al.83 was again purified with the MoBio DNA isolation kit (QIAGEN, Cat. No. 12888–100). The steps we followed are described below. Liquid nitrogen was used for effective sample grinding and the sample was transferred into 18 ml of extraction buffer [100 mM Tris–HCl (pH 8.0), 100 mM sodium EDTA (pH 8.0), 100 mM sodium phosphate (pH 8.0), 1.5 M NaCl, 1% CTAB, 10 mM MgCl2 and 1% activated charcoal] before thawing. A volume of 100 μl proteinase K (10 mg/ml) was added to the tube, mixed properly by vortexing, and incubated at 37 °C for 30 min in a water bath with gentle end-over-end inversions every 5 min. After that, 2 ml of 20% SDS was added, mixed by vortexing, and incubated at 65 °C for 2 h with gentle end-over-end inversions every 30 min. The sample was allowed to cool up to room temperature (RT), and the supernatant was collected in a fresh centrifuge tube after centrifugation at 6000×g for 10 min at RT. The supernatant was mixed with an equal volume of chloroform: isoamyl alcohol (24:1, vol/vol) by inverting the tube gently. The aqueous phase was recovered by centrifugation at 10,000×g for 10 min at RT and then precipitated with 0.6 volume of pre-chilled isopropanol at RT for one h. The pellet of crude nucleic acids was obtained by centrifugation at 12,000×g for 20 min at RT, washed with chilled 70% ethanol, and resuspended in sterile deionized water to make the final volume 700 μl. The DNA solution so obtained was purified with a MoBio DNA isolation kit, and the steps from the treatment of C4 solution onwards were followed according to the procedure of the kit. DNA was stored at –20 °C till sending for whole-metagenome sequencing to the service provider.
Whole-metagenome sequencing
The whole-genome sequencing of our metagenomic DNA was outsourced to the AgriGenome Labs Pvt. Ltd., Kerala, India, and the sequencing was done using the Illumina HiSeq 2500 Platform. The quality of the DNA was confirmed in Qubit Fluorometer and agarose gel electrophoresis before the library preparation. The Genomic DNA was fragmented using Covaris M220 for 500 bp, and the library was prepared using NEBNext Ultra DNA Library Prep Kit. The library quality was checked using Agilent Tapestation 2200. The quantity was estimated using Qubit 2.0. The libraries were sequenced in the HiSeq 2500 platform for 2 × 250 bp read length generating the required data. The FASTQ files generated by the Illumina HiSeq platform were trimmed with Cutadapt (Version 1.8.1)85 to remove the adapters.
Sequence assembly, annotation, and evaluation of potential metabolic modules
The forward and reverse DNA sequences in FASTQ format were submitted to the MAPLE Submission Data Maker (MSDM) pipeline for quality filtering, forward and reverse sequence assembly, and translation to amino acid sequences where the nucleotide sequences with a minimum base quality score of Q20, minimum 80% of quality bases in each sequence, and minimum 400 bp in each merged read were set to get high-quality amino acid (AA) sequences with a minimum cut-off length of 50 AA in FASTA format55. Genomaple ver. 2.3.2 (formerly MAPLE) is a powerful tool that can predict the metabolic functionomes from nucleotide or amino acid sequences as detailed in MAPLE references53,54,81,86. The AA sequence file so obtained was then analyzed in Genomaple ver. 2.3.2 (formerly MAPLE) server opting for the GHOST X search engine with the single-direction best hit annotation for all organisms in KEGG. The module completion ratio (MCR) and Q-value at the individual taxonomic rank (ITR) and the whole microbial community (WC) level were retrieved from MAPLE results, and the KEGG Orthology (KO) genes assigned by Genomaple were used for subsequent taxonomic binning, CAZyme and BPZyme analyses. The module information generated by MAPLE was used to create the coarse-grained metabolic maps of KEGG modules using the KEGG Atlas map87 as a reference. The metabolic map for carbohydrate and lipid metabolism was created separately for the modules having Q-values53 lower than 0.5 to understand the biomass degradation and biofuel production potentiality in emao.
