Abstract
APOBEC3 (A3) enzymes are best known for their role as antiviral restriction factors and as mutagens in cancer. Although four of them, A3A, A3B, A3F and A3G, are induced by type-1-interferon (IFN-I), their role in inflammatory conditions is unknown. We thus investigated the expression of A3, and particularly A3A and A3B because of their ability to edit cellular DNA, in Systemic Lupus Erythematosus (SLE), a chronic inflammatory disease characterized by high IFN-α serum levels. In a cohort of 57 SLE patients, A3A and A3B, but also A3C and A3G, were upregulated ~ 10 to 15-fold (> 1000-fold for A3B) compared to healthy controls, particularly in patients with flares and elevated serum IFN-α levels. Hydroxychloroquine, corticosteroids and immunosuppressive treatment did not reverse A3 levels. The A3AΔ3B polymorphism, which potentiates A3A, was detected in 14.9% of patients and in 10% of controls, and was associated with higher A3A mRNA expression. A3A and A3B mRNA levels, but not A3C or A3G, were correlated positively with dsDNA breaks and negatively with lymphopenia. Exposure of SLE PBMCs to IFN-α in culture induced massive and sustained A3A levels by 4 h and led to massive cell death. Furthermore, the rs2853669 A > G polymorphism in the telomerase reverse transcriptase (TERT) promoter, which disrupts an Ets-TCF-binding site and influences certain cancers, was highly prevalent in SLE patients, possibly contributing to lymphopenia. Taken together, these findings suggest that high baseline A3A and A3B levels may contribute to cell frailty, lymphopenia and to the generation of neoantigens in SLE patients. Targeting A3 expression could be a strategy to reverse cell death and the generation of neoantigens.
Similar content being viewed by others
Introduction
Apolipoprotein B mRNA-editing enzyme catalytic polypeptide-like 3 (APOBEC3 or A3) enzymes are a family of six cytidine deaminases (A3A-A3C, A3F-A3H) which deaminate cytidine (C) into uracil (U) on single stranded DNA (ssDNA)1,2. Uracil in genomic DNA can be read as a thymidine (T), resulting in C > T transitions. Alternatively, downstream processing of genomic uracil by the cellular Base Excision Repair (BER) machinery may correctly repair the mutation, lead to C > T transitions or C > G transversions depending on the polymerase involved, or generate double-stranded DNA breaks (DSBs)3,4,5. This latter mechanism has been shown to operate following deamination by AID, A3A and A3B6,7,8. A3A and A3B are mainly expressed in monocytes and B-cells respectively, while most other A3F and A3G predominate in T-cells9. A3 enzymes are a powerful intrinsic immune mechanism shielding cells against a range of viruses and retro-elements10,11,12,13,14. In keeping with their antiviral function, four of the seven A3 enzymes (A3A, A3B, A3F and A3G) are strongly induced by IFN-α9,15,16.
For unknown reasons A3A and A3B are able to edit chromosomal DNA8,17,18. Accordingly, clusters of C > T edits (or G > A on the opposite strand) attributed to A3A and A3B are the most common somatic mutations found in numerous cancer genomes6,7,19,20. As such, A3A and A3B are now acknowledged as DNA mutators. Recent studies suggest that A3A is the main enzyme responsible for APOBEC-induced mutations in some cancers for its higher deamination activity and propensity to generate DSBs21,22,23. Epidemiological studies have linked polymorphisms within A3A and/or A3B to different cancers24,25,26,27. Single Nucleotide Polymorphisms (SNPs) in A3A have been associated with liver, pancreas, bladder and lung cancer not related to smoking24,25,26,27. A 29 kbase deletion between exon 5 of A3A and exon 8 of A3B (A3AΔ3B) is a prevalent polymorphism in South East Asia and is almost fixed in Oceania28. This deletion generates a chimeric A3A–A3B transcript where the A3A coding sequence is unaffected but terminates with the A3B 3′ untranslated region (3′UTR). This A3AΔ3B deletion is associated with increased A3A expression28. A3AΔ3B is overrepresented in breast, ovarian and liver cancers and was proposed as a cancer-susceptibility gene6,28,29,30,31,32,33. In homozygous breast cancer carriers, it was associated with increased immune activation leading to hypermutation26.
A3 enzymes, and particulary A3A, are also involved in the catabolism of mitochondrial (mt) DNA (mtDNA) leaked into the cytoplasm in response to stress, circumventing chronic immune stimulation and apoptosis8,17,18,34,35. Nevertheless, the role of APOBEC3 deaminases in chronic inflammatory conditions has not been investigated. Chronic inflammation is a fertile ground for malignant transformation. A recent study investigating the link between dystrophic epidermolysis bullosa, a rare skin disease characterized by fragile skin and continual inflammation, and the development of squamous cell carcinoma (SCC), has unambigously identified APOBEC-mutation signatures as the most likely mechanistic cause of inflammation-driven SCC36. We hypothesized that a similar link might exist between chronic inflammatory conditions characterized by high circulating Type-I Interferon (IFN-I) such as Systemic Lupus Erythematosus (SLE) and higher cancer incidence. SLE is a chronic, disabling disease characterized by high levels of IFN-α likely associated with gain-of function in cytosolic nucleic acid sensors and sustained oxidative stress accompanied by mtDNA lesions (increased 8-oxoG or 8-OHdG). Sustained oxidative stress and mtDNA lesions lead to DNA leakage into the cytosol and induce potent IFN-I responses. Other hallmarks of SLE are profound lymphopenia, improper clearance of apoptotic and necrotic cells and circulating auto-antibodies against DNA and nuclear components37,38,39,40,41. The pathogenesis of SLE is multifactorial, involving genetic, immunological and environmental factors41. There is currently no licensed drug targeting IFN-I or the IFN-I receptor in SLE42. SLE patients have increased incidence of hematological malignancies (non-Hodgkin’s lymphoma, leukemia) and certain solid cancers (vulva and cervix, thyroid, lung, liver), but, although controversial, a decreased risk of hormonal-sensitive cancers (breast, endometrial, prostate)43,44. Some oncogenic pathways such as Akt-145 and Telomerase Reverse Transcriptase (TERT) are activated in SLE patients’ peripheral blood mononuclear cells (PBMCs)46,47,48,49. In cancer, these pathways are activated by mutations. Akt1 mutation E17K results in constitutive activation of the kinase and is considered a driver gene mutation50. Mutations in the TERT promoter generating novel binding sites for transcription factors of the E-twenty-six (Ets)/TCF family are a common mechanism of TERT reactivation51,52,53. Akt1 E17K and TERT promoter mutations conform to the A3A/A3B preferred target, i.e. TpCpW, where W stands for A or T7,21.
