Abstract
Theory predicts that self-sustained oscillations require robust delays and nonlinearities (ultrasensitivity). Delayed negative feedback loops with switch-like inhibition of transcription constitute the core of eukaryotic circadian clocks. The kinetics of core clock proteins such as PER2 in mammals and FRQ in Neurospora crassa is governed by multiple phosphorylations. We investigate how multiple, slow and random phosphorylations control delay and molecular switches. We model phosphorylations of intrinsically disordered clock proteins (IDPs) using conceptual models of sequential and distributive phosphorylations. Our models help to understand the underlying mechanisms leading to delays and ultrasensitivity. The model shows temporal and steady state switches for the free kinase and the phosphoprotein. We show that random phosphorylations and sequestration mechanisms allow high Hill coefficients required for self-sustained oscillations.
Similar content being viewed by others
Introduction
Life on earth in forms of cyanobacteria, algae , fungi, plants and animals has evolved 24 h periodicities called circadian clocks. This helps them to anticipate rhythmic environmental cues such as light, temperature and nutrients1,2,3,4,5. Circadian clocks regulate a wide variety of molecular and physiological processes6,7,8,9.
Circadian oscillators are based on a transcription-translation feedback loops (TTFLs). A delayed negative feedback loop is central to the gene regulatory network10,11,12. For example, the negative feedback loop of the fungal clock contains the negative element FREQUENCY [FRQ], which inhibits its own expression via inhibition of the circadian transcription factor White Collar Complex (WCC). FRQ is an intrinsically disordered protein (IDP) progressively hyperphosphorylated mainly by CK1a (Casein Kinase 1a). Hyperphosphorylation eventually leads to functional inactivation and degradation of FRQ allowing the WCC to reinitiate a new cycle13,14. This design principle is conserved in other frequently used model systems. Fruit flies and animal clocks are also made up of the inhibitors, kinases and transcriptional activators. In mammalian clocks, PERIOD proteins [PER1, PER2, PER3], and CRYPTOCHROME proteins [CRY1, CRY2]) inhibit their own expression. PER2 is also an IDP as FRQ and is phosphorylated by CK115,16,17.
Recent experiments suggest that many sites on both FRQ (about 100 sites) and PER2 (about 60 sites) are phosphorylated over the course of many hours at seemingly random manner18,19,20,21. Moreover their respective activators also get phosphorylated15,22,23. Phosphorylations govern nuclear translocation24, complex formations25,26, inactivation of transcription and stability27,28.
Oscillator theory predicts that self-sustained circadian clocks require long delays and nonlinearities such as switches29,30. This raises two theoretical questions: (1) How can multiple random phosphorylations produce long delays? (2) What are the underlying switch mechanisms?
According to mathematical theory a delay of negative feedback loops governs the period29,31. Under quite general assumptions the delay is in the range between a 1/4 and 1/2 of the oscillator period32,33. If gene expression and inhibitor formation last about an hour, periods of 2 to 4 h can be expected. Indeed, several TTFLs exhibit periods of a few hours including somite formation34, NF-kB rhythms35,36 and p53 pulses37. Circadian rhythms have much longer periods of about 24 h. This implies that the associate delays last at least 6 h. Several processes like transcription, translation, nuclear transport, post-translational modifications, mRNA decay, and proteasomal degradation may contribute to the needed delay in circadian rhythms15,24. It has been suggested that also the multiple phosphorylations contribute significantly to the required delay19,23,38,39.
In order to generate self-sustained rhythms (“limit cycles”) nonlinearities are necessary in addition to delays40,41. In many models switch-like inhibitions are postulated42,43. Here we explore how multiple random phosphorylation contribute to the generation of switch-like behaviour.
The role of multiple phosphorylations has been studied earlier for cell cycle dynamics44, in signaling45,46,47,48, and chronobiology49,50. Antagonistic enzymes may lead to zero-order ultrasensitivity51. Sequestration effects may enhance ultrasensitivity47,52. The identification of up to 100 phosphorylation sites on clock proteins motivated us to revisit the role of multiple random phosphorylations. Since quantitative details are not available we study conceptional models of processive and distributive phosphorylations with a focus on sequestration and amplitudes.
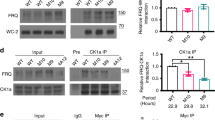
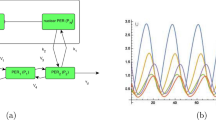
Figure 1 depicts the assumptions underlying our modelling approach: There is clear experimental evidence for a slow and random phosphorylation of the clock phosphoproteins18,28,38,53,54. Figure 1A depicts gradual phosphorylation in Neurospora. FRQ is stabilized by the FRQ-interacting RNA Helicase (FRH) and forms the FFC complex with the kinase CK1a55. Moreover, FFC interact with the transcription factor WCC13,56. FRQ undergoes slow, seemingly random multiple phosphorylations predominantly by CK1a28,38,57,58. At an overcritical phosphorylation level the complex gets inactivated in a switch-like manner23,27,59,60.
Since detailed kinetic data of these processes are missing we explore in generic models how multiple random phosphorylations can reproduce long robust delays and switch-like behaviour. Figure 1B illustrates our conceptional modelling approach: We simulate slow saturated phosphorylation (solid line) leading to a switch-like inactivation at critical phosphorylation levels.
Phosphorylation hourglass and activity switch of phosphoproteins: (A) As an example, the FRQ protein is stabilized by FRH and undergoes multiple phosphorylations by CK1a. FFC complex interacts with WCC. A phosphoswitch governs the activity status of the complex. (B) The solid line represents the simulated phosphorylations in Fig. 7 and suggests a slow and saturated phosphorylation. The dashed line represents the simulated switch induced by hyperphosphorylated protein (compare Fig. 7).
