Abstract
Recent findings suggest that changes in human odors caused by malaria infection have significant potential as diagnostic biomarkers. However, uncertainty remains regarding the specificity of such biomarkers, particularly in populations where many different pathological conditions may elicit similar symptoms. We explored the ability of volatile biomarkers to predict malaria infection status in Kenyan schoolchildren exhibiting a range of malaria-like symptoms. Using genetic algorithm models to explore data from skin volatile collections, we were able to identify malaria infection with 100% accuracy among children with fever and 75% accuracy among children with other symptoms. While we observed characteristic changes in volatile patterns driven by symptomatology, our models also identified malaria-specific biomarkers with robust predictive capability even in the presence of other pathogens that elicit similar symptoms.
Similar content being viewed by others
Introduction
The presence of disease can alter human odors, including volatile emissions from skin and breath. The potential diagnostic value of such changes in volatile chemistry has long been recognized, and volatile-based diagnostics are being actively explored for a number of diseases, including several types of cancer1,2. However, a major challenge for the development of volatile disease biomarkers is posed by the inherent variability of human volatile emissions, which are highly labile and can be influenced by a wide range of genetic, physiological, and environmental factors3. In light of this variability, there is reason to speculate that volatile biomarkers might be of particular value in diagnosing diseases caused by insect-borne pathogens, which frequently manipulate the odors of their hosts in ways that influence vector behavior and might therefore be highly conserved4,5,6. Indeed, a number of recent studies on human malaria have identified characteristic changes in the volatile emissions from the skin and breath of infected individuals7,8,9,10,11,12. Yet, uncertainty remains about the physiological bases of these volatile changes and the extent to which they are uniquely caused by malaria or products of pathological processes that might be shared with other disease states. Addressing this uncertainty has important implications for understanding the diagnostic value of such biomarkers, particularly for use in human populations where numerous pathological conditions are widespread and may give rise to similar symptomatology, as is often the case for malaria endemic regions.
A number of previous studies have documented malaria-induced changes in human skin7,8,9 and breath10,11,12 volatiles, as well as the volatile emissions of rodent malaria hosts13, while others have characterized the emissions of Plasmodium cells grown in vitro14,15,16. Effects of infection on host odors have also been reported to influence vector behavior, with several studies reporting increased mosquito attraction to hosts harboring transmissible stages of Plasmodium parasite13,17,18 and at least one showing enhanced attraction to host odor profiles in which individual compounds were manipulated to mimic the relative up- or downregulation caused by infection13.
Enhanced vector recruitment during the transmissible stage of infection is hypothesized to facilitate malaria transmission19, suggesting that the pathogen may benefit from actively manipulating host volatiles20. To the extent that Plasmodium parasites, or other vector-borne pathogens, alter host odors in ways that consistently influence vector attraction, they may also generate unique patterns of effects on host volatile profiles—possibly tailored to the olfactory responses of particular vector species—distinct from more general changes in volatile emissions that arise as mere byproducts of pathology. The presence of such unique signatures of infection might, in turn, facilitate the identification of pathogen-specific biomarkers capable of reliably predicting infection status even in populations where numerous pathological conditions elicit similar symptoms.
The identification of such robust biomarkers is of particular interest for malaria, which frequently occurs in populations affected by numerous other diseases and ailments that elicit similar symptoms, including fever, headaches, diarrhea, vomiting, etc., and might therefore plausibly be expected to have somewhat similar effects on underlying physiological processes that also influence volatile emissions. In the case of malaria, disease progression often entails alternating asymptomatic and symptomatic phases that may recur indefinitely if not treated. In a previous study, we reported differential up and down regulation of specific volatile organic compounds in both symptomatic and asymptomatic schoolchildren in Kenya7. Yet, while these characteristic changes were highly predictive of malaria infection, the specific processes by which the presence of malaria parasites alter human volatiles remain almost entirely unknown. One recent study reported that a malaria-derived isoprenoid increases the production of several monoterpenes and aldehydes in vitro21, but this does not account for the majority of volatile alterations observed by our and other studies7,8,9,10,11,12 suggesting that the underlying mechanisms are likely to be complex.
