Abstract
Haematopoietic malignancies are frequently characterized by karyotypic abnormalities. The development of targeted drugs has been pioneered with compounds against gene products of fusion genes caused by chromosomal translocations. While polysomies are equally frequent as translocations, for many of them we are lacking therapeutic approaches aimed at synthetic lethality. Here, we report two new cell lines, named MBU-7 and MBU-8, that differ in complete trisomy of chromosome18, a partial trisomy of chromosome 7 and a tetrasomy of the p-arm of chromosome 8, but otherwise share the same mutational pattern and complex karyotype. Both cell lines are divergent clones of U-937 cells and have the morphology and immunoprofile of monocytic cells. The distinct karyotypic differences between MBU-7 and MBU-8 are associated with a difference in the specific response to nucleoside analogues. Taken together, we propose the MBU-7 and MBU-8 cell lines described here as suitable in vitro models for screening and testing vulnerabilities that are associated with the disease-relevant polysomies of chromosome 7, 8 and 18.
Similar content being viewed by others
Introduction
Haematological malignancies are clonal disorders that originate from somatic mutations in haematopoietic stem and progenitor cells. These mutations often occur in epigenetic regulators and thus affect a variety of downstream processes (reviewed in Maher and Diesch et al.1). Cytogenetic alterations are frequent in leukaemia and lymphoma and are detected in 50 to 60 percent of acute myeloid leukaemia (AML) patients2. Chromosomal alterations include translocations frequently encoding fusion oncogenes, which can cause functional losses and gains that affect entire chromosomes3. Abnormal karyotypes represent powerful prognostic factors4. This has led to the inclusion of certain cytogenetic abnormalities in the WHO classification for AML5 and their analysis as part of routine diagnostics. Primarily depending on the presence of translocations and gene fusions, AML patients can be stratified into three prognostic classes: favourable, intermediate and unfavourable, according to the aggressiveness of the disease and/or poor response to treatment6. For instance, the t(15;17)(q24;q21) translocation is associated with favourable prognosis due to the availability of drugs targeting the resulting PML-RARα fusion gene7. On the other hand, other gene fusions are associated with increased leukaemogenic expression. KMT2A-MLLT3, caused by the t(9;11)(p22;q23) translocation is associated with intermediate prognosis, while the inversion of chromosome 3 leads to RPN1-MECOM (inv(3)(q21q26)) and is associated with poor prognosis (reviewed in Lagunas-Rangel et al.6). In general, complex karyotypes are usually associated with a poorer outcome. Specific chromosomal losses and gains can also be informative for prognosis; however, they are only in very few cases used to stratify patients for treatment. One example is the loss of the q-arm of chromosome 5 in myelodysplastic syndrome that is a selection criteria for lenalidomide treatment with a high rate of favourable outcomes8. Another example is monosomy 7 in AML, which is associated with a poor prognosis and thus requires aggressive chemotherapy and hematopoietic stem cell transplantation9,10. A better understanding of how cytogenetic changes relate to drug responses is needed to improve patient stratification and greater therapy success. While clinical trials routinely assess cytogenetic profiles using conventional methods, in vitro models are needed to functionally test associations between cytogenetic alterations and responses to tool compounds and drugs in the pre-clinical phase.
Matched patient-derived cell lines representing different states of aggressiveness of a disease are powerful tools. This includes the B16 melanoma cell line model11 and engineered cell lines such as the MBU series for HRAS-driven squamous cell carcinoma12. Furthermore, many leukaemia- and lymphoma-derived cell lines have been established harbouring various different cytogenetic abnormalities or particular gene mutations13. Genetic drift is often seen in cell lines during long-term cultures. Indeed, karyotypic changes in leukaemia and lymphoma cell lines can be seen as a result of increasing passage numbers14. To some extent, these changes reflect the same events of clonal evolution occurring in patients during disease progression15. Characterizing the functional effects of these chromosomal changes would help identify vulnerabilities associated with specific acquired genetic abnormalities. This has been exemplified in a recent study performed in 27 strains of the frequently used breast cancer cell line MCF7, which display a multitude of genetic variations and considerably different drug responses16. And thus, well-characterized cell lines enable screening for differences in drug sensitivity associated with clinically relevant genetic alterations.
Here, we report the characterization of two divergent leukaemia sub-clones derived from U-937 cells. The two cell lines differed in partial and complete trisomies of chromosomes 7 and 18, respectively, as well as tetrasomy 8p. These cytogenetic differences were associated with a differential drug response suggesting that this pair of cell lines can serve as a model to specifically probe for vulnerabilities associated with the clinically relevant polysomies of chromosomes 7, 8 and 18.
