Abstract
Post-translational modifications of histone proteins greatly impact gene expression and cell fate decisions in eukaryotes. To study these, it is important to develop a convenient, multiplex, and efficient method to precisely introduce mutations to histones. Because eukaryotic cells usually contain multiple copies of histone genes, it is a challenge to mutate all histones at the same time by the traditional homologous recombination method. Here, we developed a CRISPR-Cas9 based shuffle system in Saccharomyces cerevisiae, to generate point mutations on both endogenous histone H3 and H4 genes in a rapid, seamless and multiplex fashion. Using this method, we generated yeast strains containing histone triple H3–K4R–K36R–K79R mutants and histone combinatorial H3–K56Q–H4–K59A double mutants with high efficiencies (70–80%). This CRISPR-Cas9 based mutagenesis system could be an invaluable tool to the epigenetics field.
Similar content being viewed by others
Introduction
In the nucleus of eukaryotic cells, nucleosomes are basic units of chromatin and play essential roles in DNA-templated processes, including packing, transcription, replication, recombination, and repair, together with organization and maintenance of chromatin structure. A nucleosome is composed of about 147 base pairs of DNA wrapped around a histone octamer, which is consist of two copies of core histones H2A, H2B, H3, and H41,2,3. Core histones can be covalently post-translational modified by methylation, phosphorylation, acetylation, ubiquitylation, and sumoylation. These modifications impact gene expression by either directly altering nucleosome structure or recruiting histone modifiers1,4,5,6. Because there are multiple copies of histone genes in higher eukaryotic organisms, i.e. 100 copies of histone genes in Drosophila melanogaster and 64 copies of histone genes in human genome7,8, it is a challenge to generate histone site-directed mutagenesis to study histone modifications and related enzymes.
The budding yeast Saccharomyces cerevisiae has been served as a model organism to explore histone functions, because of several advantages (i) its highly-conserved histone sequences and modification functions with higher eukaryotes, (ii) its only two copies of histone genes in haploid cell stage, (iii) its ability to survive and grow with only one copy of histone genes, and (iv) its availability of powerful genetic manipulation tools. Several strategies have been carried out for systematic histone mutation libraries in S. cerevisiae: (1) transforming an episomal plasmid containing histone mutants with both endogenous copies deleted, (2) mutating one copy of endogenous histones by homologous recombination (HR) with the other copy deleted, (3) mutating both endogenous copies of histones by HR using two different selection markers9,10,11. These strategies have been very helpful to study histone functions and modifications, however additional requirements have emerged for new tools. Firstly, histone dosage effects have been discovered for some point mutations, thus previous identified crucial histone mutants by deleting one or two copies of endogenous histone may have different phenotypes with both endogenous copies mutated11,12. Secondly, multiple histone modifications have the same effect on gene expression or chromatin structure, and methods are needed for generating combinational mutagenesis on histones simultaneously. Last but not least, seamless mutagenesis is preferred for histone mutagenesis to reveal endogenous functions.
The recently developed CRISPR technology has greatly accelerated the speed of genome engineering13,14,15,16. CRISPR-Cas9 has also been implemented in histone mutagenesis in in a wide range of organisms, including human cell lines, a protozoan parasite Trypanosoma brucei, and Drosophila melanogaster7,17,18,19,20. However, there are several limitations for the current CRIPSR-Cas9 system: (i) CRISPR-Cas9 mutagenesis requires an efficient guide RNA (gRNA) targeting sequence and a following “NGG” as Protospacer Adjacent Motif (PAM) which are closed to the mutation sites; however, some targeted mutation sites do not meet such conditions. (ii) The current CRISPR-Cas9 systems performed single histone site mutation and are not capable to generate combinatorial histone mutations. In this paper, we developed a CRISPR-Cas9 based shuffle system for endogenous histone H3 and H4 combinatorial mutagenesis in Saccharomyces cerevisiae. Using this system, seamless point mutations on histone H3 and H4 in different combinations can be generated simultaneously with high efficiencies.
Results
Generating CRISPR-Cas9 based shuffle strains in Saccharomyces cerevisiae
In S. cerevisiae, the core histone genes are organized in H2A–H2B and H3–H4 gene pairs and driven by bidirectional promoters. HHT1–HHF1 and HHT2–HHF2 encode histone H3–H4 pairs on chromosome II and XIV respectively. Although histone H3 and H4 genes have corresponding identical amino acid sequences, they have 3% (8/309) and 5% (22/409) respectively differences in DNA coding and dramatically different upstream and downstream expression regulating elements. We designed a histone shuffle system for endogenous histone H3 and H4 combinatorial mutagenesis (Fig. 1). Firstly, a shuffle master strain was constructed, and endogenous H3 and H4 coding regions were replaced by synthetic H3 and H4 genes (synH3 and synH4) by the CRISPR-Cas9 system. Then, synH3 and synH4 genes were targeted by CRISPR-Cas9 and repaired by PCR fragments amplified from endogenous H3 and H4 sequences with desired mutations. To minimize the chance of homology-directed repair at undesired sites, the synH3 and synH4 share same protein sequences but different codons with endogenous H3 and H4 (see sequence alignments in Fig. 2B,C).