Taxonomic binning
The KOs assigned to the ribosomal protein module for all organisms (M91000) was extracted back from the MAPLE input file using NCBI-blast dbcmd command, which was then subjected to homology search against the non-redundant NCBI-nr protein database to assign the taxonomic identity for each sequence in GHOSTX program88 using the top hit option. GHOSTX result was manually curated to parse the species name against each sequence, which was then meganized and visualized in MEGAN89 Community Version (V6.12.5). The identified species names were uploaded to the NCBI Tree Viewer (https://www.ncbi.nlm.nih.gov/projects/treeview/) to generate the circular phylogenetic tree.
CAZyme identification
The AA sequences created by the MSDM pipeline (version 1.0)55 were used as query files in HMMR hmmscan program (version 3.2.1)90 against dbCAN database91 as a reference with an E-value threshold of 1e-5 to predict CAZymes. The AA sequences detected as CAZymes48 were retrieved back from the input file, and the duplicates were removed using some basic perl and shell commands to get the non-redundant FASTA sequences. Then the non-redundant sequences so obtained were assigned to different CAZyme categories using the Hotpep program50.
BPZyme identification
The AA sequences generated by MSDM in FASTA format were used to identify the enzymes involved in biofuel production as a query file in the HMMR phmmer program90 against BioFuelDB49 as a reference with an E-value threshold of 1e−5. The sequence homology and the corresponding EC number of non-redundant AA sequences identified as the biofuel-producing enzymes were reconfirmed using the GHOSTX homology search (top hit only) against the BioFuelDB. All the ECs were then segregated into different biofuel categories as classified by Chaudhary et al.49, and the total enzyme counts for each category were obtained.
Data availability
The nucleotide sequence raw data (DNS_R1 and DNS_R2) were submitted to the European Nucleotide Archive (ENA) and the study accession number is PRJEB45760. The amino acid sequence data generated by the MAPLE Submission Data Maker (DNS1), MAPLE assigned KOs (DNS2), KEGG metabolic modules (DNS3), and ribosomal protein-based community structure information in RMA format (DNS4) are available through figshare (https://doi.org/10.6084/m9.figshare.8868689). Any other relevant data are available from the corresponding author upon reasonable request.
References
Tamang, J.P. Diversity of fermented beverages and alcoholic drinks. in Fermented Foods and Beverages of the World (eds. Tamang J.P. & Kailasapathy, K.) 85–126 (CRC Press, Taylor & Francis Group, 2010).
Capece, A., Romaniello, R., Siesto, G. & Romano, P. Conventional and non-conventional yeasts in beer production. Fermentation 4, 38. https://doi.org/10.3390/fermentation4020038 (2018).
Cason, E. D. et al. Bacterial and fungal dynamics during the fermentation processes of Sesotho, a traditional beer of Southern Africa. Front. Microbiol. 11, 1451. https://doi.org/10.3389/fmicb.2020.01451 (2020).
McLaren, M. R., Willis, A. D. & Callahan, B. J. Consistent and correctable bias in metagenomic sequencing experiments. Elife 8, e46923. https://doi.org/10.7554/eLife.46923 (2019).
Rausch, P. et al. Comparative analysis of amplicon and metagenomic sequencing methods reveals key features in the evolution of animal metaorganisms. Microbiome. 7, 133. https://doi.org/10.1186/s40168-019-0743-1 (2019).
Mende, D. R., Sunagawa, S., Zeller, G. & Bork, P. Accurate and universal delineation of prokaryotic species. Nat. Methods. 10, 881–884. https://doi.org/10.1038/nmeth.2575 (2013).
Hodgson, H. Kocch, Bodo and Dhimal Tribes (Thomas, J. ed.). https://archive.org/details/in.ernet.dli.2015.93469 (Baptist Mission Press, 1847)
Latham, R. G. The Natural History of the Varieties of Man. https://archive.org/details/naturalhistoryof00lathuoft (John van Voorst, Paternoster Row, 1850).
Zhang, M., Yan, S., Pan, W. & Jin, L. Phylogenetic evidence for Sino-Tibetan origin in northern China in the Late Neolithic. Nature 569, 112–115. https://doi.org/10.1038/s41586-019-1153-z (2019).
Taechavasonyoo, A., Thaniyavarn, J. & Yompakdee, C. Identification of the moulds and yeasts characteristic of a superior Loogpang, starter of Thai rice-based alcoholic beverage Sato. As. J. Food Ag-Ind. 6(01), 24–38 (2013).
Chen, B., Wu, Q. & Xu, Y. Filamentous fungal diversity and community structure associated with the solid state fermentation of Chinese Maotai-flavor liquor. Int. J. Food Microbiol. 179, 80–84. https://doi.org/10.1016/j.ijfoodmicro.2014.03.011 (2014).