Results
Linear processive phosphorylations provides delays
As discussed above, PER2 and FRQ are core clock phosphoproteins with up to 100 phosphorylation sites28,53. Recent in vitro and in vivo experiments show that in Neurospora about 100 FRQ sites are phosphorylated over more than a circadian day (up to 48 h) in a seemingly random manner18. However the detailed functions of increasing phosphorylation levels in circadian timekeeping are not well understood.
Inspired by these observations, we introduce conceptual models of multiple phosphorylations. We denote the simulated clock phosphoprotein by “F” and the associated kinase by “C”. For simplicity, we start with just four phosphorylation sites. In Suppl. A1 we show the corresponding reaction scheme and the associated linear differential equations describing protein turnover and processive phosphorylation. We do not explicitly consider phosphatases throughout. Phosphatases are found to be permanently available in cells and it is basically the ratio of kinase and phosphatase which takes the reaction forward61,62. It is known that phosphatases are tightly regulated, have specificity63 and that PP1 plays a critical role for the mammalian clock64. Furthermore, the competition of kinase and phosphatase could generate zero-order ultrasensitivity51,65,66. For simplicity, we focus in our paper on kinase dynamics as a first step. Fig. 2 shows the increasing levels of different phosphorylations in the linear model. Note that we introduce just three parameters: a production Pp, phosphorylation rate k and a degradation rate kd. We have chosen parameter values that reproduce the measured full phosphorylation after about 48 h18,54. Thus we can focus on delays, amplitudes and waveforms quantified by fits of Hill-functions (see “Calculation of Hill coefficients and delays” in Materials and methods).
The curves for phosphoprotein F exhibited delays and an accumulation of fully phosphorylated F4 in a switch-like manner (Fig. 2). Therefore even linear phosphorylations alone can provide delays. A delayed switch based on multiple phosphorylations can serve as the basic hourglass mechanism hypothesized in Fig. 1. Note, that these steps required no explicit nonlinearity in the model.
Modeling a delayed switch based on multiple phosphorylations: The simulations start with unphosphorylated protein F. The normalized total number of Fk with \( \hbox {k}=1,\ldots ,4 \) phosphorylations increases with time delays and hyperphosphorylated species (such as \(\hbox {n} = 4\)) accumulate in an ultrasensitive, switch-like manner.
Nonlinear models of distributive phosphorylations enhance ultrasensitivity
Kinases bind to substrates and could phosphorylate sites while staying bound (processive mechanism). Alternatively, the kinase may bind and unbind, so that next phosphorylation first requires rebinding of a kinase molecule (distributive mechanism)61. Distributive enzyme kinetics may lead to ultrasensitive responses in protein phosphorylations65,67. Note that in these studies ultrasensitivity is quantified using input-output relations. Typically steady states of phosphorylation levels are studied as a function of ligand concentrations or kinase levels. Motivated by our temporal switch in Fig. 1, we focus on the ultrasensitive increase of phosphorylation with time. In this section we include the formation and dissociation of FC complexes (the complexes of protein F and bound kinase C) in order to study the role of enzyme sequestration.
Our models are motivated by experimental observations regarding multiple slow and partly random phosphorylations of clock proteins18,54. We emphasize, however, that the following model versions do not reflect the full complexity of clock protein interactions as phosphorylations can affect stability and complex formation in a complicated manner25,27. If the kinase stays bound to the substrate it is termed processive phosphorylation (see Suppl. A1). Dissociation of C and rebinding to another site could be represented as distributive mechanism.
The simulations below are somewhat simplistic and describe generic substrate-enzyme dynamics. For example, we start simulations with low levels of F and of recruited enzyme C. Note, that kinases under in vivo conditions are constitutively expressed. In Suppl. A9 we illustrate that the overall kinetics for constitutive kinase expression is comparable. Our simulations in Fig. 4 of gradually increasing C allow to study effects of varying kinase levels in a single simulation. Kinase C can be recruited to unphosphorylated and partially phosphorylated species of protein F (F0, F1, F2 and F3). The bound kinase can then be sequestered (F1C, F2C,..) which slows down the progressive phosphorylation kinetics. Eventually F is further phosphorylated and C dissociates, yielding the next phospho-species.
Suppl. A2 shows the turnover of F (Fk) and non-sequestered and sequestered FC complexes, C and FkC, respectively. Using mass-action kinetics, this scheme is directly translated into a system of nonlinear ordinary differential equations (ODEs)68,69,70. The equations describe 10 time-varying concentrations of (phosphorylated) F, complexes with the kinase and turnover of C. The 5 kinetic parameters in the ODEs are enumerated as Pp and Pc, k1, k2 and kd (see Suppl. A2) respectively for production of protein and kinase, rates of phosphorylation and degradation.
Figure 3 shows the production, dissociation and phosphorylation of F and the formation of FC complexes starting at zero levels. F0 denotes the unphosphorylated F protein whereas F1, F2, F3, F4 represent increasing phosphorylation levels. Ct and C denote the amounts of total and free kinase, respectively. The phosphorylated F species accumulate in a sigmoidal manner with the expected delays. There is an initial sharp peak of free enzyme C but later most of the enzyme molecules are “sequestered”. Suppl. A5 shows the Hill coefficients for phosphorylated Fk (F0...F4) calculated along the lines of47,71. In comparison to linear phosphorylations shown in Fig. 2, nonlinear phosphorylations provide higher Hill coefficients. Note, however, that the amplitudes decrease drastically for higher phosphorylation levels.
Sequestration can generate overshoots and threshold behaviour
It was shown in the preceding section that sequestration can enhance switch-like behaviour. In Fig. 3, the simulated amount of free enzyme was relatively small compared to the phosphoprotein F and thus most of the enzyme was sequestered. It is known, however, that kinase levels are quite high in Neurospora72. Consequently, we study here the effects of enlarged kinase levels. It turns out that higher kinase levels lead to overshoots and a sharp threshold.
