Abstract
Neuroendocrine tumors (NETs) of the small intestine undergo large chromosomal and methylation changes. The objective of this study was to identify methylation differences in NETs and consider how the differentially methylated genes may impact patient survival. Genome-wide methylation and chromosomal copy number variation (CNV) of NETs from the small intestine and appendix were measured. Tumors were divided into three molecular subtypes according to CNV results: chromosome 18 loss (18LOH), Multiple CNV, and No CNV. Comparison of 18LOH tumors with MultiCNV and NoCNV tumors identified 901 differentially methylated genes. Genes from the G-protein coupled receptor (GPCR) pathways are statistically overrepresented in the differentially methylated genes. One of the highlighted genes from the GPCR pathway is somatostatin (SST), a clinical target for NETs. Patient survival based on low versus high methylation in all samples identified four significant genes (p < 0.05) OR2S2, SMILR, RNU6-653P, and AC010543.1. Within the 18LOH molecular subtype tumors, survival differences were identified in high versus low methylation of 24 genes. The most significant is TRHR (p < 0.01), a GPCR with multiple FDA-approved drugs. By separating NETs into different molecular subtypes based on chromosomal changes, we find that multiple GPCRs and their ligands appear to be regulated through methylation and correlated with survival. These results suggest opportunities for better treatment strategies for NETs based on molecular features.
Similar content being viewed by others
Introduction
Well-differentiated neuroendocrine tumors (NETs), previously referred to as carcinoids, can occur anywhere in the body, with the majority occurring in the gastrointestinal tract. Small intestinal NETs makeup almost half of these gastrointestinal NETs (GINETs) and are rare, with an incidence of 0.87 per 100,000 population per year1,2,3. Since GINETs are rare cancers, the molecular mechanisms driving pathologic changes remain elusive. Small intestinal NETs arise from enterochromaffin cells, which reside alongside the epithelial layer lining the lumen of the digestive tract, predominantly in the small intestine and appendix, where they regulate intestinal motility and secretion via the production of serotonin and other peptides4,5. Excess production of serotonin by the tumor may cause carcinoid syndrome, characterized by flushing and diarrhea, which occurs with about 17% of appendiceal and 32% of small intestinal NETs6.
GINETs often express neuroendocrine markers, such as synaptophysin, chromogranin, and somatostatin receptors (SSTRs) 1 to 5, which are G-protein coupled receptors and detected by immunohistochemistry. Positron emission tomography (PET) imaging using radiolabeled SSTR ligand tracers such as 68 Ga-DOTATE and 68 Ga-DOTATOC is a highly sensitive modality to diagnose patients with local or distant well-differentiated GINETs and also to evaluate for the potential role of somatostatin analogue (SSA) therapy (octreotide or lanreotide) in patients with metastatic NETs7,8. SSAs regulate hormonal hypersecretion, notably serotonin and other vasoactive substances in tumors expressing one or more subtypes of somatostatin receptors. Currently, metastatic GINETs are treated with the standard first-line SSA therapy, but there is evidence that different molecular subtypes are associated with different progression-free survival9. A better understanding of the molecular characteristics underlying GINETs may provide guidance for understanding the biology, the prognosis, and the selection of patients for more effective and targeted treatments.
The most frequent genomic alteration in small intestinal NETs is large chromosomal deletions. Most notably, full arms of chromosomes 18 are lost in 40–80% of tumors10,11,12. Inactivating mutations in CDKN1B are found in about 8% of small intestinal NETs, but small intestinal NETs otherwise are genetically stable, and somatic mutations in individual genes are uncommon13,14. Epigenetic modifications, in particular DNA methylation, have been proposed as a mechanism for small intestinal NET development9. DNA methylation can regulate gene expression and is an established mechanism for developing multiple types of cancer15. In general, but not exclusively, hypermethylation in promoter regions tends to decrease gene expression by blocking DNA binding sites for transcription factors16,17,18.
Multiple studies have considered the clinical significance of the loss of heterozygosity in chr18 (18LOH) with inconsistent results19. Karpathakis et al. suggested that small intestinal NETs with 18LOH are associated with favorable progression free survival following resection9. Yao et al. reported the 18LOH patients to have better survival outcomes10. Kim et al. found no significant difference in overall survival between those with and without 18LOH20. Although there is evidence of methylation changes in small intestinal NETs with 18LOH, the molecular pathways are not well defined9.
The objective of this study was to identify methylation differences in small intestinal and appendiceal NETs based on genomic alteration (molecular subtype) and to consider how the differentially methylated genes may impact survival.
Materials and methods
All methods were carried out in accordance with relevant guidelines and regulations. The study was approved by the University of Utah Institutional Review Board. Informed consent was obtained from all living participants.
