Abstract
Nuclear factor-erythroid 2-related factor 2 (Nrf2) is a key transcriptional regulator of several antioxidant and anti-inflammatory enzymes. It binds to its endogenous inhibitor Kelch-like ECH-associated protein 1 (Keap1) in the cytoplasm under normal conditions. Various endogenous or environmental oxidative stresses can disrupt the Nrf2/Keap1 complex, allowing Nrf2 to translocate into the nucleus, where it induces the transcription of various cytoprotective enzymes by binding to antioxidant responsive elements. These enzymes have been reported to play a role in regulating tumour growth, angiogenesis, and chemoprevention. Invasion and migration are the most harmful aspects of cancer; they directly impacts the patients’ survival. Although the roles of Keap1/Nrf2 and their downstream genes in various cancers have been widely documented, their role in regulating cell motility still remains unclear, particularly in cancer cells. We observed that Nrf2 suppression following treatment with brusatol in non-small-cell lung cancer (NSCLC) cells with either exogenously introduced Keap1 or siNrf2 resulted in the inhibition of cell migration and invasion, with shrinking cell morphology due to decreased focal adhesions via inhibition of the RhoA–ROCK1 pathway. Nrf2 overexpression showed opposite results. Thus, the Nrf2/Keap1 pathway may affect cell motility by dysregulating the RhoA–ROCK1 signalling pathway in NSCLC.
Similar content being viewed by others
Introduction
Carcinomas, which originate from epithelial cells, constitute more than 90% of all malignant human cancers; in carcinoma patients, lymph node and peritoneal metastasis is a major cause of tumour recurrence and cancer-related death1. Cell motility is necessary for the sequential multi-step processes involved in cancer metastasis, such as invasion, intravasation, and extravasation. The ability of cancer cells to invade into the surrounding tissues is one of the major hallmarks of cancer, which requires increased cell motility driven by remodelling of the cytoskeletal system and the contact of the cells with the extracellular matrix. This acquired migratory and invasive ability of cancer cells during metastasis is similar to the epithelial to mesenchymal transition (EMT) that occurs during embryonic development, tissue remodelling, and wound healing2,3,4. Several mechanisms associated with EMT have been studied. One of the most well-studied mechanisms of EMT is the signalling pathway driven by growth factors associated with receptor tyrosine kinases (RTK) and other signalling proteins such as TFG-β, and Wnt/β-catenin-activated downstream transcriptional repressors of E-cadherin5,6. The other effectors of cell motility are modulators of other adhesion systems and activators of actin cytoskeleton remodelling, such as Rho-family GTPases7,8.
Nuclear factor-erythroid 2-related factor 2 (Nrf2) is an important transcriptional regulator of many antioxidant and anti-inflammatory enzymes. Nrf2 binds to its endogenous inhibitor, Kelch-like ECH-associated protein 1 (Keap1) in the cytoplasm under normal conditions. The Nrf2–Keap1 complex can be disrupted by various endogenous or environmental oxidative stresses, which leads to accumulation and transactivation of Nrf2. Recently, increasing number of studies have shown and continue to show that persistent Nrf2 activation due to dysregulation of the Nrf2–Keap1 pathway in various cancer cells induces cell proliferation/growth by reprogramming metabolic processes. This is associated with poor prognosis due to acquired resistance to chemotherapy9,10. However, limited studies have shown the role of Nrf2/Keap1 in the regulation of cell motility and EMT in cancer cells and hence, it remains unclear11,12,13,14,15.
Our previous study showed that overexpression of Nrf2 due to Keap1 mutation increased cell invasion and metastatic ability of EGFR tyrosine kinase inhibitor (TKI)-resistant lung cancer cells both in vitro and in vivo. We also observed suppression of RhoA, ROCK1, Snail, and β-catenin, which are all well-known markers associated with cell motility/movement and EMT16. Therefore, in this study, we further investigated the mechanism underlying the induction of cell motility by the Nrf2–Keap1 pathway in lung cancer cells.
