Abstract
DNA methylation plays important functions in gene expression regulation that is involved in individual development and various diseases. DNA methylation has been well studied in human and model organisms, but only limited data exist in companion animals like dog. Using methylation-sensitive restriction enzyme-based next generation sequencing (Canine DREAM), we obtained canine DNA methylation maps of 16 somatic tissues from two dogs. In total, we evaluated 130,861 CpG sites. The majority of CpG sites were either highly methylated (> 70%, 52.5–64.6% of all CpG sites analyzed) or unmethylated (< 30%, 22.5–28.0% of all CpG sites analyzed) which are methylation patterns similar to other species. The overall methylation status of CpG sites across the 32 methylomes were remarkably similar. However, the tissue types were clearly defined by principle component analysis and hierarchical clustering analysis with DNA methylome. We found 6416 CpG sites located closely at promoter region of genes and inverse correlation between DNA methylation and gene expression of these genes. Our study provides basic dataset for DNA methylation profiles in dogs.
Similar content being viewed by others
Introduction
DNA methylation is the conversion of cytosine to 5-methylcytosine at cytosine-guanine (CpG) dinucleotides, causing structural change in the interactions between DNA and protein(s). DNA methylation occurring at promoter regions of genes represses gene transcription1. DNA methylation at intergenic regions and gene bodies has also gained attention due to its positive association on gene expression as well2,3. In addition, global methylation at CpG loci throughout the genome is thought to reflect change in response to the environment, exposure, immune response, and the carcinogenic process4. DNA methylation is a widely recognized regulatory mechanism that is indispensable for cellular development, tissue differentiation, inactive X chromosome in female5, and in imprinting6,7.
Recently, genome-wide analyses of DNA methylation have revealed that a number of genes are unequivocally differentially methylated in a variety of normal cells as well as tumors8,9. However, genome-wide DNA methylation patterns have been well studied only in humans and rodents10,11,12. The sequencing and analysis of the dog genome13 will greatly accelerate the dog as a biomedical and spontaneous model for many diseases in humans such as tumors14. Although global hypomethylation or hypermethylation at single loci were found in dogs15,16,17,18, these studies did not search for changes in genomic location modified by DNA methylation. Few genome-wide DNA methylation studies have been conducted for the purpose of identifying differentially methylated CpG sites in diffuse large B-cell lymphoma in dogs compared to peripheral blood/lymph node from healthy control dogs based on microarray platforms19,20, however, comprehensive analysis of DNA methylation status among different normal tissues has not been investigated. Given the fact that understanding of DNA methylation pattern in dogs is limited compared to humans and rodents, we previously established Canine DREAM, which is genome-wide DNA methylation analysis in dogs based on next-generation sequencing of methylation-specific signatures created by sequential digestion with SmaI and XmaI restriction enzymes21. In this study, we aimed to construct genome-wide DNA methylation status in 16 normal cell/tissues in dogs for comprehensive understanding of DNA methylation. This study provides basic information on the dog methylome and a reference data for further study of DNA methylation and comparative animal research.
Results
CpG sites analyzed
We studied the DNA methylomes of 32 normal dog tissue samples (16 tissue types) including, lung, heart, stomach, duodenum, ileum, colon, liver, pancreas, adrenal gland, kidney, spleen, muscle, skin (white-colored), oral mucosa (white-colored and black-colored), and bone marrow. A complete list of cell and tissue types are shown in Table 1.
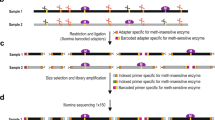
To characterize DNA methylation patterns, we employed Canine DREAM for all of 32 samples. From all 32 samples, 24.5–50.5 million unique usable sequencing reads after conservative filtering (quality filtered and aligned to the dog genome) were successfully generated for DNA methylation analyses (Table 1). We used CpG sites that had more than 20 reads (157–181 thousands CpG sites per sample) to assure quantitative ability. For all the analyses, only autosomal CpGs were considered, resulting in 130,861 common CpG sites obtained for inter-sample comparisons.
