Abstract
We investigated the ability of time-warping-based ECG-derived markers of T-wave morphology changes in time (\(d_{w}\)) and amplitude (\(d_a\)), as well as their non-linear components (\({d_w^{{\mathrm{NL}}}}\) and \({d_a^{\mathrm{NL}}}\)), and the heart rate corrected counterpart (\(d_{w,c}\)), to monitor potassium concentration (\([K^{+}]\)) changes (\(\Delta [K^+]\)) in end-stage renal disease (ESRD) patients undergoing hemodialysis (HD). We compared the performance of the proposed time-warping markers, together with other previously proposed \([K^{+}]\) markers, such as T-wave width (\(T_w\)) and T-wave slope-to-amplitude ratio (\(T_{S/A}\)), when computed from standard ECG leads as well as from principal component analysis (PCA)-based leads. 48-hour ECG recordings and a set of hourly-collected blood samples from 29 ESRD-HD patients were acquired. Values of \(d_w\), \(d_a\), \({d_w^{\mathrm{NL}}}\), \({d_a^{\mathrm{NL}}}\) and \(d_{w,c}\) were calculated by comparing the morphology of the mean warped T-waves (MWTWs) derived at each hour along the HD with that from a reference MWTW, measured at the end of the HD. From the same MWTWs \(T_w\) and \(T_{S/A}\) were also extracted. Similarly, \(\Delta [K^+]\) was calculated as the difference between the \([K^{+}]\) values at each hour and the \([K^{+}]\) reference level at the end of the HD session. We found that \(d_{w}\) and \(d_{w,c}\) showed higher correlation coefficients with \(\Delta [K^+]\) than \(T_{S/A}\)—Spearman’s (\(\rho\)) and Pearson’s (r)—and \(T_w\)—Spearman’s (\(\rho\))—in both SL and PCA approaches being the intra-patient median \(\rho \ge 0.82\) and \(r \ge 0.87\) in SL and \(\rho \ge 0.82\) and \(r \ge 0.89\) in PCA respectively. Our findings would point at \(d_{w}\) and \(d_{w,c}\) as the most suitable surrogate of \(\Delta [K^+]\), suggesting that they could be potentially useful for non-invasive monitoring of ESRD-HD patients in hospital, as well as in ambulatory settings. Therefore, the tracking of T-wave morphology variations by means of time-warping analysis could improve continuous and remote \([K^{+}]\) monitoring of ESRD-HD patients and flagging risk of \([K^{+}]\)-related cardiovascular events.
Similar content being viewed by others
Introduction
Chronic kidney disease (CKD) is defined as the presence of kidney damage, persisting for 3 months or more, irrespective of the cause1. It represents a state of progressive loss of kidney function ultimately resulting in need for renal replacement therapy such as hemodialysis (HD) or transplantation. The development of CKD and its progression to this terminal stage, called end-stage renal disease (ESRD), remains a significant source of reduced quality of life and premature mortality2. In particular, sudden cardiac death (SCD) represents an important cause of death in ESRD-HD patients3. Various risk factors may be responsible for SCD in this patient population, including left ventricular hypertrophy and fibrosis, disordered bone-mineral metabolism, HD-induced changes in electrolyte, and fluid and acid-base status, which may lead to electrocardiographic (ECG) abnormalities and ventricular arrhythmia3,4.
Recent studies have shown that blood potassium concentrations (\([K^{+}]\)) outside the physiological interval are associated with increased mortality risk5. In healthy conditions, the maintenance of \([K^{+}]\) homeostasis is ensured by normal renal activity6. However, ESRD-HD patients suffer from \([K^{+}]\) imbalance, leading to a high incidence of arrhythmic events. The pro-arrhythmic consequences of \([K^{+}]\) imbalance can be explained considering that potassium currents are involved in the repolarization process of the cardiac action potential (AP), determining membrane potential and refractoriness of the myocardium7. Therefore, even modest deviations of \([K^{+}]\) from its normal range (hypokalemia if \([K^{+}]\) < 3.5 mmol/L or hyperkalemia if \([K^{+}]\) > 5 mmol/L) may lead to hospitalisation or death in ESRD-HD patients8. Evaluation of \([K^{+}]\) levels is currently based on blood samples that require further analyses in the laboratory, limiting continuous monitoring. Non-invasive markers able to track variations in \([K^{+}]\) levels are therefore needed.
The electrocardiogram (ECG) is a non-invasive, easily accessible, and inexpensive practice that reflects the electrical activity of the heart. In particular, the T-wave reflects the spatio-temporal repolarization of the ventricle, and its analysis has been used to measure the vulnerability of a patient to ventricular arrhythmias9. This fact is of particular interest because T waves are frequently altered in ESRD-HD patients4. The QT interval is the standard index of ventricular repolarization, and it has been proposed to monitor ESRD-HD patients10. However, the effects of HD on QT interval, and its corrected version QTc, are still controversial, since several studies11 reported a prolongation during the HD sessions, but others reported opposite trend or even no changes at all12. This motivates the analysis of the overall T-wave morphology as a potential potassium level marker.
Different T-wave morphology markers have been previously reported to be correlated with \([K^{+}]\), such as the T-wave right slope13, the width of the T-wave (\(T_{w}\))14, the T-wave slope-to-amplitude ratio (\(T_{S/A}\))15, and a morphology combination score, which integrates features like T-wave asymmetry, flatness and notching16. However, these markers rely on specific local features of the T-wave rather than in the overall T-wave morphology, which may have a stronger potential in following \(\Delta [K^+]\) than indices based on local features.
A recent study reported a time-warping based methodology to quantify changes in the overall T-wave morphology17. Six indices were proposed, \(d_{w}^{u}\) and \(d_a\), reflecting morphological variations in time and amplitude, respectively, as well as their non-linear version, \(d_w^{\mathrm{NL}}\) and \(d_a^{\mathrm{NL}}\) as reported in17 and two novel markers derived from \(d_{w}^{u}\) and named \(d_{w}\) and \({d}_{w,c}\). The main goal of this study is to investigate the potential of these markers in monitoring both hypo- and hyperkalemia events excluding the variability due to the heart rate (HR) and to compare their performance against \(T_{w}\) and \(T_{S/A}\) in standard single-lead approach and by applying principal component analysis (PCA) as multilead space reduction technique. However, some of the above mentioned indices may not be robust enough for our purpose. It is the case of \(d_w\) which does not provide information about the direction of the T-wave morphological variation (i.e. if there is stretching or shortening) and has been found to be correlated with HR. Therefore, we have adapted the original methodology17 to account for hypo- and hyperkalemia, and we propose a new marker that is independent of HR, thus offering a more precise \([K^{+}]\) monitoring tool for arrhythmic risk stratification in ESRD-HD patients. Preliminary results extracted from a smaller subset of patients have been presented at Computing in Cardiology conference18,19 while the electrophysiological basis was studied in Bukhari et al.20.
The novelties of the present study with respect to the state-of-the-art are: (1) the usage of T-wave time warping analysis for non-invasive \([K^{+}]\) monitoring, together with the development of a HR correction tool for the time-warping marker, \({d}_{w,c}\); (2) the proposal of a PCA spatial transformation lead for marker extraction and (3) the validation of the proposed markers in comparison with previously published biomarkers (\(T_{w}\) and \(T_{S/A}\)) and with their extraction from standard leads.
Materials and methods
Study population
The study population included 29 patients from the Nephrology ward from Hospital Clínico Universitario Lozano Blesa (Zaragoza, Spain). Inclusion criteria were (i) 18-year-old (or older), (ii) having a diagnosed ESRD pathology and (iii) undergoing HD at least three times per week (with venous or cannula access). Table 1 shows the population characteristics. The study protocol was approved by the Aragon’s research ethics committee (CEICA, ref. PI18/003) and all patients and/or their legal guardians signed informed consent. All the procedures and all the methods were performed in accordance with the Helsinki Declaration. The database collection is still ongoing, with the current size significant enough for a pilot study21,22.