Results
Isolation of divergent cell clones that differ in trisomies 7, 8 and 18
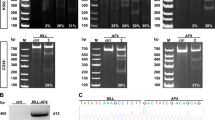
While growing leukaemia cells in suspension over periods of weeks, we noticed a change in the behaviour of the bulk population that was due to the presence and expansion of cells with unexpected characteristics. In order to characterize these cells, we first isolated and expanded single cell clones. We began to characterize two cell lines derived from the expansion of these single cell clones by conventional karyotyping (Fig. 1a). Both cell lines displayed a complex and overall highly similar karyotype. Shared alterations included der(1)t(1;5)(p22;q31.1), der(3)t(1;3)(q21;q26), der(5)t(1;5)(p22;q23.3), add(6)(p25), der(10)t(10;11)(p12;q14)/PICALM-MLLT10, del(11)(q22), der(13)t(1;13)(p32;p11.2), del(22)(q13.2q13.3) and unspecific rearrangements involving chromosomes 2q37, 3q12, 6p25, 12p12, and 16p11.1, as well as complete trisomies of chromosomes 20 and 21 (Table 1). Clear differences were the presence of trisomy 18 and a gain in material of chromosome 8 in one of the cell lines, which for this reason we termed MBU-8. We termed the other cell line MBU-7 as it had a complete trisomy 7, while in MBU-8 part of the q-arm of a third chromosome 7 was lacking. These major differences in the amount of genetic material from chromosomes 7, 8 and 18 has further been confirmed using CytoScan 750 K array-based technology (Fig. 1b and Supplementary Table S1). Interestingly, the array-based method indicated that the additional fragment of chromosome 8 contained two fused p-arms making it a partial tetrasomy 8. The array further provided additional resolution and helped to improve the karyotype formula, thus substantiating that these cell lines possess an overall complex karyotype (Supplementary Fig. S1 and Supplementary Data S1).
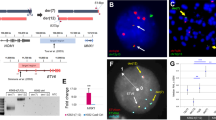
The two cell lines MBU-7 and MBU-8 differ in trisomies 8 and 18 and a partial trisomy of 7. (a) Representative karyotype images of MBU-7 and MBU-8 showing cytogenetic differences: trisomy of chromosomes 7 and 18, and tetrasomy 8p. (b) Distribution of copy number variations in chromosomes 7, 8 and 18 in MBU-7 and MBU-8 as determined by CytoScan 750 k Array analysis. Blue, gain; red, loss; purple, region of heterozygosity. (c) FISH with probes for EZH2 at 7q36 (green), the centromeric region at 7p11.1-q11.1 (blue) and KMT2E at 7q22 (red). (d) FISH with probes for IGH at 14q32.3 (green), the centromeric region at 8p11.1-q11.1 (blue) or MYC at 8q24 (red). (e) FISH with probes for BCL2 at 18q21 (orange). Arrows indicate loci that differ between MBU-7 and MBU-8. (f) Quantification of the FISH signal represented in (c–e) by the Metafer platform in MBU-7 and MBU-8 after 2–4 passages (p2-4) and 18–20 passages (p18-20).
Fluorescence in situ hybridization (FISH) confirmed these differences. While both cell lines showed three copies of the centromeric region of chromosome 7 and the locus 7q22, the locus 7q36 containing the EZH2 gene was only present twice in MBU-8 (Fig. 1c). In line with two additional copies of 8p fused at the peri-centromeric region, we detected a third signal for the centromeric region of chromosome 8 in MBU-8 but only two copies of the 8q24 portion of chromosome 8 containing the MYC gene locus (Fig. 1d). Finally, we confirmed the trisomy 18 in MBU-8 using a FISH probe targeting the locus of BCL2 on 18q21 (Fig. 1e). To examine the cytogenetic stability of MBU-7 and MBU-8 over an extended period of time, we repeated the FISH analysis after 18–20 passages and could not observe any significant differences in the chromosomes assessed (Fig. 1f). This analysis showed that both cell lines are sufficiently stable during a time window that makes them suitable for comparative experimental studies.
In summary, we isolated two cell lines with highly complex karyotypes that, at first glance, did not resemble any other previously described cell line. Importantly, the two cell lines differed in the number of copies of chromosomes 7, 8 and 18.
MBU-7 and MBU-8 are divergent subclones of U-937
As an alternative approach to identify the origin of these new cell lines, we analysed a panel of short tandem repeats (STR). Profiling of STRs is a widely used forensic technique to identify human individuals by unique signatures based on the allelic number of 16 highly conserved microsatellite sequences17. This method has found an important application in the identification of human-derived cell lines whose profiles are collected and stored in the publicly available Cellosaurus database from the Swiss Institute of Bioinformatics18. The profiles of MBU-7 and MBU-8 almost perfectly matched the U-937 cell line (Table 2). We confirmed the almost perfect match by repeating the STR analysis with U-937 cells and by visually comparing the profiles. For instance, the short tandem repeat D13S317 was present in 10 and 12 copies in MBU-7, MBU-8 and U-937 (Fig. 2a). Similarly, the repeat TH01 showed 6 and 9.3 copies in all three cell lines (Fig. 2a).
MBU-7 and MBU-8 are derived from U-937 cells. (a) Representative electropherograms of the microsatellite loci TH01 and D13S317 measured by STR profiling in U-937, MBU-7 and MBU-8. The numbers in the boxes represent the amount of microsatellites per locus. (b) Relative expression of the fusion gene PICALM-MLLT10 by RT-qPCR in SKK-1, U-937, MBU-7 and MBU-8. Data represent the mean ± SEM of three independent experiments. (c) The expression of the SKK-1 specific fusion gene ETV6-NTRK3 was analysed as in (b).
U-937 is a human haematopoietic monoblastoid cell line originally established from a 37-year old male patient with diffuse histiocytic lymphoma19, which is a rare, very aggressive type of non-Hodgkin’s lymphoma. Besides their popular use as a differentiation model, U-937 cells are well-studied for the translocation t(10;11)(p12;q14) resulting in a PICALM-MLLT10 fusion gene20. Gene fusions are a common characteristic of both myeloid and lymphoid leukemias and occur due to chromosomal translocations21. The resulting fusion genes frequently act as disease-driving oncogenes and have been investigated as part of diagnostic protocols across different leukemias due to their disease subtype specificity22,23. As shown in Fig. 2b, we could detect PICALM-MLLT10, expression in MBU-7, MBU-8 and U-937, but not in the secondary acute myeloid leukaemia (AML) cell line SKK-1 that we included as a control. Conversely, SKK-1, but not MBU-7, MBU-8 or U-937 cells expressed the ETV6-NTRK3 gene fusion (Fig. 2c) that is caused by a t(12;15)(p13;q25) translocation24.
