Abstract
The gastric colonization of human hosts by Helicobacter pylori (H. pylori) increases the risk of developing gastritis, ulcers and gastric cancer. To detect H. pylori, a nanohybrid-based BabA immunosensor is developed herein. BabA is an outer membrane protein and one of the major virulence factors of H. pylori. To design the immunosensor, an Au electrode is loaded with palladium nanoparticles (Pdnano) by electrodeposition to generate reduced graphene oxide (rGO)/poly(3,4-ethylenedioxythiophene) (PEDOT). The immobilization of these nanostructured materials imparts a large surface area and electroconductivity to bio-immune-sensing molecules (here, the BabA antigen and antibodies). After optimization, the fabricated immunosensor has the ability to detect antigens (H. pylori) in a linear range from 0.2 to 20 ng/mL with a low LOD (0.2 ng/mL). The developed immunosensor is highly specific, sensitive and reproducible. Additionally, in silico methods were employed to better understand the hybrid nanomaterials of the fabricated Pdnano/rGO/PEDOT/Au electrode. Simulations performed by molecular docking, and Metropolis Monte Carlo adsorption studies were conducted. The results revealed that the hybrid nanomaterials exhibit a stable antigen–antibody complex of BabA, yielding the lowest binding energy in relation to the electrode materials, emphasizing the functionality of the constructed electrodes in the electrochemical immunosensor.
Similar content being viewed by others
Introduction
Helicobacter pylori (H. pylori) prevalence in hosts contributes to chronic gastritis and gastric cancer. Gram-negative bacterium is the type of H. pylori that ultimately leads to the progression of ulcers and cancer by infecting the gastric mucosa1,2,3. H. pylori develops mechanisms for gastric colonization with the help of virulence factors, causes gastric ulceration and plays a major role in the progression of stomach cancer. Collectively, with H. pylori colonization, the inflammatory response of the host and the interaction of environmental and dietary factors are a few of the major causes of gastric diseases4. The most likely transmission route is individual-to-individual, but fecal–oral transmissions were also identified5,6,7. It was observed that the death rate due to stomach cancer caused by H. pylori colonization has been increasing significantly, and therefore, an immunosensor was stepwise fabricated for detecting H. pylori specific antigens in biological samples.
In the first step, studies were performed through molecular docking. Molecular docking was incorporated to predict the modes of the interactions of the nanocomposites with each other and the nanocomposite with the H. pylori biomarker. In this work, emerging computational methods such as molecular docking, Metropolis Monte Carlo (MMC), and binding energy calculations were employed to validate the energetic parameters of the immunosensor. The results revealed the adsorption behavior of the BabA antigen and its antibody complexes towards a layer-by-layer surface. Moreover, 3D structures of antigen (Ag) and antibody (Ab) biomolecules were used for docking simulations to investigate the BabA Ag–Ab interactions. The best docked model is subsequently elaborated with the hybrid electrode materials (i.e., Pdnano/rGO/PEDOT/Au) to mimic the experimental process. H. pylori BabA is used as a biomarker8. H. pylori antigen binding adhesin (BabA) protein is one of the major virulence factors of molecular docking9,10,11. Strains expressing BabA have shown a high risk of peptic ulceration and development of gastric diseases. In addition, BabA binding to host cells triggers translocation of CagA protein and promotes the pathogenicity of VacA protein12. Recently, because of their intrinsic properties, such as the antibody–antigen interactions determined by directly observation, electrochemical immunosensors have attracted considerable attention13. Thus, the requirements of the nanocomposite used in an immunosensor are very stringent: the nanocomposite should be a very good carrier and all together should not obstruct electron exchange or append proteins; the nanomaterial should have active sites and a vast surface area to immobilize many antibodies14.
Standard methods available for H. pylori detection are either invasive or noninvasive. Invasive procedures comprise histology, rapid urease tests15, polymerase chain reaction16, microbiological culture17 and biopsy-based tests. Noninvasive practices incorporate stool antigen tests18, serology and urea breath tests19. These methods, however, are time consuming, expensive and have a short shelf-life. Thus, to surmount all these downsides, immunosensors have been considered to be better because of their low LOD, high selectivity, fast response and ease of handling. Despite the numerous advancements that have been made, there is an unprecedented need for a sensor that is based on highly selective antigen–antibody interactions and exhibits good sensitivity.
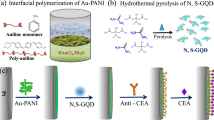
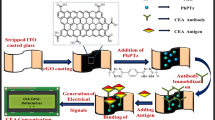
Herein, we have developed an electrochemical immunosensor using a hybrid nanomaterial probe with metal nanoparticles and a conducting polymer. The novel hybrid nanomaterials are synthesized and engineered for H. pylori detection with the incorporation of nanoscale properties, i.e., the large surface area and strong electrical conductivity of palladium nanoparticles (Pdnano), poly(3,4-ethylenedioxythiophene) (PEDOT) and reduced graphene oxide (rGO) with the BabA antigen. In addition, stability and biocompatibility are considered. Furthermore, noninvasive testing was conducted on spiked stool samples using this application. The goal of this study was to provide a promising platform for the early detection of the H. pylori antigen using label-free hybrid nanomaterial-based immunosensors (Scheme 1).
Schematic illustration of the steps involved in the preparation of the BabA Ab@Pdnano/rGO/PEDOT/Au electrode. The figures were obtained from Material Studio v.16.1.0.21 (BIOVIA, D. S. 2016. Material Studio modelling. v.16.1.0.21 ed. San Diego: Dassault Systemes) and Discovery Studio v.16.10.15350 software packages (BIOVIA, D. S. 2016. Discovery Studio modelling. v.16.10.15350 ed. San Diego: Dassault Systemes).
.