As A3A and A3B are induced by IFN-I, and because A3A contributes to catabolizing cytoplasmic mtDNA8,18,34,35, we hypothesized that these enzymes are upregulated in SLE in response to mtDNA leaked into the cytoplasm. Sustained A3A and A3B expression might however edit nuclear DNA, thereby contributing to lymphopenia during flares, but also generating neo-antigens which in turn fuel the auto-immune response against nuclear antigens and the mutational burden. In this study, we present experimental evidence that A3A, A3B, A3C and A3G mRNA expression levels are significantly upregulated in SLE patients compared to healthy controls, particularly in patients with severe disease. In the studied samples, A3 upregulation was independent of treatment. The A3AΔ3B polymorphism was detected more often in patients with severe SLE and was associated with higher A3A mRNA levels, but not with any of the other A3. A3A and A3B levels were also correlated with higher DSBs and with lower lymphocyte counts. When SLE-patient cells were exposed to IFN-α, recapitulating flares in vitro, they readily expressed massive amounts of A3A and died within 24 h. Furthermore, a common polymorphism (rs2853669 A > G) which decreases TERT expression54,55,56,57,58,59, was much more common in SLE patients than in healthy controls. These findings clearly point to a role for A3A and A3B in the pathogenesis of lupus and in inflammation-associated lymphopenia.
Results and discussion
Characteristics of the SLE patients
Patients’ baseline characteristics are described in Table 1. Mean (± standard deviation) age at sample collection was 32.9 ± 12.5 years. Mean disease duration was 8.8 ± 8. The majority of patients had a clinically and/or serologically active disease. Thirty patients suffered from severe flares, 11 from mild or moderate flares and 16 had no flares. The median (range) SELENA-SLE disease activity index (SLEDAI) score was 6 (0–32). Thirty (57%) patients had a positive Farr assay and 23 (40%) had low C3 serum level. Thirty-nine (68.4%) patients had serum auto-antibodies (14 patients had auto-antibodies against one ribonucleoprotein (RNP), SSA52/TRIM21, SSA/Ro60, SSB or Sm and 25 patients against two or more of the above nuclear auto-antigens). Eighteen (31.6%) SLE patients had no detectable auto-antibodies. Serum IFN-α levels were elevated in 33 (58%) patients, with a median of 2 IU/mL (0–201).
Treatment for SLE is based on hydroxychloroquine (HCQ), glucocorticosteroids (Prednisone (PDN)), and immunomodulatory agents (methotrexate (MTX), mycophenolate mofetil (MMF), Cyclophosphamide (Cyc), Azathioprine (AZA))60,61. In our cohort, most SLE patients were treated (treated: 47 patients, no treatment: 10 patients) (Table 1).
A3A and A3B are upregulated in SLE patients
The two APOBEC3 enzymes which edit nuclear DNA, A3A and A3B, are ISGs. We therefore first compared A3A and A3B mRNA between SLE patients and healthy controls. A3A and A3B mRNA expression were ~ 10 to 15-fold (p < 0.01) and > 1000-fold (p < 0.001) higher respectively in SLE patients than in healthy controls (Fig. 1A,B). Higher A3B levels compared to A3A have been reported previously in cancer patients; nevertheless, the deamination activity of A3A is 100-fold higher than that of A3B8,21,22,23, explaining why such high levels of A3B, but not A3A, are compatible with cell viability. Analysis of RNA-Seq data from an independent cohort of 99 active SLE patients and 18 healthy controls (GEEO: GSE72509)64 confirmed a > 2.5-fold increase in A3A expression (p < 0.0001) and a threefold increase in A3B expression (p = 0.002) (data not shown). The fold-increase in RNA-Seq ranged from 0.5 to 6 for A3A and from 1 to 100 for A3B64. The less dramatic upregulation recorded with RNA-Seq likely reflects lower sensitivity of Illumina RNA-Seq chips and/or primer design, particularly considering the high sequence homology between A3 genes.
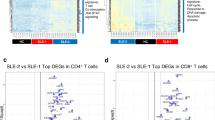
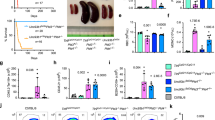
A3 relative expression in a cohort of SLE patients. A–D Relative A3A (A), A3B (B), A3C (C) A3G (D) expression (mRNA) in SLE patients and in age- and sex-matched healthy controls). (E) Relative expression of RIG-I in SLE patients and in healthy controls. F–J. A3A (F) A3B (G), A3C (H), A3G (I) and RIG-I (J) expression in relation to serum IFN-I levels (IU/mL). K–O. Correlation of expression of A3A (K), A3B (L), A3C (M), A3G (N) and RIG-I (O) with lymphocyte counts. Target gene (A3A, A3B, A3C, A3G and RIG-I) levels were normalized to the geometric mean of housekeeping genes RPL13A and GAPDH according to the Pfaffl method65. Control C1 was used as calibrator for comparison and is therefore set to 1. The mean of duplicate measures ± SEM are reported. Patients with 2 wt A3B alleles are represented by colored symbols (red circles in panels A F and K, blue triangles in panels B, G and L, green circles in panels C, H and M, purple diamonds in panels D, I and N and orange circles in panels E, J and O); patients/donors heterozygous for the A3AΔ3B deletion are represented by full black symbols in panels A though J. *p < 0.05; **p < 0.01; ***p < 0.001.
A3A and A3B expression was higher in patients with elevated serum IFN-α and was weakly correlated with IFN-α levels (for A3A: r = 0.3285, p = 0.0163 and for A3B: r = 0.3227, p = 0.0196), as expected (Fig. 1F,G). It is possible that the heterogeneity and fluctuation of symptoms associated with SLE partially contribute to the weak correlation between A3A and A3B levels and IFN-α. Although only A3A and A3B edit DNA, we also measured the expression of two other ISGs, A3G and RIG-I, as well as A3C. All were upregulated in SLE patients compared to healthy controls (A3G: p = 0.0354; A3C: p = 0.0270; and RIG-I: p = 0.0012) and particularly in patients with detectable serum IFN-I levels (Fig. 1C–E). When SLE patients with detectable IFN-α in plasma were compared to patients with no detectable IFN-α, A3A (p < 0.05), A3C (p < 0.01), A3G (p < 0.01) and RIG-I (p < 0.01) expression were also more elevated (Fig. 1H–J). The observation that all were upregulated in SLE patients could be due to the fact that SLE is a multifactorial disease featuring many immunological dysregulations. Although A3C is generally not considered an ISG because it is much less responsive to IFN-I than A3G and A3F, its expression can be induced by IFN-I in PBMCs and in hepatocytes15,16. Furthermore, it is possible that other factors account for its upregulation in SLE patients. Nevertheless, since neither A3G nor A3C edit genomic DNA, it is unlikely that they contribute to lymphopenia. In line with this view, only A3A and A3B mRNA levels, were inversely correlated with lymphocyte counts (Fig. 1K,L) while A3C, A3G, and RIG-I mRNA levels were not (Fig. 1M–O), suggesting that genomic DNA editing by A3A and A3B may play a role in lymphopenia.
