Abstract
A variety of eye-related symptoms due to the overuse of digital devices is collectively referred to as computer vision syndrome (CVS). In this study, a web-based survey about mind and body functions, including eye strain, was conducted on 1998 Japanese volunteers. To investigate the biological mechanisms behind CVS, a multi-trait genome-wide association study (GWAS), a multivariate analysis on individual-level multivariate data, was performed based on the structural equation modeling methodology assuming a causal pathway for a genetic variant to influence each symptom via a single common latent variable. Twelve loci containing lead variants with a suggestive level of significance were identified. Two loci showed relatively strong signals and were associated with TRABD2B relative to the Wnt signaling pathway and SDK1 having neuronal adhesion and immune functions, respectively. By utilizing publicly available eQTL data, colocalization between GWAS and eQTL signals for four loci was detected, and a locus on 2p25.3 showed a strong colocalization (PPH4 > 0.9) on retinal MYT1L, known to play an important role in neuronal differentiation. This study suggested that the use of multivariate questionnaire data and multi-trait GWAS can lead to biologically reasonable findings and enhance our genetic understanding of complex relationships among symptoms related to CVS.
Similar content being viewed by others
Introduction
With the growth of digital devices, there are many complaints nowadays about a variety of eye-related symptoms such as eye strain, blurred vision, and double vision, and it is collectively referred to as digital eye strain or computer vision syndrome (CVS). In addition, it is also known to be associated with non-ocular symptoms such as headaches, and shoulder and neck pain. The American Optometric Association has defined CVS as a combination of eye and vision problems linked to the use of visual display terminals (VDTs)1. The diagnosis is carried out based on a comprehensive examination including visual acuity measurements, testing for refractive error and focusing movement in addition to medical inquiries about symptoms the patient has experienced. Although CVS is associated with a variety of symptoms, Sheedy et al.2 stated that the underlying problem can be identified by the location of the symptoms and categorized into the two groups of internal and external symptoms. According to the general review by Blehm et al.3, there are four types of symptoms (asthenopic, ocular surface-related, visual, and extra-ocular) and they can be subdivided into three main potential pathophysiological causes (ocular surface mechanisms, accommodative mechanisms, and extraocular mechanisms). Numerous studies have been conducted so far to address health and safety issues for VDT users, but the efficacy of proposed treatments for minimizing CVS-related symptoms have not been adequately demonstrated yet3,4. A better understanding of the underlying physiology behind CVS is important to enable more accurate diagnosis and treatment.
The genome-wide association study (GWAS)5 has become a common genetic approach to identify potential susceptibility genes for diseases. To date, a number of GWAS have been successful in identifying the genetic pathways associated with ocular diseases such as myopia, age-related macular degeneration and glaucoma6,7. Although GWAS has brought forth an adequate amount of knowledge on the genetic etiology of ophthalmic traits, through relatively easy to recruit severe cases, more common diseases or syndromes will require an increase in sample size and well-designed analytical methods. No GWAS therefore has been reported for CVS, and we need an elaborate method to integrate multiple symptoms related to CVS, which cannot be directory measured, into a model in order to precisely evaluate genetic effects using GWAS.
With the establishment of cohorts with more detailed phenotypic information, multi-trait GWAS, which is a multivariate analysis on the same subjects, has also become possible. Multivariate methods are generally considered more powerful than univariate methods unless only one trait is associated with a genetic marker or all traits are very highly correlated8. Many methods have been proposed and applied so far, including dimensionality reduction using principal component analysis (PCA), multivariate mixed models as an extension of regression analysis, Bayesian polynomial regression and so on8,9. In recent years, methodologies using structural equation modeling (SEM) have been applied for multi-trait GWAS10,11. SEM is a multivariate statistical method in which the interactions between variables are described as simultaneous equations and their parameters are inferred12. Unlike other methods, the SEM-based method can incorporate latent variables, which are hypothetical quantities that have not been measured and can be inferred from other measured variables, into the model. A SEM-based multi-trait GWAS is attained by assuming a model structure in which a genomic variant is an exogenous predictor that influences observed variables via the latent factor, and then repeating the statistical test genome-wide for each variant. SEM has enabled investigations of complex traits such as psychology and psychiatry, in which their unmeasurable phenotypes are indexed by multiple symptoms or behaviors. Since this feature is similar to CVS, we applied a SEM-based multi-trait GWAS to CVS, which we expected increased power to identify genetic associations.