Results and discussions
Computational simulation and modeling
Computational simulations have been effectively used to observe the mechanism of nanocomposite construction and the interaction phenomena occurring in biosensor platforms20,21. In this work, the layer-by-layer electrode position method used to prepare the hybrid nanomaterials was successfully mimicked through computational modeling to reproduce the energetic contributions towards binding with the biomolecule complex. Because of the utilization of a large-scale substrate-adsorbate system, i.e., the adsorbate structures of the Ag-Ab complex as well as the substrates containing organic and inorganic nanostructures, the overall energy of each trajectory was computed with MMC. The binding energies were calculated according to Eq. (1), showing its tremendous computational capability.
Binding energy calculation
The binding energy (Eb) between the substrate and the adsorbate layers was computed according to the following equation:
where Ecomp = total energy of the complex; Eads = interaction energy between the adsorbate; and Esubs = substrate energy.
The overall energies of the reactant (i.e., layer-by-layer substrates) and the final products (i.e., substrate-adsorbate complexes) were determined with the binding energy calculations by classic and pragmatic approaches22. The whole energies were defined according to Eq. (2).
where the valence energy (Eval) includes the bond, angle, torsion and inversion energies, while the nonbond energy (Enb) includes the van der Waals, long-range correction, and electrostatic energies. A reasonable periodic cell size was employed to demonstrate the appropriate surface area. Since the hybrid nanomaterial probe contains four material elements (Au, PEDOT, rGO and Pdnano), we decided to carry out the simulations with respect to each substrate surface (Au, PEDOT/Au, rGO/PEDOT/Au and Pdnano/rGO/PEDOT/Au) (Fig. 1A (a)) and thoroughly examined their binding energy differences with respect to the Ag–Ab complex.
(A) Periodic boundary structure containing Pdnano/rGO/PEDOT/Au electrode along with the origin of each component (a) and docked structure of BabA Ag and its Ab. The Ag and Ab structures shown as line- and solid-ribbon depictions, respectively (b). The lowest energy configuration of Ag (BabA protein; green)-Ab (anti-BabA; purple) molecule onto Au electrode (c), PEDOT/Au electrode (d), rGO/PEDOT/Au electrode (e) and Pdnano/rGO/PEDOT/Au electrode (f). For the figures (c–f), the complex structures are depicted from front views using ball and stick models. The figures were obtained from Material Studio v.16.1.0.21 (BIOVIA, D. S. 2016. Material Studio modelling. v.16.1.0.21 ed. San Diego: Dassault Systemes) and Discovery Studio v.16.10.15350 software packages (BIOVIA, D. S. 2016. Discovery Studio modelling. v.16.10.15350 ed. San Diego: Dassault Systemes). (B) The lowest energy configurations of Ag (BabA protein; green)-Ab (anti-BabA; purple) molecule onto Au electrode (a), PEDOT/Au electrode (b), rGO/PEDOT/Au electrode (c) and Pdnano/rGO/PEDOT/Au electrode (d). The depictions are undertaken at the top views format. The figures were obtained from Material Studio v.16.1.0.21 (BIOVIA, D. S. 2016. Material Studio modelling. v.16.1.0.21 ed. San Diego: Dassault Systemes) and Discovery Studio v.16.10.15350 software packages (BIOVIA, D. S. 2016. Discovery Studio modelling. v.16.10.15350 ed. San Diego: Dassault Systemes).
Prior to the MMC studies, the interaction of Ag and Ab was defined through docking simulation by employing ZDOCK and ZRANK as scoring functions. The best pose used for further calculation was Pose-24, since it indicated the presence of the important amino acid residues of the Ab binding surface (i.e., Gly191, Ser234, and Ser244)23 among the top thirty binding poses. The results and scoring data are summarized in Table 1. As a preliminary step, the Ag and Ab structures were derived from PDB codes and treated using CHARMM19 for geometry optimization. The optimized structures of Ab and Ag were introduced as the receptor and ligand for docking analysis, respectively, resulting in approximately 2000 docked poses, and the best docked model, which fulfilled the active binding region of Ab, was selected from the largest clusters to perform further interaction studies with the nanomaterial surfaces (Fig. 1A (b)).
MMC simulation is a reliable approach for predicting adsorption. During the docking studies, MMC simulation was carried out on each substrate layer, which was followed by binding energy observation. By performing the calculation of single point energy, the energetics of each composite was observed and correlated with those of other materials. The optimized configuration of the whole system is depicted in Fig. 1A (c–f),B (a–d), which comprise different views, while the generated values of the binding energy are depicted in Table 2. The negative given values/binding scores demonstrate stable complex formation during the interaction studies. Indeed, the Au surface produces a direct efficient binding energy after the adsorption of the protein complex onto the electrode surface, suggesting that the metallic properties induce a physical interaction between the Au surface and the biomolecule24. On the other hand, the addition of the PEDOT polymers tends to weaken the binding interaction of the complex, which was shown by the significant increase in the binding energy to − 374.74 kJ/mol (Fig. 1A(d),B (c)). Similar evidence was shown after rGO assembled on the biosurface. The lowest binding energy was found for the fully packed system composed of Pdnano/rGO/PEDOT/Au with a binding energy of − 1386.59 kJ/mol (Fig. 1A(f),B (d)). However, the other surfaces showed higher binding energies (weaker binding interactions). Thus, the complete layered substrate efficiently releases more energy than the others. In addition, the presence of PEDOT, rGO, and Pdnano on the Au electrode contributes to the low binding energy of the complete system (i.e., strengthening the interaction with the analyte). This further suggests that assembling the hybrid nanomaterials in the immunosensing system results in the preferable interaction between the corresponding complex and analyte. Indeed, the results calculated with this experimental strategy and previous theoretical studies have an agreeable outcome, indicating that combining Pdnano with rGO can possibly improve the sensing performance to achieve a performance better than that obtained with Pdnano with rGO individually due to their excellent conductivity. The synergy of the experimental and computational results is described herein.