Analysis stages performed in this study. In panel (a) is the flow chart showing the ECG processing steps for T-wave time-warping markers extraction. The analysis starts with the original ECG, followed by a filtering step before spatial PCA analysis, to conclude with markers computation. Panel (b) shows an example of the linear and nonlinear time-warping markers for the same patient as in Fig. 5a. In particular, subpanel (i) shows both the reference (blue) and the i-th MWTW (red) while subpanel (ii) shows the warping function (red dotted line) that optimally relates the reference and studied MWTWs. Subpanel (iii) shows the MWTWs after warping and subpanel (iv) are the normalized reference and warped MWTWs.
Data collection
General information
Sex, age, concomitant therapies (e.g. assumption of anti-arrhythmic drugs), kidney disease etiology and HD treatment related information were collected for each enrolled patient, as detailed in Table 1.
Blood sample analysis
For each patient, six blood samples were taken and analysed during the HD session: the first one at the HD onset and the next three, every subsequent hour (Fig. 1, \(h_0\) to \(h_3\) in red). The 5-th blood sample was collected at the end of the HD (minute 215-th or 245-th, depending on the HD session duration) while the 6-th blood sample was taken after 48 h, immediately before the next HD session. Potassium, magnesium, calcium, urea, creatinine, bicarbonate and pH were measured from each blood test. Blood potassium concentrations values for each blood test are given in Table 2.
ECG measurements
A 48h, standard 12-lead ECG Holter recording, (H12+, Mortara Instruments, Milwaukee, WI, USA, sampling frequency of 1 kHz, amplitude resolution of 3.75 \(\upmu\)V), was obtained for each enrolled patient, starting the acquisition 5 min before the HD onset (Fig. 1, blue line). The block diagram presented in Fig. 2a describes the main steps of the whole data processing implemented in this work.
ECG pre-processing
ECG filtering
Holter ECG signals contain baseline drift and other noises, such as power-line and muscular activity (Fig. 2a). Therefore, an initial pre-processing is needed to improve the signal-to-noise ratio (SNR) and enable ECG waveform analysis. First, baseline wander was removed with a high-pass, forward-backward 6-th order Butterworth filter with 0.5 Hz cut-off frequency23 (Fig. 2a). Then, residual noise out of the T-wave band was removed with a 6-th order low-pass Butterworth filter with 40 Hz cut-off frequency.
ECG waveform detection and delineation
A wavelet-based single-lead method24 was applied to detect QRS complexes and then delineate T-wave onsets and ends in each of the 12 leads. The wavelet transform (WT) decomposes the signal in the time-scale domain, allowing its representation at different resolutions. It is, therefore, a suitable tool to analyze ECG signals, which contain patterns with different frequency content (QRS complexes, P and T-waves).
Single-lead delineation
The discrete dyadic WT is implemented in such a way that it keeps temporal resolution at different scales. The detection of the fiducial points is carried out across the adequate WT scales, attending to the dominant frequency components of each ECG wave: Q,R,S waves correspond to a simultaneous effect in scales \(2^1\)–\(2^2\), while the T and P waves affect mainly scales \(2^4\) or \(2^5\), see24 for details. ECG wave peaks correspond to zero crossings in the WT, and ECG maximum slopes correspond to WT’s maxima and minima. Depending on the number and polarity of the slopes found, a wave morphology is assigned and boundaries are located using threshold-based criteria. The onset (end) of a wave occurs before (after) the first (last) significant slope associated with the wave24.
Selection rules for multi-lead delineation
To obtain multilead peak locations, a median post-processing selection rule over the single-lead-based detected locations is used. The post-processing rules for boundaries consist of ordering the single-lead annotations and selecting as the onset (end) of a wave the first (last) annotation whose k nearest neighbours lay within a \(\delta\) ms interval24,25.
Single-lead analysis
First, we performed the analysis using the single-lead ECG, taking the T-waves from leads V3 to V6, as used in a previous study26 for \([K^+]\) estimation, and lead II being the most widely used in patient monitoring27. These T waves were further delineated by using the above motioned delineator24 and the biomarker estimation is performed as described below in section named “Time warping analysis”.
Spatial lead reduction by principal component analysis
Next, a spatial lead reduction by Principal Component Analysis (PCA) was made since it was found to be a robust spatial transformation to emphasize waveform SNR28. In this work PCA was spatially applied to the 8 independent leads, learned over the T-wave segment to mainly emphasize this waveform, and resulting in 8 principal components (PCs) or transformed leads. The coefficients defining the PCA transformation were obtained from the eigenvectors of the \(8 \times 8\) interlead auto-correlation matrix computed over the T-waves in a 10-min wide window at the end of the HD session. The correct delineation of T-waves is crucial to emphasize only T-wave energy content. The first PCA, denoted as PC1, was used for the subsequent ECG analysis, as it is the transformed lead where the T-waves have maximal energy, and thus, maximal SNR for morphological characterisation28,29. PC1 was further delineated by applying again24, and each T-wave was further low-pass filtered at 20 Hz using a 12-th order Butterworth filter to restrict shape analysis to the dominant band of the T-wave so removing remaining noisy components that could still corrupt the T-wave shape analysis.
Time-warping analysis
Two-minute ECG segments, centred on the 5-th and 35-th minutes of each available hour, were analysed. The window duration was short enough to hold the assumption of stability for both \([K^+]\) and HR values. Figure 5a shows the average RR interval for each selected i-th 2-min segments for a given patient along the ECG recording. While the blood samples (purple diamonds) were collected each hour during the HD, the warping parameters were computed every half an hour to get a more detailed view over time.
For each i-th 2-min segment, a mean warped T-wave (MWTW) was computed. First of all, the predominant T-wave polarity (e.g. upward, downward etc) within a given window, was defined as that having the highest number of occurrences. This polarity change can be physiological or induced by delineator oscillation when by-phasic to regular T-waves appears almost indistinguishable. A T-wave was considered to have inverted polarity if the magnitude of its peak had negative sign and vice-versa. Only those T-waves having the same polarity as the predominant one were considered in the following steps. Then, all these selected T-waves were aligned with respect to their gravity center and used to compute an initial MWTW17. Finally, all the T-waves were checked to find and discard the outliers, defined as having T-wave duration outside the range \(T_{dm_i} \pm 1.5\times \sigma _{d_i}\) centered at the i-th ensemble T-wave mean, \(T_{dm_i}\), and bounded function of the T-wave duration standard deviation \(\sigma _{d_i}\). Among the remaining, only those T-waves highly correlated (Pearson’s correlation coefficient > 0.98) with the previous initial MWTW were used to recalculate the final MWTW. The MWTW at the end of the HD treatment was taken as the reference, given that it is the time when the patient (a) is supposed to have recovered the normal \([K^+]\) level and (b) was discharged from hospital, being an appropriate reference for out-of-hospital ambulatory monitoring.
Since hyperkalemia has been reported to cause T-wave inversions30, any MWTWs with negative-polarity was inverted before performing the warping with the reference MWTW. Previous to warping, the two MWTWs were aligned with respect to their gravity center, so that only changes in the T-wave morphology, and not those associated with their relative delay, were quantified by the warping algorithm.
For comparison purposes, both \(T_w\)14 and \(T_{S/A}\)15 were extracted from each MWTW and their performance, with respect to T-wave time-warping based biomarkers in monitoring \([K^+]\), was assessed. This work perform a clinical study following previous analysis testing the marker by electrophysiological simulations as reported in Bukhari et al.20.