Graphic representation of CRISPR-Cas9 based shuffle system for endogenous histone H3 and H4 mutagenesis. As first step, synthetic H3 and H4 genes (synH3 and synH4) are shuffled into genome by CRISPR-Cas9 to replace endogenous H3 and H4. Then the synthetic H3 and H4 genes are shuffled-out by CRISPR-Cas9 and repaired by donors with endogenous H3 and H4 genes with desired site-directed mutations.
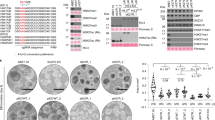
Generation of master shuffle strains with synthetic H3 and H4 by CRISPR-Cas9. (A) Generation of a master shuffle strain with synthetic H3 and H4 genes (synH3H4) through 2 rounds of CRISPR-Cas9 mediated gene replacements. The efficiencies of 2 rounds are 3/4 (75%) and 4/4 (100%) respectively, which are tested by four randomly picked colonies respectively. (B,C) The nucleotide sequence alignment of endogenous and synthetic H3 and H4. The synthetic H3 and H4 (synH3 and synH4) share same protein sequences but different DNA codons with native H3 and H4 genes. The nucleotides with grey background indicate the identical nucleotide sequences. Dark blue arrows represent gRNA sequences targeting endogenous H3 (HHT1/2); brown arrows represent gRNA sequences targeting endogenous H4 (HHF1/2); magenta arrows represent gRNA sequences targeting synH3 genes, and light blue arrows represent gRNA sequences targeting synH4 genes. The gRNAs targeting synthetic histones are used for generating Histone H3 and H4 mutants in Figs. 3 and 5.
The histone shuffle master strains with synthetic H3 and H4 (synH3H4) were generated by two steps of CRISPR-Cas9 mediated gene replacement (Fig. 2A). For the first step, endogenous H3 (HHT1/2) genes are replaced by the synH3 gene. Cas9 plasmids containing 2 gRNAs targeting two loci of H3 genes and synthetic H3 PCR donors with 40–50 bp homologous recombination (HR) arms to the upstream and downstream of both HHT1 and HHT2 genes were electroporated to yeast CEN.PK 113-5D strain. 4 colonies have been randomly selected for verification, and 75% of colonies show correct gene replacement at both endogenous loci (Fig. 2A). For the second step, endogenous H4 (HHF1/2) genes are similarly replaced by the synH4 gene with 100% efficiency (4/4 colonies) (Fig. 2A). In order to eliminate the possibility that the donor DNA fragments would be integrated into other chromosomal positions with random insertion, we performed whole genome sequencing of synH3H4 strain. And the results showed synH3 and synH4 were integrated at desired loci without any detection of other integration loci (Supplementary data).
Generation of histone H3 single and triple mutant through single transformation
Histone H3 K4, K36, and K79 are the major methylation sites highly associated active gene transcription, and these modifications are conserved from yeast to human. To determine whether our system can be applied for histone single mutation, we sought out to generate histone H3 K4R mutant first. The repairing donors were prepared in two methods: two DNA PCR fragments for repairing and replacing each synH3 gene with K4R mutation sites on the flanking HR arms (total four pieces for hht1 K4R and hht2 K4R) (Fig. 3A), and two single-piece donors containing mutation sites generated by fusion PCR (Fig. 3B). The donors and plasmids containing Cas9 and 2 gRNAs targeting synH3 were transformed into the synH3 strain. The correct gene replacement efficiencies of two strategies are at 88% and 71% with no significant difference (student T-test, p = 0.1161) (Fig. 3C, left). The two strategies for donor preparation were also tested for generating histone H3 K4R–K36R–K79R triple mutant. The efficiency of 79% was achieved by transforming two single-piece donors containing triple mutants, while the replacement efficiency of 0% was achieved by using 8-fragment donors with 40–50 bp homologues repairing arms (Fig. 3C, right). The low efficiency of the “fragment donors” strategy may be due to the unsuccessful homology direct repair (HDR) of the multiple-fragment donors.