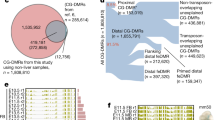
Of 130,861 CpG sites, 6416 (4.9%) sites were located closely at a promoter region of genes annotated by Ensembl Gene Predictions—version 99. 9002 (6.9%) and 55,065 (42%) sites were located at exons and introns of the genes, respectively. The remaining 60,378 CpG sites were distant from any of the above gene annotations except for 2388 sites located on non-coding RNA such as lncRNA (Fig. 1a). Clusters of CpG sites called CpG islands (CGI) have been recognized to be one of the most important methylation features of the genome and methylated differently from non-CpG island (NCGI) regions in mammals22. In this regard, 44,023 sites were in CGIs and 86,838 sites in NCGIs (Fig. 1b).
Overall status of DNA methylation in each tissue
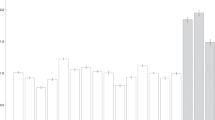
Next, we addressed overall status of DNA methylation level in all tissue types analyzed. The average DNA methylation levels for each tissue type were 59.0–65.1% with small levels of variation among tissue types. Figure 2a,b show DNA methylation fractions and the overall distribution of the DNA methylation levels. We found that the majority of CpG sites were either highly methylated (52.5–64.6% of all CpG sites analyzed) or unmethylated (22.5–28.0% of all CpG sites analyzed), and the remaining 11.9–21.8% were intermediately methylated (Table 1).
Characterization of DNA methylation patterns across cell types. (a) Violin plots of genome-wide DNA methylation levels of all CpG sites for each of the 16 methylome after averaging values for the same tissues. (b) Density plots of genome-wide DNA methylation levels of all CpG sites for each of the 32 methylomes. (c) DNA methylation level of CpG sites classified by the genomic feature. DNA methylation levels of 6416, 9002, 55,065, 2388, and 57,990 CpG sites at promoters, exons, introns, ncRNA, and other regions, respectively, were calculated first, then averaged for all cell types used in this study. The boxes signify upper and lower quartiles while the median is represented by vertical lines within the boxes. Whiskers indicate maximum and minimum values except for outliers that are shown outside of whiskers. All figures are depicted by seaborn (https://doi.org/10.5281/zenodo.592845).
Low level DNA methylation is thought to be a signature of gene regulation23,24. Therefore, we sought for the potential of each CpG site in the role in gene expression regulation by identifying the lowest DNA methylation level. We found that 53,830 CpG sites (41.1% of CpG sites analyzed) had their lowest DNA methylation level below 30%. Even though roughly 25% of CpG sites were unmethylated (calculated by the average of the percentage of unmethylated CpG sites in all cell types) for each cell type, 41.1% of the CpG sites could be unmethylated to have potential regulatory functions.
We calculated the average methylation levels for different genomic features in all cell types and found relatively lower methylation levels in promoter regions compared to exons, introns, or intergenic regions (Fig. 2c).
Tissue specificity determined by genome-wide DNA methylation patterns
To address if DNA methylome derived from different tissue types could identify tissue specificity, we calculated correlation coefficient for all the pairs (496 pairs) with samples analyzed. As expected, relatively high correlation (R = 0.84–0.99) were observed in all the pairs (Fig. 3a). However, the tissue types were clearly defined by Principal component analysis (Fig. 3b) and hierarchical clustering analysis (Fig. 3c). Interestingly, four gastrointestinal cell type methylomes (duodenum, stomach, ileum, and colon) had notable similarity and so did two epithelial cell type methylomes (oral mucosa and skin).
(a) Pairplot for correlation scores of DNA methylation levels with all CpG sites analyzed by Canine DREAM. Samples include lung, heart, stomach, duodenum, ileum, colon, liver, pancreas, adrenal gland, kidney, spleen, muscle, skin (white-colored), oral mucosa (white-colored and black-colored), and bone marrow. Also shown is (b) Principal Component Analysis (depicted by R, https://www.R-project.org/.) and (c) Unsupervised hierarchical analyses of DNA methylation levels with all CpG sites analyzed by Canine DREAM. All figures are depicted by seaborn (https://doi.org/10.5281/zenodo.592845) except for (b).