Research participants
Potential NET patient cases including both small intestine and appendix (histology codes 8240, 8241, 8243, 8244, 8246; ICDO locations 170, 171, 172, 173, 178, 179, 181) in Utah between 1999 and 2014 were identified through the Utah Cancer Registry (UCR). UCR is a Surveillance Epidemiology and End Results (SEER) registry that has collected all cancers diagnosed in Utah from 1966 to the present. Cases from subjects under 18 years old and in situ cases were excluded, leaving 552 potential NET subjects for study (Table 1). Additional cases with a diagnosis from 2014 to 2018 were identified through University of Utah Health electronic medical records and referred to the study by the Department of Surgery. UCR referred 77 cases reported to UCR by the University of Utah Health and had a pathology number on record. The Department of Surgery referred 12 additional recent cases. From these 89 cases, 47 subjects had archived FFPE blocks available, were confirmed as a well-differentiated neuroendocrine tumor by study pathologist (KA), and had sufficient tissue for methylation analysis. Confirmation by the study pathologist was essential as multiple cases were not assigned the correct histology code.
Methylation analysis of tumor DNA
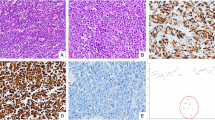
Formalin-fixed paraffin-embedded (FFPE) blocks were serially cut in 5 μm increments and mounted onto charged slides. Hematoxylin and eosin (H&E) staining of a representative slide was used to identify areas of neuroendocrine tumor (> 50% tumor) by the study pathologist (KA). Cells were microdissected from the marked tumor region, and DNA was extracted from FFPE tissue slides using a QIAamp DNA FFPE Tissue Kit [Qiagen #56404] using the manufacturer's protocol with the following modifications: (1) paraffin was removed from the slides prior to microdissection, and the tissue was added directly to Buffer ATL (2) there were two 56 °C incubations with Proteinase K: overnight, and then for an hour after supplementing with an additional 20 µL Proteinase K (3) for the elution, 2 × 25 µL of ATE was added to the center of the membrane and incubated for 5 min at room temperature before eluting.
Following extraction, DNA concentrations were measured with a Qubit™ dsDNA BR assay kit [ThermoFisher Scientific #Q32850 & #Q32856]. Bisulfite conversion was performed on 250 ng DNA per sample using a Zymo EZ DNA Methylation Kit [Zymo Research #D5001] following the manufacturer's recommended protocol for methylation microarrays. DNA quality was evaluated with an Infinium FFPE QC kit [Illumina #WG-321-1001] prior to performing the recommended FFPE DNA Restore protocol [Illumina #WG-321-1002 & Zymo #D4024]. MethylationEPIC BeadChip microarrays [Illumina #WG-317-1002] were used for genome-wide methylation profiling. The microarrays were processed with an Illumina iScan platform using the Infinium HD Methylation assay protocol with the recommended modifications for FFPE samples. The raw methylation IDAT data was analyzed using GenomeStudio (RRID:SCR_010973) v2011.1 software with Methylation Module v1.9 [Illumina].
DNA methylation data pre-processing
The methylation data were pre-processed following a pipeline of the ChAMP (RRID:SCR_012891) R package (V.2.13.5)21. Probes were filtered out using the champ.filter function with the following criteria: probes with (1) detection p > 0.01, (2) less than three beads in at least 5% of samples, (3) non-CpG cytosine, (4) polymorphic nucleotide, (5) multiple regions of the genome and (6) chromosome X or Y. Total of the 699,602 CpG sites were selected for further analysis. After the quality control, the type-II probe bias was corrected by the method of beta mixture quantile (BMIQ) normalization according to pipeline defaults.
Copy number variants using methylation data
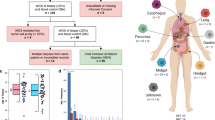
In order to identify copy number variation (CNV) in our 47 NET cases, intensity values for each probe were used to count copy numbers and determine if alterations were present. Copy number of our NETs were compared with previously published Illumina MethylationEPIC BeadChip data measured from ten normal myometrial samples in NCBI GEO database (GSM3417135-GSM3417144)22. CNV was estimated and compared from the intensity value using the champ.CNA function of ChAMP23. The whole-chromosome arm CNV was defined with the criteria of length > 80% and segment mean > ± 0.2. The 47 NETs were classified into three groups according to CNV results: Chromosome 18 loss of heterozygosity (18LOH), no chromosome arm copy number alterations (NoCNV), and multiple copy number alterations excluding those with 18LOH (MultiCNV) (Fig. 1).
Chromosomal rearrangements (molecular subtypes) of neuroendocrine tumors. The ChAMP result referring copy number variation, including loss and gain of copy number. Samples were grouped into three subgroups: loss of chromosome 18 (18LOH), no copy number variation (NoCNV) or multiple copy number variations (MultiCNV). Black indicates gain of copy number. Grey indicates loss of copy number.
Differential methylation analysis
The averaged methylation value (β-value) of all CpGs within the promoter region of a gene was compared between the three CNV groups (i.e., 18LOH vs. NoCNV, 18LOH vs. MultiCNV, and MultiCNV vs. NoCNV) with an unpaired Wilcoxon's test to identify differentially methylated genes. A parallel analysis of singular CpG sites that are differentially methylated was done (Supplementary Table S2). The promoter region was considered 1500 bp upstream and 200 bp downstream from the transcription start site (TSS). A total of 39,252 genes were tested. An FDR adjusted p value below 0.05 and change in methylation greater than 15% (Δβ > 0.15) were used to identify genes with statistically significant differential methylation levels. The pheatmap package in R was used to create the heatmap. The default method for “Euclidean” distance and the heatmap clustering method Ward.D was used (v 1.0.12)24.