Results
Nrf2 regulates the motility of NSCLC cells
To investigate whether Nrf2 affects cell motility, migration and invasion assays were performed. Wild-type Keap1-Flag or siNrf2 was introduced into A549 and H460 cells, which harbour Keap1 mutations, and hence, have a higher expression of Nrf2 compared to the cells without Keap1 mutations (Fig. 1a). These cells were then treated with brusatol, an Nrf2 inhibitor, and consequently, inhibition of cell migration and invasion was observed (Fig. 2a,Bb). The effect of brusatol was confirmed using dual luciferase assay (Fig. 1b). In addition, we observed that overexpression of Nrf2 by exogenously introducing Nrf2-EGFP into the HCC827 cells increased the migration and invasion ability of these cells (Fig. 2c,d). These results indicated that Nrf2 plays a role in regulating cell motility.
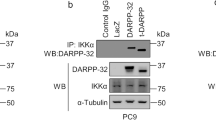
Expression levels of Nrf2 and Keap1 in NSCLC cells and inhibitory effect of brusatol on Nrf2 expression. (a) Nrf2 and Keap1 expression levels in different types of NSCLC cells were evaluated using Western blotting. (b) Inhibition of Nrf1 transcriptional activity seen after evaluation of ARE using dual luciferase assay after treatment with brusatol in H460 (10 nM) and A549 (30 nM) cells for 24 h.Expression levels of Nrf2 and Keap1 in NSCLC cells detected by western blotting.
Nrf2 regulates cell motility. Nrf2 expressed was suppressed in A549 or H460 cells following treatment with brusatol (10 nM) or transfection of Keap1-Flag or siNrf2. (a,c) Horizontal migration of these cells was compared with that of the control cells using wound healing assay for the indicated time points. (b,d) Nrf2 was overexpressed by transfecting Nrf2-EGFP into the HCC827 cells. Vertical migration was detected by transwell assay after brusatol treatment or transfection for 24 h. The degree of migration was quantified by calculating the area of migrated cells using the image processing software, Image J (Ver. 1/52n, NIH, Bethesda, MD, USA). Scale bar: 200 µm *p < 0.05, **p < 0.01, ***p < 0.001 compared with control.
RhoA–ROCK1 pathway is associated with Nrf2-induced increase in NSCLC cell motility
To investigate the molecular pathway utilised by Nrf2 in the regulation of cell motility, the levels of EMT-related proteins, such as Snail, Slug, and E-cadherin, and cytoskeleton-related RhoA–ROCK1 pathway molecules were examined by western blotting and/or qRT-PCR. Inhibition of Nrf2 after brusatol treatment resulted in a consistent decrease in the expression levels of RhoA and ROCK1 proteins in A549 and H460 cells (Fig. 3a,b). The opposite effect was seen in the HCC827 cells when Nrf2 was overexpressed by exogenously introducing Nrf2–EGFP and the Keap1 mutant, Keap1 (D236H)-Flag (Fig. 3c,d). However, the mRNA level of expression of RhoA and ROCK1 was not consistent with protein level of expression, which suggests the possibility of post-translational modification of RhoA and ROCK1. We also observed that the expression levels of EMT-related molecules after suppression or overexpression of Nrf2 varied depending on the cell lines irrespective of the presence of Nrf2 (Fig. 3a,b). These results indicated that Nrf2 regulates cell motility through the RhoA–ROCK1 pathway, and not through the EMT pathway.