In this study, a web-based survey about CVS was conducted on 1998 Japanese volunteers. To investigate the biological mechanisms behind CVS, we performed the SEM-based multi-trait GWAS for the CVS-related multiple symptoms for the first time to the best of our knowledge under the assumption of the existence of a potential phenotype behind the CVS-related symptoms.
Methods
Study population and data collection
Among customers of a direct-to-consumer (DTC) genetic testing service (MYCODE), 1998 Japanese volunteers (50.5% male, mean age 49.3 years) were recruited in August 2019 (Table 1). Customers with prior consent in writing for research recruitment were invited to participate in this study via the company website and email. Participation was voluntary and based on informed consent. The web-based questionnaire survey was conducted with the participants. A total of 14 questionnaire items covering the four categories defined by Blehm et al.3 were designed (Table 2). Each item was measured on a scale of one to seven in each of the two conditions: “When you feel like you have used your eyes” and “When you feel like you have not used your eyes very much”. These two conditions were set to simulate the answers before and after the VDT task, and the difference (delta) of these two conditional answers was used in the following analysis. In addition to that, self-reported medical history was obtained on the presence or absence of dry eye syndrome, presbyopia and other common eye diseases. This study received approval from the institutional review board of DeNA Life Science, Inc. and was implemented in accordance with the Declaration of Helsinki.
Genotyping and quality control
Participants collected their own saliva samples using the kit sent to them when they used the service before participation in this study, and DNA extraction and genotyping were performed at the laboratory of DeNA Life Science, Inc. Infinium OmniExpress-24 or Human OmniExpress-24 BeadChip (Illumina, Inc.), both of which additionally contained approximately 30,000 custom probes, was used. The stringent quality control (QC) procedures of genotyped results were applied with call rate ≥ 95%, minor allele frequency (MAF) ≥ 0.01, identity by descent < 0.1875, and Hardy–Weinberg equilibrium (p value > 1.0e−6). The gender on the declaration was confirmed to match the genotyped one. All the above procedures were performed using PLINK (ver. 1.9)13. In order to stratify the population, PCA computed the first two principal components based on a sample set of 1000 Genomes Project phase 3 (1KGP; N = 2504) and then projected study samples onto the 2D subspace. Samples falling outside the Japanese cluster were excluded (Supplementary Fig. S1). Eventually, 1966 subjects (999 male) with a total of 540,916 single nucleotide polymorphisms (SNPs) on autosomes ended up passing the QC filter. GRCh37/hg19 (GCF_000001405.13) was the reference for the genome construction information.
Statistical analysis
To implement the multi-trait GWAS based on the SEM methodology, gwsem package (ver. 0.1.17) in R, which was developed by Verhulst et al.14, was introduced. Age, sex, and the top 2 eigenvectors from PCA were added into the model as external covariates. The number of principal components to be incorporated was determined based on examination of the scree plot of their eigenvalues. After running the SEM-based multi-trait association analysis on all variants, p values were subsequently calculated, adjusting each test statistic by a genomic inflation factor. Genotype imputation was conducted on variants which were located within ± 500 kb of GWAS hits, and SEM-based GWAS was rerun on estimated genotypes. Eagle2 (ver. 2.4.1)15 was used for the phasing step referring to the East Asian population (EAS; N = 504) of 1KGP, and then imputation was performed using Minimac3 (ver. 2.0.1)16. The accuracy index of imputation performance, correlation between reference and predicted genotypes (R squared), was 0.942 ± 0.116 for common variants (MAF ≥ 0.05; see Supplementary Fig. S2). Imputed variants with R squared < 0.7 were removed for QC purpose. GWAS hits meeting a suggestive level of significance (p value < 1.0e−5) were considered lead variants, and the locus was defined as a region containing highly correlated variants (R squared > 0.6) with each lead variant in the 1KGP EAS population.