Taken together, these findings suggest that in SLE patients, the IFN-α response triggered by sustained cytosolic DNA due to oxidative mutations39 leads to persistant A3A and A3B expression among other ISGs. These deaminases may participate in DNA catabolism, but most of all, they could edit nuclear DNA. In this case, uridine resulting from cytidine deamination is removed by Uracyl DNA glycosylases (UNG), creating an apyrimidinic site. This mobilization can activate the DNA repair machinery, leading to C > T or C > G mutations which may be fixed5,6,7. Alternatively, juxtaposed apyrimidinic sites will generate DSBs causing cell death5,6. In both cases, A3-mediated deamination events may generate neoantigens, increasing the B-cell response against self-epitopes in a feed-forward loop and contributing to lymphopenia. Accordingly, A3A mRNA levels were higher in SLE patients with auto-antibodies against nuclear proteins (p < 0.05) and against dsDNA (Supplementary Fig. 1I) than in patients with no auto-antibodies, although statistical support was reached only for nuclear proteins. For A3B, mRNA levels were higher in SLE patients than in controls regardless of the presence of auto-antibodies (Supplementary Fig. 1J). For A3C and A3G, the levels of expression did not differ between SLE patients with or without auto-antibodies against nuclear antigens or against DNA (Supplementary Fig. 1 1 K and 1L). Although all A3 enzyme mRNA measured in SLE patients were upregulated, these observations are strongly suggestive of a role for A3A- and A3B-induced DSBs in cell death and the induction of auto-antibodies in the pathogenesis of SLE.
Hierarchical clustering of patients based on disease severity showed significantly exacerbated A3A, A3B and A3C expression in patients with a postive SLEDAI (Supplementary Fig. 1A to 1D), as well as with the presence or severity of flares (Supplementary Fig. 1E to 1H). Notably, hydroxychloroquine, corticosteroids and immunosuppressive treatment failed to reverse elevated A3A or A3B levels back to background levels (Supplementary Fig. 1M to 1N), while the difference in A3C and A3G mRNA levels did not differ significantly between treated SLE patients, untreated SLE patients and controls (Supplementary Fig. 1O and 1P).
In the long run, sustained exposure to A3A and A3B mutational fuel could also generate oncogenic driver events23,30,31,32, providing a direct molecular rationale for the higher prevalence of certain tumors among SLE patients. We therefore searched for the presence of subclonal oncogenic driver mutations conforming to the preferred target for A3A and A3B, i.e. TpCpW7,21,66, in 2 oncogenes, akt1 and the TERT promoter. We obtained sufficient sequences for all controls and 47 SLE patients. The threshold for distinguishing true subclonal mutations from error rate was set at 1 base call per 100 sequenced nucleotides. With these settings, we did not detect APOBEC3-mutations in akt1 (not shown) nor in the TERT promoter (Supplementary Table 1). Because SLE patients in this cohort do not have cancer, we expected to find only a handful of APOBEC3-mutations, if any. Indeed, APOBEC3-mutations are generally not detected in non-cancerous, TP53- and UNG-positive cells67. It is however not excluded that subclonal mutations may be present at frequencies below the set threshold or elsewhere in the genome.
High prevalence of the A3AΔ3B germline deletion among SLE patients
A 29-kb germline deletion spanning the A3B locus generates a chimeric A3A-UTRA3B transcript (A3AΔ3B). A3AΔ3B is associated with increased A3A mRNA stability and protein expression28 and with a higher incidence of APOBEC3-mutations and cancer6,23,29,30,31,32. Copy number evaluation of A3B was performed by ddPCR to assess the prevalence of the A3AΔ3B polymorphism among SLE patients. The A3AΔ3B polymorphism was detected at the heterozygous state in 7/47 (14.9%) SLE patients and 1/10 (10%) controls. Although patients with severe flares (4/21 (19%)) had a slightly higher incidence of the polymorphism (Fig. 2A), statistical significance was not reached. As expected, the A3AΔ3B polymorphism was associated with higher A3A expression (p < 0.05) (Fig. 2B). A3B, A3G, A3C and RIG-I expression levels were unaffected by the single A3B allele (not shown).
Prevalence and impact of the 29-kb deletion spanning A3B (A3AΔ3B) on A3A expression. (A) Prevalence of the A3AΔ3B deletion in SLE patients with severe or mild/moderate flares. (B) A3A relative expression in SLE patients with wild type A3B or with the A3AΔ3B deletion. *p < 0.05; **p < 0.01; ***p < 0.001.
High IFN-I-induced A3A expression leads to SLE-cell death
To further assess the role of detectable IFN-α and A3A in lymphopenia, freshly isolated PBMCs from SLE patients and healthy controls were put in culture with IFN-α for 24 h, mimicking flares. This setting recapitulates the fact that in SLE patients, but also in other chronic inflammatory conditions such as multiple sclerosis, flares often arise following an infectious episode, which likely triggers IFN-α. Initially, PBMCs from 5 SLE patients and 4 healthy controls were isolated and immediately exposed to IFN-α. By 4 h, IFN-α triggered a ~ tenfold increase in A3A mRNA, which was sustained throughout the time of the experiment in healthy controls (Fig. 3A). In SLE patients in contrast, IFN-α treatment increased A3A expression by more than 3 orders of magnitude (p < 0.05) (Fig. 3A). Such high A3A levels persisted 16 h post-treatment for all SLE patients, and by 24 h we could not measure A3A nor housekeaping genes, suggesting that SLE cells had died or were undergoing apoptosis (Fig. 3A). A3A and housekeeping gene mRNA could be quantified normally in cells from healthy controls treated with IFN-α, indicating that IFN-α per se did not induce massive cell death at these concentrations. In line with the high A3A mRNA levels in SLE patients, we recorded markedly higher levels of DSBs (γ-H2AX staining) in PBMCs from these SLE patients compared to healthy controls after 4 h (p = 0.0095) and 16 h (p = 0.019) in culture (Fig. 3B), indicating that higher A3A mRNA levels translate functionnally into increased DSBs and cell death.
SLE patient cells do not survive in the presence of IFN-α. (A) PBMCs from SLE or healthy controls were isolated from blood and treated with IFN-α immediately after isolation and relative A3A expression quantified by qRT-PCR and normalized to the geometric mean of housekeeping genes RPL13A and GAPDH according to the Pfaffl method65 for each sample. The fold change relative to the untreated sample is reported. (B) Untreated cells from the same donors as in A were stained for DSBs (γ-H2AX) after 4 h and 16 h in culture. (C) PBMCs from SLE or healthy controls were isolated from blood, left to rest overnight and only then treated with IFN-α. Relative A3A expression was quantified by qRT-PCR and normalized to the geometric mean of housekeeping genes RPL13A and GAPDH according to the Pfaffl method65 for each sample. The fold-change in IFN-α-treated compared to non treated control 1 is reported. *p < 0.05; **p < 0.01; ***p < 0.001.