Bhardwaj, K. N., Jain, K. K., Kumar, S. & Kuhad, R. C. Microbiological analyses of traditional alcoholic beverage (Chhang) and its starter (Balma) prepared by Bhotiya Tribe of Uttarakhand, India. Indian J. Microbiol. 56(1), 28–34. https://doi.org/10.1007/s12088-015-0560-6 (2016).
Lv, X. C., Weng, X., Zhang, W., Rao, P. F. & Ni, L. Microbial diversity of traditional fermentation starters for Hong Qu glutinous rice wine as determined by PCR-mediated DGGE. Food Control 28, 426–434. https://doi.org/10.1016/j.foodcont.2012.05.025 (2012).
Pradhan, P. & Tamang, J. P. Phenotypic and genotypic identification of bacteria isolated from traditionally prepared dry starters of the Eastern Himalayas. Front. Microbiol. 10, 1–15. https://doi.org/10.3389/fmicb.2019.02526 (2019).
Anupma, A. & Tamang, J. P. Diversity of filamentous fungi isolated from some amylase and alcohol-producing starters of India. Front. Microbiol. 11, 1–16. https://doi.org/10.3389/fmicb.2020.00905 (2020).
Sha, S. P., Suryavanshi, M. V. & Tamang, J. P. Mycobiome diversity in traditionally prepared starters for alcoholic beverages in India by high throughput sequencing method. Front. Microbiol. https://doi.org/10.3389/fmicb.2019.00348 (2019).
Ahmadsah, L. S. F., Kim, E., Jung, Y. S. & Kim, H. Y. Identification of LAB and fungi in Laru, a fermentation starter, by PCR-DGGE, SDS-PAGE, and MALDI-TOF MS. J. Microbiol. Biotechnol. 28(1), 32–39. https://doi.org/10.4014/jmb.1705.05044 (2018).
Thanh, V. N., Mai, L. T. & Tuan, D. A. Microbial diversity of traditional Vietnamese alcohol fermentation starters (banh men) as determined by PCR-mediated DGGE. Int. J. Food Microbiol. 128, 268–273. https://doi.org/10.1016/j.ijfoodmicro.2008.08.020 (2008)
Zheng, X. W. et al. Complex microbiota of a Chinese ‘Fen’ liquor fermentation starter (Fen-Daqu), revealed by culture-dependent and culture-independent methods. Food Microbiol. 31, 293–300. https://doi.org/10.1016/j.fm.2012.03.008 (2012).
Lv, X. C. et al. Microbial community structure and dynamics during the traditional brewing of Fuzhou Hong Qu glutinous rice wine as determined by culture-dependent and culture-independent techniques. Food Control. 57, 216–224. https://doi.org/10.1016/j.foodcont.2015.03.054 (2015).
Bal, J., Yun, S. H., Yeo, S. H., Kim, J. M. & Kim, D. H. Metagenomic analysis of fungal diversity in Korean traditional wheat-based fermentation starter nuruk. Food Microbiol. 60, 73–83. https://doi.org/10.1016/j.fm.2016.07.002 (2016).
Bora, S. S., Keot, J., Das, S., Sarma, K. & Barooah, M. Metagenomics analysis of microbial communities associated with a traditional rice wine starter culture (Xaj-pitha) of Assam, India. 3 Biotech. 6, 153. https://doi.org/10.1007/s13205-016-0471-1 (2016).
Hui, W. et al. Identification of microbial profile of Koji using single molecule, real-time sequencing technology. J. Food Sci. 82(5), 1193–1199. https://doi.org/10.1111/1750-3841.13699 (2017).
Sha, S. P. et al. Diversity of yeasts and molds by culture-dependent and culture-independent methods for mycobiome surveillance of traditionally prepared dried starters for the production of Indian alcoholic beverages. Front. Microbiol. 9, 2237. https://doi.org/10.3389/fmicb.2018.02237 (2018).
Buragohain, A. K., Tanti, B., Sarma, H. K., Barman, P. & Das, K. Characterization of yeast starter cultures used in household alcoholic beverage preparation by a few ethnic communities of Northeast India. Ann. Microbiol. 63, 863–869. https://doi.org/10.1007/s13213-012-0537-1 (2013).