Colocalization between GWAS summary statistics and expressed quantitative trait locus (eQTL) signals was analyzed based on a Bayesian statistical methodology proposed by Giambartolomei et al.17 We utilized GSE115828 data (N = 523)18, the only publicly available retinal tissue eQTL data from the Gene Expression Omnibus (GEO) of the National Center for Biotechnology Information as of May 2020. The whole blood eQTL data in the Genotype-Tissue Expression (GTEx ver. 7; N = 369)19 was analyzed as well. In addition, two data sources for eQTL data (GSE5335120 and hum0099.v121) were added, available from GEO and the National Bioscience Database Center and consisting of more than 100 healthy Japanese subjects. The former is whole blood data (N = 301) and the latter is eQTL data from whole blood cells and five immune cell populations (CD4+ T cells, CD8+ T cells, B cells, NK cells, and monocytes) of 105 subjects. Note that previous studies19,22 have demonstrated that a sample size of about 100 individuals is sufficient to detect major eQTL effects in various cell types collected in a single population. Posterior probability of H4 (PPH4) of colocalization was calculated for each ± 100 kb of the identified loci containing lead variants using the coloc package in R.
Results
Model specification via cause-and-effect graph
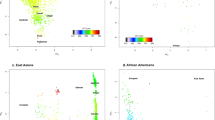
It is assumed that the 14 questionnaire items are observed symptoms due to a potential syndrome, that is CVS, and multiple items will be observed in conjunction and correlated to some extent. The results of 1998 respondents were used to calculate pairwise partial correlation coefficients between items, and positive correlations among all items were verified (Supplementary Fig. S3). The PCA calculated from the eigenvalue vectors of the partial correlation matrix showed that all items were in the same direction on the first PC, and they appeared to form clusters corresponding to the four categories to which each item belonged on the second PC, though not completely (Fig. 1a). Next, covariance selection23, a graphical modeling technique, was applied to derive a partial correlation network consisting of only well-fitting edges between items by stepwise removing edges with low partial correlation coefficients (Fig. 1b). Although q5 "watery eyes" was a singleton that did not share edges with any of the items, it could be seen as forming one large network as a whole. Since the above suggests that each of the four categories captures its own characteristics among belonging items, the items were integrated (averaged) and the representative values of each category were used in the following analysis. In addition, we constructed a factor analysis model with one common factor for all categories (Fig. 2). Using OpenMx package (ver. 2.17.3) in R to calculate goodness of model fit, the Chi-squared statistic was 22.5 (degrees of freedom = 9; p value = 7.4e−3) and the root mean square error of approximation (RMSEA) was 0.028 (confidence interval 0.00–0.045). The closer the RMSEA is to zero, the better the fit to SEM, and a value below 0.05 is desirable24. This model was considered to be robust for SEM-based multi-trait GWAS.
(a) PCA results for questionnaire items based on the partial correlation matrix. The length of arrows indicates contributions of the variables to the principal axes. The proportion of variance captured is given as a percentage for both the first and second principal components. (b) Partial correlation network estimated by covariance selection. A dense subnetwork of high partial correlated edges is called a clique, and the existence of a clique would suggest the presence of the same number of latent factors. No cliques with edges all having values greater than 0.2 were found in the network. For model optimization, gmm package (ver. 1.6-5) in R was used to minimize Akaike information criterion.
Node diagram of SEM for multi-trait GWAS. The observed variables (four symptom categories) are presented as rectangles and the latent variable as ellipses. Note that each symptom category was calculated by the average of component questionnaires. The association between the latent factor and the observed variables are estimated by the factor loadings (γk). GWAS analysis infer the strength of the causal effect (β) from each SNP to the phenotypes via the latent variable. Age and gender, as well as the first two principal components of the PCA performed for population stratification, were added as covariates. The arrows describe the relationships between the variables. Residuals (δk) and variances (σ) are drawn as double headed dashed arrows into an object.