Next, PBMCs from 7 SLE patients and 3 healthy controls were isolated, left to rest overnight before IFN-α treatment. In controls, A3A upregulation started around 8 h and reached only ~ 100-fold after 24 h (Fig. 3C). In SLE patients, IFN-α increased A3A mRNA by up to ~ 200-fold as early as 4 h post-exposure and the increase persisted at 8 and 24 h. Thus, A3A induction was much faster and stronger in SLE patients compared to healthy control cells even when cells were left to rest overnight after isolation (p < 0.05 at 8 h) (Fig. 3C).
These results nicely recapitulate the profound lymphopenia observed during flares in SLE patients, and point to A3A as a chief actor in the massive cell death that characterizes flares. They suggest that not only are basal A3A expression and DSBs persistently higher in SLE patients than in healthy individuals, but also that their cells are “primed” such that A3A expression is readily boosted to extremely high levels while it takes healthy cells much longer to trigger A3A to 100-fold lower levels. No cell could survive such high levels of A3A. This immediate and massive upregulation of A3A suggests the A3A promoter is poised in SLE cells.
High prevalence of the TERT rs2853669 A > G polymorphism in SLE patients
Although we did not detect de novo somatic mutations attributable to A3 cytidine deaminase activity in the TERT promoter of our cohort, we found a surprisingly high prevalence of the germline polymorphism rs2853669 A > G among SLE patients compared to healthy controls (p = 0.0206). Of 48 SLE patients with TERT promoter sequences, 33 (68.75%) carried the rs2853669 A > G polymorphism in the TERT promoter on both alleles. Fifteen SLE patients (31.25%) had at least one GGAA Ets/TCF binding site (5 patients were heterozygous and 10 had wild-type rs2853669 A). In contrast, only 2 of 10 healthy controls (20%) were homozygous for rs2853669 A > G while all others harbored at least one copy of the rs2853669 A allele (5 were heterozygous and 3 were homozygous) (Fig. 4A).
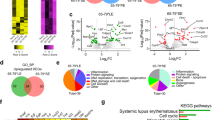
Prevalence of the TERT rs2853669 polymorphism in SLE patients. (A) Prevalence of the TERT promoter rs2853669 A/G alleles in SLE patients and controls. The number of GGAA sites at position − 245 from the TSS was assessed from the TERT promoter reads. (B) Prevalence of TERT promoter rs2853669 A/G alleles in SLE patients stratified according to serum IFN-α levels. Full black circles represent patients heterozygous for the A3AΔ3B deletion. (C) and (D) A3A (C) and A3B (D) expression levels in SLE patients based on rs2853669 G/A polymorphism. Full symbols are SLE patients and black open symbols are controls. *p < 0.05; **p < 0.01; ***p < 0.001.
The rs2853669 A > G is a common germline polymorphism which disrupts a prexisting Ets/TCF binding site located 245 bp upstream of the TERT TSS. Accordingly, the rs2853669 A > G polymorphism decreases TERT transcription in vitro. The rs2853669 A > G polymorphism has been investigated in cancers with TERT promoter mutations, where it may reverse their effect and cell immortalization54,55,56,57,58,59, but has not been investigated in inflammatory conditions to our knowledge. This germline polymorphism is not an APOBEC-mutation and accordingly, we found no relationship between the prevalence of the polymorphism and detectable IFN-α in serum (Fig. 4B) nor with A3A or A3B mRNA levels (Fig. 4C,D) or disease severity (severe or mild/moderate flares, SLEDAI > 10) (Supplementary Fig. 2A and 2B). Nevertheless, peripheral blood mononuclear subsets from SLE patients, and particularly terminally differentiated memory T cells, display shortened telomeres as a result of sustained immune activation. It was suggested that TERT activity is insufficient to compensate for telomere erosion and accelerated replicative senescence of immune cells in these patients46,47,48,49,68. Because the rs2853669 polymorphism decreases TERT activity, individuals carrying this polymorphim may have lower TERT activity and accelerated lymphocyte senescence. Although we could not test this hypothesis in our cohort, it is possible that repeated PBMC activation episodes during flares combined to a decreased inability to rescue activated immune cells from senescence could contribute to the frailty of SLE patients’ PBMCs and to lymphopenia.
Conclusions
Our results document that multiple A3 enzymes, including IFN-induced A3A, A3B and A3G, and A3C, are strongly and persistently upregulated in SLE patients despite immunosuppressive treatment, and suggest a link between elevated baseline A3A levels, the A3AΔ3B polymorphism and disease severity. Since A3A and A3B can access the nucleus, these enzymes have every chance of editing nuclear DNA leading to DSBs. In this context, exposure to IFN-α leads to an immediate and massive upregulation of A3A expression, far beyond levels compatible with DNA repair, and ultimately to cell death. It is easily conceivable how high steady state A3A together with shortened telomeres due in part to the presence of the rs2853669 A > G polymorphism, might prime cells such that any further inflammatory signal leads to impelling A3A expression and massive cell death. Massive cell death in turn leads to exposure of nuclear antigens, fueling the inflammatory response and further incrementing the generation of auto-antibodies in a vicious cycle. These findings further underscore the need to include therapies targeting interferons and/or specifically A3 enzymes in the management of patients with lupus and probably other inteferonopathies42.
Methods
Patients and patient samples
Blood samples were obtained from 57 patients diagnosed with SLE according to the 1997 American College of Rheumatology criteria for SLE classification69. SLE patients were routinely followed at the French National Referral Center for Systemic Lupus Erythematosus, Groupement Hospitalier Pitié–Salpêtrière, Paris, France. SLE clinical characteristics, the Safety of Estrogens in Lupus Erythematosus National Assessment–Systemic Lupus Erythematosus Disease Activity Index (SELENA–SLEDAI)62,63,70, and the therapeutic regimen were recorded on the day blood was drawn. The class of lupus nephritis was recorded according to ISN/RPS-200371. Routine testing to determine anti-dsDNA Ab titers Farr test (Trinity Biotech; cut-off value: 9.0 IU/mL), anti-ribonucleoprotein Abs (anti-RNP, anti-Sm, anti-SSA/Ro60, anti-Ro52/TRIM21, anti-SSB [Luminex FIDI, Theradiag]), and laboratory analyses (complement C3 levels (Optilite, Binding Site), complete blood counts, serum creatinine, proteinuria and hematuria) were run. The presence of a severe or mild/moderate lupus flare was recorded according to the SELENA-SLEDAI flare instrument62. For some analyses, patients were stratified according to the presence and severity of flares (i.e. no flare versus mild/moderate flare versus severe flare) or the SELENA-SLEDAI score as follows: SLEDAI = 0: non-active Lupus; 0 < SLEDAI ≤ 5: mild condition; 5 < SLEDAI ≤ 10: medium activity; SLEDAI > 10: severe activity. Serum-IFN-α biological activity, expressed in IU/mL, was determined by assessing the protection afforded by each patient’s serum to cultured MDBK cells challenged with vesicular stomatitis virus (VSV), as previously described72. Bioassay sensitivity (i.e. the lower limit of detection) was 2 IU/mL. Serum-IFNα activity in healthy individuals is undetectable, i.e. < 2 IU/mL72.