To explore the extent to which changes in the odors of malaria infected humans are associated with the presence of symptoms, we analyzed the volatile profiles of symptomatic children, using volatile data previously collected from a large field trial in Western Kenya7. Specifically, we compared the skin volatile profiles of symptomatic schoolchildren who tested positive for malaria to those of children presenting similar (malaria-like) symptoms, but who tested negative for malaria. We used a genetic algorithm framework to predict malaria infection and identify volatile compounds associated with disease status. Our goal was thus to discover which volatile changes might be specific to malaria, and which might be caused by infection with other illnesses or conditions.
Results and discussion
Volatile signatures differentiate symptomatic children with and without malaria infection
We first explored whether there was a clear signature of malaria infection among symptomatic children. In addition to examining differences in the volatile profiles of malaria-infected and uninfected children presenting any symptom (i.e., all symptomatic children), we made similar comparisons for non-exclusive subsets of children exhibiting the two most commonly observed symptoms in our dataset: fever and diarrhea. For each of these three symptom categories, discriminant analysis of principal components (DAPC) revealed clear separation in the overall volatile composition between individuals with and without malaria infection (Fig. 1, S1), with a permutational analysis of variance resulting in significant differences in both arms (pseudo-F1,48 = 2.96, p = 0.022) and feet (pseudo-F1,49 = 2.91, p = 0.003). These results indicate that the effect of malaria infection on volatile profiles is apparent even when directly compared to other conditions that produce similar symptoms.
We next used a genetic algorithm predictive model trained to recognize malaria infection based on a subset of the data (70%) to predict malaria infection status in the remaining test set (30%). For children exhibiting any symptom, this model was able to predict malaria status with 75% accuracy (Tables 1 and 2). Given that children included in this comparison likely exhibited highly variable physiological states (i.e., different symptomatologies arising from the presence of different pathogens and various stages of disease progression) this level of predictive accuracy indicates that the volatile signature of malaria infection is relatively robust. Our model exhibited similar accuracy (75%) in predicting malaria infection among the subset of symptomatic children with diarrhea; for children with fever, however, the model predicted infections status with 100% accuracy (based using arm volatiles) (Tables 1 and 2). It also bears noting that only five compounds were required to predict malaria among children with diarrhea, compared to greater numbers of compounds required for the other symptom categories. This is consistent with the observation that few compounds show significant alteration with malaria infection status among children with diarrhea, compared to more extensive differences observed for children with fever (Fig. 2).
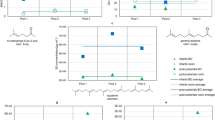
Volcano plots showing changes in individual compounds in malaria-infected symptomatic children relative to malaria-uninfected symptomatic children. Significantly up- or downregulated compounds (p < 0.05 and absolute fold change > 1.5) are shown in green. Compound IDs are listed in Table 2.
The higher level of predictive accuracy for children with fever may reflect distinct features of the pathology occurring in malaria-infected children. As Plasmodium completes its lifecycle within the host, fever is typically associated with the rupture of mature schizont cells and the release of merozoites that then reinvade red blood cells22. The resulting intermittent cyclic fever may cause unique physiological effects compared to other ailments, and may also play a role in the downregulation of volatiles, including the compounds that drive the predictive accuracy of our model (Fig. 2). Indeed, it is notable that malaria symptomatic children show decreased volatile production compared to both malaria-free febrile and asymptomatic children (Fig. 2, S3), as fever might otherwise be presumed to cause a general upregulation of volatile emissions due to increased body temperature and sweating causing increased evaporation of compounds from the skin.
Of the eight volatile compounds selected by our model to predict malaria status from the arms of children with fever (Tables 1 and 2), hexanal (C9), decane (C51) and octanal (C52), have been identified as predictors of malaria infection in several previous studies7,8,15,16. In the current study, these compounds were also important predictors of infection status among children exhibiting any symptom, as well as among the subset of children with diarrhea. Hexanal and octanal are also known to serve as host-location cues for mosquitoes, eliciting attraction or repellence depending on their concentration23,24. It is thus intriguing that the emission of these compounds appears to be specifically influenced by the presence of malaria parasites. As we have previously speculated3,7, malaria-induced changes in volatile cues that enhance transmission probability via effects on mosquito attraction19 might generate robust biomarkers of infection status.