Functional enrichment and network analysis
Functional enrichment analysis using ConsensusPathDB (RRID:SCR_002231) (Release 33, CPDB; http://cpdb.molgen.mpg.de/)25 was used to interpret the functional role of genes identified with differential methylation status. Significant pathways and GO terms were defined as having an adjusted q-value < 0.05 (p value corrected for multiple testing based on the number of pathways used). To reduce redundancy and remove potential false-positive GO terms, we used the GO-module web-tool (http://lussierlab.org/GO-Module) (v.1.3)26. Protein–protein interaction (PPI) network was constructed using StringDB V.11 (STRING, RRID:SCR_005223, http://string-db.org)27 with an interaction combined score of > 0.9, which represents the highest confidence score of the various evidence (i.e., text mining, database, and co-expression). PPI network was visualized by using Cytoscape (RRID:SCR_003032) v3.5.128.
Survival analysis
Survival in months after diagnosis was determined from death certificates (n = 16). If the individual was alive at last contact (n = 31), a study-end month was set at June 2019, the most recent contact date among research subjects.
Overall survival outcome was evaluated using Kaplan–Meier survival analysis with several comparing criteria. Analyses were conducted according to the three molecular subgroups (i.e., 18LOH, NoCNV, and MultiCNV). We also considered survival based on the methylation status of gene promoters (n = 39,252 genes including coding, noncoding, and pseudogenes). The 47 NET samples were separated into two groups according to β-value (i.e., low- and high-methylated groups) using k-means clustering with the number of clusters set at two for survival analysis. Additionally, within the 18LOH tumors (n = 19) group, survival was compared based on high versus low methylation according to the β-value using k-means clustering with the number of clusters set at two. We defined statistically significant associations as a p value < 0.05.
Results
Overview of the study cohort
Although the number of tumors tested for methylation (1999–2018) was modest, this subset was generally representative of all small intestine and appendix NET cases reported to the UCR in 1999–2014 (Table 1). None of the differences in the values reported in Table 1 reach statistical significance. In those cases tested, a larger percentage had tumor grade information available: 79% versus 43% for the cases who did not undergo testing. This is expected because the cases tested, on average, were more recent diagnoses and came from an academic institution where tumor grading was more common. Restricting the comparison to tumors that were graded, the tumors tested versus tumors of cases that were not tested were similar in the frequency of grade, with the majority (70% and 69%, respectively) being grade 1. Known risk factors of tobacco use and family history were similar between cases with and without testing as well. Survival did not significantly differ between the two groups (p = 0.76; Supplementary Fig. S1).
Genome-wide DNA methylation copy number variation profiling
Analysis of copy number variation (CNV) in the tumor, based on DNA methylation array data, showed losses on the entire chromosome 18 (18LOH) in 19 out of 47 tumors (40.4%) and gain of chromosome 5 in 5 tumors (10.6%), similar to previous reports9,10,19,20. Multiple CNV (MultiCNV) were observed in 8 tumors (17.0%), and no copy-number alternation (NoCNV) was observed in the remaining 20 tumors (42.6%), three of which are from the appendix (NoCNV_6, NoCNV_7, NoCNV_8; Fig. 1). No statistically significant differences in patient demographics or tumor characteristics were observed, but this was limited by the small numbers (Table 1). Tumors with 18LOH trend towards a more advanced stage and higher rates of tobacco use than NoCNV or MultiCNV tumors, but this difference is not significant.