Multi-trait GWAS
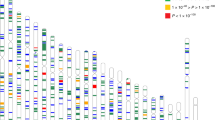
SEM is a form of graphical modeling that uses graphs to assume causalities between observed and latent variables and estimates the strength of the causalities. We introduced a latent variable into the model, where the causal effect of each variant on the questionnaire items (observed variables) was transmitted via the latent variable (Fig. 2). Comprehensive association analysis on each variant was performed to infer the strength of association between the variant and latent factor, and the statistical significance was calculated as a p value. A quantile–quartile (Q–Q) plot was generated between the actual observed p values and the p values as theoretically expected, showing no significant population stratification (Supplementary Fig. S4). The output of GWAS can be represented by a diagram called a Manhattan plot, where each variant’s strength of association (− log10 of p value) is relative to the genomic position (Fig. 3). Although there were no hits with the genome-wide significance level (5.0e−8), 12 peaks with variants at the top that met the suggestive level of significance were detected (Table 3). For each of the peaks, regional association plots were generated using LocusZoom (ver. 1.4)25, and they all formed a single peak at each close location on the genome (Supplementary Fig. S5). When GWAS was performed again with the addition of other covariates (daily VDT hours, glasses and/or contact lens use, and eye drop use), there was no noticeable change in the relative trend, although the overall signal weakened slightly.
We performed single-trait GWAS on each of the four categories using gwsem, and several peaks detected by the multi-trait GWAS were identified as well (Supplementary Fig. S6). As an example, the peak upstream of the sidekick cell adhesion molecule 1 (SDK1) gene on 7p22.2 are seen in the "ocular surface-related" and "visual" categories, while the others also show a less significant association. Furthermore, since similarly consistent peaks can be found using PLINK, which is a gold standard tool for GWAS, there is little concern about bias due to the statistical processing method.
GWAS and eQTL colocalization analysis
None of the lead variants detected from multi-trait GWAS were located on a protein coding region. We examined the influence of detected variants on gene expression levels by utilizing publicly available eQTL data sources. In order to identify genes for which the expression level is affected by each variant, we focused on colocalization between the distribution of GWAS and eQTL signals. In other words, the p value of the GWAS result for each variant was compared with that of eQTL on a genome scale to see if they both show significant p values at the same position. Calculating a posterior probability (PPH4) when GWAS and eQTL are associated and share a single causal variant, combinations of locus, genes and expression tissue with > 0.5 of PPH4 were comprehensively explored. As a result, four combinations were detected (Table 4). When using a criteria > 0.75 of PPH4 strongly suggesting GWAS and eQTL signals are due to a common causal variant26, a strong colocalization (PPH4 = 0.906) between GWAS signals at the locus (1,625,705–1,627,501) ± 100 kb on chromosome 2 and eQTL signals on the myelin transcription factor 1 like (MYT1L) gene in retinal tissue was detected. As seen in the regional association plots generated by the locuscomparer package (ver. 1.0.0)27 in R, the p values distribution in GWAS and eQTL signals showed a similar shape and the high degree of correlation was visually confirmed as well (Fig. 4). This suggests that rs9677043, which has the strongest p value for both GWAS and eQTL, is a highly possible causal variant. Note that rs9677043 is located approximately 166 kb downstream of the MYT1L gene. We analyzed the association of rs9677043 with other self-reported diseases (presbyopia, dry eye syndrome, severe myopia, cataract, and glaucoma), but no significant associations were found.
Discussion
The MYT1L gene, the expression level of which showed a strong colocalization (PPH4 > 0.9) in retina, is a paralog of MYT1 and encodes a member of the zinc finger superfamily of transcription factors. It is mainly expressed in neural tissue and is known to play an important role in neuronal differentiation by specifically suppressing the expression of non-neuronal genes28. The eQTL data showed rs9677043, detected as a possible causal variant, has a positive effect on MYT1L expression levels in retinal tissue. The path coefficient β estimated by SEM was 0.132, showing that the mutation of rs9677043 worsens the observed symptoms. This suggests that the differentiation of neurons in retinal tissue has a potential to be associated with CVS.