Das, A. J., Miyaji, T. & Deka, S. C. Amylolytic fungi in starter cakes for rice beer production. J. Gen. Appl. Microbiol. 63(4), 236–245. https://doi.org/10.2323/jgam.2016.11.004 (2017).
Parasar, D. P., Sarma, H. K. & Kotoky, J. Exploring the genealogy and phenomic divergences of indigenous domesticated yeasts cultivated by six ethnic communities of Assam, India. J. Biol. Sci. 17, 91–105. https://doi.org/10.3923/jbs.2017.91.105 (2017).
Thanh, V. N., Mai, L. T. & Tuan D. A. Microbial diversity of traditional Vietnamese alcohol fermentation starters (banh men) as determined by PCR-mediated DGGE. Int. J. Food Microbiol. 128, 268–273 (2008).
Sha, S. P. et al. Analysis of bacterial and fungal communities in Marcha and Thiat, traditionally prepared amylolytic starters of India. Sci. Rep. https://doi.org/10.1038/s41598-017-11609-y (2017).
Nath, N., Ghosh, S., Rahaman, L., Kaipeng, D. L. & Sharma, B. K. An overview of traditional rice beer of north-east india: Ethnic preparation, challenges and prospects. Indian J. Tradit. Knowl. 18(4), 744–757 (2019).
Song, H. S. S. et al. Analysis of microflora profile in Korean traditional Nuruk. J. Microbiol. Biotechnol. 23(1), 40–46. https://doi.org/10.4014/jmb.1210.10001 (2013).
Kwon, S. J. & Sohn, J. H. Analysis of microbial diversity in Nuruk using PCR-DGGE. J. Life Sci. 22(1), 110–116. https://doi.org/10.5352/jls.2012.22.1.110 (2012).
Bal, J. et al. Microflora dynamics analysis of Korean traditional wheat-based Nuruk. J. Microbiol. 52(12), 1025–1029 (2014).
Chakrabarty, J., Sharma, G. D. & Tamang, J. P. Traditional technology and product characterization of some lesser-known ehnic fermented food and beverages of North Cachar Hills district of Assam. Indian J. Tradition. Knowl. 13, 706–715 (2014).
Das, C. P. & Pandey, A. Fermentation of traditional beverages prepared by the Bhotia community of Uttaranchal Himalaya. Indian J. Tradition. Knowl. 6, 136–140 (2007).
Dung, N. T. P., Rombouts, F. M. & Nout, M. J. R. Characteristics of some traditional Vietnamese starch-based rice wine fermentation starters (men). LWT 40, 130–135 (2007).
Jeyaram, K., Singh, W. M., Capece, A. & Romano, P. Molecular identification of yeast species associated with ‘Hamei’—A traditional starter used for rice wine production in Manipur, India. Int. J. Food Microbiol. 124, 115–125 (2008).
Lv, X. C., Huang, X. L., Zhang, W., Rao, P. F. & Ni, L. Yeast diversity of traditional alcohol fermentation starters for Hong Qu glutinous rice wine brewing, revealed by culture-dependent and culture-independent methods. Food Control 34, 183–190 (2013).
Lv, X. C., Huang, Z. Q., Zhang, W., Rao, P. F. & Ni, L. Identification and characterization of filamentous fungi isolated from fermentation starters for Hong Qu glutinous rice wine brewing. J. Gen. Appl. Microbiol. 58, 33–42 (2012).
Rittiplang, J., Laopaiboon, P., Vichitpan, K. & Danvirutai, P. Lactic acid bacteria from indigeneous Loogpang samples of Northeastern Thailand and their lactic acid production ability. Thai J. Biotechnol. 44–48 (2007).
Shi, J. H. et al. Analysis of microbial consortia in the starter of Fen liquor. Lett. Appl. Microbiol. 48, 478–485 (2009).
Tamang, J. P. & Sarkar, P. K. Microflora of Marcha: An amylolytic fermentation starter. Microbios 81, 115–122 (1995).
Tamang, J. P. et al. Lactic acid bacteria in Hamei and Marcha of North East India. Indian J. Microbiol. 47, 119–125 (2007).
Tamang, J. P., Sarkar, P. K. & Hesseltine, C. W. Traditional fermented foods and beverages of Darjeeling and Sikkim—A review. J. Sci. Food Agric. 44, 375–395 (1988).