Differential DNA methylation analysis in the three subgroups
A subset of 196,354 CpG methylation sites out of a total of 699,602 CpG sites was mapped to specific promoters (39,252 genes) for analysis (Fig. 2a). Differential methylation was examined in individuals with a history of tobacco use versus those without. Hierarchical clustering of differentially methylated genes showed no difference between tumors of those who have a history of tobacco use versus those that do not. To identify differentially methylated genes related to 18LOH, intergroup comparisons were performed for 18LOH vs. MultiCNV, 18LOH vs. NoCNV, and MultiCNV vs. NoCNV (FDR < 0.05 and Δβ > 0.15); the number of differentially methylated genes are 1332, 1259, and 0, respectively when considering average promoter methylation (Fig. 2b) and 6790, 8923, and 0 respectively when considering singular CpG methylation (data not shown). An intersection of differentially methylated genes in the 18LOH group was identified; 901 genes when average promoter methylation is used and 2230 genes (2945 singular CpGs) when singular CpG methylation is used (Supplementary Table 2). The 901-gene set for average differential promoter methylation was examined further, but not the singular CpG analysis because it can have issues with independence and false positives29. Interestingly, none of the 901 genes with differential methylation reside on chromosome 18 (Supplementary Table 1) but do reside on the other 21 autosomal chromosomes. Aside from having a reduced intensity, the β-values for methylation probes on chromosome 18 in the 18LOH tumors are normal. We hypothesize that the lack of differentially regulated genes on chromosome 18 is due to hypomethylation preceding and promoting 18LOH, and once chromosome 18 is lost, methylation differences in the remaining chromosome are not observed. Another interesting observation is that 205 (22.8%) of the 901 differentially methylated genes were annotated as long noncoding RNA (lncRNA). Heatmap clustering of the 901 genes shows that these genes are predominately hypomethylated in 18LOH tumors when compared to MultiCNV and NoCNV groups. (Fig. 2c). Only 9 genes (1%) were hypermethylated in the 18LOH tumors (SST, DMBT1, RNU6-1039P, RP11-37N22.1, Lnc-FAM241A-1, RP11-153K16.1, CERS3-AS1, RP11-542M13.3, and AC025278.2; Fig. 2c). Hierarchical clustering of the tumors based on methylation of the 901 genes shows the separation of the 18LOH tumors from the other two molecular subtypes (Fig. 2d). The three neuroendocrine tumors from the appendix which were included in the study cluster with the NoCNV molecular subtype (NoCNV_6, NoCNV_7, NoCNV_8). When the analysis was repeated, removing these three appendiceal samples, very similar results were found. The coefficient of correlation was 0.962 when comparing the p value for methylation differences of the 901 genes from the original analysis of 47 tumors to the analysis of 44 tumors excluding appendiceal NETs (Supplementary Fig. 5).
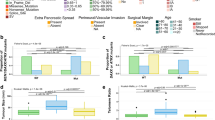
Differential methylation status between molecular subtypes. (a) A schema methylation probes within promoter region. Methylation probes between − 1500 and 200 bp from the transcript start site are considered. (b) The number of significantly differential methylation genes of each comparison. The overlapped differential methylation of 18LOH versus Multi CNV and 18LOH versus No CNV is considered as 901 genes specific to 18LOH. (c) Proportion of hypomethylated (n = 892, 99%) and hypermethylated (n = 9, 1%) genes in 18LOH tumors. (d) Heatmap of 901 differentially methylated genes for each tumor. Blue indicates hypomethylation. Red indicates hypermethylation. Tumor molecular subtypes are indicated in green (18LOH), pink (MultiCNV) or blue (NoCNV).
Functional enrichment analysis and network of significant methylation of 18LOH tumors
Overrepresentation analysis of the 901 genes, using ConsensusPathDB, revealed 12 enriched functional pathways (q-value < 0.05) (Supplementary Fig. S2). This includes multiple signaling-related pathways: GPCR signaling (including olfactory receptors), defensin and beta-defensin, neuroactive ligand-receptor interaction, vitamin D receptor, and hemostasis. Within the set of 901 differentially methylated genes, 18 Gene Ontology (GO) pathways were significantly over-represented (q-value < 0.05), including “Olfactory receptor activity” (GO:0004984), “Sensory perception of chemical stimulus” (GO:0007606), “G-protein-coupled receptor signaling pathway” (GO:0004930), “defense response to bacterium” (GO:0042742), and “humoral immune response” (GO:0006959) (Supplementary Fig. S2). The protein–protein interaction (PPI) network constructed with the STRING database identified 51 genes with direct interactions (Fig. 3). When genes from the 12 enriched functional pathways were superimposed on the PPI network, the 51 genes were functionally grouped into four subgroup pathways: GPCR downstream signaling (including olfactory receptors), neuroactive ligand-receptor interaction, beta-defensins, and hemostasis. The GPCR downstream signaling pathway is highly related to cancer progression behaviors such as proliferation, angiogenesis, and metastasis30,31,32. Multiple genes were assigned to both GPCR and neuroactive ligand-receptor interaction pathways.
Protein–protein interaction network of 901 genes with differential methylation in 18LOH group. STRING Database with input of the 901 genes predicted protein–protein interactions which are either direct (physical) or indirect (functional). Genes that met the highest confidence for interaction (> 0.9) are shown in the Fig. (51 genes). Lines connect genes that were predicted for interacting. Overlay of pathways that are overrepresented in the set of 901 genes, identified four pathways in this subset of 51 genes. Genes assigned to the pathway are indicated as follows: GPCR signaling (purple); Neuroactive ligand-receptor interaction (yellow); Beta-defensins (green) and Hemostasis (pink). For genes assigned to multiple pathways, the circle is multiple colors.
When the 47 tumors were grouped into high methylation versus low methylation for all 39,252 genes and not considering their CNV status, four genes were significantly associated with survival based on methylation status, p < 0.05; hypermethylation confers better survival for OR2S2, RNU6-653P and AC010543.1 and hypomethylation confers better survival for SMILR (Supplementary Fig. S4). Conflicting reports on survival of patients with SINETs with chromosome 18 loss led us to also consider heterogeneity in methylation within this group. The 19 tumors with 18LOH were separated into high methylation or low methylation for each of the 901 differentially methylated genes, and survival analysis was compared. A significant difference in survival (p < 0.05) was observed for 24 of the genes (Supplementary Table S1). Two are included in the PPI network (Fig. 3): TRHR (p = 0.006) and VNN2 (p = 0.050).