T-wave time warping
The method here applied was originally proposed by Ramírez et al.17. Let \({{\mathbf {f}}^i(\mathbf {t}^i)=[f^i(t^i(1)),\ldots ,f^i(t^i(N_i))]^T}\) be the MWTW of a given i-th segment, and \({\mathbf {f}^r(\mathbf {t}^r)=[f^r(t^r(1),\ldots ,f^r(t^r(N_r))]^T}\) the reference MWTW, where \({{\mathbf {t}}^i =[t^i(1),\ldots ,t^i(N_i)]^T}\) and \({{\mathbf {t}}^r =[t^r(1),\ldots ,t^r(N_r)]^T}\) with \(N_i\) and \(N_r\) being the total T-wave duration, in samples, of \({{\mathbf {t}}}^i\) and \({\mathbf {t}}^r\) respectively. Figure 2b illustrates the warping method applied between one of the i-th MWTW (red) and the reference MWTW (blue). Let \(\gamma _{i}({\mathbf {t}}^r)\) be the warping function that relates \({\mathbf {t}}^r\) and \({\mathbf {t}}^i\), such that the composition \(({\mathbf {f}}^i\circ \gamma _{i})({\mathbf {t}}^r)\) denotes the re-parametrization or time domain warping of \(\mathbf {f}^i({\mathbf {t}}^i)\) using \(\gamma _{i}({\mathbf {t}}^r)\), i.e. \((\mathbf {f}^i\circ \gamma _i)({\mathbf {t}}^r)\) represents the amplitude values of \({\mathbf {f}}^i({\mathbf {t}}^i)\) if its temporal vector was \({\mathbf {t}}^r\). The square-root slope function (SRSF) was proposed instead of the original T-waves31,32 to find the optimal warping function. This was applied by performing time-warping on the SRSFs of the T-waves, preventing the “pinching effect” in cases when T-wave amplitudes differ33. This transformation is defined as:
The optimal warping function is the one that minimizes the amplitude difference between the SRSF of \({\mathbf {f}}^r({\mathbf {t}}^r)\) and \({\mathbf {f}}^i(\gamma _i({\mathbf {t}}^r))\)32:
The dynamic programming algorithm was used to obtain the solution of this optimisation problem34. Figure 2b(ii) shows the optimal warping function between the two waves in Fig. 2b(i). The warped T-wave, \({\mathbf {f}}^i(\gamma ^*_{i}({\mathbf {t}}^r))\) is shown in Fig. 2b(iii), together with the reference T-wave, \({\mathbf {f}}^r({\mathbf {t}}^r)\).
Time warping biomarkers
The index \(d_{w}^{u}\) (corresponding to the index denoted as \(d_{w}\) in17), shown as the yellow area in Fig. 2b(ii), quantifies the amount of warping needed to optimally align the two T-waves, and is defined as the average of the absolute difference value between \(\gamma _i^*({\mathbf {t}}^r)\) and \({\mathbf {t}}^r\):
The original definition of \(d_{w}^{u}(i)\)17 was modified here to allow the marker to be signed, therefore distinguishing T-wave widenings from narrowings. This signed \(d_{w}(i)\) was defined as:
where \(s_d(i)\) was used to account for the sign of the \(d_{w}(i)\) and it was computed as:
with \(N_{r}^{u}\) being the set of T-wave up-slope samples. A positive sign means that the \({\mathbf {f}}^i({\mathbf {t}}^i)\) has to be widened to fit the \({\mathbf {f}}^r({\mathbf {t}}^r)\) and vice-versa for a negative sign.
After applying time warping between both MWTWs, the amplitude difference between \({\mathbf {f}}^r({\mathbf {t}}^r)\) and \(\mathbf {f}^i(\gamma ^*_{i}({\mathbf {t}}^r))\) is quantified as the area contained between \({\mathbf {f}}^r({\mathbf {t}}^r)\) and \(\mathbf {f}^i(\gamma ^*_{i}({\mathbf {t}}^r))\), normalized by the L2-norm of \({\mathbf {f}}^r({\mathbf {t}}^r)\):
where \(s_a (i)=\sum _{n=1}^{N_{r}} ( f^i(\gamma _i^*(t^r))- f^r(t^r))\) is used to account for the \(d_{a}(i)\) sign estimation.
Both \(d_{w}(i)\) and \(d_{a}(i)\) incorporate information from the linear and non-linear differences between both T-waves in time and amplitude domain, respectively. The non-linear components can be quantified as in17:
where \(\gamma ^*_{i,l}({\mathbf {t}}^r)\) (green line in Fig. 2b(ii)) is the best linear fitting to \(\gamma _i^*({\mathbf {t}}^r)\) according to the least absolute residual criterion35. The parameter \(d_w^{\mathrm{NL}}(i)\) quantifies the non-linear warping by computing the area of the dashed magenta region between \(\gamma ^{*}\left( {\mathbf {t}}^r\right)\) and \(\gamma ^{*}_{i,l}\left( {\mathbf {t}}^r\right)\) (in Fig. 2b(ii)). Finally, the marker \(d_{a}^{\mathrm{NL}} (i)\) quantifies the residual information in amplitude domain after normalising MWTWs (Fig. 2b(iv)).
Heart-rate-corrected T-wave warping
It is well known that T-wave duration and QT interval are strongly dependent on HR36. Although aligning the T-waves according to their gravity centre reduces most of the dependence of \(d_w(i)\) on HR, there may still be some residual dependence in T-wave morphology that should be compensated for (e.g. see Fig. 5a around hours h = 9, 12 and 43). We assume that \(d_w(i)\), as originally proposed in (4), can be modelled as the sum of two components:
where \(d_{w,HR}(i)\) is the HR dependent component and \(d_{w,c}(i)\) is the non-HR dependent component accounting for (\(K^{+}\)) induced variations and possibly others not HR related.
To estimate the corrected component \(d_{w,c}(i)\) we depart from the literature, where several formulae for HR-dependency correction of repolarization related time intervals, like the QT interval, have been developed37,38,39,40, including a variety of approaches (e.g. linear, hyperbolic, exponential models etc.) being investigated and tested in view of the complex relationship between QT interval and HR39. To derive a correction formula and estimate \(d_{w,c}(i)\), we started from a linear approximation of a hyperbolic model under small RR changes, derived similarly to the QT interval correction (QTc)38,39,
Let’s call \(RR_{r}\) the reference RR interval associated to a reference heart beat and \(RR_{i}\) the one to the i-th RR interval from one beat at the i-th segment, then
As the \(QT_{i}-QT_{r}\) difference, also \(d_{w} (i)\) is a measure of width change between the reference r and the current i-th mean T-waves from their respective observations time windows, then it is possible to extend previous relation in (11) to \(d_{w}(i)\) obtaining the HR related component
By substituting (12) in (9) we obtain
The value \(d_{w,c}(i)\) can be assumed to be non-zero mean, and uncorrelated to HR, that is:
with \(\Delta d_{w,c}(i)\) zero mean and uncorrelated to HR. Then, \(d_{w}(i)\) becomes:
where the parameters b, \(\beta\) and \(\alpha\), once jointly estimated (i.e. \({\hat{b}}\), \({\hat{\beta }}\) and \({\hat{\alpha }}\)) can be used to derive \({\hat{d}}_{w,c}(i)\) as:
Note that, \({\hat{\beta }}\) and \({\hat{\alpha }}\) cannot be assessed from (15) with a directly least square fitting, since the DC component b in (15) largely affects the results. Rather, it is possible to jointly estimate \({\hat{b}}, {\hat{\beta }}\) and \({\hat{\alpha }}\), and then use the results in (16).