In Fig. 4 the levels of phosphorylated proteins F1, F2, and F3 increase initially as in Fig. 3. After about 10 h, however, they decay to quite small values. In parallel, the fully phosphorylated protein F4 reaches high levels (see Fig. 4B). If full phosphorylation reaches saturation the amount of free kinase is increasing suddenly. Magnifications reveal that the apparent “kinks” in the time-courses are smooth curves (see Suppl. A6).
Note, that Figs. 3 and 4 refer both to our model described in Suppl. A2. The drastic differences are simply due to fivefold increase of enzyme production. The overshoot in Fig. 4 reflects the fast initial production of F0 and phosphorylation of F1, F2 and F3. Later an equilibrium is reached with lower levels of intermediate phosphorylations.
In summary, for higher kinase levels threshold behaviour arises reflecting the sequestration of enzymes by different species of phosphorylated proteins. Note, that the initial sharp increase of free protein levels is based on our somewhat artificial initial condition of zero enzyme levels. In vivo the equilibrium between free and bound proteins is reached more quickly due to the omnipresence of kinases.
Simulating nonlinear phosphorylations at higher enzyme levels: (A,B) show time-courses of phosphorylated F proteins. Graph (C) shows the concentrations of FC complexes over time. (D) shows the temporal switch of free kinase C after release from sequestration. The dynamics of total kinase Ct, not shown here, is similar as in Fig 3.
Steady state switches due to increasing enzyme levels
So far, we have characterized in Figs. 3 and 4 two specific values of enzyme production Pc (marked by arrows in Fig. 5). Figure 5 illustrates that a systematic variation of enzyme levels induces nonlinear dependencies including transitions between different regimes. For low values of Pc we get power-law increases of free kinase (exponent \(=2.85\)) and of sequestered protein (exponent \(= 4.29\)). Note, that the model contains just bilinear terms. Thus, sequential phosphorylation and sequestration leads to higher exponents as described earlier in other systems67,71,73. As discussed above the generation of such input-output switches was studied extensively over the last decades66,74,75. Even though our model can reproduce such nonlinearities, our focus are temporal switches (compare Figs. 1 to 4) relevant for circadian rhythm generation.
Inspecting Fig. 5, we find that at \(\hbox {Pc} = 3\) sequestration by increasing protein F levels flattens the curve of free kinase due to sequestration. Beyond \(\hbox {Pc} = 4\) (white part of the graph) the amount of sequestered protein starts to decline and the release of enzyme leads to more free kinase C.
In summary Fig. 5 shows that our nonlinear model generates, in addition to delayed temporal switches and threshold behaviour, interesting nonlinearities of steady states governed by sequestration.
Random phosphorylations provide large amplitudes of intermediate phosphorylations
In signaling cascades many phosphorylation sites carry specific functions for activations or complex formations. Up to 100 phosphorylations of intrinsically disordered proteins (IDPs) such as FRQ or PER2 might control cellular processes differently15. FRQ proteins have a positively charged N-terminal part and a negatively charged C-terminal part. Initial phosphorylations appear to stabilize a closed conformation whereas progressive hyperphosphorylations favours an open conformation potentially via charge repulsion60. Thus, the overall number of phosphorylated sites can govern stability, complex formation or nuclear translocation18,58.
There is some evidence that FRQ is phosphorylated by CK1a in a seemingly random opportunistic manner18,53. Hypothetically, the highly flexible FRQ protein can form almost random contacts to the active site of CK1a allowing phosphorylation72. Consequently we simulate now instead of processive phosphorylations (compare Suppl. A1) random phosphorylations as shown in Suppl. A3. Note, that for up to 100 phosphorylations the number of differently phosphorylated molecules is astronomically large and exceeds the number of FRQ molecules in a cell and even the number of bacteria on earth76.
In order to keep simulations with differential equations feasible we introduce new variables F1, F2, etc. These variables lump all F molecules with 1, 2, ... phosphorylations. In other words, Fk represents the \(\left( {\begin{array}{c}n\\ k\end{array}}\right)\) molecules with k out of n phosphorylations. These lumped variables seem reasonable since complex formation of clock proteins are influenced by the number of phosphorylations18,25. For example, an overcritical number of phosphorylations destabilizes the FFC complex and makes FRQ accessible to the E3 ligase FWD1 (F-box/WD-40 repeat-containing protein-1)18,77,78.
Our resulting nonlinear random model (see Suppl. A4) contains altogether just 2n+2 differential equation making even simulations with \(\hbox {n}=100\) phosphorylations feasible. For a direct comparison with the previous models we show in Fig. 6 simulations for \(\hbox {n}=4\). It turns out that the delayed switch with Hill coefficient around 3 are found in these simulations as well (compare Suppl. A5). Figure 6 displays a new feature of random models—the amplitudes of intermediate states do not decay monotonously. For instance, F1 and F2 have fairly high levels. This property of random models can be traced back to prefactors in the lumped equations. In other words, the combinatorial explosion of molecule types with intermediate phosphorylation numbers enhances the growth of certain Fk levels.
Figure 7 shows representative time-courses of a random model with up to 100 phosphorylations. It turns out that we find delayed temporal switches with high amplitudes in particular at intermediate phosphorylation levels.
Large-scale simulations of nonlinear, random phosphorylations: Graphs show time-courses of hypo, medium, hyperphosphorylated and fully phosphorylated F proteins for up to \(\hbox {n}=100\) phosphorylation sites. Note, that the dashed line in Fig. 1 (inactivation of F due to phosphorylations) is adapted from 1—F40 from this figure.