Generation of histone H3 single and triple mutants through a single transformation. (A) Graphic representation of CRISPR-Cas9 based mutagenesis on histone H3. The transformation was performed on synH3 strain with 2 targeting gRNAs and donors harboring mutation sites and HR regions. The single piece donors were generated through fusion PCR. (B) The mutagenesis efficiencies for generating histone H3 single and triple mutant through transformation with different formats of donors. The repairing donors can be prepared by either multiple donor fragments with flanking HR arms for mutation or two single pieces of donors containing mutation sites. The data of bar charts represent mean averages of over-all mutagenesis efficiencies of 3 biological replicates with each randomly picked 8 colonies. The error bars indicate the standard deviations of 3 biological replicates. (C) The representation of sequencing results of both Histone H3 K4R–K36R–K79R at endogenous loci. The sequence alignments were performed by SnapGene software.
To determine whether histone H3 triple mutant sensitive to methyl methanesulfonate (MMS) or hydroxyurea (HU) mediated DNA damage. Serial tenfold dilutions of OD600 = 1 were spotted onto YPD plate, YPD plate with 0.05% MMS, or YPD plate with 100 mM HU. And the results show synH3H4 strains are not sensitive to 0.05% MMS or 100 mM HU, while hht1/2 K4R–K36R–K79R mutants are sensitive to 0.05% MMS and 100 mM HU (Fig. 4). These results demonstrated that the codon differences do not affect the shuffle master synH3H4 strain sensitive to DNA damage, while the hht1/2 K4R–K36R–K79R mutants are sensitive as previously reported21.
Confirmation of MMS and HU sensitivity of synH3H4 strain and histone H3 K4R–K36R, K79R mutant. WT stain and two biological replicates of each mutant were grown in YPD overnight at 30 °C. Serial tenfold dilutions of OD600 = 1 yeast cultures were spotted onto YPD for 2 days, YPD with 0.05% MMS for 3 days, or YPD with 100 mM HU and incubated for 3 days at 30 °C.
Mutagenesis on both histone H3 and H4 through single transformation
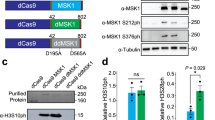
We have demonstrated that the CRISPR-Cas9 based histone shuffle system is useful for histone H3 mutagenesis with high efficiencies. Next, we sought out to determine whether this system can be applied for mutagenesis on both histone H3 and H4 with different mutation combinations (Fig. 5A,B). Histone H3 K56Q mutant and histone H4 K59A mutant have been previously shown growth defects on medium containing 200 mM HU compared to wild-type strains22. We implemented our system for generating H3 K56Q and H4 K59A on both copies of histone genes or by combination of mutants and wild-type genes (Fig. 5A,B). We first transformed single-piece donors with Cas9 plasmid containing 2 gRNAs (One gRNA targeting synH3 and the other targeting synH4). To our surprise, there was no successful gene replacement colony (Fig. 5C, left). This result suggested that either DNA cleavaged by Cas9 or HDR may not be efficient. To improve the chance of Cas9 cleavage, the Cas9 plasmid containing 4 gRNAs (with each histone synthetic gene targeted by 2 gRNAs, and gRNA sequences showed in Fig. 2B,C) was constructed and transformed with the same donors. With the help of additional gRNAs, the correct mutant efficiencies were improved to 71% for hht1/2 K56Q, hhf1/2 K59A mutant and 75% for hht1 K56Q/HHT2, HHF1/hhf2 K59A single-copy mutant. The HU sensitivity assay showed the mutations of hht1/2 K56Q, hhf1/2 K59A were sensitive to 100 mM HU, while the single-copy mutants hht1 K56Q/ HHT2, HHF1/ hhf2 K59A were less sensitive to 100 mM HU than double-copy mutants (Fig. 5D). These results demonstrated CRISPR-Cas9 based histone shuffle system is versatile for mutagenesis of histone H3 and H4 mutations with different combinations.
Mutagenesis on both histone H3 and H4 through single transformation. (A,B) Graphic representation of CRISPR-Cas9 based gene replacement for mutagenesis on both histone H3 and H4 with different combinations. By transforming different donors, the endogenous two copies of histone H3 and H4 can be repaired for identical mutants (A) or with different mutation sites (B). (C) The mutagenesis efficiencies for generating histone H3 and H4 mutants through single transformation with different gRNAs. 2 gRNAs: 1 gRNA targeting synH3 and 1 gRNA targeting synH4 and 4 gRNAs: 2 gRNAs for each synthetic histone. The data of bar charts represent mean averages of over-all mutagenesis efficiencies of 3 biological replicates with each randomly picked 8 colonies. The error bars indicate the standard deviations of 3 biological replicates. (D) Confirmation of HU sensitivity of histone H3 and H4 mutant. Two biological replicates were grown in YPD overnight at 30 °C. Serial tenfold dilutions were spotted onto YPD or YPD with 100 mM HU and incubated at 30 °C for 3 days.