To define the mutation profile of MBU-7 and MBU-8, we sequenced a previously described panel of 32 genes related to myeloid disorders25,26. Three pathogenic variants of WT1, TP53 and PTPN11 were observed in both subclones with similar variant allele frequency (Table 3). U-937 have been reported to express mutant p5327, do not express detectable levels of WT128 and harbour a PTPN11 mutation (G20R)29.
Taken together, these results suggest that MBU-7 and MBU-8 are subclones of U-937 cells. Several groups have reported different karyotypes for U-937, but reports are consistent in that U-937 cells are close to triploid30,31. Comparing these karyotypes with the karyotypes of MBU-7 and MBU-8 (Table 1) suggests that the two cell lines have diverged by acquiring additional cytogenetic alterations, such as multiple losses of genetic material. This includes the loss of the Y chromosome, a striking mosaicism given the male origin of the cell line. Overall, the loss of genetic material was more pronounced in MBU-7 cells than in MBU-8.
Monocytic morphology is accompanied by a shared immunoprofile
Phenotypically, U-937 cells resemble blast cells of the monocytic lineage19. In both MBU-7 and MBU-8, we observed a similar cell morphology resembling monocytic cells (Fig. 3a). Within both populations, cells varied in size and maintained the relationship in size between nucleus and cytoplasma. The nuclei had an irregular contour and lax chromatin with the presence of one or more evident nucleoli. Some cells displayed nuclear herniations, which are abnormal protrusions of the nucleus. The extended cytoplasm was basophilic and, in a subset of cells, presented granulation and vacuoles. Interestingly, in addition a majority of cells with the above described characteristics, both cell lines contained some particularly large cells that were multinucleated and had hyperbasophilic cytoplasm (Fig. 3a). These cells were more frequently seen in MBU-7 than MBU-8, which could be confirmed by measuring the forward and side scatter (FSC and SSC) in flow cytometry (Fig. 3b). Such large cells are often seen in leukaemia patients with complex or hyperdiploid karyotype and can indicate a more aberrant phenotype. Both cell lines were negative for myeloperoxidase, which is a marker for granulopoietic cells (data not shown). In terms of markers for the monocytic-macrophage lineage, both cell lines were positive for alpha-naphthyl butyrate esterase and negative for alpha-naphthyl acetate esterase (data not shown).
Monocytic morphology of MBU-7 and MBU-8 is accompanied by a shared immunoprofile. (a) Microscopy images of MBU-7 and MBU-8 after staining with May–Grünwald Giemsa solution. Scale bar, 10 µM. (b) Representative histogram (left) and quantification (right) of cell size of MBU-7 and MBU-8 assessed by flow cytometry (FSC, forward scatter; SSC, side scatter). Data represent the mean ± SEM of three independent experiments. Statistical analysis was performed by Student’s T-test, *p-value < 0.05. (c) Immunophenotype as determined by assessing the surface marker expression by flow cytometry in MBU-7 and MBU-8 subclones.
We next analysed the immunophenotypes and found that MBU-7 and MBU-8 have very similar surface marker expression (Fig. 3c). Specifically, they were both positive for the hematopoietic markers CD45, CD99, CD38 and CD44, as well as for the myeloid-monocytic markers CD11b, CD13 and CD15. In terms of stem cell markers, MBU-7 and MBU-8 were largely negative for CD34, HLA-DR, CD117 and CD123. Furthermore, virtually all cells expressed monocytic differentiation markers such as CD64, CD4, CD33 and CD56.
In summary, the immunoprofile and morphology of MBU-7 and MBU-8 cells associates them with the monocytic lineage. This is in line with them being derivatives of U-937 cells that, despite having originated from a histiocytic lymphoma patient, have monocytic characteristics. Indeed, U-937 are commonly used to study monocytic differentiation in vitro32.
MBU-7 and MBU-8 differ in drug response
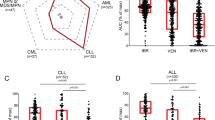
Finally, we decided to test the response of MBU-7 and MBU-8 cell lines to a panel of commonly used drugs. Specifically, we have chosen the nucleoside analogues azacitidine, decitabine, and cytarabine. Cytarabine is a DNA toxin and used as a standard chemotherapy drug in AML, acute lymphoid leukaemia (ALL) and non-Hodgkin lymphoma33. Azacitidine and decitabine can further act as DNA hypomethylating agents and are the treatment of choice for intermediate- to high-risk MDS patients not eligible for allogenic bone marrow transplantation34 and for elderly unfit AML patients35. We also included venetoclax, an inhibitor of the apoptosis regulator Bcl236, which is currently being approved for the treatment of an increasing number of haematopoietic malignancies (https://www.fda.gov/drugs/resources-information-approved-drugs/fda-approves-venetoclax-cll-and-sll). Specifically, we have determined the cell viability of MBU-7 and MBU-8 after four days of treatment. MBU-7 cells were significantly more sensitive towards the treatment with all three nucleoside analogues than MBU-8 (Fig. 4a). The biggest difference in response was observed with 0.125 μM decitabine and 0.25 μM cytarabine. Both cell lines were less sensitive to azacitidine but a pronounced difference was observed around 10 μM. In contrast, both MBU-7 and MBU-8 responded in a similar manner to venetoclax suggesting that we are not observing a general propensity of MBU-7 cells to drug-induced apoptosis (Fig. 4b).