By way of example, two genes whose methylation may have biological importance in neuroendocrine tumors are included in the large protein–protein interaction network in Fig. 3: TRHR (thyrotropin-releasing hormone receptor), a G-protein coupled receptor, and SST (somatostatin), a ligand to the SSTR G-protein coupled receptor. The TRHR promoter is hypomethylated on all six CpG methylation sites in tumors with 18LOH relative to tumors with NoCNV (p = 0.004) or with MultiCNV (p = 0.04) (Fig. 4a,b. Survival analysis of the 18LOH tumors suggests that low methylation of the TRHR gene confers poor survival p = 0.005 (Fig. 4c). Methylation of the TRHR promotor silences transcription of the gene in thyroid cancers33. This suggests that reduced TRHR expression due to promoter methylation improves survival.
Methylation status of TRHR gene promoter. (a) A plot of beta-values of methylation probes within the promoter (Y-axis) and genomic coordinates (X-axis). The promoter region (pink), exon (orange) and intron are presented a bottom line of the graph. (b) A boxplot including distributions of average beta-value of methylation probes in the promoter for each molecular subtype. (c) Survival analysis comparing 18LOH tumors with high methylation (red line; samples 18LOH_2, _4, _5, _6, _7, _9, _14, _15, _16) and low methylation (blue line; samples 18LOH_1, _3, _8, _10, _11, _12, _13, _17, _18, _19) of TRHR promoter.
The SST promoter is hypermethylated in the 18LOH tumors relative to the MultiCNV (p = 0.006) and NoCNV (p = 0.002) tumors (Fig. 5a,b). Hypermethylation of the SST promoter in gastric cancers results in decreased mRNA and protein expression, thereby reducing somatostatin's ability to inhibit tumor growth34. The somatostatin protein binds to G-protein coupled somatostatin receptors, and analogs of somatostatin are a common treatment for GINETs8.
Methylation status in the SST gene promoter. (a) A plot of beta-values of methylation probes within the promoter (Y-axis) and genomic coordinates (X-axis). The promoter region (pink), exon (orange), intron, CpG island (green), and shore (dark orange) are presented a bottom line of the graph. (b) A boxplot including distributions of average beta-value of methylation probes in the promoter for each molecular subtype.
Discussion
In this study, we investigated differential DNA methylation status in well-differentiated small intestinal and appendiceal NETs with different molecular subtypes. We found 901 genes with differentially methylated promoter regions in the study's 18LOH NETs, and these genes were enriched in tumor-related pathways, including “GPCR downstream signaling”, “neuroactive ligand-receptor interaction”, “beta-defensins”, and “hemostasis”. Most of the genes were hypomethylated, broadly suggesting transcriptional activation of these genes. None of the 901 genes resided on chromosome 18, suggesting that hypomethylation of chromosome 18 may contribute to its loss in NETs. Chromosomal instability in tumors is promoted by DNA hypomethylation and is observed in multiple tumor types including pancreatic NETs35,36. The lncRNA, which made up 22.8% of the differentially methylated genes, has been shown to regulate cancer genes and represent additional disease pathways37,38,39,40. We examined the influence of tobacco use on methylation in GINET tumors. However, differences were not observed when comparing tumors from patients with and without an indication of tobacco use41. This may be due to the small number of samples, timing of use (current versus past), and/or the quantity (pack-years)42.
The GPCR pathway represents a large family of cell-surface molecules regulating the signal transmission for multiple cellular functions. GPCRs are also the most common class of therapeutic targets, with approximately 700 FDA-approved drugs targeting 128 GPCRs43,44. Pertinent to the findings of this study, GPCRs play multiple roles in cancer development and have been the focus of past studies to define differences in neuroendocrine tumors, primarily those originating from the small intestine versus pancreas and lung45,46. Our significantly enriched GPCR pathways suggest that expression of GPCR pathways is controlled through methylation events, which drives tumor progression a subset of GINETs46,47. Being neuroendocrine tumors, it makes sense that the “neuroactive ligand-receptor interaction” pathway is involved, and accordingly, is part of the GPCR downstream signaling pathway. We describe hypermethylation of one such neuroactive ligand, the SST gene encoding somatostatin, in 18LOH tumors relative to NoCNV and MultiCNV. Somatostatin is the ligand for SSTRs, G-protein coupled somatostatin receptors that are overexpressed in a subset of GINETs and the target of somatostatin analog therapies. Examining the clinical response to somatostatin analog therapies relative to SST methylation state may provide guidance on more targeted treatment strategies.
Our survival analysis with the three molecular subgroups based on chromosomal changes showed there was no significant difference between subgroups (p = 0.41; Supplementary Fig. S3). However, the ability to detect such differences is limited by the sample size. Survival analysis based on promoter methylation did identify four associated genes OR2S2, SMILR, RNU6-653P, and AC010543, although very little is known about their biologic function, and nothing is published related to cancer. We also identified 24 genes with methylation levels associated with survival outcomes, specifically in 18LOH tumors, potentially explaining why there is disagreement in the literature on survival outcomes for 18LOH tumors. The most significant association was with TRHR. Hypomethylation of the TRHR promoter may be associated with 18LOH tumor progression and survival outcomes. The mechanism behind this survival difference is not obvious but may represent a constellation of methylation events that are clinically relevant, potentially as a pharmacologic target for small intestinal NET patients.