Classification of CpG sites by DNA methylation patterns across all samples
The overall distributions of methylated or unmethylated CpG sites from the DNA methylation data of 32 samples were similar, however, genome-wide DNA methylation patterns were variable enough to identify difference of tissue types, indicating that methylation level could be either stable or variable across the samples. To address this hypothesis, we classified CpG sites into constitutively methylated (M), unmethylated (U), and intermediately methylated (I) using 70% and 30% as cutoffs. This resulted in 63,790 (48.7%) of CpG sites either being constitutively methylated (39,116 CpG sites, 29.9%) or unmethylated (24,674 CpG sites, 18.9%). Constitutively intermediately methylated CpG sites were found only in 1.1% (1466 CpG sites) of all CpG sites analyzed.
The majority (18,112 CpG sites, 73.4% of 24,674 CpG sites) of constitutively unmethylated CpG sites were in CGIs whereas the majority of constitutively methylated CpG sites (29,800 CpG sites, 76.2% of 39,116 CpG sites) and intermediately methylated CpG sites (1176 CpG sites, 80.2% of 1466 CpG sites) were in NCGIs (Fig. 4a). These findings agreed well with generally appreciated DNA methylation patterns regarding CGI22.
Percentage of constitutively unmethylated (U), methylated (M), and intermediately methylated (I) CpGs analyzed in this study in each category of (a) CpG islands and (b) genomic features. (c) Density scatterplot of CpG-wise DNA methylation level differences (x axis) and CpG median methylation (y axis) across the 32 samples depicted by seaborn (https://doi.org/10.5281/zenodo.592845). Coloring indicates CpG density from low (sparse) to high (dense). The blue box highlights variable CpG sites.
Notably, of 24,674 CpG constitutively unmethylated CpG sites, 4777 (19.4%) sites were located closely at promoter region of genes, which was four times higher frequency than that of all CpG sites analyzed. 3898 (15.8%) and 8387 (34.0%) sites were located at exon and intron of genes, respectively (Fig. 4b).
Next, we sought for variable (V) CpG sites across the samples by utilizing definition of having a gap of at least 40% between the third highest and the third lowest methylation values. As a result, 22,385 (17.1%) CpG sites were identified to be variable in our dataset, of which a substantial number of these sites located at intron (9981 CpG sites) or intergenic regions (10,521 CpG sites), leaving only 449 (2.0%) CpG sites located at promoter regions.
DNA methylation level correlated with gene expression
DNA methylation at promoter is known to correlates with gene expression silencing4. Of 130,861 CpG sites, 6416 (4.9%) sites were located closely at promoter region of genes, which corresponded to 7517 transcripts. We obtained RNA-seq data of the same samples and integrated available gene expression data to observe correlation between differential gene expression and DNA methylation. As a result, we could utilize 3783 CpG sites that were located at promoter region of these transcripts. We plotted differential expression and methylation levels between each pair of all samples after averaging anatomically similar tissues (stomach, duodenum, ileum, and colon were combined intro GI tract. Likewise, skin and oral mucosa samples were combined into Epithelial) (Fig. 5). All of 90 scatterplots showed negative correlation between the two parameters; hypermethylation with gene expression decrease and hypomethylation with gene expression increase between assigned different tissues (r = − 0.12 to − 0.02, Pearson’s correlation).
Integration of methylation and expression data. Starburst plot of the 3783 CpG sites that were located at promoter region of transcripts whose gene expression data were available depicted by seaborn (https://doi.org/10.5281/zenodo.592845). Log fold change in DNA methylation (x-axis) and gene expression (y-axis) are plotted for every comparison between each tissue. When the RPKM for a transcript is less than 0.5, the value was adjusted to 0.5. Log fold change were calculated from the value of tissues shown in a row subtracted by tissues in a column. Pearson’s correlation coefficient and p-values were shown in left lower and right upper of each panel, respectively.