Nano-electrochemical deposition of Pdnano over the rGO/PEDOT/Au electrode
Figure 2 depicts the electrodeposition of Pdnano over the rGO/PEDOT/Au electrode at the specified scan rate (50 mV/s) by adjusting the voltage from − 0.25 to + 1.2 V for 10 cycles. Ten cycles of CV were repeated for the electrodeposition of Pdnano to achieve reproducible curves. In first few cycles, peaks were not differentiable because the added precursor was not completely dissolved. Here, CV demonstrates the characteristic current features of the decrease in Pd size at a voltage of 0.5 V, Pd oxide formation at 0.24 V and 0.62 V (at the same time), and hydrogen adsorption and desorption between 0.1 and − 0.1 V. Pdnano accumulated on the modified electrode while the potential was scanned in the negative direction, and the peak at − 0.02 V indicates the process of reducing protons that are adsorbed on the Pd surface to hydrogen. The Pdnano particles deposited on the surface of the electrode are further oxidized to Pd2+ to create a layer of Pd oxide (0.24 V)25,26. During the reverse potential scan, the size of the synthesized Pd oxides further decreased, leading to Pdnano particles capable of hydrogen adsorption. The enhancement of peaks was observed during this repeated cycling process, confirming that the process of Pdnano deposition on the Au electrode occurred. These characterizations clearly indicate the successive synthesis of a Pdnano layer over the nanocomposite-modified Au electrode.
The established procedure used to reduce the PdCl42− complex to Pd is as follows:
Nanocharacterization of the various steps involved in the immunoassembly
Electron microscopic characterization of nanocomposites
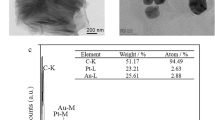
Figure 3A shows the morphological study results obtained by SEM for the (a) bare Au electrode and (b) PEDOT, (c) rGO/PEDOT, (d) Pdnano/rGO/PEDOT and (e) BabA Ab@Pdnano/rGO/PEDOT-modified Au electrode. The Au electrode surface was a smooth and uniform layer of a bare Au electrode (Fig. 3A (a)). Figure 3A (b) reveals the granular structure of PEDOT on the surface of the Au electrode, while the rGO film shows a wrinkled and crumpled surface (image c). As shown in Fig. 3A (d), a spherical structure with uniformly distributed nanospheres was observed for Pdnano on the rGO/PEDOT-modified Au electrode surface. Pdnano almost covered and was well distributed on the entire graphene surface of the rGO/PEDOT/Au electrode. Figure 3A (e) depicts the globule-like structure of BabA Ab, which formed clusters and was distributed evenly on the surface of the nanocomposite modified electrode. Figure 3A (f) shows the EDX spectrum of carbon, oxygen, gold and palladium. The composition of these elements in the spectrum (Au = 4.88%, C = 60.38%, O = 33.47%, and Pd = 1.27%) confirms that these elements are present in the nanocomposite-modified electrode.
Visualization of Pdnano through transmission electron microscopy
Figure 3B shows transmission electron microscopy (TEM) images of nanoparticle clusters of palladium, showing their shapes and size distributions. Round or oval shaped particles with 20 nm sizes were determined.
The electrochemical characterizations of the amplified strategies
To achieve both oxidation and reduction peaks, cyclic voltammetry was performed, however, square wave voltammetry and differential pulse voltammetry were performed to obtain only oxidation peaks. Figure 4A displays the CV analysis of the (a) bare Au electrode and (b) PEDOT, (c) rGO/PEDOT, (d) Pdnano/rGO/PEDOT and (e) BabA Ab@Pdnano/rGO/PEDOT-modified Au electrode. Slightly compressed redox peaks were observed for the Au electrode (curve a). Curve (b) shows the deposition of rGO, demonstrating that the material is reductive in nature. After the electropolymerization of PEDOT onto the modified electrode, there is slight improvement in the cathodic and anodic peaks (curve c). The CV curve shows the rate of electron transfer and redox peak current of the nanocomposites (Pdnano/rGO/PEDOT-modified Au electrode), which is significantly higher because of the good electrical conductivity achieved by combining both Pdnano, PEDOT and rGO14. Once BabA Ab was immobilized on the outer surface of the hybrid electrode, the current response decreased. The decrease in current is attributed to the fact that BabA Ab forms a layer that blocks the transfer of electrons, interrupting the redox reactions27,28. After analyzing the CV studies, it was revealed that the immunosensor fabrication process was effectively completed.
(A) CV curves of bare Au electrode, PEDOT/Au electrode, rGO/PEDOT/Au electrode, Pdnano/rGO/PEDOT/Au electrode and BabA Ab@ Pdnano/rGO/PEDOT/Au electrode obtained with 5 mM [Fe(CN)6]3−/4− electrolyte containing 0.5 mM sodium phosphate buffer (pH 7.5) at 20 mVs−1. (B) Electrochemical impedance plots (Nyquist plots) of the different electrodes bare Au electrode, PEDOT/Au electrde, rGO/PEDOT/Au electrode, Pdnano/rGO/PEDOT/Au electrode and BabA Ab@Pdnano/rGO/PEDOT/Au electrode obtained with 5 mM [Fe(CN)6]3−/4− electrolyte containing 0.5 mM sodium phosphate buffer (pH 7.5). The frequency range is between 10 Hz to 100 kHz with amplitude of 5 mV.
The charge transfer resistance (Rct) of the interface of the nanocomposite-coated Au electrode was monitored (Fig. 4B). The observed semicircular graph at the high-frequency region is attributed to the restricted phase of e- transfer, and the diameter is equivalent to Rct. The resistance of the interface to transmit charges from the electrolyte to the electrode is associated with surface variation. Therefore, a nanomaterial with a higher conductivity will have a lower resistance. Comparing the larger semicircle observed at the high-frequency region of the bare Au electrode (curve a) shows that the PEDOT and rGO/PEDOT/Au electrode exhibited a lower resistance than the bare Au electrode, because rGO and PEDOT had a higher conductivity (curves b and c). This shows the successful fabrication of rGO/PEDOT. In addition, a decrease in resistance was further observed for Pdnano decorated with rGO/PEDOT on the electrode due to the high conductivity of the metal nanoparticles (curve d). The insulating Ab layer on the Pdnano/rGO/PEDOT/Au electrode (curve e) further blocks the transfer of electrons, resulting in a higher resistance. The BabA Ab layer on the modified electrode promotes the obstruction of electron and mass transfer. In addition, modification with BabA Ab insulates the conductive electrode surface, causing an increase in the resistance by inhibiting oxidation and reduction reactions on the electrode surface29. The change in the Rct of the electrode assembly confirms the successful immobilization of BabA Ab onto the modified Au electrode, exhibiting results that differed from the cyclic voltammetry results shown in Fig. 4A.