MBU-7 and MBU-8 display different drug sensitivity. (a) Percentage of live MBU-7 or MBU-8 cells after 4 days of treatment with indicated concentrations of azacitidine, decitabine or cytarabine. (b) Percentage of live MBU-7 or MBU-8 cells after 4 days of treatment with indicated concentrations of venetoclax. (a,b) Data represent the mean ± SEM of four independent experiments. Statistical analysis was performed using ANOVA. *p-value < 0.01. ns non-significant.
Taken together, the cytogenetic differences of MBU-7 and MBU-8 are associated with a differential response to a subset of drugs commonly used for the treatment of haematological diseases.
Discussion
In our study, we characterized the two cell lines MBU-7 and MBU-8 that we isolated as clones from a suspension culture. The profile of small tandem repeats identified both cell lines as likely derivatives of U-937 cells. This was consistent with a shared mutational profile and a morphology and immunoprofile of monocytic cells. Although the karyotypes of MBU-7 and MBU-8 were different to U-937, they contained some characteristic features shared with U-937 cells such as the translocation t(10;11)(p12;q14) resulting in a PICALM-MLLT10 fusion gene20. Chromosomal instability and loss of heterozygosity has been previously attributed to U-93737. Indeed, subclones of U-937 with differences in both small tandem repeats and chromosomal aberrations have been identified following long periods in culture38,39. This can explain the divergence in karyotypes of MBU-7 and MBU-8 when compared to U-937. More specifically, U-937 cells have previously been described to have a near triploid karyotype31, while MBU-7 and MBU-8 only share trisomies of chromosomes 3, 20 and 21 but differ in complete or partial polysomies of 7, 8 and 18. U-937 cells are derived from a male donor19. However, Y chromosome is known to be frequently lost during cell culture38. Occasionally this leaves U-937 cells with two X-chromosomes31, which we also observed in MBU-7 and MBU-8. Taken together, our results suggest that MBU-7 and MBU-8 are clonal cell lines with divergent karyotypes that spontaneously derived over time from U-937 cells.
MBU-7 and MBU-8 have highly similar characteristics but differ in trisomies 7 and 18 and tetrasomy 8p in the context of a shared complex karyotype. Gains of material of chromosome 8 are frequent in myeloid malignancies. Trisomy 8 is the most frequent cytogenetic alteration in AML occurring in 10 to 15% of patients40,41. This includes MDS, where trisomy 8 represents the sole genetic aberration in 11% of cases42. Patients with trisomy of chromosome 8 are included in the intermediate-risk group and their prognosis thought to be defined by accompanying aberrations42. Trisomy 8 is frequently associated with mutations in driver genes including RUNX1, ASXL1, DNMT3A43, TET244, and IDH1 and IDH245. AML with tetrasomy of chromosome 8 has poor prognosis46. To our knowledge, tetrasomies of only the p-arm of chromosome 8 have not yet been reported in myeloid diseases. However, it needs to be pointed out that the partial tetrasomy 8 can easily be mistaken to be trisomy 8. Indeed, this would have been our interpretation of conventional karyotyping and FISH data in the absence of quantitative data from the CytoScan array. Importantly, the q-arm carries several cancer-relevant genes such as FGFR1, KAT6A and PCM1 (Supplementary Table S1).
In conclusion, MBU-7 and MBU-8 differ in discrete gains of genetic material of chromosomes 7, 8, and 18. The clinical significance of these differences remains to be determined in future studies.
Abnormal and additional chromosomes may have consequences for disease outcome due to changes in gene expression. Compared to MBU-7, MBU-8 had an additional copy of chromosome 18, two additional copies of the p-arm of chromosome 8 but lacked part of a third copy of chromosome 7. We wondered whether these differences in chromosomes would be associated with differences in the response to drugs. Interestingly, our results showed a clear difference between MBU-7 and MBU-8 in the response towards the nucleoside analogues azacitidine, decitabine and cytarabine, with MBU-7 being significantly more sensitive. In this respect, it is interesting to note that 20% of MDS patients that relapsed after decitabine- treatment had acquired additional cytogenetic abnormalities including trisomy 847. Taken together, these observations might indicate a possible link between gains in chromosome 8 and the response of cells to decitabine and related drugs.
Overexpression of the BCL2 gene on chromosome 18 contributes to MDS by blocking cell death48. Even though MBU-7 and MBU-8 differed in the copy number of the BCL2 gene, they were equally sensitive to the Bcl2-inhibitor venetoclax. From this observation we draw two conclusions: First, MBU-7 cells do not have a general hypersensitivity to treatments that induce cell death. Second, gene copy number of drug targets is not a reliable predictor of drug response. Potential synthetic lethality needs to be assessed on the functional level. The combination of well-characterized cell lines with powerful genetic loss of function screens has the potential to identify novel vulnerabilities associated with distinct genetic and cytogenetic abnormalities in close to unbiased manner49.
We propose the here described cell lines MBU-7 and MBU-8 as suitable models for screening for vulnerabilities that are associated with the disease-relevant gains in chromosome 7, 8 and 18.
Methods
Cell lines and cell culture
The human cell lines U-937 and SKK-1 were obtained from the Leibniz-Institute DSMZ—German Collection of Microorganisms and Cell Cultures (Braunschweig, Germany). We obtained SKK-1 cells as part of a collaboration with Prof. Hans Drexler that included their detailed cytogenetic and genetic characterization50. MBU-7 and MBU-8 have been isolated as single-cell clones and are described here in great detail. Both cell lines will be deposited at DSMZ. All cell lines were cultured in RPMI 1640 (ThermoFisher Scientific, Waltham, MA) supplemented with 10%, 1% Pen/Strep, and 1% l-glutamine at 37 °C in 5% CO2. All cell lines were routinely analysed for the presence of mycoplasma.