This estimate can be further approximated linearly for small RR changes. Denoting \(\Delta RR(i)=RR_{i} - RR_{r}\), \(RR_{i}\) can be expressed as \(RR_{i}= RR_{r}+ \Delta RR(i)\) and by replacing this in the right side of (12):
Operating on the terms and under the assumption that \(\Delta RR(i) \ll RR_{r}\) , \((\frac{ \Delta RR(i)}{RR_{r}}) \ll 1\) and by using the Taylor’s series expansion, we have
where b, \(\alpha\), \(\beta\) and \((RR_{r})^{(\alpha -1)}\) are constant values; then placing:
the actual \(d_{w}(i)\) dependency with RR will be:
From the geometrical point of view, b and c can be estimated as the zero-crossing and the slope, respectively, of the least-squares line fit to the \(d_{w}(i)\) values (in a \(\Delta RR(i)\) vs. \(d_{w}(i)\) graph). Then, the \(d_{w}(i)\) component that does not dependent on the RR, meaning it is assumed not correlated, can assessed as:
where \({\hat{c}}\) is the estimated slope from the Holter recording, see Fig. 3, \({\hat{d}}_{w,c}(i)\) is then the corrected estimated of \(d_{w,c}(i)\), with \(RR_{i}\) and \(RR_{r}\) the mean RR interval from the i-th studied segment and the reference windows respectively and \({\hat{c}}\) parameter is estimated for every patient during the time course of the Holter recording. When the linear approximation presented above cannot be assumed, \({\hat{b}}\), \({\hat{\beta }}\), and \({\hat{\alpha }}\) can be jointly estimated from the model in (15), and use the (16) as the corrected estimate.
An example of the estimated \({\hat{d}}_{w,c}(i)\) is given in Fig. 5a where both \({\hat{d}}_{w,c}(i)\) and \(d_w(i)\) where displayed. Notice how the proposed correction formula removed the HR-dependency, for example around h = 12.
Table 3 provides an overview of the morphology markers studied in this work.
Potassium concentration variations \(\Delta [K^+]\)
The proposed biomarkers have been compared with the relative variations in \([K^+]\) (denoted as \(\Delta [K^+](h)\)) with respect to a reference \([K^+]\) that was taken at the end of the HD:
being \([K^{+}]_{h}\) the concentration at the h-th hour during the HD and \([K^{+}]_{r}\) the concentration at the end of the treatment. An example of the \(\Delta [K^+](h)\) evolution is shown in Fig. 5a (purple diamonds).
Statistical analyses
Results are presented as median and interquartile range (IQR). Spearman rank correlation coefficient (\(\rho\)) and Pearson correlation (r) were used for correlation analysis between \(\Delta [K^+]\) and the proposed biomarker, giving information about both the monotonic relation and the strength of the association between the time warping based biomarkers and \([K^+]\) changes and then providing a more complete characterisation. The average duration of the ECG recordings was 44 h mainly due to electrode detachment or early battery exhaustion. For this reason, correlation coefficients were computed using the first five values of \(\Delta [K^{+}](h)\) throughout the HD and the warping markers evaluated at the corresponding i-th segment points (\(h=(i-1)/2\) where i=1, 3, 5, 7, 9 or i=1, 3, 5, 7, 8 depending on the HD duration). All statistical analyses were performed using MATLAB version R2018b.
Results
In this study, ECG signals and \([K^+]\) from 29 ESRD-HD patients were investigated. An example of \(d_{w}\) and \({\hat{d}}_{w,c}\) time evolution for a particular patient, in PCA approach, was provided in Fig. 3. \(\Delta RR\) was represented on the x-axis in both panels, while \(d_{w}\) and \({\hat{d}}_{w,c}\) were shown on the y-axis in panel (a) and panel (b), respectively. The least-square fitting line (red line) was depicted in both panels. Spearman’s correlation coefficients (\(\rho\)) and p-values were also showed in each panel. High and significant correlation (\(\rho =-\,0.90\) and p-value \(<0.001\)) was found between \(\Delta RR\) and \(d_{w}\). However, after correcting for the HR-dependency, \(\rho =0.03\) and p-value \(=0.76\).
Correlation between \([K^+]\) and mean HR expressed in beats per minute (bpm) have also been computed and the results are presented in Table 2, with a Spearman’s correlation coefficient median (IQR) values of 0.10 (1.35), and a median p-value of p=0.33. These values were 0.09 (1.45), p = 0.22 for Pearson’s correlation coefficient.
Table 4 shows the intra-patient Spearman’s (\(\rho\)) and Pearson’s (r) correlation coefficients computed between the relative variations in \([K^+]\) (denoted as \(\Delta [K^+]\)) with respect to a reference \([K^+]\) that was taken at the end of the HD and the time-warping parameters. In both single-lead and PCA approaches, the highest median Spearman’s and Pearson’s correlation coefficients were found for \(d_{w}^{u}\), \(d_{w}\) and \({d}_{w,c}\) being \(\rho \ge 0.82\) and \(r \ge 0.86\) for single-lead analysis and \(\rho \ge 0.82\) and \(r \ge 0.89\) in PCA.
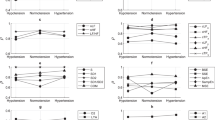
Boxplots in Fig. 4 show the distributions of \(\Delta [K^{+}]\) and the proposed PCA-based time-warping descriptors during HD. Figure 5b shows the average time evolution of PCA-based \(d_{w}^{u}\), \(d_w\), \({\hat{d}}_{w,c}\) and \(d_{w}^{\mathrm{NL}}\) in the studied population along the monitoring period, while the evolution of \(d_a\) and \(d_{a}^{\mathrm{NL}}\) is shown Fig. 5c.
Scatterplot showing the values of both \(d_w\) panel (a) and \({\hat{d}}_{w,c}\) panel (b) with respect to \(\Delta RR\) for a given patient in PCA approach. Spearman’s correlation coefficients (\(\rho\)) and p-values for both \(d_w\) and \({\hat{d}}_{w,c}\) are shown on top of each panel, while the least-square fitting regression lines are plotted in red.
Boxplots showing the distribution of \(\Delta [K^{+}]\) (blue) and all the described PCA-based time-warping biomarkers (\(d_{w}^{u}\), \(d_a\), \(d_w^{\mathrm{NL}}\), \(d_a^{\mathrm{NL}}\), \(d_{w}\), and \({\hat{d}}_{w,c}\)) (red), computed at different time points from the beginning, \(h_0\), to the end, \(h_4\), of the HD session (\(h_0\) to \(h_4\) in Fig. 1). \(\Delta [K^{+}]\) was computed as in (23). + denotes outliers.
PCA-based time-warping markers and RR interval time trends. An example for a given patient undergoing 4h-long HD therapy, is depicted in panel (a) with the evolution of \(d_{w}\) (filled green squares), \({\hat{d}}_{w,c}\) (filled orange squares), both referring to the left vertical scale, and the average RR intervals (unfilled dark red squares, referring to the right vertical scale). \(\Delta [K^{+}]\) relative variations with respect to the concentration at HD end (purple diamonds) are expressed in mmol/L. Time is expressed in hours from the beginning of the treatment onward. Each square denotes the mean RR interval in a 2-min wide segment used to compute the warping parameters, while the highlighted blue square corresponds to the reference segment at the end of HD. The filled red square denotes the time-point from which the studied MWTW in Fig. 2b was selected. Note that for this patient, the Holter recording did not reach the planned 48h. Panel (b) shows the median and IQR for each observing i-th segment, computed by using the values from all the available patients for \(d_{w}^{u}\), \(d_w\) and \({\hat{d}}_{w,c}\) and \(d_{w}^{\mathrm{NL}}\) (this latter refers to the right axis, while the others to the left). Time trends (expressed as median and IQR) for \(d_{a}\) and \(d_{a}^{\mathrm{NL}}\), referring to the left and right axes respectively are in panel (c). Panels (b) and (c) give an overview of the time evolution for these biomarkers along the ECG acquisition from the HD beginning onward. Only the first 44 h were depicted being that the average ECG duration in our database.