Optimization of immunoassay parameters
Immunosensing behavior with various concentrations of the BabA antigen
Figure 5A displays the graph (CV) of various concentrations of BabA Ag added to the BabA Ab@Pdnano/rGO/PEDOT/Au electrode; the scanning rate was 20 mV/s, the potential ranged from − 0.2 V to + 0.4 V in PBS at a pH of 7.5 (0.5 mM), and the electrolyte was 5 mM potassium ferrocyanide/ferricyanide. As depicted in the figure, the current response decrease is proportionally to the increasing BabA Ag concentration. At the optimum concentration, Ag could interact with the Ab immobilized on the modified electrode, and the formed Ag-Ab complex serves as an inert kinetic obstacle to e- transmission in the ferricyanide mediator. As shown in the graphical image, the electrochemical response decreased with increasing the BabA Ag concentration from the lowest concentration (0.2 ng/mL) to the highest concentration (20 ng/mL), indicating that the LOD is 0.2 ng/mL (S/N = 3).
(A) Biosensing response studies (CV) of BabA Ab@Pdnano/rGO/PEDOT/Au electrode with different standard BabA Ag concentrations at 20 mVs−1 between -0.2 and + 0.4 V in [Fe(CN)6]3−/4− electrolyte containing 0.5 mM sodium phosphate buffer (pH 7.5). (B) Standard curve of BabA Ab@Pdnano/rGO/PEDOT/Au electrode with different BabA Ag concentrations at 20 mVs−1 between -0.2 and + 0.4 V in 5 mM [Fe(CN)6]3−/4− electrolyte containing 0.5 mM sodium phosphate buffer (pH 7.5).
Figure 5B shows an image of a standard plot of the fabricated immunosensor based on the result obtained in Figure 5A. In the concentration range, an excellent linear correlation is exhibited. The current shift of the formed immunosensor is linear for Ag concentrations ranging from 0.2 to 20 ng/mL. The linear equation, correlation coefficient, linear range and sensitivity were calculated from the calibration curve. The sensitivity of 0.375 mA/ng mL−1 was calculated.
The developed BabA Ab@Pdnano/rGO/PEDOT/Au electrode immunosensor has shown good linearity and a low limit of detection. This sensor is more advanced than previously reported sensors30,31. Previously, the working range of the sensor was reported to be between 0.72–7.92 μg/mL.
Study of the scan rate of the immunosensor
The standard CV graphs in Fig. 6A show the electrochemical studies of the modified Au electrode in PBS with a pH of 7.5 (0.5 mM), which were performed using 5 mM potassium ferrocyanide/ferricyanide as the electrolyte, 10 to 100 mV/s scanning rates and a potential range from − 0.2 V to + 0.4 V. Anodic (oxidation) and cathodic (reduction) peaks were obtained at similar potentials, while the current increased by increasing the scan rate, supporting the high stability of the immunosensor. The enhancement of the peak current with increasing the scan rate demonstrated the linearity of the diffusion-controlled mechanism. In particular, the peak-to-peak separation was enhanced with increasing the scanning frequency up to 100 mV/s.
(A) The CV of BabA Ab@Pdnano/rGO/PEDOT/Au electrode scanned in [Fe(CN)6]3−/4− electrolyte containing 0.5 mM sodium phosphate buffer (pH 7.5) with 0.2 ng/mL BabA Ag at different scan rates from 10 to 120 m V s-1 between -0.2 and + 0.4 V. (B) 3D representation of the cyclic voltammogram data of BabA Ab@Pdnano/rGO/PEDOT/Au electrode scanned in 5 mM [Fe(CN)6]3−/4− electrolyte containing 0.5 mM sodium phosphate buffer (pH 7.5) with 0.2 ng/mL BabA Ag at different scan rates from 10 to 100 m V s-1. (C) The dependency of peak currents on variation of peak oxidation (Ipa) and reduction (Ipc) as a function of square root of scan rate.
Figure 6B represents a 3D graph of the sensor with different scanning rates. It is observed that with the increase in the scanning rate, there is also an increase in the current response. To verify the reproducibility, each experiment was replicated three times.
Figure 6C suggests that the maximum redox peak current is reached from 10 to 100 mV/s, showing the direct proportionality with the square root of the scan rate. The regression coefficient (R2) values and the linear formula of the peak current with respect to the square root of the scanning rate were expressed (0.932 and 0.992 for anodic and cathodic currents, respectively) for the Pdnano/rGO/PEDOT/Au electrode.
Effect of pH, temperature and time of incubation
The optimization of the diagnosing conditions was developed by conducting experiments with the immunosensor for three parameters, including pH, temperature and incubation time.
Figure S1 depicts the electrochemical performance of the immunosensor in phosphate buffers with different pH values from 6 to 8.5. The working electrode was dipped into a 5 mM potassium ferrocyanide/ferricyanide electrolyte containing 5 mL of phosphate buffer solution (with pH values ranging from 6 to 8.5) and 0.2 ng/mL BabA Ag. The current response decreased by changing the pH range from 6.0 to 7.5. The low current value represents the strong binding or optimum binding of Ag-Ab over the working electrode. Furthermore, the electrochemical response/current was increased (due to the decrease in Ag-Ab binding) with increasing the pH value up to 8.5. This occurred because of the denaturation of Ab and loss in its activity or stability due to the extremely acidic or basic surrounding. Thus, pH 7.5 was chosen as the most favorable pH for the present immunoelectrode.