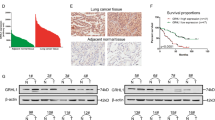
Molecules from the RhoA-ROCK1 pathway, and not the EMT-related molecules, are associated with Nrf2-induced cell motility. The expression levels of Nrf2, RhoA, Rock1, and/or EMT markers were detected using (a,c) western blotting or (b,d) qRT-PCR. (a,b) Nrf2 was suppressed by treating the A549 or H460 cells with brusatol (10 nM) or transfection of Keap1-Flag or siNrf2. (c,d) Nrf2 was overexpressed by transfecting HCC827 cells with Nrf2-EGFP or Keap1 mutant (D236H)-Flag for 24 h.*p < 0.05, **p < 0.01, ***p < 0.001 compared with control.
Stability and/or activity of RhoA is associated with Nrf2-induced increase in NSCLC cell motility
Since it has been shown that Nrf2 controls the RhoA–ROCK1 pathway, the mechanism how Nrf2 affects the RohA–ROCK1 signal axis beyond changes in total protein was investigated. H460 and A549 cells treated with MG132 to inhibit proteasomal degradation. The result showed RhoA and ROCK1 expression was recovered by treatment of MG132 with brusatol in H460 indicating Nrf2 controls RhoA–ROCK1 pathway through proteosomal degradation, but not in A549 cells (Fig. 4a). Besides, the effect of Nrf2 on the activity of RhoA was evaluated using GST pulldown assay. Brusatol-mediated inhibition of Nrf2 induced down-regulation of the active RhoA-GTP form in the A549, but not in H460 cells (Fig. 4b), and overexpression of Nrf2 by exogenously introducing Nrf2 resulted in a rise in the RhoA-GTP levels in HCC827 cells (Fig. 4c). These findings suggest that Nrf2 can regulate the RhoA–ROCK1 pathway through increased level of RhoA-GTP induced by the stability of RhoA and/or activity of RhoA-GTP in NSCLC cells, which is associated with protein level of RhoA.
Nrf2 increases the stability or activity of RhoA. (a) Cells were treated with MG132 with or without brusatol for 3 h and then harvested for western blotting. (b,c) Activation of RhoA was measured using RhoA Pull-down activation assay. RhoA-GTP was detected by western blotting. A549 and H460 cells were treated with brusatol (15 nM and 10 nM) for 24 h. HCC827 was transfected by exogenously introduced Nrf2-EGFP for 24 h. The densitometry quantification of the western blot determined using Image J software (Ver. 1/52n, NIH).
Nrf2 regulates the formation of actin stress fibres
To demonstrate the effect of Nrf2 on cell motility, we observed the morphology and distribution of the F-actin cytoskeleton along with focal adhesions (Fig. 5). F-actin was stained with rhodamine phalloidin (red) and the focal adhesions were detected by vinculin staining after the inhibition of Nrf2 via brusatol treatment in the H460 and A549 cells. Results showed the presence of diffused short rod-like F-actin (red) with reduced focal adhesion (yellow) with increased expression of vinculin, throughout the cytoplasm and the appearance of shrinking cell morphology, while the control cells exhibited noticeably stretched stress fibre formations. This indicated the down-regulation of the formation of stress fibres, as well as a significant decrease in the formation of focal adhesions.
Nrf2 is associated with the formation of F-actin stress fibres with focal adhesion. Representative images of cells stained with vinculin (green) with rhodamine phalloidin (red) after brusatol treatment (30 nM and 10 nM for A549 and H460 cells, respectively). White circled cell enlarged. Nuclei were counterstained with DAPI (blue). Images were obtained with a confocal laser scanning microscope (LSM 780, ZEISS) and visualised using the 100× oil immersion objective (EC Plan-Neofluar 100×/1.3 oil). Intensity of fluorescence from the captured image of cells was quantified using ImageJ software (Ver. 1/52n, NIH).