Looking at other identified loci, Chr1:48,233,438–48,233,438 and Chr7:3,319,615–3,715,120 have relatively strong GWAS signals (p value < 1.0e−6), and they were associated with the TraB domain containing 2B (TRABD2B) and SDK1 genes according to the position on the genome, respectively. TRABD2B is a metalloproteinase having the function of a negative regulator of the Wnt signaling pathway, which has been reported to regulate the transdifferentiation of retinal nerves into the ciliary body29. On the other hand, SDK1 encodes a protein in the immunoglobulin superfamily taking a critical part of the immune response, and in addition, is thought to work in selective synapse formation between retinal neurons because of its ability to adhere specifically to another SDK1 on the surface of other cells30. All of these genes are associated with retinal nerves or a wide range of neural tissues. As chronic inflammation is reported to be one factor in neuronal dysfunction in the retina31,32, immune response also might provide clues to investigate the physiology of CVS.
We will describe the limitations of this study from the following four of viewpoints: (1) self-reported questionnaires via a web browser, (2) sufficiency of the level of statistical significance, (3) impact of racial differences in eQTL analysis, (4) need for validation study using different sample groups.
There is no doubt that web-based questionnaires are less clinically reliable than diagnosis by a doctor. Furthermore, it has been pointed out that there are some problems, such as the bias caused by the web-mediated process, compared to traditional laboratory questionnaire interviews33. On the other hand, one of the DTC genetic testing services, 23andMe in the US, has uncovered numerous biological findings utilizing self-reported medical history and web-based questionnaires from a large group of customers34. This is a new research approach that is driven by mass data and this study is a continuation of that trend.
The ordinary GWAS has established a very strict criteria—a genome-wide significance level. This study instead used the suggestive significance level to give a biological interpretation. The main reasons for the low statistical power may be the subjective way in which the trait was assessed, the wide spectrum of traits, and the small number of samples. Considering the original purpose of GWAS, we should find its meaning and value in the discovery of unknown important molecular mechanisms or new markers for diagnosis. In fact, other studies have led to an extended list of associated variants, which can provide a resource for functional studies, by using more exploratory criteria35. This requires sufficiently narrowing the window for the possibility of false positives, and this study used colocalization with eQTL to give additional validity to biological certainty.
It has been noted that the expression levels of many genes are variable between human populations, and it is mainly explained by differences in genotype frequencies20. We used HaploReg (ver. 4.1)36 to check rs9677043 detected from the European retinal eQTL data, and the MAF in the European and East Asian populations were 0.53 and 0.47, respectively. Although there is no large difference compared to this study (0.464), it is desirable to obtain retinal tissue eQTL data from the Japanese population, considering the fact that transcriptional regulation at the gene level cannot be explained by a single variant alone.
GWAS can be positioned as a “forward genetic” experiment to identify the genetic basis of a trait and provide an opportunity for exploratory hypothesis generation. In order to strengthen the hypothesis, it is ideal to conduct external validation on different sample groups, and this study is no exception. On the other hand, the common study design of examining the functional role of genetic variation is to knock out a specific gene in a model organism. Such an experiment based on “reverse genetics” is necessary for hypothesis verification. In recent years, national biobanks and DTC genetic testing services provide a large-scale cohort linked to genetic and rich phenotypic information, mainly obtained from electronic medical records. An approach of extending GWAS to the entire set of phenotypic entities is called the phenome-wide association study (PheWAS)37, and it allows “reverse genetic” experiments to be virtually conducted on human subjects38,39. The proof-of-principle for the hypotheses generated in this study is expected by making use of cohorts in the future. Besides, the Mendelian randomization approach is an extension of PheWAS to test for causal relationships between phenotypes, and the SEM methodology has been used in many cases40,41.