Following differentially methylated regions identified in human, the differential methylation has been linked to tissue specific gene expression (Zhang et al. 2013; Lowdon et al. 2014). Therefore, we hypothesized uniquely methylated or unmethylated CpG sites were hallmarks of gene expression in different cell types. In this regard, we extracted CpG sites that were uniquely unmethylated in only one out of 11 tissue types for the following analysis. After the tissues were ranked by their DNA methylation level for individual CpG sites, we defined uniquely unmethylated sites as being only one tissue showing < 30% methylation with a gap of at least 30% between the lowest and the second lowest methylation values. As a result, we identified 1690 uniquely unmethylated sites and found 94 transcripts with these sites at promoter regions. To analyze the relationship between DNA methylation and gene expression for different tissue types, we utilized RNA-seq data for the same samples and found representative 10 transcripts with significantly lower gene expression in the tissue with the second highest DNA methylation levels (Table 2). One of the significantly correlated genes, PKP3, was known to play a role in cellular desmosome-dependent adhesion and was found to be highly expressed in epithelial tissue as assumed from the function of the gene.
Discussion
We previously reported Canine DREAM, which is a genome-wide DNA methylation analysis of the dog genome21. We utilized Canine DREAM in this study for a variety of normal tissues to construct DNA methylation profiles in dogs and provided basic information for improving our understanding. We analyzed 130,861 CpG sites detected in all samples, which is much more than generally reported DNA methylation studies with strategy that targeted genes of interest, suggesting that Canine DREAM can provide a more in-depth view of DNA methylation status than single-locus studies17,25. Compared to genome-wide DNA methylation analysis based on a customized or human microarray platforms19,20, analyses with next-generation sequencing followed by mapping to the dog genome will provide flexibility to conduct DNA methylation studies without concern of interspecies difference.
The average DNA methylation levels of all the CpG sites as well as overall distribution of the DNA methylation levels analyzed for each tissue type were quite similar with small levels of variation among tissue types. These findings in the dog were consistent with other species. In this study, the 16 dog tissues showed similar global methylation with correlation coefficients ranging from 0.84 to 0.99. Previously, ten bovine tissues showed correlations ranging from 0.93 to 0.9826. The pig study with closely related tissues yielded slightly higher correlations (> 0.95)27.
Particularly, 29.9% of CpG sites were constitutively methylated regardless of the cell types whereas 18.9% of CpG sites were constitutively unmethylated. The fact that approximately half of CpG sites are stable in all the 32 samples analyzed in this study is consistent with the fact that DNA methylation is a stable mark across different dog cell types. The majority (73.4%) of constitutively unmethylated CpGs were in CGI, and the majority of constitutively methylated CpG sites (76.2%) and intermediately methylated CpG sites (80.2%) were in NCGIs, which is consistent with the notion that the CGIs were generally less methylated than the NCGIs22 and also that methylated NCGIs are suggested to suppress unnecessary retrotransposon expression28.
Although overall DNA methylation status among tissue types was similar, principal component analysis and hierarchical clustering analysis with all 32 methylome data showed clear differences indicating tissue-specific DNA methylation patterns. This is consistent with the reports of human and bovine methylome where cell/tissue types were also separated clearly by similar analyses26,29. These results suggest that DNA methylation patterns are profoundly involved in tissue differentiation across species. In any given cell type, approximately the same percentage of CpG sites were found methylated and unmethylated, however, methylation status of CpG sites in a certain cell type could be cell-type dependent.
Since our findings indicated different DNA methylome depending on tissue types, we sought for tissue-specific DNA methylation that were associated with gene expression regulation. DNA methylation in promoter region of genes is associated with gene silencing30. This was supported by our results that DNA methylation in the promoter regions showed largely negative correlation with gene expression. In addition, we filtered uniquely unmethylated CpG sites at promoter regions in only one tissue type to identify exclusive expression characteristics. As a result, 0 (Adrenal gland)–434 (Pancreas) uniquely methylated CpG sites were found and this variable numbers of uniquely unmethylated CpG sites could partly be attributed to the tissue characteristics in this study.