Selection of tumors from the cancer registry included ICDO codes for the small intestine as well as the appendix because we believe there is cross-over in the etiology of these tumors. Three of the 47 tumors examined for methylation were from the appendix. All three were classified as NoCNV subtype, and hierarchical clustering of their methylation profile was fully integrated with the other NoCNV samples from the small intestine, suggesting a similar etiology. Additionally, we have observed neuroendocrine tumors of the appendix in families with multiple NETs of the small intestine, which suggests there is overlap in the genetic etiology.
Limitations of the study included a small sample size, methylation arrays that were not run on matched normal tissues, and a lack of treatment data for the NET patients. Also, the methylation changes were not validated with RNA expression profiling but rather relied on published results in different tissues and frequently observed (but not exclusive) negative correlation between promoter methylation and gene expression in other studies18. This study lays the foundation for future work to address these limitations. Strengths are that the samples tested were representative of all tumors reported in the state for that period of time. The frequency of molecular subtypes was similar to previous reports9,11,12,13.
In conclusion, by separating small intestine and appendiceal NETs into different molecular subtypes based on chromosomal changes, we were able to define major pathways that are differentially methylated. Two relevant pathways were the GPCR and the overlapping neuroactive ligand-receptor interaction. One gene that is differentially hypermethylated in 18LOH tumors is SST, encoding somatostatin. Somatostatin analogs are a primary treatment for GINETs, targeting overexpressed somatostatin receptors on the tumors. Somatostatins are also used for GINET visualization, radiotherapy, and repressing carcinoid-syndrome side effects. GPCR may be important for survival in small intestinal NETs and are targetable by drugs in some cases. In future work, methylation status in these genes can be explored as a prognostic and/or predictive biomarker to predict responses to SSA's and for careful selection of patients for appropriate treatments. Future studies that incorporate detailed histologic characterization, GPCR expression, and treatments will be important. The molecular characterization of GINETs can lead to novel prognostic and predictive biomarkers to better inform treatments and may improve the survival of patients with GINETs.
References
Modlin, I. M., Lye, K. D. & Kidd, M. A 5-decade analysis of 13,715 carcinoid tumors. Cancer 97, 934–959. https://doi.org/10.1002/cncr.11105 (2003).
DiSario, J. A., Burt, R. W., Vargas, H. & McWhorter, W. P. Small bowel cancer: epidemiological and clinical characteristics from a population-based registry. Am. J. Gastroenterol. 89, 699–701 (1994).
Kharazmi, E., Pukkala, E., Sundquist, K. & Hemminki, K. Familial risk of small intestinal carcinoid and adenocarcinoma. Clin. Gastroenterol. Hepatol. 11, 944–949. https://doi.org/10.1016/j.cgh.2013.02.025 (2013).
Bertrand, P. P. & Bertrand, R. L. Serotonin release and uptake in the gastrointestinal tract. Auton. Neurosci. 153, 47–57. https://doi.org/10.1016/j.autneu.2009.08.002 (2010).
Graeme-Cook, F. Surgical Pathology of the GI Tract, Liver, Biliary Tract, and Pancreas (2nd Edition) (eds Odze, R. D. & Goldblum, J. R.) 653–680 (W.B. Saunders, 2009).
Halperin, D. M. et al. Frequency of carcinoid syndrome at neuroendocrine tumour diagnosis: a population-based study. Lancet Oncol. 18, 525–534. https://doi.org/10.1016/S1470-2045(17)30110-9 (2017).
National Comprehensive Cancer Network. Clinical Practice Guidelines in Oncology Neuroendocrine and Adrenal Tumors (version 1.2019). (2019).
Cives, M., Soares, H. P. & Strosberg, J. Will clinical heterogeneity of neuroendocrine tumors impact their management in the future? Lessons from recent trials. Curr. Opin. Oncol. 28, 359–366. https://doi.org/10.1097/CCO.0000000000000299 (2016).
Karpathakis, A. et al. Prognostic impact of novel molecular subtypes of small intestinal neuroendocrine tumor. Clin. Cancer Res. 22, 250–258. https://doi.org/10.1158/1078-0432.CCR-15-0373 (2016).
Yao, J. et al. Genomic profiling of NETs: a comprehensive analysis of the RADIANT trials. Endocr. Relat. Cancer 26, 391–403. https://doi.org/10.1530/ERC-18-0332 (2019).
Kulke, M. H. et al. High-resolution analysis of genetic alterations in small bowel carcinoid tumors reveals areas of recurrent amplification and loss. Genes Chromosom. Cancer 47, 591–603. https://doi.org/10.1002/gcc.20561 (2008).