Discussions
In this study, we developed a convenient, seamless, and efficient CRISPR-Cas9 based shuffle system to precisely introduce multiple mutations to histones H3 and H4 in Saccharomyces cerevisiae. We demonstrated this method by generating yeast histone triple H3–K4R–K36R–K79R mutants and histone H3–K56Q–H4–K59A double mutants with high efficiencies (70–80%). This system is also capable to mutate two copies of histones with any combinatorial mutations.
Although CRISPR-Cas9 has been applied to generate protein point mutations, it still needs to consider the limitation of gRNA closed to the desired mutation sites. Our method provides an alternative two-step method: firstly, replacing the original gene with a synthetic copy (codons changed) by CRISPR/Cas9, then targeting the synthetic copy using CRISPR/Cas9 and repair back to the endogenous gene with desired mutations.
We also found that increasing gRNA numbers to target a multiple-copy gene can improve gene replacement efficiencies. One possible explanation is that a single double strain break (DSB) site would be easily repaired by the other uncleavaged gene copies, while multiple DSB sites on single gene would trigger homology directed repair by donor DNA fragments when other gene copies also cleavaged. This finding can provide strategies for other multiple-copy gene editing.
For the future studies, it is useful to generate synthetic master strain with histone H2A, H2B, H3, and H4. Thus, any combinatorial mutations on histone genes can be generated through a single transformation.
Material and methods
Strains and broth media
Escherichia coli Top10 was used for vector cloning. The yeast strain used in this work was Saccharomyces cerevisiae CEN.PK 113-5D (MAT a MAL2-8c SUC2 ura3-52). E. coli top10 cells were grown on LB-Agar or LB broth with 50 mg/L Amp antibiotics for plasmid construction. Yeast strains were grown in YPD media with 2% glucose before the transformation. Transformants were plated on synthetic complete (SC) media minus uracil plates to select yeast cells. SC-5-FOA agar plates were employed to drop-out Cas9 plasmids.
Plasmid construction
The SynH3 and synH4 coding sequences were designed according to the codon usage of S. cerevisiae, synthesized by Sangon Biotech (Shanghai) Co., Ltd., and cloned into pUC57 vector. The pCas vector with multiple gRNA was constructed as the previous report23.
Donor DNA preparation
Donors used in this work were obtained by PCR reaction and purified by gel extraction. we cloned HHT1–HHF1, and HHT2–HHF2 pair genes with their upstream and downstream sequences on pUC18 vectors. The repair donors were generated by PCR or fusion PCR with indicated primers amplification from the plasmid and purified by gel extraction.
Yeast transformation and mutant identification
Yeast transformation was carried out using the electroporation as the previous report23. Genome DNA of strains survived on the auxotrophic plate was extracted for PCR reaction. The genotype of H3 and H4 of mutants were identified with PCR amplification and followed DNA sequencing. And genome DNA of Saccharomyces cerevisiae CEN.PK 113-5D was used as a control. H3 and H4 sequences of mutants were amplified by primers on ~ 100 bp upper and lower sites for sequencing analysis. The mutation rate was calculated as the ratio of mutants to total colonies tested with 3 biological replicates.
DNA damage sensitivity assays
WT stain and two biological replicates of each mutant were grown in YPD overnight at 30 °C. Yeast culture with diluted at OD600 = 1 as the starting point. Serial tenfold dilutions of yeast culture were spotted onto YPD for 2 days, YPD with 0.05% MMS for 3 days, or YPD with 100 mM HU and incubated for 3 days at 30 °C.
References
Luger, K., Mader, A. W., Richmond, R. K., Sargent, D. F. & Richmond, T. J. Crystal structure of the nucleosome core particle at 2.8 angstrom resolution. Nature 389(6648), 251–260 (1997).
Luger, K. & Hansen, J. C. Nucleosome and chromatin fiber dynamics. Curr. Opin. Struct. Biol. 15(2), 188–196 (2005).
MacAlpine, D. M. & Almouzni, G. Chromatin and DNA replication. Cold Spring Harb. Perspect. Biol. 5(8), a010207 (2013).
Iwasaki, W. et al. Contribution of histone N-terminal tails to the structure and stability of nucleosomes. FEBS Open Biol. 3, 363–369 (2013).