Note, that the widely varying amplitudes can not be directly related to experimental data. Nevertheless, there are generic features of the model that do not depend on sensitivity to chosen parameter values (compare also Fig. 8): hypophosphorylated proteins increase quickly up to medium levels. Levels of medium phosphorylations (F30...F40) exhibit ultrasensitive increases to fairly large amplitudes. Fully phosphorylated proteins are found only at tiny levels.
In Fig. 8A we use a logarithmic scale to cover these drastic amplitude differences. The graph reveals that for different parameters intermediate levels of phosphorylations exhibit quite large amplitudes. The steps reflect changes in waveforms (see Fig. 7) due to sequestration effects.
In addition to large amplitudes mediators of cellular switches require robust delays and ultrasensitivity. The graphs in Fig. 8B,C show that intermediate phosphorylation levels obey these properties for different parameters constellations.
In order to test the robustness of these features we analyzed tenfold reductions of the degradation rate (red) and a tenfold increase of the enzyme/substrate ratio (green). It turns out that the main properties—huge amplitudes at intermediate phosphorylation levels, long delays, and Hill coefficients above 3—are robust with respect to large parameter variations.
Intermediate phosphorylation levels exhibit large amplitudes, delays and ultrasensitivity: (A) shows log10 values of amplitude for phosphorylated species. (B) depicts delays and (C) shows Hill coefficients for phosphorylated species. Note, that Hill coefficients for most values of our \(\hbox {k}<35\) exceed 6 and are not visible.
Combining transcriptional–translational feedback loops with multiple phosphorylations
So far we discussed conceptional models of phosphorylations leading to delays and ultrasensitivity. Phosphorylated clock proteins are core elements of transcriptional–translational feedback loops (TTFLs) leading to self-sustained oscillations1,10. Such feedback loops are often modeled via Goodwin oscillators including gene X, protein Y, and inhibitor42,79,80. Most of these models, however, require ad hoc Hill coefficients40.
In Suppl. A8 we present a fusion model that combines a Goodwin oscillator with distributive random phosphorylation. Along the lines of50 the negative feedback is realized by increasing phosphorylation levels. Gene expression is driven only by “Fact”, i.e. by the sum of active protein forms. The fast turnover of Fk provides the required negative feedback.
Figure 9 shows self-sustained oscillations of our fusion model. The separation of time scales allows limit cycles even for few phosphorylations. The number of phosphorylations controls the period. It turns out that the distinction of active forms and inactive forms of clock proteins can contribute to rhythm generation as discussed also in previous studies25,50,78,81.
Discussion
Self-sustained circadian oscillations require long delays and nonlinearities (“switches”)29,30. Recent experiments suggest that slow and seemingly random phosphorylations of intrinsically disordered clock proteins control stability and function of clock protein complexes such as FFC in Neurospora and PER:CRY in mammals18,25,82,83. Since only few detailed quantitative data are available we compared several generic models describing linear processive phosphorylation, nonlinear distributive phosphorylation, and random phosphorylation. We find that long delays are robustly achieved by long reaction chain lengths and slow degradation (Figs. 2, 3 and 4). Sequestration enhances the formation of temporal switches with high Hill coefficients (Suppl. A5). Thus our simulations support our hypothesis sketched in Fig. 1 that multiple random phosphorylation can provide robust delayed switches.
Our theoretical approach is based on a few experimental observations that help to constrain our generic models. Interestingly, most of the data suggest quite similar principles in the Neurospora clock and mammalian clocks. In both cases intrinsically disordered proteins (FRQ and PER, respectively) with multiple random phosphorylations are central players within the negative transcriptional-translational feedback loop21, 72,84,85. Interestingly, the control of the delay via phosphorylations is also a target of pharmacological interventions86,87. Below we focus on the Neurospora clock.
The half-life of the FRQ protein is about 3–5 h88. About 100 phosphorylation sites have been identified using isotope labelling and mass spectrometry. The priming-independent phosphorylation of non-consensus sites on FRQ by CK1a seems to be slower than five sites per hour62. Priming by other kinases is relatively fast and appears to be less essential for the principle function of the core clock. However, other kinases might be relevant for entrainment and temperature compensation not discussed in this paper18.
Figure 1 illustrates the important role of multiple phosphorylations. FRQ has a positively net charged N-terminal part, a negatively net charged C-terminal part, and a central part involved in proteasomal degradation by FWD160,77,88. Initial phosphorylation of the C-terminal part early in the circadian day has a stabilizing effect18,89. Subsequent phosphorylation of the N-terminal part destabilized the FFC complex allowing degradation. Thus, if charge repulsion were to govern stability and function, the number of phosphorylations would play a central role.
These experimental findings are reviewed in recent publications18,23,89 and provide the framework of our models. We do not fit individual parameters to the sparse quantitative data but we adapt the models design to the observations listed above. For example, the number of phosphorylations, the degradation rates, and the central role of casein kinase are consistent with the data. Since quantitative details of binding and unbinding of CK1a to FRQ are not known, we simulated two mechanisms: linear processive phosphorylations (Suppl. A1 and Fig 2) and distributive phosphorylations (Suppl. A2, A3, Figures 3, 4, 5, 6, 7 and 8). At a first glance, the distributive mechanism seems not consistent with Fig. 1 showing a relatively stable complex involving FRQ and CK1a. Indeed, CK1a is first recruited to the FRQ-CK1a-domains (FCD1 and 2) of FRQ89. Then the active site of bound CK1a can phosphorylate step by step the FRQ protein. This could happen at the same FRQ molecule or after dissociation and rebinding at another FRQ molecule60. Such reoccurring binding events could be simulated by our approach using the distributive mechanism. We start our simulations with zero levels of protein and enzyme. This allows to follow the kinetics of complex formations, sequestration, and phosphorylations.