Hashemi, J. et al. Copy number alterations in small intestinal neuroendocrine tumors determined by array comparative genomic hybridization. BMC Cancer 13, 505. https://doi.org/10.1186/1471-2407-13-505 (2013).
Francis, J. M. et al. Somatic mutation of CDKN1B in small intestine neuroendocrine tumors. Nat. Genet. 45, 1483–1486. https://doi.org/10.1038/ng.2821 (2013).
Banck, M. S. et al. The genomic landscape of small intestine neuroendocrine tumors. J. Clin. Invest. 123, 2502–2508. https://doi.org/10.1172/JCI67963 (2013).
Wajed, S. A., Laird, P. W. & DeMeester, T. R. DNA methylation: an alternative pathway to cancer. Ann. Surg. 234, 10–20. https://doi.org/10.1097/00000658-200107000-00003 (2001).
Esteller, M. Epigenetics in cancer. N. Engl. J. Med. 358, 1148–1159. https://doi.org/10.1056/NEJMra072067 (2008).
Rauluseviciute, I., Drablos, F. & Rye, M. B. DNA hypermethylation associated with upregulated gene expression in prostate cancer demonstrates the diversity of epigenetic regulation. BMC Med. Genom. 13, 6. https://doi.org/10.1186/s12920-020-0657-6 (2020).
Spainhour, J. C., Lim, H. S., Yi, S. V. & Qiu, P. Correlation patterns between DNA methylation and gene expression in The Cancer Genome Atlas. Cancer Inform. 18, 1176935119828776. https://doi.org/10.1177/1176935119828776 (2019).
Samsom, K. G. et al. Molecular prognostic factors in small-intestinal neuroendocrine tumours. Endocr. Connect. 8, 906–922. https://doi.org/10.1530/EC-19-0206 (2019).
Kim, D. H. et al. Allelic alterations in well-differentiated neuroendocrine tumors (carcinoid tumors) identified by genome-wide single nucleotide polymorphism analysis and comparison with pancreatic endocrine tumors. Gene Chromosome Cancer 47, 84–92. https://doi.org/10.1002/gcc.20510 (2008).
Tian, Y. et al. ChAMP: updated methylation analysis pipeline for Illumina BeadChips. Bioinformatics 33, 3982–3984. https://doi.org/10.1093/bioinformatics/btx513 (2017).
George, J. W. et al. Integrated epigenome, exome, and transcriptome analyses reveal molecular subtypes and homeotic transformation in uterine fibroids. Cell Rep. 29, 4069–4085. https://doi.org/10.1016/j.celrep.2019.11.077 (2019).
Feber, A. et al. Using high-density DNA methylation arrays to profile copy number alterations. Genome Biol. 15, R30. https://doi.org/10.1186/gb-2014-15-2-r30 (2014).
Clifford, H., Wessely, F., Pendurthi, S. & Emes, R. D. Comparison of clustering methods for investigation of genome-wide methylation array data. Front. Genet. 2, 88. https://doi.org/10.3389/fgene.2011.00088 (2011).
Kamburov, A., Stelzl, U., Lehrach, H. & Herwig, R. The ConsensusPathDB interaction database: 2013 update. Nucleic Acids Res. 41, D793-800. https://doi.org/10.1093/nar/gks1055 (2013).
Yang, X., Li, J., Lee, Y. & Lussier, Y. A. GO-Module: functional synthesis and improved interpretation of Gene Ontology patterns. Bioinformatics 27, 1444–1446. https://doi.org/10.1093/bioinformatics/btr142 (2011).
Szklarczyk, D. et al. The STRING database in 2017: quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res. 45, D362–D368. https://doi.org/10.1093/nar/gkw937 (2017).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504. https://doi.org/10.1101/gr.1239303 (2003).
Chatterjee, A., Rodger, E. J., Morison, I. M., Eccles, M. R. & Stockwell, P. A. Tools and strategies for analysis of genome-wide and gene-specific DNA methylation patterns. Methods Mol. Biol. 1537, 249–277. https://doi.org/10.1007/978-1-4939-6685-1_15 (2017).
Gad, A. A. & Balenga, N. The emerging role of adhesion GPCRs in cancer. ACS Pharmacol. Transl. Sci. 3, 29–42. https://doi.org/10.1021/acsptsci.9b00093 (2020).
Singh, A., Nunes, J. J. & Ateeq, B. Role and therapeutic potential of G-protein coupled receptors in breast cancer progression and metastases. Eur. J. Pharmacol. 763, 178–183. https://doi.org/10.1016/j.ejphar.2015.05.011 (2015).
Khalil, B. D. et al. GPCR signaling mediates tumor metastasis via PI3Kbeta. Cancer Res. 76, 2944–2953. https://doi.org/10.1158/0008-5472.CAN-15-1675 (2016).
Xing, M. et al. Methylation of the thyroid-stimulating hormone receptor gene in epithelial thyroid tumors: a marker of malignancy and a cause of gene silencing. Cancer Res. 63, 2316–2321 (2003).
Shi, X., Li, X., Chen, L. & Wang, C. Analysis of somatostatin receptors and somatostatin promoter methylation in human gastric cancer. Oncol. Lett. 6, 1794–1798. https://doi.org/10.3892/ol.2013.1614 (2013).