Teramoto, Y., Yoshida, S. & Ueda, S. Characteristics of a rice beer (Zutho) and a yeast isolated from the fermented product in Nagaland, India. World J. Microbiol. Biotechnol. 18, 813–816 (2002).
Tsuyoshi, N., Fudou, R., Yamanaka, S., Kozaki, M., Tamang, N., Thapa, S. & Tamang, J. P. Identification of yeast strains isolated from marcha in Sikkim, a microbial starter for amylolytic fermentation. Int. J. Food Microbiol. 99, 135–146 (2005).
Wu, Q., Chen, L. & Xu, Y. Yeast community associated with the solid-state fermentation of traditional Chinese Maotai-flavor liquor. Int. J. Food Microbiol. 166, 323–330 (2013).
Lombard, V., Golaconda Ramulu, H., Drula, E., Coutinho, P. M. & Henrissat, B. The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res. 42, D490–D495. https://doi.org/10.1093/nar/gkt1178 (2014).
Chaudhary, N., Gupta, A., Gupta, S. & Sharma, V. K. BioFuelDB: A database and prediction server of enzymes involved in biofuels production. PeerJ 5, e3497. https://doi.org/10.7717/peerj.3497 (2017).
Busk, P. K., Pilgaard, B., Lezyk, M. J., Meyer, A. S. & Lange, L. Homology to peptide pattern for annotation of carbohydrate-active enzymes and prediction of function. BMC Bioinform. 18, 214. https://doi.org/10.1186/s12859-017-1625-9 (2017).
Janusz, G. et al. Lignin degradation: Microorganisms, enzymes involved, genomes analysis and evolution. FEMS Microbiol. Rev. 41(6), 941–962. https://doi.org/10.1093/femsre/fux049G (2017).
Bredon, M., Dittmer, J., Noël, C., Moumen, B. & Bouchon, D. Lignocellulose degradation at the holobiont level: Teamwork in a keystone soil invertebrate. Microbiome. 6, 162. https://doi.org/10.1186/s40168-018-0536-y (2018).
Takami, H. et al. An automated system for evaluation of the potential functionome: MAPLE version 2.1.0. DNA Res. 23, 467–475. https://doi.org/10.1093/dnares/dsw030 (2016).
Arai, W. et al. Maple 2.3.0: An improved system for evaluating the functionomes of genomes and metagenomes. Biosci. Biotechnol. Biochem. 82, 1515–1517. https://doi.org/10.1080/09168451.2018.1476122 (2018)
Takami, H. MAPLE enables functional assessment of microbiota in various environments. in Marine Metagenomics-Technological Aspects and Applications. (eds. Gojobori, T., Wada, T., Kobayashi, T., Mineta, K.) 85–119 (Springer, 2019).
Kanehisa, M., Furumichi, M., Sato, Y., Ishiguro-Watanabe, M. & Tanabe, M. KEGG: Integrating viruses and cellular organisms. Nucleic Acids Res. 49(D1), D545–D551. https://doi.org/10.1093/nar/gkaa970 (2020).
Courchesne, N. M. D., Parisien, A., Wang, B. & Lan, C. Q. Enhancement of lipid production using biochemical, genetic and transcription factor engineering approaches. J. Biotechnol. 141, 31–41. https://doi.org/10.1016/j.jbiotec.2009.02.018 (2009).
Phulara, S. C., Chaturvedi, P. & Gupta, P. Isoprenoid-based biofuels: Homologous expression and heterologous expression in prokaryotes. Appl. Environ. Microbiol. 82(19), 5730–5740. https://doi.org/10.1128/AEM.01192-16 (2016).
Borah, A. Ethanol Production Performance Assay of Non-Saccharomyces Fungi Obtained from Starter Culture Used by Bodo Tribe in Rice-Beer Fermentation (Thesis Submitted to Gauhati University, 2017).
Deka, A. K., Handique, P. & Deka, D. C. Antioxidant-activity and physicochemical indices of the rice beer used by the Bodo community in north-east India. J. Am. Soc. Brew. Chem. https://doi.org/10.1080/03610470.2018.1424400 (2018).
Das, S., Deb, D., Adak, A. & Khan, M. J. Exploring the microbiota and metabolites of traditional rice beer varieties of Assam and their functionalities. 3 Biotech 9, 174. https://doi.org/10.1007/s13205-019-1702-z (2019).