The presence of symptoms alters volatile profiles in malaria-free children
To explore how the presence of symptoms itself influences volatile emissions, we next used predictive models, similar to those described above, to examine differences between the volatile profiles of symptomatic and asymptomatic children who tested negative for malaria infection. Here our models were able to predict the presence of symptoms with 60–80% accuracy across the three symptom categories described above (fever, diarrhea, any symptom) and using either foot or arm volatiles (Table 3). Several compounds that were important predictors for the presence of symptoms in these models were also important predictors of malaria infection status among symptomatic children in our previous analyses, including C8 (octane), C9 (hexanal) and C17 (4-hydroxy-4-methylpentan-2-one), (Table 1).
Comparing the volatile emissions of malaria-free children exhibiting any symptom to those without symptoms revealed significant downregulation of several compounds (Fig. 3), with ethylbenzene (C20) showing the greatest downregulation in the presence of symptoms for both arm and foot volatiles. When focusing on individual symptoms, malaria-free children with diarrhea showed no significant changes in compound levels compared to those without symptoms, whereas febrile children without malaria showed an upregulation of several foot volatile compounds (Fig. 3), in contrast to the downregulation of compounds seen in febrile children with malaria compared to malaria-free children without symptoms (Fig. S3). The only compound downregulated in febrile children without malaria was C17 (4-hydroxy-4-methylpentan-2-one) (Fig. 3), which was further downregulated in malaria-infected children compared to febrile children without malaria (Fig. 2). Among upregulated compounds in malaria-free febrile children, C9 (hexanal) was previously reported to be downregulated in malaria-infected vs uninfected children7. The overlap in compounds identified as important predictors of malaria infection and of symptoms in malaria-free children (but with different patterns of up and down regulation) is intriguing and may reflect the influences of different pathological conditions on the same underlying physiological processes.
Volcano plots showing changes in individual compounds in malaria-uninfected symptomatic children relative to malaria-uninfected asymptomatic children. Significantly up- or downregulated compounds (p < 0.05 and absolute fold change > 1.5) are shown in green. Compound IDs are listed in Table 2.
Overlap in volatile biomarkers for malaria and other diseases
As noted above, the use of volatile diagnostics is under investigation or optimization for a number of diseases, including various types of cancer and pneumonia, and many studies have examined the volatiles in breath that change dependent on disease status1,2,25. In the case of lung cancer, a large-scale trial for diagnostic detection using VOCs is currently underway26. For many other diseases, however, significant challenges remain to be overcome prior to the implementation of large-scale trials, including the identification of a robust and reproducible set of candidate biomarkers3, which can be complicated by the absence of standardized methods for volatile collection and analysis27. In the case of malaria, several previous studies have reported successful prediction of infection status based on analysis of VOCs; however, there is considerable variation in the predictive compounds identified, likely owing to divergent methodologies across studies. Two compounds, hexanal and nonanal, have consistently been found informative with respect to with malaria status, each reported in three separate studies from skin and breath volatiles as well as in the prediction here of malaria status in symptomatic children (Table 4). Among compounds identified as predictors of malaria infection in the current study, hexanal and nonanal, along. with decane and octanal, have previously been implicated as disease predictors in studies on other diseases in addition to malaria (Table 4). Hexanal, nonanal, octanal and have been identified in the breath of lung and (along with decane) breast cancer patients, while nonanal has also been identified as a predictor of colorectal cancer and pneumonia2,25. A further comparison of the key predictive compounds from this study and those identified for other diseases reveals several compounds that are specific to the differentiation of infection status for malaria, including octane, 4-hydroxy-4-methylpentan-2-one, o-xylene and 2-ethylhexan-1-ol. This specificity may be related to the efficacy of these compounds in distinguishing malaria infections form other conditions that give rise to similar symptoms; however, this remains speculative given that many of the other diseases for which volatile diagnostics have been studied do not display similar symptoms. Finally, the recurrence of certain compounds such as hexanal and nonanal as predictors of multiple diseases may, again, reflect the influences of different pathological conditions on the same underlying processes, but with disease-specific patterns of effects on volatile emissions that can give rise to distinct signatures.