Our conceptual models could be extended in future studies by more detailed features of protein dynamics18. For example, the number of phosphorylations influences stability and dissociation constants implying dependencies of model parameters on the number of phosphorylations k89. In our Suppl. A7 we show that step by step decreasing and increasing phosphorylation rates and varying stability do not change our main results. Such simulations are a first step towards more detailed incorporation of the effects of specific phosphorylations86. In future studies, stabilizing effects of FRH55, interactions with WCC23, and degradation assisted by FWD178 could be incorporated.
We emphasize, that our focus on multiple random phosphorylations neglects other essentials of the TTFL modeled in detail elsewhere26,80,90,91. Nevertheless, delayed switch-like behavior due to slow random multiple phosphorylation seems to be central element in circadian rhythm generation. A delayed switch due to multiple phosphorylation is a robust design principle that could be relevant also in other biological systems such as ligand specificity, nuclear import, DNA binding in T-cells61,92,93, timing of critical transitions in cell cycle44, regulation of sleep-wake homeostasis in mice94, Familial advanced sleep-phase syndrome (FASPS) in humans24, phototactic sensitivity in green algae95, and reproductive fitness in cyanobacteria96.
Materials and methods
Numerics
All the simulations have been performed on a Spyder Python 3.4 platform. Simulations resulting in Figs 1, 2, 3, 4, 5, 6, 7 and 8 have been obtained by numerically solving the ordinary differential equations provided in Suppl. A1 to A4 via the odeint function from the integrate module of the Scientific Python (SciPy) package with a constant stepsize \(\hbox {dt} = 0.01 \,\hbox {h}\). The Matplotlib library has been used to generate figures97. Codes are available upon request.
Calculation of Hill coefficients and delays
An ultrasensitive response is often sigmoidal and the curve can be well approximated by the Hill equation. The effective Hill equation is defined for the temporal curves in Figs 2, 3 and 6 (see Eq. 1). Note, that the Hill-function is typically used to study input–output relations X versus Y. In our case we quantify temporal switches, i.e. X represents time and Y are the corresponding protein concentrations. The effective Hill coefficient n is related to the effective time 90% (ET90) and 10% (ET10) ratio by Eq. (2). Here K is the effective time when 50% (or ET50) of total phosphorylation is achieved.
Delays are calculated as the time when 50% of the maximal value is achieved for each phosphoprotein Fk as given in Eq. (3).
References
Wever, R. Zum Mechanismus der biologischen 24-Stunden-Periodik. Kybernetik 2, 214–231 (1963).
Konopka, R. J. & Benzer, S. Clock mutants of Drosophila melanogaster. Proc. Natl. Acad. Sci. 68, 2112–2116 (1971).
Nakajima, M. et al. Reconstitution of circadian oscillation of cyanobacterial kaic phosphorylation in vitro. Science 308, 414–415 (2005).
Mittag, M., Kiaulehn, S. & Johnson, C. H. The circadian clock in Chlamydomonas reinhardtii. What is it for? What is it similar to?. Plant Physiol. 137, 399–409 (2005).
Dunlap, J., Loros, J. & DeCoursey, P. Chronobiology: Biological Timekeeping. (2004). https://psycnet.apa.org/record/2003-06316-000.
Relógio, A. et al. Ras-mediated deregulation of the circadian clock in cancer. PLoS Genet. 10, e1004338. https://doi.org/10.1371/journal.pgen.1004338 (2014).
Dibner, C., Schibler, U. & Albrecht, U. The mammalian circadian timing system: organization and coordination of central and peripheral clocks. Annu. Rev. Physiol. 72, 517–549 (2010).
Panda, S., Hogenesch, J. B. & Kay, S. A. Circadian rhythms from flies to human. Nature 417, 329–335 (2002).
Reddy, A. B. & O’Neill, J. S. Healthy clocks, healthy body, healthy mind. Trends Cell Biol. 20, 36–44 (2010).
Hardin, P. E., Hall, J. C. & Rosbash, M. Feedback of the Drosophila period gene product on circadian cycling of its messenger RNA levels. Nature 343, 536–540 (1990). https://www.nature.com/articles/343536a0.pdf.
Aronson, B. D., Johnson, K. A., Loros, J. J. & Dunlap, J. C. Negative feedback defining a circadian clock: autoregulation of the clock gene frequency. Science 263, 1578–1584 (1994).
Duong, H. A., Robles, M. S., Knutti, D. & Weitz, C. J. A molecular mechanism for circadian clock negative feedback. Science 332, 1436–1439 (2011).
Brunner, M. & Káldi, K. Interlocked feedback loops of the circadian clock of Neurospora crassa. Mol. Microbiol. 68, 255–262. https://doi.org/10.1111/j.1365-2958.2008.06148.x (2008).
Loros, J. J. & Dunlap, J. C. Genetic and molecular analysis of circadian rhythms in Neurospora. Annu. Rev. Physiol. 63, 757–794. https://doi.org/10.1146/annurev.physiol.63.1.757 (2001).
Partch, C. L., Green, C. B. & Takahashi, J. S. Molecular architecture of the mammalian circadian clock. Trends Cell Biol. 24, 90–99 (2014).
Reppert, S. M. & Weaver, D. R. Molecular analysis of mammalian circadian rhythms. Annu. Rev. Physiol. 63, 647–676 (2001).
Brown, S. A., Kowalska, E. & Dallmann, R. (re) inventing the circadian feedback loop. Dev. Cell 22, 477–487 (2012).
Diernfellner, A. C. & Brunner, M. Phosphorylation timers in the neurospora crassa circadian clock. J. Mol. Biol. 432, 3449–3465 (2020).
Lee, C., Etchegaray, J.-P., Cagampang, F. R., Loudon, A. S. & Reppert, S. M. Posttranslational mechanisms regulate the mammalian circadian clock. Cell 107, 855–867 (2001).