Last of all, we will discuss the benefits of the use of SEM for multi-trait GWAS. The path coefficients estimated by SEM reflect the strength of each causal relationship and quantify the influence of a variant on the target phenotype via intermediate variables (latent factors) and predefined causal paths. For instance, the positive path coefficient from an intermediate variable to an observed variable suggests that an increase in the intermediate variable directly leads to an increase in the observed variable. In the case of this study, the coefficients (γ) from the four categories (asthenopic, ocular surface-related, visual, and extra-ocular) to the intermediate variable were 0.842 ± 0.024, 0.657 ± 0.018, 0.787 ± 0.027, and 0.823 ± 0.022, respectively, and “asthenopic” and “extra-ocular” were found to be relatively strong symptoms for CVS. Thus, we believe that SEM has the potential to enhance our understanding of complex phenotypes, such as syndromes, by genetically unraveling the relationship between observed symptoms. SEM is expected to be a core technique for performing a series of association analyses from GWAS to PheWAS.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding authors on reasonable request.
References
American Optometric Association. Computer Vision Syndrome [Internet]. https://www.aoa.org/patients-and-public/caring-for-your-vision/protecting-your-vision/computer-vision-syndrome (2019).
Sheedy, J. E., Hayes, J. N. & Engle, J. Is all asthenopia the same?. Optom. Vis. Sci. 80, 732–739. https://doi.org/10.1097/00006324-200311000-00008 (2003).
Blehm, C., Vishnu, S., Khattak, A., Mitra, S. & Yee, R. W. Computer vision syndrome: A review. Surv. Ophthalmol. 50, 253–262. https://doi.org/10.1016/j.survophthal.2005.02.008 (2005).
Rosenfield, M. Computer vision syndrome: A review of ocular causes and potential treatments. Ophthalm. Physiol. Opt. 31, 502–515. https://doi.org/10.1111/j.1475-1313.2011.00834.x (2011).
Hirschhorn, J. N. & Daly, M. J. Genome-wide association studies for common diseases and complex traits. Nat. Rev. Genet. 6, 95–108. https://doi.org/10.1038/nrg1521 (2005).
Cooke Bailey, J. N. et al. Advances in the genomics of common eye diseases. Hum. Mol. Genet. 22, 59–65. https://doi.org/10.1093/hmg/ddt396 (2013).
Mackey, D. A. & Hewitt, A. W. Genome-wide association study success in ophthalmology. Curr. Opin. Ophthalmol. 25, 386–393. https://doi.org/10.1097/ICU.0000000000000090 (2014).
Galesloot, T. E., van Steen, K., Kiemeney, L. A., Janss, L. L. & Vermeulen, S. H. A comparison of multivariate genome-wide association methods. PLoS One 9, e95923. https://doi.org/10.1371/journal.pone.0095923 (2014).
Hackinger, S. & Zeggini, E. Statistical methods to detect pleiotropy in human complex traits. Open Biol. https://doi.org/10.1098/rsob.170125 (2017).
Jeon, S., Shin, J. Y., Yee, J., Park, T. & Park, M. Structural equation modeling for hypertension and type 2 diabetes based on multiple SNPs and multiple phenotypes. PLoS One 14, e0217189. https://doi.org/10.1371/journal.pone.0217189 (2019).
Momen, M. et al. Including phenotypic causal networks in genome-wide association studies using mixed effects structural equation models. Front Genet. 9, 455. https://doi.org/10.3389/fgene.2018.00455 (2018).
Li, R. et al. Structural model analysis of multiple quantitative traits. PLoS Genet. 2, e114. https://doi.org/10.1371/journal.pgen.0020114 (2006).
Chang, C. C. et al. Second-generation PLINK: Rising to the challenge of larger and richer datasets. Gigascience 4, 7. https://doi.org/10.1186/s13742-015-0047-8 (2015).
Verhulst, B., Maes, H. H. & Neale, M. C. GW-SEM: A statistical package to conduct genome-wide structural equation modeling. Behav. Genet. 47, 345–359. https://doi.org/10.1007/s10519-017-9842-6 (2017).
Loh, P. R. et al. Reference-based phasing using the Haplotype Reference Consortium panel. Nat. Genet. 48, 1443–1448. https://doi.org/10.1038/ng.3679 (2016).