Discussion
Repolarization abnormalities play a fundamental role in the genesis of arrhythmic events and the risk increases in patients at ESRD with imbalance in \([K^{+}]\)41. In this work, two previously reported potassium estimators, \(T_{w}\)14 and \(T_{S/A}\)15, four warping-based ECG-derived biomarkers for \([K^{+}]\) monitoring proposed in Ramírez et al.17, \(d_{w}^{u}\), \(d_a\), \(d_w^{\mathrm{NL}}\), \(d_a^{\mathrm{NL}}\), and the here proposed modified versions \(d_{w}\) and \({d}_{w,c}\) were tested as bloodless indices for \([K^{+}]\) variations in ESRD-HD patients computed from standard leads as well as in a PCA-derived lead. The most promising results in terms of correlation were obtained for markers \(d_{w}^{u}\), \(d_{w}\), and \({d}_{w,c}\), leading to the highest median intra-patient \(\rho \ge 0.82\) and \(r \ge 0.87\) in single-lead and \(\rho \ge 0.82\) and \(r \ge 0.89\) in PCA lead respectively, evidencing high monotonic and linear association with \([K^{+}]\) and making them a promising non-invasive indices for blood \([K^{+}]\) monitoring.
The signed biomarker \(d_{w}\) followed a similar time-course as the unsigned \(d_{w}^{u}\) during the whole monitoring period, showing a similar distribution in Fig. 5a, as a result of the fact that the sign computed as in (5) is positive in roughly all the patients. That can be explained by the fact that the T-wave morphology in hyperkalemia is usually more peaked and shorter in time than a T-wave from regular \([K^{+}]\) concentrations, as happens at the end of HD, where the reference has been taken42,43. Therefore, all the other MWTWs needed to be shrunk in amplitude and widened in time duration during the warping procedure to fit the reference one, and this is given by a positive signed \(d_{w}\). However, other external factors, such as the potassium removal rates44 or the dialysate potassium level45,46, might also have played a role in altering ventricular repolarization activity.
The warping algorithm is applied over the MWTWs computed from different observing windows with different HRs, as is evident in Fig. 5a. Therefore, a corrected version of the \(d_{w}\), derived similarly to the QT correction formula38,39, was proposed since the HR influences this marker as pointed out in Ramírez et al.17, and can be observed in Fig. 5a as an example around hours h\(=\)9, 12 and 43. A large number of models have been proposed for the computation of QTc values independent of HR37,38,39,40. However, a previous study38 found that the linear regression model fits better than any other model to the relationship between QT and the RR intervals. Also, for small RR variations, in section “Heart-rate-corrected T-wave warping” it is shown that hyperbolic QT to RR dependency becomes linear. Therefore, we used a linear model to derive an HR-corrected index, \({d}_{w,c}\). This approach was used to estimate the \(d_{w}\) component strictly related to \([K^{+}]\) removing its relation with HR as showed in Fig. 3, where the HR-dependency, clearly visible in panel (a), was cancelled after the correction, panel (b). Comparing the results for \(d_{w}^{u}\), \(d_{w}\) and \({\hat{d}}_{w,c}\), all of them have proved to be highly correlated with \([K^{+}]\) variations. However, it is important to remember that \(d_{w}^{u}\) (and so \(d_{w}\)) is biased by the HR effects as previously described17, while \({\hat{d}}_{w,c}\) is no longer dependent on it, possibly being responsible for the lower IQR in the correlation, 0.25, as compared to 0.35 and 0.36 for \(d_{w}\) and \(d_{w}^{u}\), respectively (see Table 4, PCA column). It should also be noted that the small differences between the \(\rho\) and r computed for \({\hat{d}}_{w,c}\) and \(d_w\) could be due to the low HR variations observed during HD, but larger HR variations, and consequently a higher impact of the correction, are expected in ambulatory monitoring.
In addition, we investigated the correlation between HR and \([K^{+}]\) finding no significant correlation between the two. To place these results into a proper context, in a previous study on computer-based models it was found that the heart rate in ESRD-HD patients is influenced by the combined effects of \([K^{+}]\)), calcium and pH47. In particular, it was observed that when \([K^{+}]\) is between 3 and 4 mmol/L, the HR sensitivity is about 10 bpm/mmol of \([K^{+}]\)47. As can be seen in Table 2, the median \([K^{+}]\) falls within the above mentioned range from \(h_1\) to \(h_4\) (i.e., almost during the entire HD session) but the range of HR variations is much bigger than the 10 bpm mentioned in Severi et al.47 (minimum IQR = 17 bpm at \(h_3\)) so anticipating the obtained low and insignificant median correlations coefficients between HR and \([K^+]\). This result suggests that HR variations are a poor indicator of \(\Delta [K^+]\), at least in our dataset. Moreover, this finding gives validity to the proposal of the \({\hat{d}}_{w,c}\) presented in this work, which assumes no correlation between HR and \([K^{+}]\)-related changes on T-wave morphology.
In a previous study20 a comparison of \(T_w\), \(T_{S/A}\), \(d_{w}\), \(d_w^{\mathrm{NL}}\), \(d_a\) and \(d_a^{\mathrm{NL}}\), based on a electrophysiological simulation of ECG under hyperkalemia, was performed, and tested on a subset of the ESRD-HD patients described in Materials and Methods section. In that study it is shown that similar results were obtained in terms of Pearson’s correlation coefficient between \([K^{+}]\) and \(T_w\), \(T_{S/A}\), and time-warping based markers, also showing a high correlation with HR. In the present work we extended the analysis by increasing the sample size, introducing the proposed correction by HR, and comparing with single lead recordings. According to the Spearman’s (\(\rho\)) and Pearson’s (r) correlation coefficients, the median values are higher, and the IQR smaller, for \(d_{w}^{u}\), \(d_{w}\) and \({d}_{w,c}\) than for \(T_{S/A}\) using both PCA and standard single-lead approaches. For \(T_w\), we found similar r median absolute values, as compared to \(d_{w}^{u}\),\(d_{w}\) and \({d}_{w,c}\), while Spearman’s correlation values were much lower for \(T_w\) when using PCA and several single-leads (e.g. lead V4 and V5). This shows that the proposed time-warping based markers present either higher correlation or stronger monotonic relationship with \(\Delta [K^+]\) than \(T_w\), and \(T_{S/A}\), making them more suitable for \([K^{+}]\) monitoring purposes.
When comparing single-lead and PCA-based analysis, and according to Spearman’s (\(\rho\)) and Pearson’s (r) correlation coefficients in Table 4, leads II, V3 and V5 are those showing the highest median and the smallest IQR values in a single-lead approach. Moreover, for \(d_a\) and \(d_a^{\mathrm{NL}}\), leads V3 and V5 show stronger \(\rho\) and r correlation with \(\Delta [K^+]\) than the PCA lead, with \(\rho \ge 0.45\) and \(r \ge 0.62\). However, median values for \(\rho\) and r computed between \(\Delta [K^+]\) and \(d_{w}^{u}\), \(d_{w}\) and \({d}_{w,c}\), are similar (\(\rho \ge 0.82\) and \(r \ge 0.89\) for the PCA lead and \(\rho \ge 0.82\) and \(r \ge 0.86\) for the standard single leads) showing no evident benefit in using one or the other strategy. Nevertheless, a clear advantage of extracting \(d_w^{\mathrm{NL}}\) from the PCA lead rather than from the standard single leads can be observed when comparing the medians both from \(\rho\) and r. In the same line, both \(T_w\) and \(T_{S/A}\) showed higher r computed from the PCA lead than from standard leads. This improvement when analysing biomarkers estimated from specific local features of the T-wave, as \(T_w\) and \(T_{S/A}\), which is not that evident when analyzing global T-wave waveform based biomarkers, like those proposed in this work, can be explained by the improved SNR obtained by using PCA, leading to a more robust definition of the T-wave delineation marks. This is in agreement with several studies reporting the benefits of PCA as an intermediate step when addressing problems related with noise reduction28. Therefore, the use of PCA from multilead ECG is recommended as a more robust alternative to single-lead analysis.