RT-qPCR
For real-time quantitative PCR (RT-qPCR), RNA was extracted using the Maxwell RSC simplyRNA Cells Kit (Promega, Madison, Wisconsin) and reverse transcribed with the First strand cDNA synthesis kit (ThermoFisher Scientific). The cDNA was PCR amplified in triplicate using the Fast SYBR green dye on the Applied Biosystems QuantStudio 7 Flex Real-Time PCR System. Relative expression was determined using SKK-1 or U-937 as reference samples, and GAPDH as internal control. The sequences of primers were: GAPDH, forward primer 5′-CGACCACTTTGTCAAGCTCA-3′, reverse primer 5′-TCTTACTCCTTGGAGGCCAT-3′; ETV6-NTRK3, forward primer 5′-CATTCTTCCACCCTGGAAAC-3′, reverse primer 5′-GGCTCCCTCACCCAGTTCTC-3′; PICALM-MLLT10, forward primer 5′-TGAGACCTCCAAACCCCTTT-3′, reverse primer 5′-TCGGCACCATTACCTTCTTC-3′.
Identifier test
Cells were identified by comparing their short tandem repeat (STR) profiles to their corresponding profiles on the searchable cell line database, Cellosaurus (web.expasy.org/cellosaurus). Genomic DNA was extracted from collected cells using PureLink Genomic DNA Mini Kit (ThermoFisher Scientific) and processed by the Genomics facility (IGTP) using the CLA IdentiFiler Plus PCR Amplification kit (ThermoFisher Scientific) according to the manufacturer’s instructions. Following the PCR, capillary electrophoresis was performed with the amplified microsatellite loci fragments and the allelic ladder, GeneScan 500 LIZ Size Standard, using the Applied Biosystems 3130xl Genetic Analyzer to discriminate alleles that differ by single nucleotides. GeneMapper software (ThermoFisher Scientific) was then used to call alleles according to size and quality based on the selected allelic ladder.
Cytogenetics
Cytogenetic studies, karyotype and fluorescent in situ hybridization (FISH), were carried out by standard procedures in single cell expanded clones of MBU-7 and MBU-8, at the Cytogenetics Platform, Josep Carreras Leukaemia Research Institute and Institut Català d'Oncologia (ICO).
Karyotyping
Conventional G-banding metaphases were obtained by standard procedures. In brief, mitotic cells were arrested in metaphase using Colcemid, lysed in hypotonic solution (KCl, 0.075 M), the metaphases fixed with Carnoy’s solution (3 parts of methanol: 1 part of acid acetic glacial) and the G band pattern achieved with Wright dye. Karyotypes for 20 metaphases were analysed for each cell line and described following the International System for Human Cytogenetic Nomenclature (ISCN 2020)51.
Fluorescence in situ hybridization (FISH)
Cells were fixed with Carnoy’s solution and the samples and 5 µL probe simultaneously denatured by heating the slides on a hotplate at 75 °C (2 min). The sides were then incubated in a humidified chamber at 37 °C overnight, washed in 0.4 × SSC (pH 7.0) at 72 °C (2 min), washed in 2 × SSC, 0.05% Tween-20 (pH 7.0) at room temperature (30 s), rinsed in distilled water to avoid crystal formation, air dried. Finally, the cells were counterstained with DAPI and analysed under a fluorescent microscope. Different probes were used to detect several chromosomal abnormalities in MBU-7 and MBU-8. The XL 7q22/7q36 Deletion Probe (MetaSystems, Germany) detects deletions in the long arm of chromosome 7. The IGH/MYC/CEP 8 tri-color dual fusion probe (Abbott, USA) detects the t(8;14)(q24;q32) reciprocal translocation involving the IGH and MYC gene regions. The BCL2 break apart probe (Abbott, USA) detects chromosomal rearrangements at the BCL2 locus on chromosome 18q21. 500 cells per probe have been analysed using the Metafer platform (MetaSystems). The standardised cut-off for probes analyses with this automated system was > 10%.
Cytoscan array
Genomic DNA was extracted from collected cells using PureLink Genomic DNA Mini Kit (ThermoFisher Scientific) and processed by the Microarray Unit of the IJC. Genomic microarrays were performed with CytoScan 750 K Array from ThermoFisher. Standard protocol and QC guidelines supplied by the manufacturer were followed. The Affymetrix 450 fluidics station and GeneChip Scanner 3000 7G were used to wash, stain and scan the arrays. SNP-array data analysis was performed with Chromosome Analysis Suite Version 4.2 (ThermoFisher Scientific). Data was analyzed using annotations of genome version GRCh37 (hg19). Detailed visual data analysis was performed in all samples, in addition to software-reported alterations. Only copy number (CN) segments with a minimum of 20 SNP/CN-altered markers. Furthermore, germline abnormalities were excluded by comparing findings with control Databases. SNP array data has been deposited in the GEO database under accession number GSE184764.
Cytology
Cells were immobilized on glass slides at a concentration by cytospin centrifugation (ThermoFisher Scientific) at 300 rpm for 10 min. Once dried, cells were stained with May-Grünwald Giemsa solution (Merck, New Jersey). Cells were further stained with Myeloperoxidase, alpha-naphthyl butyrate esterase and alpha-naphthyl acetate esterase. Cells were mounted with DPX Mountant for histology (Sigma-Aldrich, Misuri) and examined using the microscope Olympus AX70 TRF.