Das, S. et al. Next-generation genotype imputation service and methods. Nat. Genet. 48, 1284–1287. https://doi.org/10.1038/ng.3656 (2016).
Giambartolomei, C. et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet. 10, e1004383. https://doi.org/10.1371/journal.pgen.1004383 (2014).
Ratnapriya, R. et al. Retinal transcriptome and eQTL analyses identify genes associated with age-related macular degeneration. Nat. Genet. 51, 606–610. https://doi.org/10.1038/s41588-019-0351-9 (2019).
Consortium, G. T. Human genomics. The Genotype-Tissue Expression (GTEx) pilot analysis: Multitissue gene regulation in humans. Science 348, 648–660. https://doi.org/10.1126/science.1262110 (2015).
Narahara, M. et al. Large-scale East-Asian eQTL mapping reveals novel candidate genes for LD mapping and the genomic landscape of transcriptional effects of sequence variants. PLoS One 9, e100924. https://doi.org/10.1371/journal.pone.0100924 (2014).
Ishigaki, K. et al. Polygenic burdens on cell-specific pathways underlie the risk of rheumatoid arthritis. Nat. Genet. 49, 1120–1125. https://doi.org/10.1038/ng.3885 (2017).
Lappalainen, T. et al. Transcriptome and genome sequencing uncovers functional variation in humans. Nature 501, 506–511. https://doi.org/10.1038/nature12531 (2013).
Dempster, A. P. Covariance selection. Biometrics 28, 157–175 (1972).
Chen, F., Curran, P. J., Bollen, K. A., Kirby, J. & Paxton, P. An empirical evaluation of the use of fixed cutoff points in RMSEA test statistic in structural equation models. Sociol. Methods Res 36, 462–494. https://doi.org/10.1177/0049124108314720 (2008).
Pruim, R. J. et al. LocusZoom: Regional visualization of genome-wide association scan results. Bioinformatics 26, 2336–2337. https://doi.org/10.1093/bioinformatics/btq419 (2010).
Franceschini, N. et al. GWAS and colocalization analyses implicate carotid intima-media thickness and carotid plaque loci in cardiovascular outcomes. Nat. Commun. 9, 5141. https://doi.org/10.1038/s41467-018-07340-5 (2018).
Liu, B., Gloudemans, M. J., Rao, A. S., Ingelsson, E. & Montgomery, S. B. Abundant associations with gene expression complicate GWAS follow-up. Nat. Genet. 51, 768–769. https://doi.org/10.1038/s41588-019-0404-0 (2019).
Mall, M. et al. Myt1l safeguards neuronal identity by actively repressing many non-neuronal fates. Nature 544, 245–249. https://doi.org/10.1038/nature21722 (2017).
Liu, H. et al. Ciliary margin transdifferentiation from neural retina is controlled by canonical Wnt signaling. Dev. Biol. 308, 54–67. https://doi.org/10.1016/j.ydbio.2007.04.052 (2007).
Goodman, K. M. et al. Molecular basis of sidekick-mediated cell-cell adhesion and specificity. Elife https://doi.org/10.7554/eLife.19058 (2016).
Parmeggiani, F. et al. Mechanism of inflammation in age-related macular degeneration. Mediators Inflamm. 2012, 546786. https://doi.org/10.1155/2012/546786 (2012).
Thomas, C. N., Berry, M., Logan, A., Blanch, R. J. & Ahmed, Z. Caspases in retinal ganglion cell death and axon regeneration. Cell Death Discov. 3, 17032. https://doi.org/10.1038/cddiscovery.2017.32 (2017).
Bilder, R. M. et al. Phenomics: The systematic study of phenotypes on a genome-wide scale. Neuroscience 164, 30–42. https://doi.org/10.1016/j.neuroscience.2009.01.027 (2009).
Eriksson, N. et al. Web-based, participant-driven studies yield novel genetic associations for common traits. PLoS Genet. 6, e1000993. https://doi.org/10.1371/journal.pgen.1000993 (2010).