As already described, ECGs recorded under hyperkalemic conditions commonly present T-waves which are taller than those for normal levels of blood \([K^{+}]\). The marker \(d_a\) was designed to capture the amplitude variability between \({\mathbf {f}}^i({\mathbf {t}}^i)\) and \({\mathbf {f}}^r({\mathbf {t}}^r)\) after warping. However, from its original definition, \(d_a\) can take both positive and negative values, producing positive intra-patient \(\rho\) and r between \(\Delta [K^+]\) and \(d_a\) in \(62\%\) of the cases and negative in \(38\%\) of the cases. This explains the low median inter-patient correlation (\(\rho =0.21\)) and the wide IQR (1.20). However, if its absolute value was considered then correlation between \(\Delta [K^+]\) and \(d_a\) becomes larger being \(\rho =0.70\) (0.60) and \(r=0.75\) (0.31).
Cellular AP duration can be affected in a heterogeneous manner by \([K^{+}]\) fluctuations since ion channel expression is heterogeneous throughout the ventricles48 and this can result in non-linear changes on T-wave morphology. According to their definition17, \(d_{w}^{\mathrm{NL}}\) and \(d_{a}^{\mathrm{NL}}\) were designed to quantify the inhomogeneous morphological variations during ventricular repolarization; this fact might explain their correlation with \(\Delta [K^+]\). Both of them showed a remarkable sensitivity to the variations of ventricular spatio-temporal dispersion independently from changes in HR17, meaning that they did not need a HR correction as was done for \(d_{w}\). This last point, particularly for \(d_a^{\mathrm{NL}}\), in addition to its also high correlation with \(\Delta [K^+]\), make it an interesting T-wave descriptor for blood \([K^{+}]\) assessment.
Results of the present work, show high accuracy of the \(d_{w}\) \({d}_{w,c}\) T-wave based time-warping biomarkers for \([K^{+}]\) monitoring, in contrast with the findings in a recent study published by Regolisti et al.49 where lower correlation values were reported for other ECG-derived markers. The explanation for this discrepancy could be tracked back to the biomarkers the authors took as reference for their research, from Dillon et al.13 and Corsi et al.15, which are a) focused on specific features of the T-waves and b) measured in absolute value and not in relative terms to a reference which can personalize the biomarker as here presented. Nevertheless, time warping analysis was applied in recent studies19, 20 to investigate both hypo- and hyperkalemia on a extremely heterogeneous pool of simulated cases, performing equally well in all the cases and proving its adaptability, at least in silico simulated ECGs, probably due to its personalize profile as being comparative with a reference. Therefore, we hypothesized that the proposed methodology for \([K^{+}]\) monitoring could be applied to investigate a wide range of populations and experimental sets.
Other recent studies developed deep-learning models and tested them on large datasets to screen for hyperkalemia in patients with CKD, reporting a more robust recognition of severe hyperkalemia, thus potentially reducing the risk of sudden cardiac death in those patients50,51. However, in our study we aim, not just to detect hyperkalemia, but also in continuous \([K^+]\) quantification. Extension of the database will open the door to use deep learning techniques, which will allow proper comparison with the methodology here proposed.
The main limitation of this work concerns the reduced amount of blood tests (only six) took from each patient during the 48-h ECG recording, which provided an accurate representation of the \([K^{+}]\) time evolution only during HD, but not during the ambulatory period. On this basis, it was possible to investigate the relationship between the proposed markers and the \(\Delta [K^+]\) only during the therapy but not through the remaining hours. Moreover, the course of HD procedures can be accompanied by ischaemia52,53,54, which can also be associated with changes in the T-wave, independently from HR, therefore reflected on \(d_{w,c}\). Elucidation of its impact in monitoring normal-life outside the HD process needs to be explored in a dedicated study. Also postural or body position changes (BPC) are known to affect the T-wave morphology and mainly the T-wave amplitude55, which can influence \(d_a\) related biomarkers. The fact that those markers were measured on PCA lead could attenuate the effect of BPC on T-wave, however a specific study should precisely establish the impact of this aspect on the biomarker, particularly when applied in a non controlled scenario. Assessment of the performance of the proposed descriptors in patients with hypokalemia remains to be studied. In addition, the methodology proposed in this work needs to be validated in: (a) a larger population; and (b) subsequent HD sessions on the same population, which would allow to quantify the accuracy in \([K^{+}]\) estimation.
Beyond the previous considerations, \(d_{w}\), \({\hat{d}}_{w,c}\) and \(d_{a}^{\mathrm{NL}}\) showed potential value in monitoring changes in blood potassium concentration. All of them showed a time-course similar to that of \([K^{+}]\) in ESRD-HD patients as described in literature45,46: a rapid decline during the HD with a fast rebound just after the end of the therapy, followed by a steady increase in the remaining hours before the next HD session (see Fig. 5b,c). Therefore, it is possible to consider the biomarkers use for continuous \([K^{+}]\) monitoring, also in other pathologies as in patients suffering heart failure56, where hyperkalemia also increases the risk for SCD.
As a future study, we propose to replicate the same analysis over a larger population and assess the correlation between the proposed time-warping markers and alterations in other electrolytes beyond \([K^{+}]\), like calcium, magnesium, or related to the rate and amount of \([K^{+}]\) removal44,45,46. These variations could alter the ECG and it would be interesting to analyse their relationship with the induced T-wave changes. Then, as mentioned above, there is a need to test the proposed indices and investigate their robustness against these possible confounders. Other alternatives for lead space reduction, such as Periodic Component Analysis57, can be explored to check whether the T-wave shape is better emphasized with a criterion different from just the energy maximisation considered by PCA. A study to test the extrapolation power of the current findings, which refer to \([K^+]\) levels at \(h_4\), to subsequent HD sessions should also be performed to assess their clinical long-term validity.
Conclusions
In this study, time-warping markers (\(d_{w}\), \(d_a\)), their non linear components (\(d_{w}^{\mathrm{NL}}\) and \(d_{a}^{\mathrm{NL}}\)) and a HR-corrected version \(d_{w,c}\), in all cases personalised making it relative to reference point, were studied in ESRD patients undergoing HD and evaluated as estimator of \([K^{+}]\) changes over time. Among the analyzed biomarkers and methods (i.e. standard single-lead or PCA transformed lead approaches), the proposed PCA-based \(d_{w}\) and its HR corrected version, \({d}_{w,c}\), achieved better results than the previously proposed \(T_{S/A}\) in terms of both Spearman’s and Pearson’s correlation with \(\Delta [K^+]\), and showed higher monotonic relationship than \(T_w\), thus making the proposed time-warping markers a valid and more accurate alternative to the currently available tools for \([K^{+}]\) ambulatory monitoring of ESRD-HD patients. In conclusion, this study proposes new markers for robust quantitative ambulatory monitoring of \([K^{+}]\) from the ECG. The proposed markers could improve routine \([K^{+}]\) monitoring without the need of invasive blood test, both in hospital and in an ambulatory scenario. A comprehensive validation is needed to corroborate the extrapolation power of these results.