Discussion
Cell migration plays a crucial role in many types of human diseases including cancer, where dysregulation of cell migration can lead to metastasis, resulting in the poor prognosis of cancer patients. Investigation of genes or pathways that regulate cell migration and metastasis will facilitate the understanding of this process and the development of novel diagnostic and treatment strategies for cancer. Nrf2 is a transcription factor that has been known to be a master regulator of the cytoprotective response towards oxidative and electrophilic stresses. Under basal conditions, Nrf2 is localised in the cytoplasm by a cluster of proteins that degrade it quickly. Under oxidative stress, instead of being degraded, Nrf2 is translocated to the nucleus, where it binds to DNA promoters and initiates the transcription of several anti-oxidant genes and their proteins in healthy normal cells17,18. In contrast to the cytoprotective role of Nrf2 in normal cells, accumulating evidence suggests that persistently high activation of Nrf2 in various type of cancers, including lung cancer, is related with progression, metastasis, and resistance against chemotherapy and radiotherapy, which leads to poor prognosis16,19,20. The Keap1–Nrf2 pathway is therefore, considered to be a novel oncogenic signalling pathway and an attractive target for developing anti-cancer therapeutics. However, although the role of Keap–Nrf2 is well studied, studies on the role of Nrf2 in cell motility, EMT, and the underlying signalling pathways in cancer cells are scarce11,12,13,14,15.
In this study, we further investigated how Nrf2 regulates motility in NSCLC cells based on our previous findings, which showed that high activation of Nrf2 due to Keap1 mutation promotes migration ability both in vitro and in vivo in EGFR-TKI resistant lung cancer cells. Well-known EMT makers such as Snail, Slug, and E-cadherin, were examined in Nrf2-overexpressing or -knockdown NSCLC cells. Results showed that the expression of these molecules was not consistent, which might explain why previously published reports have shown controversial and cell-type specific results21,22,23,24. Other molecules associated with cytoskeletal remodelling such as RhoA and ROCK1 were also examined and we observed that overexpression of Nrf2 up-regulated, and suppression of Nrf2 down-regulated the RhoA–ROCK1 pathway. The RhoA/ROCK1 pathway plays an important role in cell movement and morphogenesis, which is associated with invasion and metastasis, as cellular cytoskeletal regulation and reprogramming is one of its main functions8,25. RhoA, which is a member of the Rho family of GTPases, facilitates the polymerisation of actin (F-actin formation) to form stress fibres, which are antiparallel actin filaments that are crosslinked by myosin, as well as the activation of myosin to trigger cell contractility. Binding of GTP-bound active RhoA activates ROCK1, which induces actin-myosin contraction by stimulating the phosphorylation of the myosin light chain directly. Additionally, the contractile force of the actin-myosin network is used to pull on the extracellular matrix (ECM) at the focal adhesion sites, while ECM stiffness can facilitate the formation of focal adhesions to allow cell movement. Increased level of RhoA-GTP was verified by the increase in the levels of RhoA stability and/or activity of RhoA-GTP following Nrf2 overexpression and decrease in RhoA-GTP levels after Nrf2 suppression in NSCLC cells. Moreover, immunostaining of F-actin with vinculin after Nrf2 inhibition by brusatol resulted in the increased expression of vinculin, which is known to inhibit cellular motility26 and formation of diffused short rod-like F-actin with reduced focal adhesions throughout the cytoplasm, along with the appearance of shrunken cell morphology, while the control cells exhibited noticeable formation of stretched stress fibres, indicating that Nrf2 activated the formation of actin stress fibres, leading to increased cell motility27. Similar results were reported in breast and gastric cancer cells, where it was seen that Nrf2/Keap1 promoted cell migration by upregulating the RhoA/ROCK1 pathway11,28. In lung adenocarcinoma cells, it has been reported that Keap1–Nrf2 interactions suppressed cell motility by targeting S100P15. In addition, this report suggested the possibility of an association of RhoA in the Nrf2/Keap1 pathway with the motility of lung cancer cells, as significant morphological and cytoskeletal changes were noticed due to the manipulation of Keap1 and suppression of RhoA activity by Keap1 overexpression in Cl1 lung cancer cells15. These results are consistent with ours. Taken together, we conclude that the Nrf2–Keap1 complex regulates cancer cell motility through the RhoA–ROCK1 signalling pathway, and not via EMT related molecules, in lung cancer cells.