Nelson, C. P. et al. Association analyses based on false discovery rate implicate new loci for coronary artery disease. Nat. Genet. 49, 1385–1391. https://doi.org/10.1038/ng.3913 (2017).
Ward, L. D. & Kellis, M. HaploReg: A resource for exploring chromatin states, conservation, and regulatory motif alterations within sets of genetically linked variants. Nucleic Acids Res. 40, D930-934. https://doi.org/10.1093/nar/gkr917 (2012).
Denny, J. C. et al. PheWAS: Demonstrating the feasibility of a phenome-wide scan to discover gene-disease associations. Bioinformatics 26, 1205–1210. https://doi.org/10.1093/bioinformatics/btq126 (2010).
Roden, D. M. Phenome-wide association studies: A new method for functional genomics in humans. J. Physiol. 595, 4109–4115. https://doi.org/10.1113/JP273122 (2017).
Hebbring, S. Genomic and phenomic research in the 21st century. Trends Genet. 35, 29–41. https://doi.org/10.1016/j.tig.2018.09.007 (2019).
Evans, D. M. & DaveySmith, G. Mendelian randomization: New applications in the coming age of hypothesis-free causality. Annu. Rev. Genom. Hum. Genet. 16, 327–350. https://doi.org/10.1146/annurev-genom-090314-050016 (2015).
Pingault, J. B. et al. Using genetic data to strengthen causal inference in observational research. Nat. Rev. Genet. 19, 566–580. https://doi.org/10.1038/s41576-018-0020-3 (2018).
Sakane, Y. et al. Development and validation of the Dry Eye-Related Quality-of-Life Score questionnaire. JAMA Ophthalmol. 131, 1331–1338. https://doi.org/10.1001/jamaophthalmol.2013.4503 (2013).
Ishikawa, S., Takeuchi, M. & Kato, N. The combination of strip meniscometry and dry eye-related quality-of-life score is useful for dry eye screening during health checkup: Cross-sectional study. Medicine (Baltimore) 97, e12969. https://doi.org/10.1097/MD.0000000000012969 (2018).
The Vision Council. Digital Eye Strain [Internet]. https://www.thevisioncouncil.org/content/digital-eye-strain.
Coles-Brennan, C., Sulley, A. & Young, G. Management of digital eye strain. Clin. Exp. Optom. 102, 18–29. https://doi.org/10.1111/cxo.12798 (2019).
Ozawa, Y. et al. Bilberry extract supplementation for preventing eye fatigue in video display terminal workers. J. Nutr. Health Aging 19, 548–554. https://doi.org/10.1007/s12603-014-0573-6 (2015).
McLaren, W. et al. The Ensembl variant effect predictor. Genome Biol. 17, 122. https://doi.org/10.1186/s13059-016-0974-4 (2016).
Acknowledgements
The authors thank all participants enrolled in the present study. No funding bodies had any role in study design, data collection and analysis, decision to publish, or preparation of the manuscript. The authors alone are responsible for the content and writing of the paper.
Funding
This study was funded by Kirin Holdings Company, Limited and DeNA Life Science, Inc. No funding bodies had any role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
K.Y., Y.M., K.Konomi, D.F. and K.Kobayashi designed research; K.Y. and Y.M. performed research; K.Y., Y.M., S.I. and M.T. analyzed data; K.Y. and Y.M. wrote the paper; and Y.M., K.Kobayashi and M.T. conceived and supervised the research. All authors contributed to the discussion and were involved in editing the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
This study was funded by Kirin Holdings Company, Limited and DeNA Life Science, Inc. K.Y., S.I., K.Kobayashi, and M.T. are employees of DeNA Life Science, Inc. Y.M. and D.F. are employees of Kirin Holdings Company, Limited. The authors declare no other conflict of interest associated with this manuscript.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yoshimura, K., Morita, Y., Konomi, K. et al. A web-based survey on various symptoms of computer vision syndrome and the genetic understanding based on a multi-trait genome-wide association study. Sci Rep 11, 9446 (2021). https://doi.org/10.1038/s41598-021-88827-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-88827-y
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.