The study was approved by CPP Ile-de-France VI Ethics Committee. Samples were collected between November 2015 and October 2016. Ten sex-matched healthy controls whose blood was collected between June 15 and July 26, 2016 by the ICAReB platform at Institut Pasteur (Paris, France) were used for all experiments except for cell culture. All the participants gave written informed consent in the frame of the healthy volunteers CoSImmGEn cohort (Clinical trials NCT 03,925,272), which was approved by the CPP Ile-de-France I Ethics Committee (Jan 18, 2011). For cell culture experiments, blood from healthy volunteer blood donors from the Croix-Rouge Luxembourg was used. Written informed consent was provided by all patients and healthy donors. The research was carried out in compliance with the Helsinki Declaration.
Blood (2 × 7 mL) was collected on EDTA or PaxGene. Peripheral blood mononuclear cells (PBMCs) were isolated immediately by centrifugation, washed, split in two and dry pellets were stored at − 20 °C for DNA extraction and at − 80 °C for RNA extraction until use.
RNA isolation and RT-qPCR
Total RNA was extracted from frozen PBMC pellets using Trizol. One microgram of total RNA was reverse transcribed with the QuantiTect reverse transcription kit (Qiagen). qPCR for A3A, A3B, A3C and A3G, RPL13A and GAPDH was performed in duplicate using Takyon Rox dTT Blue 2X Master Mix (Eurogentec), primers and Universal Probe Library probes as described in16 or the Applied Biosystems kit for GAPDH. For RIG-I, the following primers were used: FRIG-I: 5′-CTTTTTCTCAAGTTCCTGTTGGA-3′ and RRIG-I:5′-TCCCAACTTTCAATGGCTTC-3′, with UPL probe #79 (Roche). All genes of interest were normalized to the geometric mean of housekeeping genes RPL13A and GAPDH according to the Pfaffl method65 and the amplification efficiencies for these primers16. A3A, A3B, A3C, A3G and RIG-I relative expression levels were calculated by comparison with Healthy Donor C1.
Next generation sequencing (NGS)
DNA was extracted from frozen PBMC pellets using the Epicentre kit and 1 μg of genomic DNA was amplified using Platinum HiFi Taq (Invitrogen). The Telomerase Reverse Trancriptase (TERT) core promoter was amplified using outer primers hTERT-out-F: 5′-AGTGGATTCGCGGGCACAGA73 and hTERT-out-R: 5′-GGCTTCCCACGTGCGCAGCAGGA74 and nested PCR primers : hTERT-in-F : 5′-GCACCCGTCCTGCCCCTTCACCT and hTERT-in-R : 5′-CAGCAGGACGCAGCGCTGCCTGA, spanning mutations C228T and C250T. Akt1 intron 1 + exon 2, spanning mutation E17K, was amplified using primers Akt1-F: 5′-GCTGCCTGGCGAAGGTCTGACG and Akt1-R: 5′-CCTTGTAGCCAATGAAGGTGCC. PCR settings for both genes were: 5 min denaturation at 94 °C followed by 40 amplification cycles (94 °C 1 min, 63 °C 1 min and 68 °C 1 min) and a 10 min final extension at 68 °C. PCR products for the 2 genes were obtained for all controls and for 47 SLE patients for the TERT promoter and for 46 patients for Akt1. PCR products were gel purified, adaptors were added according to standard procedures for Illumina NextSeq 500 sequencing. For data processing, quality control was performed using FASTQC. All sequences were cut-off at a minimum quality PHRED score of 20. A minimum read length of 125 bp was selected. Reads were mapped against TERT (AF097365) and Akt1 (NG_012188) reference sequences using the Geneious Software (V11.0.2) and Bowtie2 with standard settings for high sensitivity. SNPs were detected with the Geneious Software SNP detection tool with the following parameters: minimum coverage 500, minimum variant frequency 0.01, maximum variant p-value 1e-6 and minimum strand bias p-value 1e-5. Four SLE patients were excluded from further analyses because of insufficient reads mapping to the genes of interest.
Detection of A3AΔ3B by ddPCR
A3B copy number was determined using the droplet digital PCR system (BioRad laboratories) in a duplex PCR targeting A3B and reference gene RPP30 (RNase P, 2 copies per diploid genome75) for normalization. The A3B target region falls within the 29,5 kb deletion and within an 800 bp-fragment strictly specific to A3B. The following primers were used to generate a 60 bp fragment: F1_3B: 5′-GGCTGGACTCGCAGTCAC and R1_3B: AACAGCAGGGCTTAGGAACA together with A3B–specific probe UPL33 (Roche). Reaction mixtures (20 μL) comprised 10 μL 2× ddPCR Supermix (with dUTP; Bio–Rad), 900 nM of each A3B–specific PCR primer, 250 nM A3B–specific probe (UPL33) and 2 µL of 20× PrimePCR ddPCR reference assay (RNase P-HEX Bio-Rad). Genomic DNA (25 ng in 5 µL) was added to the ddPCR mixture and directly digested at room temperature with 3U of HaeIII (Thermo Scientific). Droplets were generated with a QX100 Droplet Generator (Bio–Rad) by mixing 20 μL of the assay reaction with 70 μL of droplet oil into the QX200 DG cartridge (Bio-Rad) according to the manufacturer’s recommendations, and were carefully transferred into a PCR plate. Thermal cycling conditions in a Bio-Rad iCycler (slow ramp rate, 2 °C per second) consisted in 10 min at 95 °C, 40 cycles of a two-step thermal profile of 15 s at 94 °C and 60 s at 59 °C, and a final step at 98 °C for 10 min. Droplets were counted using the QX200 Droplet Reader and A3B copy number assigned using QuantaSoft software (Bio-Rad). Each 96 well plate included 8 randomly distributed `no template’ control reactions to check for the absence of contamination.
Cell culture
Blood samples (30–35 mL) from an additional 12 patients with SLE (10 with inactive disease and 2 with active disease) and from 7 healthy donors from Croix Rouge Luxembourg, were collected on EDTA. No clinical data was available for these patients and healthy controls. PBMCs were isolated immediately by Lymphoprep density gradient centrifugation (Axis-Shield, Oslo, Norway), washed and seeded in 24-well plates (106 cells/well) in RPMI 1640 medium supplemented with 10% FCS, 2 mM L-Glutamine, 50 µg/mL Penicillin and 50 µg/mL Streptomycin and 10 U/mL interleukin 2 (IL-2, Roche). Cells were treated with 750 U/mL IFN-α (PBL biomedical laboratories) either immediately for 4, 16 or 24 h (5 SLE patients and 4 healthy controls) or put to rest overnight and treated with IFN-α the following morning for 4, 8 or 24 h (7 SLE patients and 3 healthy controls). At the indicated time points, cells were harvested, washed and dry pellets were frozen at − 80 °C. RNA extraction and relative A3A and A3B expression levels (mRNA) were measured as for the other samples and normalized to the geometric mean of RPL13A and GAPDH according to the Pfaffl method65 for each sample (IFN-α-treated compared to the corresponding non treated control).