Immunophenotypic analysis
Immunostaining and flow cytometry analyses were performed at the Clinical Haematology Department at ICO-Hospital Germans Trias i Pujol according to standard procedures with the Navios cytometer (Beckman Coulter, California). The same antibodies are used for diagnostic purposes (see Supplementary Table S2).
NGS targeted gene panel
Targeted NGS including 32 genes related to myeloid disorders was used. Briefly, DNA was extracted from cell lines and library was prepared using the QIAseq FX DNA library kit (Qiagen, Germany). Target regions were captured and amplified by hybridizing the DNA library with the Myeloid Solution Capture Kit (Sophia Genetics, Switzerland) and, finally, sequenced using MiSeq platform (Illumina, California). The results were analyzed using Sophia DDM platform25,26 and interpreted using the classification described in Sukhai et al.52.
Flow cytometric analysis of viability and apoptosis
Following 4 days’ incubation with the indicated inhibitors, cell viability was assessed by flow cytometry of cells stained with 1 μg/mL DAPI (4′,6-Diamidino-2-phenylindole dihydrochloride) (ThermoFisher Scientific), and 100 μM MitoTracker Red CMXRos (ThermoFisher Scientific), using the LSR Fortessa cytometer (BD, New Jersey). Statistical analysis (ANOVA test) was performed using GraphPad Prism software (version 6).
Data availability
The CytoScan array data generated in this study have been deposited in the GEO database under accession code GSE184764. Reviewer access: GSE184764: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE184764. Password: yrmvqicerjulbel.
References
Maher, M., Diesch, J., Le Pannérer, M.-M. & Buschbeck, M. Epigenetics in a spectrum of myeloid diseases and its exploitation for therapy. Cancers https://doi.org/10.3390/cancers13071746 (2021).
Mrózek, K., Heerema, N. A. & Bloomfield, C. D. Cytogenetics in acute leukemia. Blood Rev. 18, 115–136. https://doi.org/10.1016/S0268-960X(03)00040-7 (2004).
Rabbitts, T. H. Chromosomal translocations in human cancer. Nature 372, 143–149. https://doi.org/10.1038/372143a0 (1994).
Daneshbod, Y., Kohan, L., Taghadosi, V., Weinberg, O. K. & Arber, D. A. Prognostic significance of complex karyotypes in acute myeloid leukemia. Curr. Treat. Opt. Oncol. 20, 15–15. https://doi.org/10.1007/s11864-019-0612-y (2019).
Swerdlow, S. H. et al. WHO Classification of Tumours of Haematopoietic and Lymphoid Tissues, vol. 2 4th ed. 188. (International Agency for Research on Cancer, 2017).
Lagunas-Rangel, F. A., Chávez-Valencia, V., Gómez-Guijosa, M. & Cortes-Penagos, C. Acute myeloid leukemia-genetic alterations and their clinical prognosis. Int. J. Hematol.-Oncol. Stem Cell Res. 11, 328–339 (2017).
Jing, Y. The PML-RARα fusion protein and targeted therapy for acute promyelocytic leukemia. Leuk. Lymphoma 45, 639–648. https://doi.org/10.1080/10428190310001609933 (2004).
List, A. et al. Lenalidomide in the myelodysplastic syndrome with chromosome 5q deletion. N. Engl. J. Med. 355, 1456–1465. https://doi.org/10.1056/NEJMoa061292 (2006).
Vali, S. et al. Predictive analysis on prognostic impact of monosomy 7 in AML and identified therapy options for this cohort. Blood 132, 1539–1539. https://doi.org/10.1182/blood-2018-99-116603 (2018).
Locatelli, F. & Strahm, B. How I treat myelodysplastic syndromes of childhood. Blood 131, 1406–1414. https://doi.org/10.1182/blood-2017-09-765214 (2018).
Kapoor, A. et al. The histone variant macroH2A suppresses melanoma progression through regulation of CDK8. Nature 468, 1105–1109. https://doi.org/10.1038/nature09590 (2010).
Cantariño, N. et al. A cellular model reflecting the phenotypic heterogeneity of mutant HRAS driven squamous cell carcinoma. Int. J. Cancer 139, 1106–1116. https://doi.org/10.1002/ijc.30139 (2016).
Kennedy, J. A. & Barabé, F. Investigating human leukemogenesis: From cell lines to in vivo models of human leukemia. Leukemia 22, 2029–2040. https://doi.org/10.1038/leu.2008.206 (2008).
MacLeod, R., Dirks, W. G., Reid, Y. A., Hay, R. J. & Drexler, H. G. Identity of original and late passage Dami megakaryocytes with HEL erythroleukemia cells shown by combined cytogenetics and DNA fingerprinting. Leukemia 11, 2032–2038. https://doi.org/10.1038/sj.leu.2400868 (1997).
Steensma, D. P. et al. Clonal hematopoiesis of indeterminate potential and its distinction from myelodysplastic syndromes. Blood 126, 9–16. https://doi.org/10.1182/blood-2015-03-631747 (2015).
Ben-David, U. et al. Genetic and transcriptional evolution alters cancer cell line drug response. Nature 560, 325–330. https://doi.org/10.1038/s41586-018-0409-3 (2018).
Fregeau, C. J. & Fourney, R. M. DNA typing with fluorescently tagged short tandem repeats—A sensitive and accurate approach to human identification. Biotechniques 15, 100 (1993).