Widespread colocalization between transcription factor binding and variably methylated CpG sites outside promoter regions were reported in humans24,29. Although 22,385 variably methylated CpG sites were found in this study, the majority (> 91%) of these sites located at intron or intergenic regions and could not be analyzed systematically in this regard due to lack of these data in the dog genome. It is no wonder that those variably methylated CpG sites that did overlap with nothing on currently available database might harbor roles in regulation of transcription factor binding which would be identified by increasing amount of transcription factor and histone modification ChIP-seq data in the future.
We expect that more variably and uniquely methylated CpG sites will be identified if more cell types are sequenced and analyzed in the future. We would clearly suggest that DNA methylation plays a role in the regulation of cell or tissue type-specific gene functions. The data and results provided in this study will be useful in the research field of veterinary medicine as well as human medicine. Our methylation data would also be helpful in interpreting the epigenomic status in a variety of cells and conditions such as tumors. It is important to remind that Canine DREAM only analyzed a small part of the dog genome, and more extensive studies such as whole genome bisulfite sequencing are needed to confirm these findings. Nevertheless, our data and many more single-CpG-resolution DNA methylome data available in the future will provide greater insights into the knowledge of epigenetics research field.
Methods
Dogs
Two female Beagles 58-week and 73-week-old were used as healthy controls. These dogs were healthy, had no clinical signs, no abnormalities in urinalysis, hematological examination, or blood biochemical analysis. No parasites or pathogenic bacteria were detected in fecal samples. Food was withheld from each dog for 12 h. Dogs then were euthanized and 16 tissues including, lung, heart, stomach, duodenum, ileum, colon, liver, pancreas, adrenal gland, kidney, spleen, muscle, skin, oral, oral (pigmented), and bone marrow were obtained. The use of dogs in this study was approved by the Animal Care Committee of the University of Tokyo (Approval No. P17-064), and the protocol was carried out in compliance with the ARRIVE guidelines and the American Veterinary Medical Association (AVMA) Guidelines for the Euthanasia of Animals (2020). All specimens for Canine DREAM was stored at − 80 °C.
Digital restriction enzyme analysis of methylation (DREAM)
Genome-wide DNA methylation analysis using next-generation sequencing was performed as previously21 with genomic DNA (2 μg) extracted from the above samples. We used the University of California, Santa Cruz (UCSC) definition of CpG islands31. Promoter regions are defined as being located within 1 kb from transcription start sites of given genes. Autosomal CpG sites were initially grouped into four biologically motivated categories based on their distribution of DNA methylation values across the 32 samples: constitutively unmethylated (U) if all values were below 30%, intermediately methylated (I) if at least 30 libraries had values between 30 and 70%, constitutively methylated (M) if all values were above 70%.
RNA-Seq analysis
Total RNA was extracted by the NucleoSpin RNA (Macherey–Nagel, Duren, Germany) for each tissue. RNA integrity number of all RNA is confirmed to be more than 7.0. The RNA-Seq library was constructed by SMART-Seq v4 (Takara Bio) and Nextera XT DNA Library Kit (Illumina). Sequencing was performed with Illumina NovaSeq 6000 according to the manufacture’s software. Quality filtering for sequence reads were performed using Trim Galore (version 0.6.4) with the Phred cutoff score of 30. We used 32–42 million reads after quality control of sequenced reads. Transcripts per million (TPM) was calculated to evaluate gene expression by using Kalisto (version 0.46.2) with default settings and dog gene annotation data sets (CanFam3.1).
Data analysis and visualization
Violin plots, density plots, boxplots, and Starburst plots were drawn and visualized with Seaborn on Python 3.732. Hierarchical clustering analysis was performed with the agglomeration method ‘ward’ where the distance was calculated with the Euclidean and visualized with Seaborn on Python 3.7. Principal component analysis was performed using R with a package ggfortify 0.4.1033. Pearson’s correlation coefficient was calculated for associations between DNA methylation and gene expression levels.
Institutional animal care and use committee (IACUC) or other approval declaration
This study was approved by the Animal ethical committee of The University of Tokyo.