Flow Cytometry
Cells from SLE patients or healthy controls cultured for 4 and 16 h without any stimulus were washed with cold PBS, fixed, permeabilized with methanol and stained with an AF647-conjugated mouse antibody against γ-H2AX (BD Pharmingen, clone N1-431) or AF647-conjugated Isotype control. Cells were analyzed using the MACSQuant cytometer and FlowJo version 10.6 (BD Bioscience).
Statistical analyses
Statistical analyses were performed using GraphPad Prims v5 (GraphPad Software, San Diego, CA, USA). Mean/Median ± SEM were calculated. Groups were compared using a Mann–Whitney t-test for pairs and a Kruskal–Wallis with Dunn’s Multiple Comparison post test for more than two groups. Groups were considered to differ statistically if p < 0.05. Correlations between groups were tested using a Spearman test.
Ethical clearance
The study was approved by the CPP Ile-de-France VI and CPP Ile-de-France I Ethics Committees for SLE patients and healthy volunteers CoSImmGEn cohort respectiviely. All donors provided written informed consent for this study. For cell culture experiments, blood from healthy volunteer blood donors from the Croix-Rouge Luxembourg was used. Written informed consent was provided by all donors. The research was carried out in compliance with the Helsinki Declaration.
References
Jarmuz, A. et al. An anthropoid-specific locus of orphan C to U RNA-editing enzymes on chromosome 22. Genomics 79, 285–296. https://doi.org/10.1006/geno.2002.6718 (2002).
Harris, R. S., Petersen-Mahrt, S. K. & Neuberger, M. S. RNA editing enzyme APOBEC1 and some of its homologs can act as DNA mutators. Mol. Cell 10, 1247–1253 (2002).
Kavli, B. et al. hUNG2 is the major repair enzyme for removal of uracil from U: A matches, U: G mismatches, and U in single-stranded DNA, with hSMUG1 as a broad specificity backup. J. Biol. Chem. 277, 39926–39936. https://doi.org/10.1074/jbc.M207107200 (2002).
Pettersen, H. S. et al. Uracil-DNA glycosylases SMUG1 and UNG2 coordinate the initial steps of base excision repair by distinct mechanisms. Nucleic Acids Res. 35, 3879–3892. https://doi.org/10.1093/nar/gkm372 (2007).
Henderson, S. & Fenton, T. APOBEC3 genes: retroviral restriction factors to cancer drivers. Trends Mol. Med. 21, 274–284. https://doi.org/10.1016/j.molmed.2015.02.007 (2015).
Taylor, B. J. et al. DNA deaminases induce break-associated mutation showers with implication of APOBEC3B and 3A in breast cancer kataegis. Elife 2, e00534. https://doi.org/10.7554/eLife.00534 (2013).
Leonard, B. et al. APOBEC3B upregulation and genomic mutation patterns in serous ovarian carcinoma. Cancer Res. 73, 7222–7231. https://doi.org/10.1158/0008-5472.CAN-13-1753 (2013).
Landry, S., Narvaiza, I., Linfesty, D. C. & Weitzman, M. D. APOBEC3A can activate the DNA damage response and cause cell-cycle arrest. EMBO Rep. 12, 444–450. https://doi.org/10.1038/embor.2011.46 (2011).
Koning, F. A. et al. Defining APOBEC3 expression patterns in human tissues and hematopoietic cell subsets. J. Virol. 83, 9474–9485. https://doi.org/10.1128/JVI.01089-09 (2009).
Sheehy, A. M., Gaddis, N. C., Choi, J. D. & Malim, M. H. Isolation of a human gene that inhibits HIV-1 infection and is suppressed by the viral Vif protein. Nature 418, 646–650. https://doi.org/10.1038/nature00939 (2002).
Mangeat, B. et al. Broad antiretroviral defence by human APOBEC3G through lethal editing of nascent reverse transcripts. Nature 424, 99–103. https://doi.org/10.1038/nature01709 (2003).
Harris, R. S., Sheehy, A. M., Craig, H. M., Malim, M. H. & Neuberger, M. S. DNA deamination: not just a trigger for antibody diversification but also a mechanism for defense against retroviruses. Nat. Immunol. 4, 641–643. https://doi.org/10.1038/ni0703-641 (2003).
Lecossier, D., Bouchonnet, F., Clavel, F. & Hance, A. J. Hypermutation of HIV-1 DNA in the absence of the Vif protein. Science 300, 1112. https://doi.org/10.1126/science.1083338 (2003).
Harris, R. S. & Dudley, J. P. APOBECs and virus restriction. Virology 479–480C, 131–145. https://doi.org/10.1016/j.virol.2015.03.012 (2015).
Bonvin, M. et al. Interferon-inducible expression of APOBEC3 editing enzymes in human hepatocytes and inhibition of hepatitis B virus replication. Hepatology 43, 1364–1374. https://doi.org/10.1002/hep.21187 (2006).
Refsland, E. W. et al. Quantitative profiling of the full APOBEC3 mRNA repertoire in lymphocytes and tissues: implications for HIV-1 restriction. Nucleic Acids Res. 38, 4274–4284. https://doi.org/10.1093/nar/gkq174 (2010).
Stenglein, M. D., Burns, M. B., Li, M., Lengyel, J. & Harris, R. S. APOBEC3 proteins mediate the clearance of foreign DNA from human cells. Nat. Struct. Mol. Biol. 17, 222–229. https://doi.org/10.1038/nsmb.1744 (2010).
Mussil, B. et al. Human APOBEC3A isoforms translocate to the nucleus and induce DNA double strand breaks leading to cell stress and death. PLoS ONE 8, e73641. https://doi.org/10.1371/journal.pone.0073641 (2013).
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421. https://doi.org/10.1038/nature12477 (2013).
Roberts, S. A. et al. An APOBEC cytidine deaminase mutagenesis pattern is widespread in human cancers. Nat. Genet. 45, 970–976. https://doi.org/10.1038/ng.2702 (2013).
Chan, K. et al. An APOBEC3A hypermutation signature is distinguishable from the signature of background mutagenesis by APOBEC3B in human cancers. Nat. Genet. 47, 1067–1072. https://doi.org/10.1038/ng.3378 (2015).
Cortez, L. M. et al. APOBEC3A is a prominent cytidine deaminase in breast cancer. PLoS Genet. 15, e1008545. https://doi.org/10.1371/journal.pgen.1008545 (2019).