Bairoch, A. The cellosaurus, a cell-line knowledge resource. J. Biomol. Tech. JBT 29, 25–38. https://doi.org/10.7171/jbt.18-2902-002 (2018).
Sundström, C. & Nilsson, K. Establishment and characterization of a human histiocytic lymphoma cell line (U-937). Int. J. Cancer 17, 565–577. https://doi.org/10.1002/ijc.2910170504 (1976).
Dreyling, M. H. et al. The t(10;11)(p13;q14) in the U937 cell line results in the fusion of the AF10 gene and CALM, encoding a new member of the AP-3 clathrin assembly protein family. Proc. Natl. Acad. Sci. U. S. A. 93, 4804–4809. https://doi.org/10.1073/pnas.93.10.4804 (1996).
Zhang, Y. M. & Rowley, J. D. Chromatin structural elements and chromosomal translocations in leukemia. DNA Repair 5, 1282–1297. https://doi.org/10.1016/j.dnarep.2006.05.020 (2006).
Laforêt, M. P. et al. Design and feasibility of a novel, rapid, and simple fluorescence 26-plex RT-PCR assay for simultaneous detection of 24 fusion transcripts in adult acute myeloid leukemia. J. Mol. Diagn. 15, 186–195. https://doi.org/10.1016/j.jmoldx.2012.11.004 (2013).
Strefford, J. C. et al. The characterisation of the lymphoma cell line U937, using comparative genomic hybridisation and multi-plex FISH. Cytogenet. Genome Res. 94, 9–14. https://doi.org/10.1159/000048774 (2001).
Jawhar, M. et al. Cytogenetically cryptic ZMYM2-FLT3 and DIAPH1-PDGFRB gene fusions in myeloid neoplasms with eosinophilia. Leukemia 31, 2271–2273. https://doi.org/10.1038/leu.2017.240 (2017).
Espasa, A. et al. Chronic myelomonocytic leukemia and blastic plasmacytoid dendritic cell neoplasm. A case report and systematic review. Cytom. Part B Clin. Cytom. 100, 292–295. https://doi.org/10.1002/cyto.b.21932 (2021).
Huguet, M. et al. Polycythemia vera evolution to chronic myelomocytic leukemia: The prognostic value of next generation sequencing. HemaSphere 4, e466. https://doi.org/10.1097/hs9.0000000000000466 (2020).
Sugimoto, K. et al. Frequent mutations in the p53 gene in human myeloid leukemia cell lines. Blood 79, 2378–2383 (1992).
Baird, P. N. & Simmons, P. J. Expression of the Wilms’ tumor gene (WT1) in normal hemopoiesis. Exp. Hematol. 25, 312–320 (1997).
Chen, L. et al. Mutated Ptpn11 alters leukemic stem cell frequency and reduces the sensitivity of acute myeloid leukemia cells to Mcl1 inhibition. Leukemia 29, 1290–1300. https://doi.org/10.1038/leu.2015.18 (2015).
Drexler, H.G. et al. Guide to Leukemia-Lymphoma Cell Lines on CD. Blood 106. 4340–4340. https://doi.org/10.1182/blood.V106.11.4340.4340. (2005)
MacKinnon, R. N., Peverall, J., Campbell, L. J. & Wall, M. Detailed molecular cytogenetic characterisation of the myeloid cell line U937 reveals the fate of homologous chromosomes and shows that centromere capture is a feature of genome instability. Mol. Cytogenet. 13, 50. https://doi.org/10.1186/s13039-020-00517-y (2020).
Harris, P. & Ralph, P. Human leukemic models of myelomonocytic development: A review of the HL-60 and U937 cell lines. J. Leukoc. Biol. 37, 407–422. https://doi.org/10.1002/jlb.37.4.407 (1985).
Galmarini, C. M., Jordheim, L. & Dumontet, C. Pyrimidine nucleoside analogs in cancer treatment. Expert Rev. Anticancer Ther. 3, 717–728. https://doi.org/10.1586/14737140.3.5.717 (2003).
Diesch, J., Zwick, A., Garz, A.-K., Buschbeck, M. & Götze, K. S. A clinical-molecular update on azanucleoside-based therapy for the treatment of hematologic cancers. Clin. Epigenet. https://doi.org/10.1186/s13148-016-0237-y (2016).
Webster, J. & Pratz, K. Acute myeloid leukemia in the elderly: Therapeutic options and choice. Leuk. Lymphoma 59, 1–14. https://doi.org/10.1080/10428194.2017.1330956 (2017).
Pan, R. et al. Selective BCL-2 inhibition by ABT-199 causes on-target cell death in acute myeloid leukemia. Cancer Discov. 4, 362–375. https://doi.org/10.1158/2159-8290.cd-13-0609 (2014).
Stacey, G. N., Bolton, B. J., Morgan, D., Clark, S. A. & Doyle, A. Multilocus DNA fingerprint analysis of cellbanks: Stability studies and culture identification in human B-lymphoblastoid and mammalian cell lines. Cytotechnology 8, 13–20. https://doi.org/10.1007/BF02540025 (1992).
Lee, J. Y. et al. Molecular cytogenetic analysis of the monoblastic cell line U937. karyotype clarification by G-banding, whole chromosome painting, microdissection and reverse painting, and comparative genomic hybridization. Cancer Genet. Cytogenet. 137, 124–132. https://doi.org/10.1016/s0165-4608(02)00565-4 (2002).
Parson, W. et al. Cancer cell line identification by short tandem repeat profiling: power and limitations. FASEB J. 19, 1–18. https://doi.org/10.1096/fj.04-3062fje (2005).