Figure S2 demonstrates the effect of incubation temperature (from 10 to 40 °C) on the immunoelectrode in an electrolyte solution of 5 mM potassium ferrocyanide/ferricyanide and PBS (pH 7.5) containing 0.2 ng/mL BabA Ag. Once the temperature was increased from 10 °C to 30 °C, the current response gradually decreased, and later on, the electrochemical response increased up to 40 °C. At temperatures between 10 and 30 °C, the optimal binding of the immunocomplex (Ab-Ag binding) occurred, and thereby, the current response diminished. However, at high temperatures (more than 30 °C), the irreversible denaturation of proteins (BabA Ab and BabA Ag) may occur over the working electrode. Hence, 30 °C was selected as the optimum temperature for the present immunoelectrode.
Figure S3 shows the effect of incubation time on the immunosensor in a 5 mM potassium ferrocyanide/ferricyanide electrolyte solution containing PBS (pH 7.5) and 0.2 ng/mL BabA Ag. Once the BabA antibody on the electrode surface encounters BabA Ag in the solution with a short incubation time, the interaction of the Ag–Ab immunocomplex could not occur. As a result, the current response decreases with the increase in the incubation time up to 30 min and gradually stabilizes. After 30 min of incubation, the uniformity and similarity in the pattern of the current response (Figure S3) shows saturation in the binding of the Ag-Ab complex.
Real sample analysis
To perform the clinical assessment of H. pylori, human stool samples (n = 5) were retrieved from Bio-Diagnostics Laboratory, New Delhi. To estimate the precision, the relative standard deviation (RSD) procedure was adopted so that the BabA antigen was tested 5 times in the samples of human stools. The accuracy of the present sensor was tested using the standard approach by applying a 1.0 ng/mL standardized BabA Ag solution to the relevant samples through a recovery test. The precision was calculated within and between batch processes by a repeatability method. Within the batch processes, the analyte was added 7 times at regular intervals of 1 h, and between batch processes, the analyte was added to the incubation mixture at regular intervals of 24 h; the response was evaluated. The average recovery was estimated five times by conducting the same experiments. Consequently, a 2.1% RSD and 98.5% recovery were calculated.
The selectivity of the immunosensor was examined with various interferents. The study was conducted using solutions of CagA, VacA, alpha-fetoprotein (AFP), BSA, glucose, triglyceride and ascorbic acid containing 1.0 ng/mL interfering compounds, and then, detection in PBS was performed by the immunosensor (pH 7.5). The findings are revealed in Fig. 7. Compared to the electrochemical response caused by BabA Ag, the current variations caused by other antigens were less than 20%. Up to the concentrations of the interfering agents, which were five times higher than those of BabA, the high specificity of the immunosensor interface was still retained.
Storage stability and reproducibility of the BabA Ab@Pdnano/rGO/PEDOT/Au electrode
The shelf-life of the BabA Ab@Pdnano/rGO/PEDOT/Au electrode was tested with a 0.2 ng/mL standard BabA Ag solution in PBS at a regular interval of 7 days until the 9th week. A stable current value was noticed, and the activity of the immunoelectrode was calculated. The immunoelectrode preserved up to 70 percent of its activity; after it was used 20 times, the electrode was stored at 4 °C for 8–9 weeks. After testing, the electrode surface was washed with phosphate buffer solution. Figure S4 shows the decrease in the immunoelectrode activity by the denaturation of the BabA protein, while the BabA Ab@Pdnano/rGO/PEDOT/Au electrode's instability was retained.
The reliability of the immunosensor was evaluated by performing a reproducibility study on the BabA Ab@Pdnano/rGO/PEDOT/Au electrode. The reproducibility was examined by using the same concentration of BabA antigen (0.2 ng/mL) with five equally prepared working electrodes. As illustrated in Fig. S5, the 5 individually designed immunosensors had roughly identical current responses, and the interassay coefficient of variance was 5.5%. This RSD value reveals the good repeatability of the proposed immunosensor.
To summarize, we have successfully established the mechanism for constructing, optimizing and implementing a fast, quick, easy and highly selective label-free immunosensing device for sensitive H. pylori detection. The experiments were designed to develop a noninvasive method of H. pylori detection. Initially, the interaction characteristics of the electrode system were successfully determined through computational simulations. The layer-by-layer electrode assembly (Pdnano/rGO/PEDOT/Au electrode) generates a stable complex formation with the BabA Ag-Ab complex (− 1386.59 kJ/mol). In addition, the binding energy calculation confirms the capability of the hybrid nanomaterial-based electrode (Pdnano/rGO/PEDOT/Au electrode) to enhance the immunosensing performance towards the targeted biomolecule.
The modified immunosensor, as suggested by the hybrid nanomaterials, had an excellent current response to BabA Ag, which showed a broad linearity and an LOD of 0.2 ng/mL. In addition, the developed immunosensor has good specificity, stability and reproducibility. Furthermore, the application was evaluated in stool samples, showing that the sensor is easy to handle and selective for H. pylori diagnosis.
Materials and methods
Chemicals and reagents
Biologics SP-150 was used for all the electrochemical analyses (CV and impedance). 3,4-Ethylenedioxythiophene (EDOT), K2PdCl4, ethylene glycol, poly(N-vinyl-2-pyrrolidone) (PVP) and BSA were procured from Sigma Aldrich, USA. The Bab A antibody and antigens were purified at the Max von Pettenkofer Institute, Munich, Germany. Liquor ammonia, graphite, methanol, K3Fe(CN)6, and K4Fe(CN)6 were obtained from SRL, Mumbai, India. Pure 23 carat gold wire (with a 0.3 cm2 geometric area) was acquired from a local shop in New Delhi, India. All the solutions were prepared using deionized water (DI).