Narasimamurthy, R. et al. CK1\(\delta\)/\(\epsilon\) protein kinase primes the PER2 circadian phosphoswitch. Proc. Natl. Acad. Sci. 115, 5986–5991 (2018). https://www.pnas.org/content/115/23/5986.short.
Gallego, M. & Virshup, D. M. Post-translational modifications regulate the ticking of the circadian clock. Nat. Rev. Mol. Cell Biol. 8, 139–148 (2007).
Sancar, G., Sancar, C., Brunner, M. & Schafmeier, T. Activity of the circadian transcription factor white collar complex is modulated by phosphorylation of sp-motifs. FEBS Lett. 583, 1833–1840 (2009).
Wang, B., Kettenbach, A. N., Zhou, X., Loros, J. J. & Dunlap, J. C. The phospho-code determining circadian feedback loop closure and output in neurospora. Mol. Cell 74, 771–784 (2019).
Vanselow, K. et al. Differential effects of PER2 phosphorylation: molecular basis for the human familial advanced sleep phase syndrome (FASPS). Genes Dev. 20, 2660–2672 (2006).
Aryal, R. P. et al. Macromolecular assemblies of the mammalian circadian clock. Mol. Cell 67, 770–782 (2017).
Upadhyay, A., Brunner, M. & Herzel, H. An inactivation switch enables rhythms in a neurospora clock model. Int. J. Mol. Sci. 20, 2985 (2019).
Schafmeier, T. et al. Transcriptional feedback of Neurospora circadian clock gene by phosphorylation-dependent inactivation of its transcription factor. Cell 122, 235–246 (2005). http://www.ncbi.nlm.nih.gov/pubmed/16051148http://linkinghub.elsevier.com/retrieve/pii/S0092867405005933.
Tang, C.-T. et al. Setting the pace of the neurospora circadian clock by multiple independent frq phosphorylation events. Proc. Natl. Acad. Sci. 106, 10722–10727 (2009).
Glass, L. & Mackey, M. C. From Clocks to Chaos: The Rhythms of Life (Princeton University Press, Princeton, 1988).
Korenčič, A. et al. The interplay of cis-regulatory elements rules circadian rhythms in mouse liver. PloS One 7 (2012).
Bordyugov, G., Westermark, P. O., Korenčič, A., Bernard, S. & Herzel, H. Mathematical Modeling in Chronobiology. , Vol. 217, 335–357. https://doi.org/10.1007/978-3-642-25950-0_14 (Springer, Berlin, 2013).
MacDonald, N. Biological delay systems: Linear stability theory (Cambridge University Press, Cambridge, 2008).
Bernard, S., Čajavec, B., Pujo-Menjouet, L., Mackey, M. C. & Herzel, H. Modelling transcriptional feedback loops: The role of gro/tle1 in hes1 oscillations. Philos. Trans. R. Soc. A 364, 1155–1170 (2006).
Aulehla, A. & Pourquie, O. Oscillating signaling pathways during embryonic development. Curr. Opin. Cell Biol. 20, 632–637 (2008). https://www.sciencedirect.com/science/article/pii/S0955067408001622.
Hoffmann, A., Levchenko, A., Scott, M. L. & Baltimore, D. The i\(\kappa\)b-nf-\(\kappa\)b signaling module: Temporal control and selective gene activation. Science 298, 1241–1245 (2002).
Ashall, L. et al. Pulsatile stimulation determines timing and specificity of nf-\(\kappa\)b-dependent transcription. Science 324, 242–246 (2009).
Purvis, J. E. et al. p53 dynamics control cell fate. Science 336, 1440–1444 (2012).
Diernfellner, A., Querfurth, C., Salazar, C., Höfer, T. & Brunner, M. Phosphorylation modulates rapid nucleocytoplasmic shuttling and cytoplasmic accumulation of Neurospora clock protein FRQ on a circadian time scale. Genes Dev. 23, 2192–2200 (2009). http://genesdev.cshlp.org/content/23/18/2192.short.
Toh, K. L. et al. An hper2 phosphorylation site mutation in familial advanced sleep phase syndrome. Science 291, 1040–1043 (2001).
Griffith, J. Mathematics of cellular control processes I. Negative feedback to one gene. J. Theor. Biol. 20, 202–208 (1968).
Ananthasubramaniam, B. & Herzel, H. Positive feedback promotes oscillations in negative feedback loops. PLoS One 9, e104761 (2014).
Goodwin, B. C. Oscillatory behavior in enzymatic control processes. Adv. Enzym. Regul. 3, 425–438 (1965). http://www.ncbi.nlm.nih.gov/pubmed/5861813.
Gonze, D. & Ruoff, P. The goodwin oscillator and its legacy. Acta Biotheor. 1, 1–18. https://doi.org/10.1007/s10441-020-09379-8 (2020).
Deshaies, R. J. & Ferrell, J. E. Jr. Multisite phosphorylation and the countdown to s phase. Cell 107, 819–822 (2001).
Salazar, C. & Höfer, T. Kinetic models of phosphorylation cycles: A systematic approach using the rapid-equilibrium approximation for protein-protein interactions. BioSystems 83, 195–206 (2006).
Ferrell, J. E. & Bhatt, R. R. Mechanistic studies of the dual phosphorylation of mitogen-activated protein kinase. J. Biol. Chem. 272, 19008–19016 (1997).
Legewie, S., Blüthgen, N. & Herzel, H. Quantitative analysis of ultrasensitive responses. FEBS J. 272, 4071–4079. https://doi.org/10.1111/j.1742-4658.2005.04818.x (2005).
Markevich, N. I., Hoek, J. B. & Kholodenko, B. N. Signaling switches and bistability arising from multisite phosphorylation in protein kinase cascades. J. Cell Biol. 164, 353–359 (2004).