References
KDIGO. Definition and classification of CKD. Kidney Int. Suppl. 3, 16–92. https://doi.org/10.1038/kisup.2012.64 (2013).
Grassmann, A., Gioberge, S., Moeller, S. & Brown, G. ESRD patients in 2004: Global overview of patient numbers, treatment modalities and associated trends. Nephrol. Dial. Transplant. 20, 2587–2593. https://doi.org/10.1093/ndt/gfi159 (2005).
Makar, M. S. & Pun, P. H. Sudden cardiac death among hemodialysis patients. Am. J. Kidney Dis. 69, 684–695. https://doi.org/10.1053/j.ajkd.2016.12.006 (2017).
Secemsky, E. A. et al. High prevalence of cardiac autonomic dysfunction and T-wave alternans in dialysis patients. Heart Rhythm 8, 592–598. https://doi.org/10.1016/j.hrthm.2010.11.041 (2010).
Pitt, B. & Rossignol, P. The association between serum potassium and mortality in patients with hypertension: A wake-up call. Eur. Heart J. 38, 113–115. https://doi.org/10.1093/eurheartj/ehw209 (2016).
Palmer, B. F. Regulation of potassium homeostasis. Clin. J. Am. Soc. Nephrol. 10, 1050–1060. https://doi.org/10.2215/CJN.08580813 (2015).
Ravens, U. & Cerbai, E. Role of potassium currents in cardiac arrhythmias. EP Europace 10, 1133–1137. https://doi.org/10.1093/europace/eun193 (2008).
Jain, N. et al. Predictors of hyperkalemia and death in patients with cardiac and renal disease. Am. J. Cardiol. 109, 1510–1513. https://doi.org/10.1016/j.amjcard.2012.01.367 (2012).
Antzelevitch, C. Role of spatial dispersion of repolarization in inherited and acquired sudden cardiac death syndromes. Am. J. Physiol. Heart Circ. Physiol. 293, H2024–H2038. https://doi.org/10.1152/ajpheart.00355.2007 (2007).
Algra, A., Tijssen, J. G., Roelandt, J. R., Pool, J. & Lubsen, J. QTc prolongation measured by standard 12-lead electrocardiography is an independent risk factor for sudden death due to cardiac arrest. Circulation 83, 1888–1894. https://doi.org/10.1161/01.CIR.83.6.1888 (1991).
Covic, A. et al. Haemodialysis increases QTc interval but not QTc dispersion in ESRD patients without manifest cardiac disease. Nephrol. Dial. Transplant. 17, 2170–2177. https://doi.org/10.1093/ndt/17.12.2170 (2002).
Severi, S. et al. Cardiac response to hemodialysis with different cardiovascular tolerance: Heart rate variability and QT interval analysis. Hemodial. Int. 10, 287–293. https://doi.org/10.1111/j.1542-4758.2006.00110.x (2006).
Dillon, J. J. et al. Noninvasive potassium determination using a mathematically processed ECG: Proof of concept for novel “blood-less, blood test. J. Electrocardiol. 48, 12–18. https://doi.org/10.1016/j.jelectrocard.2014.10.002 (2016).
Frohnert, P. P., Gluliani, E. R., Friedberg, M., Johnson, W. J. & Tauxe, W. N. Statistical investigation of correlations between serum potassium levels and electrocardiographic findings in patients on intermittent hemodialysis therapy. Circulation 41, 667–676. https://doi.org/10.1161/01.CIR.41.4.667 (1970).
Corsi, C. et al. Noninvasive quantification of blood potassium concentration from ECG in hemodialysis patients. Sci. Rep. 7, 42492. https://doi.org/10.1038/srep42492 (2017).
Krogager, M. L. et al. The relationship between serum potassium concentrations and electrocardiographic characteristics in 163,547 individuals from primary care. J. Electrocardiol. 57, 104–111. https://doi.org/10.1016/j.jelectrocard.2019.09.005 (2019).
Ramírez, J., Orini, M., Tucker, J. D., Pueyo, E. & Laguna, P. Variability of ventricular repolarization dispersion quantified by time-warping the morphology of the T-wave. IEEE Trans. Biomed. Eng. 64, 1619–1630. https://doi.org/10.1109/TBME.2016.2614899 (2017).
Palmieri, F. et al. T-wave morphology changes as surrogate for blood potassium concentration in hemodialysis patients. 2019 Comput. Cardiol. https://doi.org/10.23919/CinC49843.2019.9005904 (2019).
Bukhari, H. A. et al. Transmural ventricular heterogeneities play a major role in determining T-wave morphology at different extracellular potassium levels. 2019 Comput. Cardiol. https://doi.org/10.23919/CinC49843.2019.9005944 (2019).
Bukhari, H. A. et al. Characterization of T-wave amplitude, duration and morphology changes during hemodialysis: Relationship with serum electrolyte levels and heart rate. IEEE Trans. Biomed. Eng. https://doi.org/10.1109/TBME.2020.3043844 (2020).
Browne, R. H. On the use of a pilot study for sample size determination. Stat. Med. 14, 1933–1940. https://doi.org/10.1002/sim.4780141709 (1995).
Kieser, M. & Wassmer, G. On the use of the upper confidence limit for the variance from a pilot sample for sample size determination. Biometr. J. 38, 941–949. https://doi.org/10.1002/bimj.4710380806 (1996).
Sörnmo, L. & Laguna, P. Electrocardiogram (ECG) signal processing. In Wiley Encyclopedia of Biomedical Engineering (ed. Akay, M.) (Wiley, Hoboken, 2006).
Martínez, J. P., Almeida, R., Olmos, S., Rocha, A. P. & Laguna, P. A wavelet-based ECG delineator: Evaluation on standard databases. IEEE Trans. Biomed. Eng. 51, 570–581. https://doi.org/10.1109/TBME.2003.821031 (2004).
Laguna, P., Jané, R. & Caminal, P. Automatic detection of wave boundaries in multilead ECG signals: Validation with the CSE database. Comput. Biomed. Res. 27, 45–60. https://doi.org/10.1006/cbmr.1994.1006 (1994).
Attia, Z. I. et al. Novel bloodless potassium determination using a signal-processed single-lead ECG. J. Am. Heart. Assoc. 5, e002746. https://doi.org/10.1161/JAHA.115.002746 (2016).
Yoon, D. et al. Quantitative evaluation of the relationship between t-wave-based features and serum potassium level in real-world clinical practice. BioMed. Res. Int. https://doi.org/10.1155/2018/3054316 (2018).
Castells, F., Laguna, P., Sörnmo, L., Bollmann, A. & Millet Roig, J. Principal component analysis in ECG signal processing. EURASIP J. Adv. Signal Proc. 2007, 074580. https://doi.org/10.1155/2007/74580 (2007).
Ramírez, J., Mincholé, A., Laguna, P. & Pueyo, E. Characterization of cardiac repolarization response to heart rate changes provoked by a tilt test. 2012 Comput. Cardiol. 39, 673–679 (2012).
Montford, J. R. & Linas, S. How dangerous is hyperkalemia? JASN 28, 3155–3165. https://doi.org/10.1681/ASN.2016121344 (2017).
Srivastava, A., Wu, W., Kurtek, S., Klassen, E. & Marron, J. S. Registration of functional data using fisher-rao metric. Preprint at arXiv:1103.3817 (2011).
Tucker, J. D., Srivastava, A. & Wu, W. Generative models for functional data using phase and amplitude separation. Comput. Stat. Data Anal. 61, 50–66. https://doi.org/10.1016/j.csda.2012.12.001 (2013).
Ramsay, J. O. & Li, X. Curvature registration. J. R. Stat. Soc. Ser. B Stat. Methodol. 61, 351–363. https://doi.org/10.1111/1467-9868.00129 (1998).