Methods
Cell culture and reagents
The human NSCLC cells cell lines HCC827, H460, and A549 were purchased from American Type Culture Collection (ATCC, USA). These cells were cultured in RPMI1640 (Welgene, Daegu, South Korea) supplemented with 10% foetal bovine serum (FBS) (Welgene, Daegu, South Korea) and 1% penicillin/streptomycin at 37 ℃ in a humidified atmosphere containing 5% CO2. Brusatol (Carbosynth, UK) was purchased and dissolved in dimethyl sulfoxide (DMSO) (D2650, Sigma Aldrich, USA). MG132 was purchased from CalBiochem (SanDiego, CA, USA). Nrf2-EGFP and Keap1-Flag vectors were purchased from Addgene (http://www.addgene.org). The oligo ribonucleotide sequences of human Nrf2 siRNA (siNrf2) were as follows: 5′-UCUGACUCCGGCAUUUCACUTT-3′ (sense) and 5′-AGUGAAAUGCCGGAGUCAGATT-3′ (antisense) (Shanghai GenePharma Co. Ltd., China).
Cell migration assay
Cell migration was measured using wound healing assay. Cells were seeded into 6-well plates and cultured to a confluent monolayer for 24 h. The cell monolayer was scratched using a 200-µl sterile pipette tip and washed twice with phosphate buffered saline (PBS) to remove the detached cells. Following hours of culture in RMPI-1640 supplemented with 1% serum to minimize cell proliferation during the period of assay29. Initial images of four independent areas after each scratch were acquired at time zero. Each scratch was examined, the images of the same location were captured, and the healed area was measured after the indicated time. The degree of migration was quantified by calculating the area of migrated cells using the image processing software, Image J (Ver. 1/52n, NIH, Bethesda, MD, USA).
Transwell invasion assay
The invasiveness of tumour cells was evaluated using an invasion assay with Transwell devices (CT-3422, 8 μm pore size, 6.5 mm diameter, Corning Life Science, USA) coated with Matrigel (BD, 356230, 100 μg/ml, 15 µl/well). Cells (8 × 104) were placed in the upper chamber of the Transwell device in serum-free media. Media containing 10% FBS was added as a chemoattractant in the lower chamber of each well. Non-invading cells were removed using cotton swabs. The remaining cells were fixed and stained by Hemacolor rapid staining solutions (Merck, Kenilworth, NJ, USA) for 3 min. The numbers of invasive cells were counted from five representative fields on the membrane.
RNA isolation and quantitative real time polymerase chain reaction (qRT-PCR)
Total RNA was isolated from the cells using TRIzol reagent (Invitrogen, Carlsbad, CA, USA). cDNA was synthesised from the total RNA using the reverse transcription kit (LaboPass, Cosmo Genetech, Seoul, South Korea) according to the manufacturer’s instructions. qRT-PCR was carried out using gene specific primers with SYBR green Q master mix (Labopass, Cosmo Genetech, Seoul, South Korea) on an ABI7500 real-time PCR system (Applied Biosystems, Warrington, UK). The oligonucleotide primers that were used for the qRT-PCR are described in Table 1. The Ct values of the target genes were normalised to an endogenous reference gene (GAPDH). Each gene was analysed in duplicate and the analysis was repeated via three independent experiments.