Human ethics approval declaration
Authors declare human ethics approval was not needed for this study.
Data availability
The dataset generated and/or analysed during the current study are underway to be submitted to Gene Expression Omnibus (GEO) and will be available once it is accepted.
Abbreviations
- CGI:
-
CpG islands
- DREAM:
-
Digital restriction enzyme of DNA methylation analysis
References
Razin, A. & Cedar, H. DNA methylation and gene expression. Microbiol. Rev. 55, 451–458 (1991).
Jones, P. A. Functions of DNA methylation: Islands, start sites, gene bodies and beyond. Nat. Rev. Genet. 13, 484–492. https://doi.org/10.1038/nrg3230 (2012).
Yang, X. et al. Gene body methylation can alter gene expression and is a therapeutic target in cancer. Cancer Cell 26, 577–590. https://doi.org/10.1016/j.ccr.2014.07.028 (2014).
Jones, P. A. & Baylin, S. B. The epigenomics of cancer. Cell 128, 683–692. https://doi.org/10.1016/j.cell.2007.01.029 (2007).
Heard, E., Clerc, P. & Avner, P. X-chromosome inactivation in mammals. Annu. Rev. Genet. 31, 571–610. https://doi.org/10.1146/annurev.genet.31.1.571 (1997).
Barlow, D. P. Gametic imprinting in mammals. Science 270, 1610–1613 (1995).
Ferguson-Smith, A. C. Genomic imprinting: The emergence of an epigenetic paradigm. Nat. Rev. Genet. 12, 565–575. https://doi.org/10.1038/nrg3032 (2011).
Figueroa, M. E. et al. MDS and secondary AML display unique patterns and abundance of aberrant DNA methylation. Blood 114, 3448–3458. https://doi.org/10.1182/blood-2009-01-200519 (2009).
Noushmehr, H. et al. Identification of a CpG island methylator phenotype that defines a distinct subgroup of glioma. Cancer Cell 17, 510–522. https://doi.org/10.1016/j.ccr.2010.03.017 (2010).
Lister, R. et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462, 315–322. https://doi.org/10.1038/nature08514 (2009).
Schultz, M. D. et al. Human body epigenome maps reveal noncanonical DNA methylation variation. Nature 523, 212–216. https://doi.org/10.1038/nature14465 (2015).
Habibi, E. et al. Whole-genome bisulfite sequencing of two distinct interconvertible DNA methylomes of mouse embryonic stem cells. Cell Stem Cell 13, 360–369. https://doi.org/10.1016/j.stem.2013.06.002 (2013).
Kirkness, E. F. et al. The dog genome: Survey sequencing and comparative analysis. Science 301, 1898–1903. https://doi.org/10.1126/science.1086432 (2003).
Paoloni, M. & Khanna, C. Translation of new cancer treatments from pet dogs to humans. Nat. Rev. Cancer 8, 147–156. https://doi.org/10.1038/nrc2273 (2008).
Morimoto, C. Y. et al. Evaluation of the global DNA methylation in canine mast cell tumour samples by immunostaining of 5-methyl cytosine. Vet. Comp. Oncol. https://doi.org/10.1111/vco.12241 (2016).
Yamaya, Y., Sugiya, H. & Watari, T. Methylation of free-floating deoxyribonucleic acid fragments in the bronchoalveolar lavage fluid of dogs with chronic bronchitis exposed to environmental tobacco smoke. Ir. Vet. J. 68, 7. https://doi.org/10.1186/s13620-015-0035-4 (2015).
Ferraresso, S. et al. Epigenetic silencing of TFPI-2 in canine diffuse large B-cell lymphoma. PLoS ONE 9, e92707. https://doi.org/10.1371/journal.pone.0092707 (2014).
Tomiyasu, H. et al. Evaluation of DNA methylation profiles of the CpG island of the ABCB1 gene in dogs with lymphoma. Am. J. Vet. Res. 75, 835–841. https://doi.org/10.2460/ajvr.75.9.835 (2014).