Roper, N. et al. APOBEC Mutagenesis and copy-number alterations are drivers of proteogenomic tumor evolution and heterogeneity in metastatic thoracic tumors. Cell Rep. 26, 2651–2666. https://doi.org/10.1016/j.celrep.2019.02.028 (2019).
Golka, K. et al. Genetic variants in urinary bladder cancer: collective power of the “wimp SNPs”. Arch. Toxicol. 85, 539–554. https://doi.org/10.1007/s00204-011-0676-3 (2011).
Schwender, H. et al. Distinct SNP combinations confer susceptibility to urinary bladder cancer in smokers and non-smokers. PLoS ONE 7, e51880. https://doi.org/10.1371/journal.pone.0051880 (2012).
Cescon, D. W., Haibe-Kains, B. & Mak, T. W. APOBEC3B expression in breast cancer reflects cellular proliferation, while a deletion polymorphism is associated with immune activation. Proc. Natl. Acad. Sci. USA 112, 2841–2846. https://doi.org/10.1073/pnas.1424869112 (2015).
Matsuda, K. et al. Genome-wide association study identified SNP on 15q24 associated with bladder cancer risk in Japanese population. Hum. Mol. Genet. 24, 1177–1184. https://doi.org/10.1093/hmg/ddu512 (2015).
Caval, V., Suspene, R., Shapira, M., Vartanian, J. P. & Wain-Hobson, S. A prevalent cancer susceptibility APOBEC3A hybrid allele bearing APOBEC3B 3’UTR enhances chromosomal DNA damage. Nat. Commun. 5, 5129. https://doi.org/10.1038/ncomms6129 (2014).
Xuan, D. et al. APOBEC3 deletion polymorphism is associated with breast cancer risk among women of European ancestry. Carcinogenesis 34, 2240–2243. https://doi.org/10.1093/carcin/bgt185 (2013).
Nik-Zainal, S. et al. Association of a germline copy number polymorphism of APOBEC3A and APOBEC3B with burden of putative APOBEC-dependent mutations in breast cancer. Nat. Genet. 46, 487–491. https://doi.org/10.1038/ng.2955 (2014).
Chen, Z. et al. Integrative genomic analyses of APOBEC-mutational signature, expression and germline deletion of APOBEC3 genes, and immunogenicity in multiple cancer types. BMC Med. Genomics 12, 131. https://doi.org/10.1186/s12920-019-0579-3 (2019).
Hashemi, M., Moazeni-Roodi, A. & Taheri, M. Association of APOBEC3 deletion with cancer risk: A meta-analysis of 26 225 cases and 37 201 controls. Asia Pac. J. Clin. Oncol. 15, 275–287. https://doi.org/10.1111/ajco.13107 (2019).
Kim, Y. S. et al. Clinical implications of APOBEC3A and 3B expression in patients with breast cancer. PLoS ONE 15, e0230261. https://doi.org/10.1371/journal.pone.0230261 (2020).
Suspene, R. et al. Self-cytoplasmic DNA upregulates the mutator enzyme APOBEC3A leading to chromosomal DNA damage. Nucleic Acids Res. 45, 3231–3241. https://doi.org/10.1093/nar/gkx001 (2017).
Mussil, B. et al. Genotoxic stress increases cytoplasmic mitochondrial DNA editing by human APOBEC3 mutator enzymes at a single cell level. Sci. Rep. 9, 3109. https://doi.org/10.1038/s41598-019-39245-8 (2019).
Cho, R. J. et al. APOBEC mutation drives early-onset squamous cell carcinomas in recessive dystrophic epidermolysis bullosa. Sci. Transl. Med. https://doi.org/10.1126/scitranslmed.aas9668 (2018).
Gaipl, U. S. et al. Clearance of apoptotic cells in human SLE. Curr. Dir. Autoimmun 9, 173–187. https://doi.org/10.1159/000090781 (2006).
Niewold, T. B., Hua, J., Lehman, T. J., Harley, J. B. & Crow, M. K. High serum IFN-alpha activity is a heritable risk factor for systemic lupus erythematosus. Genes Immun. 8, 492–502. https://doi.org/10.1038/sj.gene.6364408 (2007).
Lopez-Lopez, L. et al. Mitochondrial DNA damage is associated with damage accrual and disease duration in patients with systemic lupus erythematosus. Lupus 23, 1133–1141. https://doi.org/10.1177/0961203314537697 (2014).
Crow, M. K. Type I interferon in the pathogenesis of lupus. J. Immunol. 192, 5459–5468. https://doi.org/10.4049/jimmunol.1002795 (2014).
Arneth, B. Systemic lupus erythematosus and DNA degradation and elimination defects. Front. Immunol. 10, 1697. https://doi.org/10.3389/fimmu.2019.01697 (2019).
Mathian, A., Hie, M., Cohen-Aubart, F. & Amoura, Z. Targeting interferons in systemic lupus erythematosus: current and future prospects. Drugs 75, 835–846. https://doi.org/10.1007/s40265-015-0394-x (2015).
Kiss, E., Kovacs, L. & Szodoray, P. Malignancies in systemic lupus erythematosus. Autoimmun Rev. 9, 195–199. https://doi.org/10.1016/j.autrev.2009.07.004 (2010).
Bernatsky, S. et al. Cancer risk in systemic lupus: an updated international multi-centre cohort study. J. Autoimmun. 42, 130–135. https://doi.org/10.1016/j.jaut.2012.12.009 (2013).
Garcia-Rodriguez, S. et al. Altered AKT1 and MAPK1 gene expression on peripheral blood mononuclear cells and correlation with T-helper-transcription factors in systemic lupus erythematosus patients. Med. Inflamm. 2012, 495934. https://doi.org/10.1155/2012/495934 (2012).
Honda, M. et al. Telomere shortening and decreased replicative potential, contrasted by continued proliferation of telomerase-positive CD8+CD28(lo) T cells in patients with systemic lupus erythematosus. Clin. Immunol. 99, 211–221. https://doi.org/10.1006/clim.2001.5023 (2001).
Kurosaka, D. et al. Telomerase activity and telomere length of peripheral blood mononuclear cells in SLE patients. Lupus 12, 591–599. https://doi.org/10.1191/0961203303lu426oa (2003).
Fritsch, R. D. et al. Abnormal differentiation of memory T cells in systemic lupus erythematosus. Arthr. Rheum. 54, 2184–2197. https://doi.org/10.1002/art.21943 (2006).
Wu, C. H., Hsieh, S. C., Li, K. J., Lu, M. C. & Yu, C. L. Premature telomere shortening in polymorphonuclear neutrophils from patients with systemic lupus erythematosus is related to the lupus disease activity. Lupus 16, 265–272. https://doi.org/10.1177/0961203307077155 (2007).
Rudolph, M. et al. AKT1 (E17K) mutation profiling in breast cancer: prevalence, concurrent oncogenic alterations, and blood-based detection. BMC Cancer 16, 622. https://doi.org/10.1186/s12885-016-2626-1 (2016).