Western blot analysis
Proteins from the cell lysates were separated using SDS-PAGE and transferred onto Immobilon-P PVDF membranes (IPVH00010, Millipore, USA). These membranes were subsequently probed with the appropriate primary antibodies and incubated with the corresponding goat anti-mouse Ig G (7076) or goat anti-rabbit Ig G (7074) secondary antibodies conjugated with horseradish peroxidase (HRP). Chemiluminescence was detected using the enhanced chemiluminescence (ECL) system (Translab, South Korea) according to the manufacturer’s instructions. Secondary antibodies were purchased from Cell Signaling technology, Inc. The primary antibodies against Snail (3879), Slug (9585), E-cadherin (3195), β-catenin (9582), and Keap1 (7705) were purchased from Cell Signaling technology. Antibodies against ROCK1 (sc-5560), RhoA (sc-179), and β-actin (sc-47778) were purchased from Santa Cruz Biotechnology. Antibodies against Nrf2 (ab62352) were purchased from Abcam. Anti-Flag antibodies (F1804) were purchased from Sigma Aldrich. The result of gels images were cropped and full-length gels and blots are included in the Supplementary Figures.
GST-Rhotekin-RBD pull-down assay
The activation of RhoA was measured using the RhoA Pull-down Activation Assay Biochem kit (BK036, Cytosekeleton, USA) according to the manufacturer’s protocol. Cells were harvested and washed with PBS and then lysed in ice-cold cell lysis buffer containing 1× protease Inhibitor Cocktail. The lysates were centrifuged at 10,000 rpm for 2 min at 4 ℃. The supernatant (500 µg) was incubated with GST-Rhotekin-RBD to detect RhoA-GTP. The beads were washed thrice with washing buffer. The bound proteins were eluted with 2× Laemmli sample buffer and loaded onto an SDS-PAGE gel. The primary anti-RhoA antibody was provided in the kit.
Immunofluorescence
Cells (2 × 102 cells/well) were plated in a 6-well-plate containing coverslips in 2 ml of culture media and cultured overnight. The cells were washed twice with PBS and then fixed with 4% formaldehyde in PBS at room temperature (RT) for 30 min. The fixed cells were washed thrice with PBS for 5 min. After washing, the cells were permeabilised with 0.1% TritonX-100 in PBS for 15 min, followed by three washes with PBS. The fixed cells were blocked with 3% (w/v) BSA, stained with an anti-Vinculin antibody (Abcam, CA, Cat No. ab129002) and rhodamine phalloidin (R415, Invitrogen, USA) overnight at 4 ℃ in the dark, followed by incubation with Fluor488-conjugated secondary antibodies (Invitrogen, CA, Cat No. A21202) after three washes with PBS. Each coverslip was mounted with a medium containing the nuclear stain DAPI (Vectashield) (H 1200, Vector Laboratories, USA). The fluorescence images were obtained with a Laser Scanning Confocal Microscope (LSM 780, ZEISS, Germany) and visualised with the 100× oil immersion objective lenses (EC Plan-Neofluar 100×/1.3 oil).
Luciferase reporter assay
Cells (2 × 105 cells/well) were seeded into a 6-well plate and cultured overnight. The cells were then transfected with the luciferase reporter vector pGL4.37-ARE-Luc using Gene-Fect Transfection Reagent (TLC-001, TransLab, South Korea) according to the manufacturer’s instructions. The transfected cells, with or without brusatol treatment for 24 h, were harvested and lysed. Luciferase assays were performed on the lysate using the Dual-Luciferase Reporter Assay Systems (E1960, Promega, USA). The activity of the firefly luciferase was normalised to Renilla luciferase activity and was assessed using the Enspire Multi Plate Reader (PerkinElmer, USA).
Statistical analysis
Results were indicated as the means ± standard deviations (SDs) from at least three independent experiments. Comparisons between groups were performed using a two-tailed Student’s t-test or ANOVA test. If p-value obtained by one-way ANOVA is < 0.05, the p-values between the groups are compared with post-test, Bonferroni, and Tukey HSD. p-values ≤ 0.05 were considered statistically significant.
References
Birchmeier, C., Birchmeier, W. & Brand-Saberi, B. Epithelial-mesenchymal transitions in cancer progression. Acta Anat. (Basel) 156, 217–226. https://doi.org/10.1159/000147848 (1996).