Nanomaterial modeling
All the nanomaterial models involving the Au cluster, PEDOT, rGO and Pdnano were constructed using Materials Studio32. The BabA antigen and antibody were designed using Discovery Studio 201733 software packages (Accelrys Inc., San Diego, CA, USA). These software accurately mimic the experimental electrodes. Specifically, the Au cluster, used as an archetypical electrode, was built with its (111) facet being the most preferential orientation34,35 and was composed of a 56 × 49 unit cell with a C-plane orientation. On the other hand, the PEDOT polymer was constructed using the build polymer function provided by the software. In the simulated polymer model, the fifty PEDOT chains were built by using an optimized single chain consisting of 20 monomers (EDOT; 3,4-ethylenedioxythiophene). For rGO, the structure was initially constructed from the graphite crystal structure, which turned into a single nonperiodic layer of graphene containing 7,266 carbon atoms. According to previous experimental reports36,37,38, the ratio of carbon to oxygen of rGO is higher than that of GO, which is approximately 12:1. For this purpose, the oxygen functional groups were grafted randomly on top of the basal points of the graphene sheet layer. The rGO layer was adjusted by adding hydrogen atoms and subsequently optimized with similar protocols for the remaining layers. Furthermore, the Pdnano particles were constructed with the (111) plane, which is its most energetically favorable facet according to theoretical39,40 and experimental studies41. The Pd(111) cluster was built along with the 58 × 51-unit cell and C-plane orientations to facilitate the periodic boundary box conditions with the following parameters; a = 160.3 Å, b = 140.4 Å, c = 259.5 Å. Subsequently, the substrate layers of the studied nanostructure were created using the build layer function, with a highly accurate lattice mismatch of less than 5%. For the first layer, the Au(111) substrate was used, and for the second layer, the PEDOT polymer was modified on the surface of the Au(111) cluster to build PEDOT/Au layers. Subsequently, rGO was affixed in a planar fashion onto the PEDOT/Au surface to build the rGO/PEDOT/Au layer, with rGO acting as the third layer. Finally, Pd(111) was added to the rGO/PEDOT/Au layer to computationally fabricate the complete scheme of the Pdnano/rGO/PEDOT/Au layer, as depicted in Fig. 1A (a).
For the biomolecules, molecular docking was essential to search the binding modes within Ag and Ab. Docking studies were performed to predict the interaction between the receptor and the ligand complex, which is somehow difficult to determine experimentally. The crystal structures of Ag and Ab were derived from the protein data bank with PDB codes of 5F7N and 5F7K, respectively22. For the docking simulation, Ag and Ab were selected and subsequently prepared as the input structures using ZDOCK and ZRANK modules. ZDOCK employs shape-complementary scoring methods advanced by the electrostatic and desolvation energy terms, while for ZRANK, the docked poses were reranked, resulting in an accurate bound complex structure. The best model of the largest cluster that fulfills the active binding region of Ab (Fig. 1(b)) was selected with respect to the binding energy investigations along with the nanomaterial structures.
Metropolis Monte Carlo simulations
Metropolis Monte Carlo (MMC) adsorption studies were applied for bulk preparations. These studies were carried out to identify the lowest energy configurations of the adsorbate on the surface of the selected substrate while a gradual decrease in temperature occurred. In this analysis, the temperature was controlled automatically with three temperature cycles during the simulated annealing process. The simulated substrate and adsorbate were the electrode layers and the Ag-Ab complexes, respectively. The overall adsorption studies were performed by using the Universal force field42,43, which corresponds to a force field across the periodic table of elements and is reliably applied in structures having metals and organic systems44,45. The energy and gradient tolerances were set to 0.001 kcal/mol and 0.5 kcal/mol/Å, while the current method allocated the charges. To measure the nonbonding electrostatic and van der Waals interactions in the MMC simulations, the group-based summation process is used for the energy parameters. These parameters were applied for all the substrate layers (rather the layer-by-layer system) with similar adsorbates, i.e., the Ag–Ab binding pose.
The total energy of the complex was calculated as follows: after the optimum configuration of the substrate-adsorbate system was determined, the single point energy calculation was carried out using the COMPASS force field, which accurately computes the total energy of systems. Thereafter, a similar calculation was also applied for the adsorbate molecule once the substrate was removed and vice versa.
Reduced graphene oxide (rGO) preparation
Hummer’s method was used to prepare 0.1 g of dried GO powder46,47; the dried GO powder was dispersed in 150 mL of DI water by sonicating for 25 to 30 min to prepare a brown uniform solution. Dropwise addition of 1.0 mL of hydrazine monohydrate to the GO solution was performed, and the resulting solution was further immersed in an oil bath (80 °C). Once immersed, the reaction mixture was stirred for 12 h. The color of the liquid changed from brown to black. Black particles settled in the solution within a few minutes. A black powder was collected in a vacuum oven at 60 °C by vacuum filtration and was further dried for 24 h48.
Interface design of the immunoelectrode
Electropolymerization of PEDOT over the Au electrode
An alumina slurry (0.05 μM) was applied to cleaning the Au electrode, which was later immersed in a piranha solution (H2O2 (30%) and concentrated H2SO4, 3:1 proportion (v/v)) for ten minutes and subsequently cleaned by ultrasonication with DI water. The Au electrode was further cycled electrochemically in H2SO4 (1.0 M) solution until a stable gold oxide formation/reduction CV graph was achieved. The electropolymerization of EDOT on the Au electrode was carried out in an aqueous deaerated EDOT monomer solution (0.1% w/v) containing NaPSS (0.7% w/v). To completely dissolve the monomers of EDOT, the solubility of the aqueous deaerated solution should be less than the EDOT solubility (15 mM) in H2O at 25 °C. A cyclic voltammetry study was then performed at a scan rate of 20 mV/s for 10 cycles between − 0.9 and 1.2 V49.
Electrochemical deposition of rGO on the PEDOT-modified Au electrode
The film electrodeposition process was performed by submerging the electrode in a solution containing rGO (10 mL), and then, polymerization was achieved by applying 10 cycles from 0 V to − 1 V at a scan rate of 50 mV/s. The polymer-deposited electrode was rinsed with DI water after electropolymerization.
Electrochemical deposition of Pdnano
Pdnano was deposited through an electrochemical method (cyclic voltammetry). The modified rGO/PEDOT/Au electrode was plunged into a mixture that contained 1 × 10−3 M K2PdCl4 supplemented with 0.5 M H2SO4. The voltage was fixed for 10 cycles (from − 0.25 V to 1.2 V at a scan rate of 50 mV/s). After the Pdnano/rGO/PEDOT/Au electrode was prepared, it was cleaned with DI water and air dried at room temperature.