Bell, R. J. et al. Cancer. The transcription factor GABP selectively binds and activates the mutant TERT promoter in cancer. Science 348, 1036–1039. https://doi.org/10.1126/science.aab0015 (2015).
Akincilar, S. C. et al. Long-Range Chromatin Interactions Drive Mutant TERT Promoter Activation. Cancer Discov. 6, 1276–1291. https://doi.org/10.1158/2159-8290.CD-16-0177 (2016).
Barthel, F. P. et al. Systematic analysis of telomere length and somatic alterations in 31 cancer types. Nat. Genet. 49, 349–357. https://doi.org/10.1038/ng.3781 (2017).
Hsu, C. P., Hsu, N. Y., Lee, L. W. & Ko, J. L. Ets2 binding site single nucleotide polymorphism at the hTERT gene promoter–effect on telomerase expression and telomere length maintenance in non-small cell lung cancer. Eur. J. Cancer 42, 1466–1474. https://doi.org/10.1016/j.ejca.2006.02.014 (2006).
Rachakonda, P. S. et al. TERT promoter mutations in bladder cancer affect patient survival and disease recurrence through modification by a common polymorphism. Proc. Natl. Acad. Sci. USA 110, 17426–17431. https://doi.org/10.1073/pnas.1310522110 (2013).
Hosen, I. et al. Mutations in TERT promoter and FGFR3 and telomere length in bladder cancer. Int. J. Cancer 137, 1621–1629. https://doi.org/10.1002/ijc.29526 (2015).
Batista, R. et al. The prognostic impact of TERT promoter mutations in glioblastomas is modified by the rs2853669 single nucleotide polymorphism. Int. J. Cancer 139, 414–423. https://doi.org/10.1002/ijc.30057 (2016).
Simon, M. et al. TERT promoter mutations: a novel independent prognostic factor in primary glioblastomas. Neuro Oncol. 17, 45–52. https://doi.org/10.1093/neuonc/nou158 (2015).
Hafezi, F. & Perez Bercoff, D. The Solo Play of TERT Promoter Mutations. Cells https://doi.org/10.3390/cells9030749 (2020).
Amoura, Z. Protocole National de Diagnostic et de Soins Lupus Systémique (2020).
Fanouriakis, A. et al. 2019 update of the EULAR recommendations for the management of systemic lupus erythematosus. Ann. Rheum. Dis. 78, 736–745. https://doi.org/10.1136/annrheumdis-2019-215089 (2019).
Buyon, J. P. et al. The effect of combined estrogen and progesterone hormone replacement therapy on disease activity in systemic lupus erythematosus: a randomized trial. Ann. Intern. Med. 142, 953–962. https://doi.org/10.7326/0003-4819-142-12_part_1-200506210-00004 (2005).
Petri, M. et al. Combined oral contraceptives in women with systemic lupus erythematosus. N. Engl. J. Med. 353, 2550–2558. https://doi.org/10.1056/NEJMoa051135 (2005).
Hung, T. et al. The Ro60 autoantigen binds endogenous retroelements and regulates inflammatory gene expression. Science 350, 455–459. https://doi.org/10.1126/science.aac7442 (2015).
Pfaffl, M. W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 29, e45 (2001).
Nik-Zainal, S. et al. Mutational processes molding the genomes of 21 breast cancers. Cell 149, 979–993. https://doi.org/10.1016/j.cell.2012.04.024 (2012).
Suspene, R. et al. Somatic hypermutation of human mitochondrial and nuclear DNA by APOBEC3 cytidine deaminases, a pathway for DNA catabolism. Proc. Natl. Acad. Sci. USA 108, 4858–4863. https://doi.org/10.1073/pnas.1009687108 (2011).
Georgin-Lavialle, S. et al. The telomere/telomerase system in autoimmune and systemic immune-mediated diseases. Autoimmun. Rev. 9, 646–651. https://doi.org/10.1016/j.autrev.2010.04.004 (2010).
Hochberg, M. C. Updating the American College of Rheumatology revised criteria for the classification of systemic lupus erythematosus. Arthritis Rheum. 40, 1725. https://doi.org/10.1002/art.1780400928 (1997).
Bombardier, C., Gladman, D. D., Urowitz, M. B., Caron, D. & Chang, C. H. Derivation of the SLEDAI. A disease activity index for lupus patients. The Committee on Prognosis Studies in SLE. Arthritis Rheum. 35, 630–640. https://doi.org/10.1002/art.1780350606 (1992).
Weening, J. J. et al. The classification of glomerulonephritis in systemic lupus erythematosus revisited. Kidney Int. 65, 521–530. https://doi.org/10.1111/j.1523-1755.2004.00443.x (2004).
Lebon, P., Ponsot, G., Aicardi, J., Goutieres, F. & Arthuis, M. Early intrathecal synthesis of interferon in herpes encephalitis. Biomedicine 31, 267–271 (1979).
Liu, X. et al. Highly prevalent TERT promoter mutations in aggressive thyroid cancers. Endocr. Relat. Cancer 20, 603–610. https://doi.org/10.1530/ERC-13-0210 (2013).
Liu, T. et al. The activating TERT promoter mutation C228T is recurrent in subsets of adrenal tumors. Endocr. Relat. Cancer 21, 427–434. https://doi.org/10.1530/ERC-14-0016 (2014).
Hindson, B. J. et al. High-throughput droplet digital PCR system for absolute quantitation of DNA copy number. Anal. Chem. 83, 8604–8610. https://doi.org/10.1021/ac202028g (2011).
Funding
This work was supported by Fonds National de la Recherche du Luxembourg (INTER/Mobility/16/1154722/APO-SLE), by Ministère de l’Education et de la Recherche du Luxembourg and by Institut Pasteur Paris. FH was supported by FNR-PRIDE scheme (PRIDE/11012546/NEXTIMMUNE).
Author information
Authors and Affiliations
Contributions
D.P.-B., J.P.V., H.L. and S.W.H. designed the study and drafted the manuscript. D.P.-B. and H.L. analyzed A3 expression and editing in SLE patient samples. A.M. and Z.A. followed and selected patients, provided samples and participated in data analysis. M.L., F.H. and D.P.-B. performed S.L.E. cell cultures. V.T. and V.C. screened A3AΔ3B by ddPCR; H.B., M.V. and O.H. performed, curated and analzyed N.G.S. sequences; A.P. analyzed SLE RNASeq data and RS participated in sample treatment and RNA exctractions. All authors listed made a substantial, direct and intellectual contribution to the work, and approved it for publication.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Perez-Bercoff, D., Laude, H., Lemaire, M. et al. Sustained high expression of multiple APOBEC3 cytidine deaminases in systemic lupus erythematosus. Sci Rep 11, 7893 (2021). https://doi.org/10.1038/s41598-021-87024-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-87024-1
This article is cited by
-
Learning single-cell perturbation responses using neural optimal transport
Nature Methods (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.