Kalluri, R. & Weinberg, R. A. The basics of epithelial-mesenchymal transition. J. Clin. Investig. 119, 1420–1428. https://doi.org/10.1172/jci39104 (2009).
Wu, Y. & Zhou, B. P. New insights of epithelial-mesenchymal transition in cancer metastasis. Acta Biochim. Biophys. Sin. (Shanghai) 40, 643–650. https://doi.org/10.1111/j.1745-7270.2008.00443.x (2008).
Pastushenko, I. & Blanpain, C. EMT transition states during tumor progression and metastasis. Trends Cell Biol. 29, 212–226. https://doi.org/10.1016/j.tcb.2018.12.001 (2019).
Iwatsuki, M. et al. Epithelial-mesenchymal transition in cancer development and its clinical significance. Cancer Sci. 101, 293–299. https://doi.org/10.1111/j.1349-7006.2009.01419.x (2010).
Katsuno, Y., Lamouille, S. & Derynck, R. TGF-beta signaling and epithelial-mesenchymal transition in cancer progression. Curr. Opin. Oncol. 25, 76–84. https://doi.org/10.1097/CCO.0b013e32835b6371 (2013).
Hanna, S. & El-Sibai, M. Signaling networks of Rho GTPases in cell motility. Cell Signal. 25, 1955–1961. https://doi.org/10.1016/j.cellsig.2013.04.009 (2013).
Ridley, A. J. Rho GTPase signalling in cell migration. Curr. Opin. Cell Biol. 36, 103–112. https://doi.org/10.1016/j.ceb.2015.08.005 (2015).
Jaramillo, M. C. & Zhang, D. D. The emerging role of the Nrf2-Keap1 signaling pathway in cancer. Genes Dev. 27, 2179–2191. https://doi.org/10.1101/gad.225680.113 (2013).
de la Vega, M. R., Chapman, E. & Zhang, D. D. NRF2 and the hallmarks of cancer. Cancer Cell 34, 21–43. https://doi.org/10.1016/j.ccell.2018.03.022 (2018).
Zhang, C. et al. NRF2 promotes breast cancer cell proliferation and metastasis by increasing RhoA/ROCK pathway signal transduction. Oncotarget 7, 73593–73606. https://doi.org/10.18632/oncotarget.12435 (2016).
Zhang, H. S. et al. Nrf2 promotes breast cancer cell migration via up-regulation of G6PD/HIF-1alpha/Notch1 axis. J. Cell. Mol. Med. 23, 3451–3463. https://doi.org/10.1111/jcmm.14241 (2019).
Li, D. et al. Targeting the NRF-2/RHOA/ROCK signaling pathway with a novel aziridonin, YD0514, to suppress breast cancer progression and lung metastasis. Cancer Lett. 424, 97–108. https://doi.org/10.1016/j.canlet.2018.03.029 (2018).
Rachakonda, G. et al. Increased cell migration and plasticity in Nrf2-deficient cancer cell lines. Oncogene 29, 3703–3714. https://doi.org/10.1038/onc.2010.118 (2010).
Chien, M. H. et al. Keap1-Nrf2 interaction suppresses cell motility in lung adenocarcinomas by targeting the S100P protein. Clin. Cancer Res. 21, 4719–4732. https://doi.org/10.1158/1078-0432.Ccr-14-2880 (2015).
Park, S. H. et al. Resistance to gefitinib and cross-resistance to irreversible EGFR-TKIs mediated by disruption of the Keap1-Nrf2 pathway in human lung cancer cells. FASEB J. 32, 5862–5873. https://doi.org/10.1096/fj.201800011R (2018).
Reddy, N. M. et al. Genetic dissection of the Nrf2-dependent redox signaling-regulated transcriptional programs of cell proliferation and cytoprotection. Physiol. Genomics 32, 74–81. https://doi.org/10.1152/physiolgenomics.00126.2007 (2007).