BabA (antibody) immuno-interaction with the Pdnano/rGO/PEDOT/Au electrode
The surface of the immunoelectrode was covered by 1.0 μL of the BabA antibody (1:1000 dilution) and was then incubated at 4 °C for 12 h. The unbound antigens were removed by slowly immersing the electrode in PBS buffer (0.1 M, pH 7.5). The BabA Ab@Pdnano/rGO/PEDOT/Au electrode was subsequently processed with PBS containing 1% BSA for one hour to block nonreactive and nonspecific areas. Eventually, the working electrode was washed, dried and used for further characterizations and experimentation 50.
H. pylori detection in real samples
Five H. pylori-positive stool samples were collected from Bio-diagnostics Laboratory, New Delhi, India. The presence of bacterial infections in the patients was diagnosed for the 5 stool samples by a commercially available rapid urease test, and it was further confirmed by evaluation through a clinical pathologist. To prepare the stool samples, 0.1 g of stool was immersed in 0.5 ml of PBS (0.1 M, pH 7.2) and vortexed for 15 s. The suspension was centrifuged for 5 min at 5000 rpm. The formed pellets were discarded, and the clear supernatant was used in the experiment.
Analytical properties of the immunosensor
The LOD of the BabA Ab@ Pdnano/rGO/PEDOT/Au electrode is elucidated as the minimum quantity of BabA Ag required to provide the current value of the background (blank) + 3 times the SD of the blank. The LOD is calculated by using both the estimated limit of the blank (LoB) and sample test replicates known to have a low analyte concentration. The limit of quantification (LoQ) is similar to the LOD or much higher51. The minimum concentration required was 0.2 ng/mL.
References
Damasceno, J. P. L., Rodrigues, R. P., Gonçalves, R. D. C. R. & Kitagawa, R. R. Anti-helicobacter pylori activity of isocoumarin paepalantine: morphological and molecular docking analysis. Molecules 22, 786 (2017).
Steiger, C. et al. Ingestible electronics for diagnostics and therapy. Nat. Rev. Mater. 4, 83–98 (2019).
Malaty, H. M. Epidemiology of Helicobacter pylori infection. Best Pract. Res. Clin. Gastroenterol. 21, 205–214 (2007).
Ahmed, F. et al. Advances in research progress of H. pylori. Br. J. Pharm. Res. 21, 1–8 (2018).
5Plummer, M., Franceschi, S. & Muñoz, N. Epidemiology of Gastric Cancer, 311–326 (IARC scientific publications, 2004).
Vafaee, F. et al. A data-driven, knowledge-based approach to biomarker discovery: application to circulating microRNA markers of colorectal cancer prognosis. NPJ Syst. Biol. Appl. 4, 20 (2018).
Niu, Q. et al. Immune response in H. pylori-associated gastritis and gastric cancer. Gastroenterol. Res. Pract. 2020, 9342563 (2020).
Epplein, M. et al. Prospective study of Helicobacter pylori biomarkers for gastric cancer risk among Chinese men. Cancer Epidemiol. Prev. Biomark. 21, 2185–2192 (2012).
Kao, C.-Y., Sheu, B.-S. & Wu, J.-J. Helicobacter pylori infection: an overview of bacterial virulence factors and pathogenesis. Biomed. J. 39, 14–23 (2016).
Dunne, C. et al. Binding of Helicobacter pylori to human gastric mucins correlates with binding of TFF1. Microorganisms 6, 44 (2018).
Skoog, E. C. et al. BabA dependent binding of Helicobacter pylori to human gastric mucins cause aggregation that inhibits proliferation and is regulated via ArsS. Sci. Rep. 7, 40656 (2017).
Ansari, S. & Yamaoka, Y. Helicobacter pylori virulence factors exploiting gastric colonization and its pathogenicity. Toxins 11, 677 (2019).
Wang, H. et al. Label-free immunosensor based on Pd nanoplates for amperometric immunoassay of alpha-fetoprotein. Biosens. Bioelectron. 53, 305–309 (2014).
Zhou, C. et al. A sensitive label–free amperometric immunosensor for alpha-fetoprotein based on gold nanorods with different aspect ratio. Sci. Rep. 5, 9939 (2015).
Moon, S. W. et al. United rapid urease test is superior than separate test in detecting Helicobacter pylori at the gastric antrum and body specimens. Clin. Endosc. 45, 392 (2012).
Ontsira Ngoyi, E. N. et al. Molecular detection of Helicobacter pylori and its antimicrobial resistance in Brazzaville, Congo. Helicobacter 20, 316–320 (2015).
Binh, T. T. et al. Molecular epidemiology of Helicobacter pylori infection in a minor ethnic group of Vietnam: a multiethnic, population-based study. Int. J. Mol. Sci. 19, 708 (2018).
Leal, Y. A. et al. Utility of stool sample-based tests for the diagnosis of Helicobacter pylori infection in children. J. Pediatr. Gastroenterol. Nutr. 52, 718–728 (2011).
Garza-González, E., Perez-Perez, G. I., Maldonado-Garza, H. J. & Bosques-Padilla, F. J. A review of Helicobacter pylori diagnosis, treatment, and methods to detect eradication. World J. Gastroenterol. WJG 20, 1438 (2014).
Bathinapatla, A., Kanchi, S., Singh, P., Sabela, M. I. & Bisetty, K. An ultrasensitive performance enhanced novel cytochrome c biosensor for the detection of rebaudioside A. Biosens. Bioelectron. 77, 116–123 (2016).
Ashaduzzaman, M. et al. On/off-switchable LSPR nano-immunoassay for troponin-T. Sci. Rep. 7, 44027 (2017).
Moonens, K. et al. Structural insights into polymorphic ABO glycan binding by Helicobacter pylori. Cell Host Microbe 19, 55–66 (2016).
Bogdan, D. & Morari, C. Electronic properties of DNA nucleosides adsorbed on a Au (100) surface. J. Phys. Chem. C 116, 7351–7359 (2012).