Bertsekas, D. P. Dynamic Programming and Optimal Control (Athena Scientific, Belmont, 2005).
Holland, P. W. & Welsch, R. E. Robust regression using iteratively reweighted least-squares. Commun. Stat. Theory Methods 6, 813–827. https://doi.org/10.1080/03610927708827533 (1977).
di Bernardo, D. & Murray, A. T wave shape changes with heart rate: A computer model analysis. 2000 Comput. Cardiol. 27, 151–154. https://doi.org/10.1109/CIC.2000.898478 (2000).
Hnatkova, K. & Malik, M. Optimum formulae for heart rate correction of the QT interval. Pacing Clin. Electrophysiol. 22, 1683–1687. https://doi.org/10.1111/j.1540-8159.1999.tb00390.x (1999).
Pueyo, E. et al. Characterization QT interval adaptation to RR interval changes and its use as a risk-stratifier of arrhytmic mortality in amiodarone-treated survivors of acute myocardial infarction. IEEE Trans. Biomed. Eng. 51, 1511–1520. https://doi.org/10.1109/TBME.2004.828050 (2004).
Luo, S., Michler, K., Johnston, P. & Macfarlane, P. W. A comparison of commonly used QT correction formulae: The effect of heart rate on the QTc of normal ECGs. J. Electrocardiol. 37, 81–90. https://doi.org/10.1016/j.jelectrocard.2004.08.030 (2004).
Vandenberk, B. et al. Which QT correction formulae to use for QT monitoring? J. Am. Heart. Assoc. 5, e003264. https://doi.org/10.1161/JAHA.116.003264 (2016).
Ajam, F. et al. Cardiac arrhythmias in patients with end stage renal disease (ESRD) on hemodialysis: Recent update and brief literature review. Am. J. Internal Med. 7, 22–26. https://doi.org/10.11648/j.ajim.20190701.16 (2019).
Webster, A., Brady, W. & Morris, F. Recognising signs of danger: ECG changes resulting from an abnormal serum potassium concentration. Case Rep. 17, 74–77. https://doi.org/10.1136/emj.19.1.74 (2002).
Levis, J. T. ECG diagnosis: Hyperkalemia. Perm J. 17, 69. https://doi.org/10.7812/TPP/12-088 (2013).
Severi, S., Vecchietti, S., Cavalcanti, S., Mancini, E. & Santoro, A. Electrocardiographic changes during hemodiafiltration with different potassium removal rates. Blood Purif. 21, 381–388. https://doi.org/10.1159/000073440 (2003).
Blumberg, A., Roser, H. W., Zehnder, C. & Müller-Brand, J. Plasma potassium in patients with terminal renal failure during and after haemodialysis; relationship with dialytic potassium removal and total body potassium. Nephrol. Dial. Transplant. 12, 1629–1634. https://doi.org/10.1093/ndt/12.8.1629 (1997).
Pun, P. H. & Middleton, J. P. Dialysate potassium, dialysate magnesium, and hemodialysis risk. J. Am. Soc. Nephrol. 28, 3441–3451. https://doi.org/10.1681/ASN.2017060640 (2017).
Severi, S. & Cavalcanti, S. Electrolyte and pH dependence of heart rate during hemodialysis: A computer model analysis. Artif. Organs 24, 245–260. https://doi.org/10.1046/j.1525-1594.2000.06480.x (2000).
Poulikakos, D. et al. Sudden cardiac death in dialysis: Arrhythmic mechanisms and the value of non-invasive electrophysiology. Front. Physiol. https://doi.org/10.3389/fphys.2019.00144 (2019).
Regolisti, G. et al. Electrocardiographic T wave alterations and prediction of hyperkalemia in patients with acute kidney injury. Intern. Emerg. Med. 15, 463–472. https://doi.org/10.1007/s11739-019-02217-x (2020).
Galloway, C. D. et al. Development and validation of a deep-learning model to screen for hyperkalemia from the electrocardiogram. JAMA Cardiol. 4, 428–436. https://doi.org/10.1001/jamacardio.2019.0640 (2019).
Lin, C. S. et al. A deep-learning algorithm (ECG12Net) for detecting hypokalemia and hyperkalemia by electrocardiography: Algorithm development. JMIR Med. Inform. 8, e15931. https://doi.org/10.2196/15931 (2020).
Kremastinos, D. et al. Electrocardiographic T wave alterations and prediction of hyperkalemia in patients with acute kidney injury. Nephron 60, 164–170. https://doi.org/10.1159/000186733 (1992).
Kremastinos, D., Jha, V., Bali, H. K., Sakhuja, V. & Sapru, R. P. Cardiac arrhythmias and silent myocardial ischemia during hemodialysis. Renal Fail. 22, 355–368. https://doi.org/10.1081/JDI-100100879 (2000).
Flythe, J. E., Kimmel, S. E. & Brunelli, S. M. Rapid fluid removal during dialysis is associated with cardiovascular morbidity and mortality. Kidney Int. 79, 250–257. https://doi.org/10.1038/ki.2010.383 (2011).
García, J. E., Astrom, M., Mendive, J., Laguna, P. & Sörnmo, L. ECG-based detection of body position changes in ischemia monitoring. IEEE Trans. Biomed. Eng. 50, 677–685. https://doi.org/10.1109/TBME.2003.812208 (2003).
Crespo-Leiro, M. G. et al. Hyperkalemia in heart failure patients in spain and its impact on guidelines and recommendations: ESC-EORP-HFA heart failure long-term registry. Rev. Esp. Cardiol. Engl. Ed. 73, 313–323. https://doi.org/10.1016/j.rec.2019.05.015 (2020).
Monasterio, V., Clifford, G. D., Laguna, P. & Martínez, J. P. A multilead scheme based on periodic component analysis for T-wave alternans analysis in the ECG. Ann. Biomed. Eng. 38, 2532–2541. https://doi.org/10.1007/s10439-010-0029-z (2010).
Acknowledgements
This work was funded by Products & Technology S.L. (Castellbisbal, Barcelona, Spain), and by AGAUR, Generalitat de Catalunya (Spain) grant DI001-2018 and CIBER-BBN ref: “DEKOALE”. The work was also supported by project PID2019-104881RB-I00, and PID2019-105674RB-I00 funded by Spanish Ministry of Science and Innovation (MICINN) and FEDER, by Gobierno de Aragón (project LMP124-18 and Reference Group Biomedical Signal Interpretation and Computational Simulation (BSICoS) T39-20R) co-funded by FEDER 2014-2020 “Building Europe from Aragón”, and by European Research Council (ERC) through project ERC-StG 638284. The computation was performed at the High Performance computing platform of the NANBIOSIS ICTS. J. Ramírez would like to thank the support from the Marie Sklodowska-Curie Grant No. 786833. A. Martín-Yebra is a “Juan de la Cierva” fellow (project FJC2018-037369-I).
Author information
Authors and Affiliations
Contributions
Each of the authors gave substantial contributions to the study design, and data interpretation and discussion. In particular F.P. developed the algorithms, analysed the data and drafted the manuscript; F.P., P.G., A.M.-Y., H.A.B., E.P., J.P.M., J.R. and P.L. structured and discussed the methodology; J.E.R. and B.B. collected the data; D.F., J.E.R., and P.L. conceived the study. All authors critically reviewed the article for important intellectual content and gave final approval of the submitted version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Palmieri, F., Gomis, P., Ferreira, D. et al. Monitoring blood potassium concentration in hemodialysis patients by quantifying T-wave morphology dynamics. Sci Rep 11, 3883 (2021). https://doi.org/10.1038/s41598-021-82935-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-82935-5
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.