Davey, C. A., Sargent, D. F., Luger, K., Maeder, A. W. & Richmond, T. J. Solvent mediated interactions in the structure of the nucleosome core particle at 1.9 angstrom resolution. J. Mol. Biol. 319(5), 1097–1113 (2002).
Dorigo, B. et al. Nucleosome arrays reveal the two-start organization of the chromatin fiber. Science 306(5701), 1571–1573 (2004).
Zhang, W. et al. Probing the function of metazoan histones with a systematic library of H3 and H4 mutants. Dev. Cell 48(3), 406-419 e5 (2019).
Fitch, D. H., Strausbaugh, L. D. & Barrett, V. On the origins of tandemly repeated genes: Does histone gene copy number in Drosophila reflect chromosomal location?. Chromosoma 99(2), 118–124 (1990).
Dai, J. et al. Probing nucleosome function: A highly versatile library of synthetic histone H3 and H4 mutants. Cell 134(6), 1066–1078 (2008).
Nakanishi, S. et al. A comprehensive library of histone mutants identifies nucleosomal residues required for H3K4 methylation. Nat. Struct. Mol. Biol. 15(8), 881–888 (2008).
Jiang, S. et al. Construction of comprehensive dosage-matching core histone mutant libraries for Saccharomyces cerevisiae. Genetics 207(4), 1263–1273 (2017).
Cross, S. L. & Smith, M. M. Comparison of the structure and cell cycle expression of mRNAs encoded by two histone H3–H4 loci in Saccharomyces cerevisiae. Mol. Cell. Biol. 8(2), 945–954 (1988).
Stovicek, V., Holkenbrink, C. & Borodina, I. CRISPR/Cas system for yeast genome engineering: Advances and applications. FEMS Yeast Res. 17(5), fox030 (2017).
Khan, S. et al. CRISPR/Cas9: The Jedi against the dark empire of diseases. J. Biomed. Sci. 25(1), 29 (2018).
Bortesi, L. & Fischer, R. The CRISPR/Cas9 system for plant genome editing and beyond. Biotechnol. Adv. 33(1), 41–52 (2015).
Zhao, D. et al. Development of a fast and easy method for Escherichia coli genome editing with CRISPR/Cas9. Microb. Cell Fact. 15(1), 205 (2016).
Vasquez, J. J., Wedel, C., Cosentino, R. O. & Siegel, T. N. Exploiting CRISPR-Cas9 technology to investigate individual histone modifications. Nucleic Acids Res. 46(18), e106 (2018).
Pulecio, J., Verma, N., Mejia-Ramirez, E., Huangfu, D. & Raya, A. CRISPR/Cas9-based engineering of the epigenome. Cell Stem Cell 21(4), 431–447 (2017).
Leung, J. W. C., Emery, L. E. & Miller, K. M. CRISPR/Cas9 gene editing of human histone H2A variant H2AX and MacroH2A. Methods Mol. Biol. 1832, 255–269 (2018).
Harutyunyan, A. S. et al. H3K27M induces defective chromatin spread of PRC2-mediated repressive H3K27me2/me3 and is essential for glioma tumorigenesis. Nat. Commun. 10(1), 1262 (2019).
Zhou, Z. et al. Independent manipulation of histone H3 modifications in individual nucleosomes reveals the contributions of sister histones to transcription. eLife 6, e30178 (2017).
Hyland, E. M. et al. Insights into the role of histone H3 and histone H4 core modifiable residues in Saccharomyces cerevisiae. Mol. Cell. Biol. 25(22), 10060–10070 (2005).
Zhang, Y. et al. A gRNA-tRNA array for CRISPR-Cas9 based rapid multiplexed genome editing in Saccharomyces cerevisiae. Nat. Commun. 10(1), 1053 (2019).
Acknowledgements
This work was supported by Outstanding Talent Introduction Program from College of Veterinary Medicine, China Agricultural University, Beijing 100193, China, National Key Research and Development Program of China (2018YFD0500600), National Natural Science Foundation of China (31572442 and 31200941), and the Fundamental Research Funds for the Central Universities (106112016CDJXY290002).
Author information
Authors and Affiliations
Contributions
Y.F., G.M., R.Z. and Y.Z. designed the research; Y.F. carried out the experiment; Y.F., Z.Z, G.M. and Y.Z. analyzed data; Y.Z. wrote the paper. G.M., R.Z. and Y.Z. supervised the research.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fu, Y., Zhu, Z., Meng, G. et al. A CRISPR-Cas9 based shuffle system for endogenous histone H3 and H4 combinatorial mutagenesis. Sci Rep 11, 3298 (2021). https://doi.org/10.1038/s41598-021-82774-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-82774-4
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.