Conclusion
Volatile biomarkers hold significant promise for the development of non-invasive techniques for disease diagnosis1. However, extensive variation in human volatile emissions, including that due to the presence of many different diseases and ailments within human populations, poses a significant challenge. While the robustness of malaria biomarkers across varying genetic and environmental backgrounds still needs to be assessed in large-scale trials before they could enter clinical use, the current findings indicate that volatile biomarkers can identify malaria infection even in the face of variation elicited by the presence of other symptomatic conditions. Our predictive models identified malaria cases with 100% accuracy for children with fever and with 75% accuracy for other symptoms. We also identified specific compounds that are important predictors of malaria infection among symptomatic children, as well as compounds that are more generally indicative of the presence of symptoms. These results suggest that, while some changes in human volatile profiles are broadly associated with the presence of symptomatic disease, malaria elicits specific changes in key compounds that provide a unique signature of infection.
Methods
Ethics approval and participant selection
This study was approved by The Pennsylvania State University (IRB #41,529), ETH Zürich (EK2015-N-59), and the Kenya Medical Research Institute (SERU 391) and all experiments were performed in accordance with relevant guidelines and regulations. Before sample collection, the study and consent form were explained to parents/guardians and their written informed consent was obtained.
Participant exclusion criteria included (1) receipt of antimalarial medication during the previous 2 wk; (2) chronic disease, such as HIV; (3) not signing (or having a parent sign) the consent form; and (4) refusal of malaria treatment.
Symptom categories
Our analyses were performed on data derived from skin volatile samples (1 h collections from arms and feet) collected from students at 41 primary schools near Mbita Point, Kenya between 2013 and 20167. Symptoms were self-reported in an initial interview using a standardized questionnaire; symptoms indicative of malaria included fever, abdominal pain, rash, diarrhea, vomiting and body aches. For our analysis, we sorted children into the categories (1) any symptom (2) fever, (3) diarrhea and (4) asymptomatic. The any symptom category comprised children with fever, diarrhea, abdominal pain or vomiting; the fever and diarrhea categories were non-exclusive subsets of the any symptom category. Abdominal pain and vomiting were not analyzed independently due to low numbers of children with these symptoms. Malaria infection status was initially assessed via rapid diagnostic testing (SD Bioline), then confirmed by light microscopy and PCR. For the current study, children in the malaria infected category tested positive by both microscopy and PCR.
The dataset used for this study comprised volatile profiles for 114 children. Once categorized into symptoms for the below analyses, numbers of children in each category were as follows: malaria infected children with any symptom = 29, fever = 18, diarrhea = 11. Malaria-free children with any symptom = 22, fever = 11, diarrhea = 9, asymptomatic = 16.
Volatile data
Arm and foot volatiles were collected simultaneously for 1 h by enclosing the arm (from wrist to above the elbow) or foot (to below the knee) in a teflon bag, pushing filtered air through an entry port (arm: 1.1 L/min, foot: 1.8 L/min) and pulling it through an exit port (arm: 0.8 L/min, foot: 1.1 L/min) where it was collected on an adsorbent HaySepQ filter. Compounds trapped on filters were then eluted with 150ul dichloromethane and analyzed by GC–MS (Full methods: De Moraes 20187).
Statistical analyses
Discriminant Analysis of Principal Components (DAPC)41,42 was used to compare the separation of healthy from malaria-infected individuals. These group differences were tested using a permutational analysis of variance (PERMANOVA) with the Euclidean similarity on the scaled data43.
We used a Genetic Algorithm (GA) to predict disease condition and to select one or more sets of compounds for each of the symptom subsets to be assessed as a potential biomarker for malaria. Each subset was randomly split into 70% for model training and 30% for validation. Using the training set, each model was trained five times using a repeated tenfold cross-validation. Genetic Algorithms are feature selection procedures that are conceptually based on the principle of evolution by natural selection44. They have been used as a promising multivariate approach in the analysis of metabolomics datasets45,46. The algorithm works by evolving initial sets of variables (chromosomes) from a random population via cycles of replication, recombination and mutation of the fittest chromosomes. The iterations were repeated for 100 generations, with a population size of 50 candidate solutions for each model and crossover and mutation probabilities of 0.8 and 0.1, respectively. The performance of the fittest GA model was used to predict the samples in the validation set. To complement the predictive models, a differential analysis between infection statuses was performed using the limma R package47. The GA models were formulated with the caret R package48.