Eden, A., Gaudet, F., Waghmare, A. & Jaenisch, R. Chromosomal instability and tumors promoted by DNA hypomethylation. Science 300, 455. https://doi.org/10.1126/science.1083557 (2003).
Marinoni, I. et al. Hypo-methylation mediates chromosomal instability in pancreatic NET. Endocr. Relat. Cancer 24, 137–146. https://doi.org/10.1530/ERC-16-0554 (2017).
Chen, B. H. et al. New insights into long noncoding RNAs and pseudogenes in prognosis of renal cell carcinoma. Cancer Cell Int. https://doi.org/10.1186/s12935-018-0652-6 (2018).
Wu, C. et al. Identification of cancer-related potential biomarkers based on lncRNA-pseudogene-mRNA competitive networks. FEBS Lett. 592, 973–986. https://doi.org/10.1002/1873-3468.13011 (2018).
Stewart, G. L. et al. Aberrant expression of pseudogene-derived lncRNAs as an alternative mechanism of cancer gene regulation in lung adenocarcinoma. Front. Genet. 10, 138. https://doi.org/10.3389/fgene.2019.00138 (2019).
Lian, Y. et al. The pseudogene derived from long non-coding RNA DUXAP10 promotes colorectal cancer cell growth through epigenetically silencing of p21 and PTEN. Sci. Rep. 7, 7312. https://doi.org/10.1038/s41598-017-07954-7 (2017).
Curtin, K. et al. Associations of tobacco and alcohol use with risk of neuroendocrine tumors of the small intestine in Utah. Cancer Epidemiol. Biomark. Prev. 28, 1998–2004. https://doi.org/10.1158/1055-9965.EPI-19-0465 (2019).
Breitling, L. P., Yang, R., Korn, B., Burwinkel, B. & Brenner, H. Tobacco-smoking-related differential DNA methylation: 27K discovery and replication. Am. J. Hum. Genet. 88, 450–457. https://doi.org/10.1016/j.ajhg.2011.03.003 (2011).
Cornwell, A. C. & Feigin, M. E. Unintended effects of GPCR-targeted drugs on the cancer phenotype. Trends Pharmacol. Sci. 41, 1006–1022. https://doi.org/10.1016/j.tips.2020.10.001 (2020).
Sriram, K. & Insel, P. A. G protein-coupled receptors as targets for approved drugs: How many targets and how many drugs?. Mol. Pharmacol. 93, 251–258. https://doi.org/10.1124/mol.117.111062 (2018).
Dorsam, R. T. & Gutkind, J. S. G-protein-coupled receptors and cancer. Nat. Rev. Cancer 7, 79–94. https://doi.org/10.1038/nrc2069 (2007).
Cui, T. et al. Olfactory receptor 51E1 protein as a potential novel tissue biomarker for small intestine neuroendocrine carcinomas. Eur. J. Endocrinol. 168, 253–261. https://doi.org/10.1530/EJE-12-0814 (2013).
Drozdov, I. et al. Gene network inference and biochemical assessment delineates GPCR pathways and CREB targets in small intestinal neuroendocrine neoplasia. PLoS ONE 6, e22457. https://doi.org/10.1371/journal.pone.0022457 (2011).
Acknowledgements
We gratefully acknowledge support from the Center for High Performance Computing and the Genomics Core Facility at the University of Utah and the Utah Population Database. We would like to acknowledge Megan Keener for assisting with sample acquisition and Joshua Unger and Greyson Hullinger for processing the tumor samples.
Funding
This work was supported by Grants from the National Cancer Institute at the National Institutes of Health, R21CA205796 (DWN) and P30CA042014 and Huntsman Cancer Foundation. The Utah Cancer Registry was funded by the National Cancer Institute’s SEER Program, Contract No. HHSN261201800016I, the US Centers for Disease Control and Prevention’s National Program of Cancer Registries, Cooperative Agreement No. NU58DP006320, with additional support from the University of Utah and Huntsman Cancer Foundation. Support for the computational resources at the Center for High Performance Computing at the University of Utah were partially funded by the NIH Shared Instrumentation Grant 1S10OD021644-01A1.
Author information
Authors and Affiliations
Contributions
S.B., K.A., A.S., and D.N. wrote the main manuscript. S.B. prepared Figs. 1, 2, 3, 4 and 5 and supplemental tables. D.N. prepared Table 1. S.B., K.A., A.S., K.C., A.C. and D.N. generated data and conducted analysis. L.C.A., R.T. and D.N. conceptualized the study. All authors revised and reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Byun, S., Affolter, K.E., Snow, A.K. et al. Differential methylation of G-protein coupled receptor signaling genes in gastrointestinal neuroendocrine tumors. Sci Rep 11, 12303 (2021). https://doi.org/10.1038/s41598-021-91934-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-91934-5
This article is cited by
-
Hypermethylated GPR135 gene expression is a favorable independent prognostic factor in nasopharyngeal carcinoma
Holistic Integrative Oncology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.