Grimwade, D. et al. Refinement of cytogenetic classification in acute myeloid leukemia: Determination of prognostic significance of rare recurring chromosomal abnormalities among 5876 younger adult patients treated in the United Kingdom Medical Research Council trials. Blood 116, 354–365. https://doi.org/10.1182/blood-2009-11-254441 (2010).
Hemsing, A. L., Hovland, R., Tsykunova, G. & Reikvam, H. Trisomy 8 in acute myeloid leukemia. Expert Rev. Hematol. 12, 947–958. https://doi.org/10.1080/17474086.2019.1657400 (2019).
Paulsson, K. & Johansson, B. Trisomy 8 as the sole chromosomal aberration in acute myeloid leukemia and myelodysplastic syndromes. Pathol. Biol. (Paris) 55, 37–48. https://doi.org/10.1016/j.patbio.2006.04.007 (2007).
Ribeiro, A. F. T. et al. Mutant DNMT3A: A marker of poor prognosis in acute myeloid leukemia. Blood 119, 5824–5831. https://doi.org/10.1182/blood-2011-07-367961 (2012).
Chou, W. C. et al. TET2 mutation is an unfavorable prognostic factor in acute myeloid leukemia patients with intermediate-risk cytogenetics. Blood 118, 3803–3810. https://doi.org/10.1182/blood-2011-02-339747 (2011).
Chou, W.-C. et al. Distinct clinical and biological features of de novo acute myeloid leukemia with additional sex comb-like 1 (ASXL1) mutations. Blood 116, 4086–4094. https://doi.org/10.1182/blood-2010-05-283291 (2010).
Tsirigotis, P. et al. Acute myelogenous leukemia with tetrasomy 8 is a disease with a poor prognosis. Cancer Genet. Cytogenet. 161, 78–81. https://doi.org/10.1016/j.cancergencyto.2004.12.016 (2005).
Qin, T. et al. Mechanisms of resistance to decitabine in the myelodysplastic syndrome. PLoS One. https://doi.org/10.1371/journal.pone.0023372 (2011).
Parker, J. E. et al. The role of apoptosis, proliferation, and the Bcl-2-related proteins in the myelodysplastic syndromes and acute myeloid leukemia secondary to MDS. Blood 96, 3932–3938. https://doi.org/10.1182/blood.V96.12.3932 (2000).
Huang, A., Garraway, L. A., Ashworth, A. & Weber, B. Synthetic lethality as an engine for cancer drug target discovery. Nat. Rev. Drug Discov. 19, 23–38. https://doi.org/10.1038/s41573-019-0046-z (2020).
Palau, A. et al. Immunophenotypic, cytogenetic, and mutational characterization of cell lines derived from myelodysplastic syndrome patients after progression to acute myeloid leukemia. Genes Chromosomes Cancer 56, 243–252. https://doi.org/10.1002/gcc.22430 (2017).
Nomenclature, I. S. C. O. H. C., McGowan-Jordan, J., Hastings, R. J. & Moore, S. ISCN 2020: An International System for Human Cytogenomic Nomenclature (2020). (Karger, 2020).
Sukhai, M. A. et al. A classification system for clinical relevance of somatic variants identified in molecular profiling of cancer. Genet. Med. 18, 128–136. https://doi.org/10.1038/gim.2015.47 (2016).
Acknowledgements
We would like to thank Elisa Orna Montero for helping with the cytology images; all the members of the Cytogenetics Platform, Haematology Laboratory Department at Germans Trias i Pujol Hospital and Catalan Institute of Oncology for providing the karyotype techniques and images; Raquel Casquero and other members of the Buschbeck lab for their valuable help and discussions; the facilities of IGTP and IJC for providing excellent technical support.
Funding
This project was supported by the FEDER/Ministerio de Ciencia e Innovación—Agencia Estatal de Investigación through the grant PIE16/00011 RESPONSE (to MB), the grant Sara Borrell CD17/00084 (to JD), and the Deutsche José Carreras Leukämie Stiftung DJCLS 14R/2018 (to MB). MM was a fellow through the Marie Skłodowska Curie Training network ‘ChroMe’ H2020-MSCA-ITN-2015-675610 (awarded to MB). Research in the Buschbeck lab is further supported by the following grants: MINECO Grant RTI2018-094005-B-I00 (to MB), the Marie Skłodowska Curie Training network ‘INTERCEPT-MDS’ H2020-MSCA-ITN-2015-953407 (to MB); AGAUR 2017-SGR-305 (to MB) and Fundació La Marató de TV3 257/C/2019 (to MB). Research at the IJC is generously supported by the ‘La Caixa’ Foundation, the Fundació Internacional Josep Carreras, Celgene Spain and the CERCA Programme/Generalitat de Catalunya.
Author information
Authors and Affiliations
Contributions
Conceptualization: M.M. and M.B. Data curation: M.M., J.D., Ma.M. Funding acquisition: M.B. Performed the experiments: J.D., M.M.L.P., M.M., M.C., S.V., A.M.T., I.G., A.M.L. Analysed the data: A.M.T., M.S., L.Z., I.G., Ma.M., K.S., A.M.L. Project administration: M.B. Writing—original draft: M.M., J.D. and M.B. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Maher, M., Diesch, J., Le Pannérer, MM. et al. Divergent leukaemia subclones as cellular models for testing vulnerabilities associated with gains in chromosomes 7, 8 or 18. Sci Rep 11, 21145 (2021). https://doi.org/10.1038/s41598-021-00623-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-00623-w
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.