Data availability
All relevant data reported in this paper have been deposited in ETH Zurich’s Research Collection, http://hdl.handle.net/20.500.11850/458605 (https://doi.org/10.3929/ethz-b-000458605).
References
Buljubasic, F. & Buchbauer, G. The scent of human diseases: a review on specific volatile organic compounds as diagnostic biomarkers. Flavour Fragr. J. 30, 5–25. https://doi.org/10.1002/ffj.3219 (2015).
Oakley-Girvan, I. & Davis, S. W. Breath based volatile organic compounds in the detection of breast, lung, and colorectal cancers: a systematic review. Cancer Biomark 21, 29–39. https://doi.org/10.3233/cbm-170177 (2017).
Stanczyk, N. M., De Moraes, C. M. & Mescher, M. C. Can we use human odors to diagnose malaria?. Future Microbiol. 14, 5–9. https://doi.org/10.2217/fmb-2018-0312 (2018).
Hughes, D. P., Brofeur, J., and Thomas, F. Host Manipulation by Parasites. (Oxford University Press, 2012).
Hurd, H. Manipulation of medically important insect vectors by their parasites. Annu. Rev. Entomol. 48, 141–161. https://doi.org/10.1146/annurev.ento.48.091801.112722 (2003).
Lefevre, T. & Thomas, F. Behind the scene, something else is pulling the strings: emphasizing parasitic manipulation in vector-borne diseases. Infect. Genet. Evolut. J. Mol. Epidemiol. Evolut. Genet. Infect. Dis. 8, 504–519. https://doi.org/10.1016/j.meegid.2007.05.008 (2008).
De Moraes, C. M. et al. Volatile biomarkers of symptomatic and asymptomatic malaria infection in humans. Proc Natl Acad Sci U S A (2018).
Robinson, A. et al. Plasmodium-associated changes in human odor attract mosquitoes. Proc Natl Acad Sci U S A (2018).
de Boer, J. G. et al. Odours of Plasmodium falciparum-infected participants influence mosquito-host interactions. Sci. Rep. 7, 9283. https://doi.org/10.1038/s41598-017-08978-9 (2017).
Schaber, C. L. et al. Breathprinting reveals malaria-associated biomarkers and mosquito attractants. J. Infect. Dis. https://doi.org/10.1093/infdis/jiy1072 (2018).
Berna, A. Z. et al. Analysis of breath specimens for biomarkers of plasmodium falciparum infection. J. Infect. Dis. 212, 1120–1128. https://doi.org/10.1093/infdis/jiv176 (2015).
Berna, A. Z. et al. Diurnal variation in expired breath volatiles in malaria-infected and healthy volunteers. J. Breath Res. 12, 046014. https://doi.org/10.1088/1752-7163/aadbbb (2018).
De Moraes, C. M. et al. Malaria-induced changes in host odors enhance mosquito attraction. Proc. Natl. Acad. Sci. USA 111, 11079–11084. https://doi.org/10.1073/pnas.1405617111 (2014).
Kelly, M. et al. Malaria parasites produce volatile mosquito attractants. mBiol https://doi.org/10.1128/mBio.00235-15 (2015).
Correa, R., Coronado, L. M., Garrido, A. C., Durant-Archibold, A. A. & Spadafora, C. Volatile organic compounds associated with Plasmodium falciparum infection in vitro. Parasit Vectors 10, 215. https://doi.org/10.1186/s13071-017-2157-x (2017).
Capuano, R. et al. Simultaneous Proton Transfer Reaction-Mass Spectrometry and electronic nose study of the volatile compounds released by Plasmodium falciparum infected red blood cells in vitro. Sci. Rep. 9, 12360. https://doi.org/10.1038/s41598-019-48732-x (2019).