Masuda, S. et al. Mutation of a per2 phosphodegron perturbs the circadian phosphoswitch. Proc. Natl. Acad. Sci. 117, 10888–10896 (2020).
Gonze, D. & Abou-Jaoudé, W. The Goodwin model: Behind the Hill function. PLoS One 8, e69573 (2013). http://journals.plos.org/plosone/article/file?id=10.1371/journal.pone.0069573&type=printable
Goldbeter, A. & Koshland, D. Ultrasensitivity in biochemical systems controlled by covalent modification. Interplay between zero-order and multistep effects. J. Biol. Chem. 259, 14441–14447 (1984). http://www.jbc.org/content/259/23/14441.short.
Ode, K. L. & Ueda, H. R. Design principles of phosphorylation-dependent timekeeping in eukaryotic circadian clocks. Cold Spring Harb. Perspect Biol. 10, a028357 (2018).
Baker, C. L., Kettenbach, A. N., Loros, J. J., Gerber, S. A. & Dunlap, J. C. Quantitative proteomics reveals a dynamic interactome and phase-specific phosphorylation in the neurospora circadian clock. Mol. Cell 34, 354–363 (2009).
Marzoll, D. The Role of Neurospora Casein Kinase 1a in Circadian Timekeeping. (PhD Dissertation, Heidelberg University, 2019).
Lauinger, L., Diernfellner, A., Falk, S. & Brunner, M. The RNA helicase FRH is an ATP-dependent regulator of CK1a in the circadian clock of Neurospora crassa. Nat. Commun. 5, 3598 (2014). https://www.nature.com/articles/ncomms4598.
Merrow, M., Brunner, M. & Roenneberg, T. Assignment of circadian function for the Neurospora clock gene frequency. Nature 399, 584 (1999). https://www.nature.com/articles/21190.pdf.
Tataroglu, O. & Schafmeier, T. Of switches and hourglasses: Regulation of subcellular traffic in circadian clocks by phosphorylation. EMBO Rep. 11, 927–935 (2010). http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=2999865&tool=pmcentrez&rendertype=abstract.
Diernfellner, A. & Schafmeier, T. Phosphorylations: Making the Neurosporacrassa circadian clock tick. FEBS Lett. 585, 1461–1466 (2011). https://www.sciencedirect.com/science/article/pii/S0014579311002158.
Schafmeier, T., Káldi, K., Diernfellner, A., Mohr, C. & Brunner, M. Phosphorylation-dependent maturation of Neurospora circadian clock protein from a nuclear repressor toward a cytoplasmic activator. Genes Dev. 20, 297–306 (2006). http://genesdev.cshlp.org/content/20/3/297.short.
Querfurth, C. et al. Circadian conformational change of the Neurospora clock protein FREQUENCY triggered by clustered hyperphosphorylation of a basic domain. Mol. Cell 43, 713–722 (2011). https://www.sciencedirect.com/science/article/pii/S1097276511006253.
Salazar, C. & Höfer, T. Multisite protein phosphorylation-from molecular mechanisms to kinetic models. FEBS J. 276, 3177–3198 (2009).
Diernfellner, A. C., Lauinger, L., Shostak, A. & Brunner, M. A pathway linking translation stress to checkpoint kinase 2 signaling in neurospora crassa. Proc. Natl. Acad. Sci. 116, 17271–17279 (2019).
Ueki, Y. et al. A consensus binding motif for the pp4 protein phosphatase. Mol. Cell 76, 953–964 (2019).
Lee, H.-M. et al. The period of the circadian oscillator is primarily determined by the balance between casein kinase 1 and protein phosphatase 1. Proc. Natl. Acad. Sci. 108, 16451–16456 (2011).
Ferrell, J. E. et al. Ultrasensitivity part II: Multisite phosphorylation, stoichiometric inhibitors, and positive feedback. Trends Biochem. Sci. 39, 556–569 (2014).
Blüthgen, N. et al. Effects of sequestration on signal transduction cascades. FEBS J. 273, 895–906 (2006).
Legewie, S., Schoeberl, B., Blüthgen, N. & Herzel, H. Competing docking interactions can bring about bistability in the mapk cascade. Biophys. J. 93, 2279–2288 (2007).
Klipp, E., Liebermeister, W., Wierling, C. & Kowald, A. Systems biology : a textbook 2nd edn. (Wiley-Blackwell, London, 2016).
Kim, J. K. Mathematical Modeling and Analysis of Cellular Clocks. (Ph.D. Thesis, University of Michigan, 2013).
Ingalls, B. Mathematical modeling in systems biology: An introduction (MIT Press, London, 2013).
Ferrell, J. E. & Ha, S. H. Ultrasensitivity part I: Michaelian responses and zero-order ultrasensitivity. Trends Biochem. Sci. 39, 496–503 (2014). https://www.sciencedirect.com/science/article/pii/S0968000414001431.
Querfurth, C. et al. Posttranslational regulation of neurospora circadian clock by ck1a-dependent phosphorylation Vol. 72, 177–183 (Cold Spring Harbor Laboratory Press, New York, 2007).
Clodong, S. et al. Functioning and robustness of a bacterial circadian clock. Mol. Syst. Biol. 3 (2007).
Thomson, M. & Gunawardena, J. Unlimited multistability in multisite phosphorylation systems. Nature 460, 274–277 (2009).
Legewie, S., Blüthgen, N., Schäfer, R. & Herzel, H. Ultrasensitization: switch-like regulation of cellular signaling by transcriptional induction. PLoS Comput. Biol. 1 (2005).
Whitman, W. B., Coleman, D. C. & Wiebe, W. J. Prokaryotes: The unseen majority. Proc. Natl. Acad. Sci. 95, 6578–6583 (1998).