Dung, N. T. P., Rombouts, F. M. & Nout, M. J. R. Functionality of selected strains of moulds and yeasts from Vietnamese rice wine starters. Food Microbiol. 23(4), 331–340. https://doi.org/10.1016/j.fm.2005.05.002 (2006).
Limtong, S., Sintara, S. & Suwannarit, P. Yeast diversity in Thai traditional alcoholic starter (Loog-Pang). Kasetsart J. (Nat. Sci.) 36, 149–158 (2002).
Pina, C., Antonio, J. & Hogg, T. Inferring ethanol tolerance of Saccharomyces and non-Saccharomyces yeasts by progressive inactivation. Biotechnol. Lett. 26, 1521–1527. https://doi.org/10.1007/s10529-005-1787-9 (2005).
Pina, C., Santos, C., Couto, J. A. & Hogg, T. Ethanol tolerance of five non-Saccharomyces wine yeasts in comparison with a strain of Saccharomyces cerevisiae - Influence of different culture conditions. Food Microbiol. 21(4), 439–447. https://doi.org/10.1016/j.fm.2003.10.009 (2004).
Jamai, L., Ettayebi, K., Yamani, J. E. & Ettayebi, M. Production of ethanol from starch by free and immobilized Candida tropicalis in the presence of α-amylase. Bioresour. Technol. 98(14), 2765–2770. https://doi.org/10.1016/j.biortech.2006.09.057 (2007).
Steensels, J. & Verstrepen, K. J. Taming wild yeast: Potential of conventional and nonconventional yeasts in industrial fermentations. Annu. Rev. Microbiol. 68, 61–80. https://doi.org/10.1146/annurev-micro-091213-113025 (2014).
Casey, G. P. & Ingledew, W. M. M. Ethanol tolerance in yeasts. Crit. Rev. Microbiol. (1986). 13(3), 219–280. https://doi.org/10.3109/10408418609108739 (1986).
Ghareib, M., Youssef, K. A. & Khalil, A. A. Ethanol tolerance of Saccharomyces cerevisiae and its relationship to lipid content and composition. Folia Microbiol. (Praha) 33, 447–452. https://doi.org/10.1007/BF02925769 (1988).
Riles, L. & Fay, J. C. Genetic basis of variation in heat and ethanol tolerance in Saccharomyces cerevisiae. G3 Genes Genomes Gene 9(1), 179–188. https://doi.org/10.1534/g3.118.200566 (2019).
Yamada, R. et al. Direct and efficient ethanol production from high-yielding rice using a Saccharomyces cerevisiae strain that express amylases. Enzyme Microb. Technol. https://doi.org/10.1016/j.enzmictec.2011.01.002 (2011).
Hollerová, I. & Kubizniaková, P. Monitoring Gram positive bacterial contamination in Czech breweries. J. Inst. Brew. 107, 355–358. https://doi.org/10.1002/j.2050-0416.2001.tb00104.x (2001).
Bokulich, N. A. & Bamforth, C. W. The microbiology of malting and brewing. Microbiol. Mol. Biol. Rev. 77(2), 157–172. https://doi.org/10.1016/B978-0-12-809633-8.13014-6 (2013).
Ennahar, S., Sonomoto, K. & Ishizaki, A. Class IIa bacteriocins from lactic acid bacteria: Antibacterial activity and food preservation. J. Biosci. Bioeng. 87, 705–716. https://doi.org/10.1016/S1389-1723(99)80142-X (1999).
Mokoena, M. P. Lactic acid bacteria and their bacteriocins: Classification, biosynthesis and applications against uropathogens: A mini-review. Molecules 22(8), 1255. https://doi.org/10.3390/molecules22081255 (2017).
Ghosh, S. et al. Community-wise evaluation of rice beer prepared by some ethnic tribes of Tripura. J. Ethn. Foods. https://doi.org/10.1016/j.jef.2016.12.001 (2016).
Miller, M. Monitoring Acids and pH in Wine Making. eBook. http://www.gencowinemakers.com/docs/Acids%20Presentation.pdf (2019).
Sakamoto, K. & Konings, W. N. Beer spoilage bacteria and hop resistance. Int. J. Food Microbiol. 89(2), 105–124. https://doi.org/10.1016/S0168-1605(03)00153-3 (2003).