Lacroix, R., Mukabana, W. R., Gouagna, L. C. & Koella, J. C. Malaria infection increases attractiveness of humans to mosquitoes. PLoS Biol. 3, e298. https://doi.org/10.1371/journal.pbio.0030298 (2005).
Busula, A. O. et al. Gametocytemia and attractiveness of Plasmodium falciparum–infected Kenyan children to Anopheles gambiae mosquitoes. J. Infect. Dis. https://doi.org/10.1093/infdis/jix214 (2017).
Cator, L. J., Lynch, P. A., Thomas, M. B. & Read, A. F. Alterations in mosquito behaviour by malaria parasites: potential impact on force of infection. Malar J. 13, 11. https://doi.org/10.1186/1475-2875-13-164 (2014).
Cator, L. J., Lynch, P. A., Read, A. F. & Thomas, M. B. Do malaria parasites manipulate mosquitoes?. Trends Parasitol. 28, 466–470. https://doi.org/10.1016/j.pt.2012.08.004 (2012).
Emami, S. N. et al. A key malaria metabolite modulates vector blood seeking, feeding, and susceptibility to infection. Science 355, 1076–1080. https://doi.org/10.1126/science.aah4563 (2017).
Crutcher, J. & Hoffman, S. in Medical Microbiology (ed S. Baron) Ch. 83, (1996).
Nyasembe, V. O., Teal, P. E., Mukabana, W. R., Tumlinson, J. H. & Torto, B. Behavioural response of the malaria vector Anopheles gambiae to host plant volatiles and synthetic blends. Parasit Vectors 5, 234. https://doi.org/10.1186/1756-3305-5-234 (2012).
Takken, W., Knols, B. G., Takken, W., editors. Olfaction in vector-host interactions. Vol. 2 (Wageningen Academic Publishers, 2010).
Pouline, M. O. et al. The potential role of exhaled breath analysis in the diagnostic process of pneumonia—a systematic review. J. Breath Res. 12, 024001 (2018).
Lung Cancer Indication Detection. https://clinicaltrials.gov/ct2/show/NCT02612532
Hanna, G. B., Boshier, P. R., Markar, S. R. & Romano, A. Accuracy and methodologic challenges of volatile organic compound-based exhaled breath tests for cancer diagnosis: a systematic review and meta-analysis. JAMA Oncol. 5, e182815–e182815. https://doi.org/10.1001/jamaoncol.2018.2815 (2019).
Fuchs, P., Loeseken, C., Schubert, J. K. & Miekisch, W. Breath gas aldehydes as biomarkers of lung cancer. Int. J. Cancer 126, 2663–2670. https://doi.org/10.1002/ijc.24970 (2010).
Schumer, E. M. et al. High sensitivity for lung cancer detection using analysis of exhaled carbonyl compounds. J. Thorac. Cardiovasc. Surg. 150, 1517–1524. https://doi.org/10.1016/j.jtcvs.2015.08.092 (2015).
Filipiak, W. et al. Release of volatile organic compounds (VOCs) from the lung cancer cell line CALU-1 in vitro. Cancer Cell Int. 8, 17. https://doi.org/10.1186/1475-2867-8-17 (2008).
Li, J. et al. Investigation of potential breath biomarkers for the early diagnosis of breast cancer using gas chromatography–mass spectrometry. Clin. Chim. Acta 436, 59–67. https://doi.org/10.1016/j.cca.2014.04.030 (2014).
Phillips, M. et al. Volatile biomarkers in the breath of women with breast cancer. J. Breath Res. 4, 026003. https://doi.org/10.1088/1752-7155/4/2/026003 (2010).
Altomare, D. F. et al. Exhaled volatile organic compounds identify patients with colorectal cancer. BJS (British Journal of Surgery) 100, 144–150. https://doi.org/10.1002/bjs.8942 (2013).
Peng, G. et al. Detection of lung, breast, colorectal, and prostate cancers from exhaled breath using a single array of nanosensors. Br. J. Cancer 103, 542–551. https://doi.org/10.1038/sj.bjc.6605810 (2010).