He, Q. et al. Fwd1-mediated degradation of frequency in neurospora establishes a conserved mechanism for circadian clock regulation. EMBO J. 22, 4421–4430 (2003).
Larrondo, L. F., Olivares-Yañez, C., Baker, C. L., Loros, J. J. & Dunlap, J. C. Decoupling circadian clock protein turnover from circadian period determination. Science 347, 1257277 (2015). http://science.sciencemag.org/content/347/6221/1257277.short.
Ruoff, P., Loros, J. J. & Dunlap, J. C. The relationship between FRQ-protein stability and temperature compensation in the Neurospora circadian clock. Proc. Natl. Acad. Sci. 102, 17681–17686 (2005). https://www.pnas.org/content/102/49/17681.short.
Gonze, D., Leloup, J.-C. & Goldbeter, A. Theoretical models for circadian rhythms in Neurospora and Drosophila. C. R. Acad. Sci. 323, 57–67 (2000). https://www.sciencedirect.com/science/article/pii/S0764446900001116.
Zhou, M., Kim, J., Eng, G., Forger, D. & Virshup, D. A Period2 phosphoswitch regulates and temperature compensates circadian period. Mol. Cell 60, 77–88 (2015). https://www.sciencedirect.com/science/article/pii/S1097276515006681.
Philpott, J. M. et al. Casein kinase 1 dynamics underlie substrate selectivity and the per2 circadian phosphoswitch. Elife 9, e52343 (2020).
Virshup, D. M. et al. CK1\(\delta\)/\(\epsilon\) protein kinase primes the PER2 circadian phosphoswitch. Proc. Natl. Acad. Sci. 115, 5986–5991 (2018). https://www.pnas.org/content/115/23/5986.short.
Hurley, J. M., Larrondo, L. F., Loros, J. J. & Dunlap, J. C. Conserved RNA helicase FRH acts nonenzymatically to support the intrinsically disordered neurospora clock protein FRQ. Mol. Cell 52, 832–843 (2013). https://www.sciencedirect.com/science/article/pii/S1097276513008307.
Schlosser, A., Vanselow, J. T. & Kramer, A. Mapping of phosphorylation sites by a multi-protease approach with specific phosphopeptide enrichment and nanolc- ms/ms analysis. Anal. Chem. 77, 5243–5250 (2005).
Kim, D. W. et al. Systems approach reveals photosensitivity and per 2 level as determinants of clock-modulator efficacy. Mol. Syst. Biol. 15, e8838 (2019).
Meng, Q.-J. et al. Entrainment of disrupted circadian behavior through inhibition of casein kinase 1 (ck1) enzymes. Proc. Natl. Acad. Sci. 107, 15240–15245 (2010).
Görl, M. et al. A pest-like element in frequency determines the length of the circadian period in neurospora crassa. EMBO J. 20, 7074–7084 (2001).
Liu, X. et al. Frq-ck1 interaction determines the period of circadian rhythms in neurospora. Nat. Commun. 10, 1–13 (2019).
François, P. A model for the Neurospora circadian clock. Biophys. J. 88, 2369–2383 (2005). https://www.sciencedirect.com/science/article/pii/S0006349505732951.
Hong, C. I., Jolma, I. W., Loros, J. J., Dunlap, J. C. & Ruoff, P. Simulating dark expressions and interactions of frq and wc-1 in the Neurospora circadian clock. Biophys. J. 94, 1221–1232 (2008).
Gabriel, C. H. et al. Identification of novel nuclear factor of activated t cell (nfat)-associated proteins in t cells. J. Biol. Chem. 291, 24172–24187 (2016).
Thurley, K., Wu, L. F. & Altschuler, S. J. Modeling cell-to-cell communication networks using response-time distributions. Cell Syst. 6, 355–367 (2018).
Brüning, F. et al. Sleep-wake cycles drive daily dynamics of synaptic phosphorylation. Science 366, 3617 (2019).
Böhm, M. et al. Channelrhodopsin-1 phosphorylation changes with phototactic behavior and responds to physiological stimuli in chlamydomonas. Plant Cell 31, 886–910 (2019).
Ouyang, Y., Andersson, C. R., Kondo, T., Golden, S. S. & Johnson, C. H. Resonating circadian clocks enhance fitness in cyanobacteria. Proc. Natl. Acad. Sci. 95, 8660–8664 (1998).
Hunter, J. D. Matplotlib: A 2D graphics environment. Comput. Sci. Eng. 9, 90–95 (2007).
Acknowledgements
The authors are thankful to Christoph Schmal and Bharath Ananthsubramaniam for fruitful discussions. We thank the anonymous reviewers for their helpful suggestions.
Funding
Open Access funding enabled and organized by Projekt DEAL. This research was funded by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation)—Project Number 278001972 - TRR 186. We also acknowledge the Open Access Publication Fund of Charité—Universitätsmedizin Berlin.
Author information
Authors and Affiliations
Contributions
Conceptualization by M.B. and H.H.; methodology by A.U. and H.H.; software by A.U.; validation by A.U. and H.H.; formal analysis by A.U.; investigation, A.U. and H.H.; resources by H.H.; data curation by M.B., D.M. and A.D.; writing—original draft preparation by A.U. and H.H.; writing—review and editing, A.U., H.H., D.M., A.D. and M.B.; visualization by A.U. and H.H.; supervision by M.B. and H.H.; project administration by M.B. and H.H.; funding acquisition by M.B. and H.H.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Upadhyay, A., Marzoll, D., Diernfellner, A. et al. Multiple random phosphorylations in clock proteins provide long delays and switches. Sci Rep 10, 22224 (2020). https://doi.org/10.1038/s41598-020-79277-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-79277-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.