Vriesekoop, F., Krahl, M., Hucker, B. & Menz, G. 125th Anniversary review: Bacteria in brewing: The good, the bad and the ugly. J. Inst. Brew. 118, 335–345. https://doi.org/10.1002/jib.49 (2012).
Cadete, R. M. & Rosa, C. A. The yeasts of the genus Spathaspora: Potential candidates for second-generation biofuel production. Yeast 35, 191–199. https://doi.org/10.1002/yea.3279 (2018).
Takami, H., Taniguchi, T., Moriya, Y., Kuwahara, T., Kanehisa, M. & Goto, S. Evaluation method for the potential functionome harboured in the genome and metagenome. BMC Genomics 13, 699. http://www.biomedcentral.com/1471-2164/13/699 (2012).
Gryganskyi, A. P. & Muszewska, A. Whole genome sequencing and the Zygomycota. Fungal Genom. Biol. 4, 1. https://doi.org/10.4172/2165-8056.1000e116 (2014).
Zhou, J., Bruns, M. A. & Tiedje, J. M. DNA recovery from soils of diverse composition. Appl. Environ. Microbiol. 62, 316–322. https://doi.org/10.1128/aem.62.2.316-322.1996 (1996).
Sharma, S., Sharma, K. K. & Kuhad, R. C. An efficient and economical method for extraction of DNA amenable to biotechnological manipulations, from diverse soils and sediments. J. Appl. Microbiol. 116, 923–933. https://doi.org/10.1111/jam.12420 (2013).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17(1), 10–12 (2011). (Nurk, S., Meleshko, D., Korobeynikov, A. &. Pevzner, P.A. metaSPAdes: A new versatile metagenomic assembler. Genome Res. 27(5), 824–834. https://doi.org/10.1101/gr.213959.116, 2017).
Okubo, T. et al. The physiological potential of anammox bacteria as revealed by their core genome structure. DNA Res. 1, 1–12. https://doi.org/10.1093/dnares/dsaa028 (2020).
Okuda, S. et al. KEGG Atlas mapping for global analysis of metabolic pathways. Nucleic Acids Res. 36(12), W423–W426. https://doi.org/10.1093/nar/gkn282 (2008).
Suzuki, S., Kakuta, M., Ishida, T. & Akiyama, Y. GHOSTX: An improved sequence homology search algorithm using a query suffix array and a database suffix array. PLoS ONE 9(8), e103833. https://doi.org/10.1371/journal.pone.0103833 (2014).
Huson, D.H. et al. MEGAN community edition—Interactive exploration and analysis of large-scale microbiome sequencing data. PLoS Comput. Biol. https://doi.org/10.1371/journal.pcbi.1004957 (2016).
Mistry, J., Finn, R. D., Eddy, S. R., Bateman, A. & Punta, M. Challenges in homology search: HMMER3 and convergent evolution of coiled-coil regions. Nucleic Acids Res. 41(12), e121. https://doi.org/10.1093/nar/gkt263 (2013).
Yin, Y. et al. DbCAN: A web resource for automated carbohydrate-active enzyme annotation. Nucleic Acids Res. https://doi.org/10.1093/nar/gks479 (2012).
Acknowledgements
The authors thank Prof. Kishore K. Wary, Dr. T.S. Rana, and J.S. Chandrani for their valuable feedback and suggestions. This research was supported by the DST, Govt of India under the SERB scheme (SB/EMEQ-443/2014) to D.N. We thank the Department of Botany, Gauhati University, Guwahati, and the Yokohama Institute (JAMSTEC), Japan for research facilities. DN is also thankful to the DBT, Govt of India for financial support to set up a sophisticated research laboratory under the Unit of Excellence scheme (BT/408/NE/U-Excel/2013), and also for overseas fellowship to visit JAMSTEC for data analysis under the DBT-Associateship program. The authors are thankful to the local Bodo women who shared their traditional material and information for our research.
Author information
Authors and Affiliations
Contributions
D.N. conceived and designed experiments. N.B. and A.B. contributed to sample collection and DNA isolation. H.T. directed metagenomic data analysis. D.N. and O.T. performed computational work. D.N. wrote and H.T. revised the manuscript. All authors reviewed the manuscript .
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Narzary, D., Boro, N., Borah, A. et al. Community structure and metabolic potentials of the traditional rice beer starter ‘emao’. Sci Rep 11, 14628 (2021). https://doi.org/10.1038/s41598-021-94059-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-94059-x
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.