Schmutzhard, J. et al. Pilot study: Volatile organic compounds as a diagnostic marker for head and neck tumors. Head Neck 30, 743–749. https://doi.org/10.1002/hed.20779 (2008).
Hakim, M. et al. Diagnosis of head-and-neck cancer from exhaled breath. Br. J. Cancer 104, 1649–1655. https://doi.org/10.1038/bjc.2011.128 (2011).
Filipiak, W. et al. Molecular analysis of volatile metabolites released specifically by staphylococcus aureus and pseudomonas aeruginosa. BMC Microbiol. 12, 113. https://doi.org/10.1186/1471-2180-12-113 (2012).
Fowler, S. J., Basanta-Sanchez, M., Xu, Y., Goodacre, R. & Dark, P. M. Surveillance for lower airway pathogens in mechanically ventilated patients by metabolomic analysis of exhaled breath: a case-control study. Thorax 70, 320. https://doi.org/10.1136/thoraxjnl-2014-206273 (2015).
Schnabel, R. et al. Analysis of volatile organic compounds in exhaled breath to diagnose ventilator-associated pneumonia. Sci. Rep. 5, 17179. https://doi.org/10.1038/srep17179 (2015).
van Oort, P. et al. Exhaled breath metabolomics for the diagnosis of pneumonia in intubated and mechanically ventilated ICU–patients. Europ. Respirat. J. 50, OA4653. https://doi.org/10.1183/1393003.congress-2017.OA4653 (2017).
Jombart, T. adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24, 1403–1405. https://doi.org/10.1093/bioinformatics/btn129 (2008).
Jombart, T. & Ahmed, I. Adegenet 1.3–1: new tools for the analysis of genome-wide SNP data. Bioinformatics 27, 3070–3071. https://doi.org/10.1093/bioinformatics/btr521 (2011).
Oksanen, J. vegan: Community Ecology package. https://CRAN.R-project.org/package=vegan (2019).
Mitchell, M. An introduction to genetic algorithms. (MIT Press, 1996).
Goodacre, R. Making sense of the metabolome using evolutionary computation: seeing the wood with the trees. J. Exp. Bot. 56, 245–254. https://doi.org/10.1093/jxb/eri043 (2004).
Zou, W. & Tolstikov, V. V. Pattern recognition and pathway analysis with genetic algorithms in mass spectrometry based metabolomics. Algorithms https://doi.org/10.3390/a2020638 (2009).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 43, e47–e47. https://doi.org/10.1093/nar/gkv007 (2015).
Caret: Classification and regression training v. R package version 6.0–21 (CRAN: Wien, Austria, 2014).
Acknowledgements
We thank Caroline Wanjiku, James W. Sims, Heike S Betz, Andrew F. Read and Baldwyn Torto for the collection and analysis of the original dataset and/or the facilitation of the project, as well as everyone involved with the project at the International Centre of Insect Physiology and Ecology’s Thomas Odhiambo campus in Mbita, Kenya. Funding was provided by Bill and Melinda Gates Foundation Grant OPP1060415, and ETH Zürich. Predictive models were performed on the ETH Euler cluster.
Author information
Authors and Affiliations
Contributions
Conceived and designed the research: C.M.D.M., H.P., N.M.S., and M.C.M. Analyzed the data: H.P. Wrote the paper: N.M.S., H.P., C.M.D.M., and M.C.M.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pulido, H., Stanczyk, N.M., De Moraes, C.M. et al. A unique volatile signature distinguishes malaria infection from other conditions that cause similar symptoms. Sci Rep 11, 13928 (2021). https://doi.org/10.1038/s41598-021-92962-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-92962-x
This article is cited by
-
The volatilome signatures of Plasmodium falciparum parasites during the intraerythrocytic development cycle in vitro under exposure to artemisinin drug
Scientific Reports (2023)
-
A review on the significance of body temperature interpretation for early infectious disease diagnosis
Artificial Intelligence Review (2023)
-
No skin off your back: the sampling and extraction of sebum for metabolomics
Metabolomics (2023)
-
Manipulation by Plasmodium Parasites of Anopheles Mosquito Behavior and Human Odors
Acta Parasitologica (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.