Niture, S. K., Khatri, R. & Jaiswal, A. K. Regulation of Nrf2-an update. Free Radic. Biol. Med. 66, 36–44. https://doi.org/10.1016/j.freeradbiomed.2013.02.008 (2014).
Tian, Y. et al. Emerging roles of Nrf2 signal in non-small cell lung cancer. J. Hematol. Oncol. 9, 14. https://doi.org/10.1186/s13045-016-0246-5 (2016).
Yamadori, T. et al. Molecular mechanisms for the regulation of Nrf2-mediated cell proliferation in non-small-cell lung cancers. Oncogene 31, 4768–4777. https://doi.org/10.1038/onc.2011.628 (2012).
Bocci, F. et al. NRF2 activates a partial epithelial-mesenchymal transition and is maximally present in a hybrid epithelial/mesenchymal phenotype. Integr. Biol. (Camb) 11, 251–263. https://doi.org/10.1093/intbio/zyz021 (2019).
Zhang, Z. et al. Nrf2 antioxidant pathway suppresses Numb-mediated epithelial-mesenchymal transition during pulmonary fibrosis. Cell Death Dis. 9, 83. https://doi.org/10.1038/s41419-017-0198-x (2018).
Zhou, W. et al. Nrf2 inhibits epithelial-mesenchymal transition by suppressing snail expression during pulmonary fibrosis. Sci. Rep. 6, 38646. https://doi.org/10.1038/srep38646 (2016).
Qu, J. et al. Downregulation of HMGB1 is required for the protective role of Nrf2 in EMT-mediated PF. J. Cell. Physiol. 234, 8862–8872. https://doi.org/10.1002/jcp.27548 (2019).
O’Connor, K. & Chen, M. Dynamic functions of RhoA in tumor cell migration and invasion. Small GTPases 4, 141–147. https://doi.org/10.4161/sgtp.25131 (2013).
Rodriguez Fernandez, J. L. et al. Overexpression of vinculin suppresses cell motility in BALB/c 3T3 cells. Cell. Motil. Cytoskelet. 22, 127–134 (1992).
Pellegrin, S. & Mellor, H. Actin stress fibres. J. Cell Sci. 120, 3491–3499. https://doi.org/10.1242/jcs.018473 (2007).
Matsuoka, T. & Yashiro, M. Rho/ROCK signaling in motility and metastasis of gastric cancer. World J. Gastroenterol. 20, 13756–13766. https://doi.org/10.3748/wjg.v20.i38.13756 (2014).
Rodriguez, L. G. et al. Wound-healing assay. Methods Mol. Biol. 294, 23–29. https://doi.org/10.1385/1-59259-860-9:023 (2005).
Acknowledgements
This work was supported by the Basic Science Research Program Grants (NRF- 2017R1A2B4003233 and NRF-2019R1A2C1083909) from the National Research Foundation of Korea, which is funded by the Ministry of Education, Science, and Technology, South Korea. We thank the Confocal Microscope core facility at the Seoul Center, KBSI.
Author information
Authors and Affiliations
Contributions
J.L. conceived and designed the experiments; E.S.K., D.K. and S.-H.K. conducted the experiments; E.S.K., D.K., S.-H.K. and J.L. analysed and interpreted the results. E.S.K. and J.L. wrote the manuscript and all authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ko, E., Kim, D., Min, D.W. et al. Nrf2 regulates cell motility through RhoA–ROCK1 signalling in non-small-cell lung cancer cells. Sci Rep 11, 1247 (2021). https://doi.org/10.1038/s41598-021-81021-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-81021-0
This article is cited by
-
Cordyceps sinensis relieves non-small cell lung cancer by inhibiting the MAPK pathway
Chinese Medicine (2024)
-
Fluorouracil exacerbates alpha-crystallin B chain—mediated cell migration in triple-negative breast cancer cell lines
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.