Ferraresso, S. et al. DNA methylation profiling reveals common signatures of tumorigenesis and defines epigenetic prognostic subtypes of canine diffuse large B-cell lymphoma. Sci. Rep. 7, 11591. https://doi.org/10.1038/s41598-017-11724-w (2017).
Epiphanio, T. M. F. et al. Global DNA methylation of peripheral blood leukocytes from dogs bearing multicentric non-Hodgkin lymphomas and healthy dogs: A comparative study. PLoS ONE 14, e0211898. https://doi.org/10.1371/journal.pone.0211898 (2019).
Yamazaki, J., Jelinek, J., Hisamoto, S., Tsukamoto, A. & Inaba, M. Dynamic changes in DNA methylation patterns in canine lymphoma cell lines demonstrated by genome-wide quantitative DNA methylation analysis. Vet. J. 231, 48–54. https://doi.org/10.1016/j.tvjl.2017.11.007 (2018).
Bird, A. DNA methylation patterns and epigenetic memory. Genes Dev. 16, 6–21. https://doi.org/10.1101/gad.947102 (2002).
Tsankov, A. M. et al. Transcription factor binding dynamics during human ES cell differentiation. Nature 518, 344–349. https://doi.org/10.1038/nature14233 (2015).
Stadler, M. B. et al. DNA-binding factors shape the mouse methylome at distal regulatory regions. Nature 480, 490–495. https://doi.org/10.1038/nature10716 (2011).
Bryan, J. N. et al. Hypermethylation of the DLC1 CpG island does not alter gene expression in canine lymphoma. BMC Genet. 10, 73. https://doi.org/10.1186/1471-2156-10-73 (2009).
Zhou, Y. et al. Reduced representation bisulphite sequencing of ten bovine somatic tissues reveals DNA methylation patterns and their impacts on gene expression. BMC Genomics 17, 779. https://doi.org/10.1186/s12864-016-3116-1 (2016).
Choi, M. et al. Genome-wide analysis of DNA methylation in pigs using reduced representation bisulfite sequencing. DNA Res. 22, 343–355. https://doi.org/10.1093/dnares/dsv017 (2015).
Zhou, W., Liang, G., Molloy, P. L. & Jones, P. A. DNA methylation enables transposable element-driven genome expansion. Proc. Natl. Acad. Sci. U.S.A. 117, 19359–19366. https://doi.org/10.1073/pnas.1921719117 (2020).
Gu, J. et al. Mapping of variable DNA methylation across multiple cell types defines a dynamic regulatory landscape of the human genome. G3 (Bethesda) 6, 973–986. https://doi.org/10.1534/g3.115.025437 (2016).
Varley, K. E. et al. Dynamic DNA methylation across diverse human cell lines and tissues. Genome Res. 23, 555–567. https://doi.org/10.1101/gr.147942.112 (2013).
Gardiner-Garden, M. & Frommer, M. CpG islands in vertebrate genomes. J. Mol. Biol. 196, 261–282 (1987).
Michael Waskom and the seaborn development team. mwaskom/seaborn. Series. https://doi.org/10.5281/zenodo.592845 (2020).
Tang, Y., Horikoshi, M. & Li, W. ggfortify: Unified Interface to Visualize Statistical Result of Popular R Packages. Series 2016. https://journal.r-project.org/. Accessed on 03 August 2020.
Author information
Authors and Affiliations
Contributions
J.Y. designed the study. S.M. provided resources. J.Y., Y.M., T.I. and K.W. conducted the experiment(s). J.Y., Y.M., J.J. and J.Y. analysed the data. J.Y., Y.M. and J.J. wrote and edited the manuscript. G.I. and M.T. supervised project administration. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yamazaki, J., Matsumoto, Y., Jelinek, J. et al. DNA methylation landscape of 16 canine somatic tissues by methylation-sensitive restriction enzyme-based next generation sequencing. Sci Rep 11, 10005 (2021). https://doi.org/10.1038/s41598-021-89279-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-89279-0
This article is cited by
-
Development of a prognostic model for personalized prediction of colon adenocarcinoma (COAD) patient outcomes using methylation-driven genes
Journal of Applied Genetics (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.