Tang, S. & Zhu, J. Structural and electronic properties of Pd-decorated graphene oxides and their effects on the adsorption of nitrogen oxides: insights from density functional calculations. RSC Adv. 4, 23084–23096 (2014).
Pan, W., Zhang, X., Ma, H. & Zhang, J. Electrochemical synthesis, voltammetric behavior, and electrocatalytic activity of Pd nanoparticles. J. Phys. Chem. C 112, 2456–2461 (2008).
Chiou, Y.-J. et al. Synthesis and electrocatalytical application of hybrid Pd/metal oxides/MWCNTs. Int. J. Electrochem. 2018, 8416268 (2018).
Zhao, Y. et al. Dual amplified electrochemical immunosensor for highly sensitive detection of Pantoea stewartii sbusp. stewartii. ACS Appl. Mater. Interfaces 6, 21178–21183 (2014).
Chauhan, N., Chawla, S., Pundir, C. & Jain, U. An electrochemical sensor for detection of neurotransmitter-acetylcholine using metal nanoparticles, 2D material and conducting polymer modified electrode. Biosens. Bioelectron. 89, 377–383 (2017).
Putzbach, W. & Ronkainen, N. J. Immobilization techniques in the fabrication of nanomaterial-based electrochemical biosensors: a review. Sensors 13, 4811–4840 (2013).
Ly, S. Y., Yoo, H.-S. & Choa, S. H. Diagnosis of Helicobacter pylori bacterial infections using a voltammetric biosensor. J. Microbiol. Methods 87, 44–48 (2011).
Shahrashoob, M. et al. Detection of Helicobacter pylori genome with an optical biosensor based on hybridization of urease gene with a gold nanoparticles-labeled probe. J. Appl. Spectrosc. 83, 322–329 (2016).
Biovia, D. S. 2016. Material Studio modelling. v.16.1.0.21 ed. San Diego: Dassault Systemes.
Biovia, D. S. 2016. Discovery Studio modelling. v.16.10.15350 ed. San Diego: Dassault Systemes.
Grönbeck, H., Curioni, A. & Andreoni, W. Thiols and disulfides on the Au (111) surface: the headgroup−gold interaction. J. Am. Chem. Soc. 122, 3839–3842 (2000).
Wang, Y. et al. Spatially resolved electronic and vibronic properties of single diamondoid molecules. Nat. Mater. 7, 38 (2008).
Mattevi, C. et al. Evolution of electrical, chemical, and structural properties of transparent and conducting chemically derived graphene thin films. Adv. Funct. Mater. 19, 2577–2583 (2009).
Pei, S. & Cheng, H.-M. The reduction of graphene oxide. Carbon 50, 3210–3228 (2012).
Tiwari, A. & SyvÃ, M. Graphene Materials: Fundamentals and Emerging Applications (Wiley, Hoboken, 2015).
Li, M., Guo, W., Jiang, R., Zhao, L. & Shan, H. Decomposition of ethanol on Pd (111): a density functional theory study. Langmuir 26, 1879–1888 (2009).
Wang, S., Vorotnikov, V. & Vlachos, D. G. Coverage-induced conformational effects on activity and selectivity: hydrogenation and decarbonylation of furfural on Pd (111). ACS Catal. 5, 104–112 (2014).
Davis, J. & Barteau, M. Polymerization and decarbonylation reactions of aldehydes on the Pd (111) surface. J. Am. Chem. Soc. 111, 1782–1792 (1989).
Rappé, A. K., Casewit, C. J., Colwell, K., GoddardIii, W. & Skiff, W. UFF, a full periodic table force field for molecular mechanics and molecular dynamics simulations. J. Am. Chem. Soc. 114, 10024–10035 (1992).
Rappe, A., Colwell, K. & Casewit, C. Application of a universal force field to metal complexes. Inorg. Chem. 32, 3438–3450 (1993).
Bristow, J. K., Tiana, D. & Walsh, A. Transferable force field for metal–organic frameworks from first-principles: BTW-FF. J. Chem. Theory Comput. 10, 4644–4652 (2014).
Coupry, D. E., Addicoat, M. A. & Heine, T. Extension of the universal force field for metal–organic frameworks. J. Chem. Theory Comput. 12, 5215–5225 (2016).
Hummers, W. S. & Offeman, R. Functionalized graphene and graphene oxide: materials synthesis and electronic applications. J. Am. Chem. Soc. 80, 1339–1339 (1958).
Jain, U., Gupta, S. & Chauhan, N. Detection of glycated hemoglobin with voltammetric sensing amplified by 3D-structured nanocomposites. Int. J. Biol. Macromol. 101, 896–903 (2017).
Guo, C. X. & Li, C. M. A self-assembled hierarchical nanostructure comprising carbon spheres and graphene nanosheets for enhanced supercapacitor performance. Energy Environ. Sci. 4, 4504–4507 (2011).
Castagnola, V., Bayon, C., Descamps, E. & Bergaud, C. Morphology and conductivity of PEDOT layers produced by different electrochemical routes. Synth. Met. 189, 7–16 (2014).
Geng, P. et al. Self-assembled monolayers-based immunosensor for detection of Escherichia coli using electrochemical impedance spectroscopy. Electrochim. Acta 53, 4663–4668 (2008).
Armbruster, D. A. & Pry, T. Limit of blank, limit of detection and limit of quantitation. Clin. Biochem. Rev. 29, S49 (2008).
Acknowledgements
The authors of this manuscript are thankful to Dr. Wolfgang Fischer and Ms. Evelyn Weiss of the Max von Pettenkofer Institute of Hygiene and Medical Microbiology, Ludwig Maximilians University, Munich, Germany for providing the BabA antibody and BabA antigen for the present analysis. Furthermore, the authors are obliged to Dr. Rainer Haas for his valuable motivation and help. This work was supported by DST, the Govt. of India (F.No. TDP/BDTD/33/2019). The authors are also grateful to the Centre for High-Performance Computing (CHPC), Cape Town, South Africa for providing computational power and the facilities necessary for the research work.
Author information
Authors and Affiliations
Contributions
Ms. S.G., Dr. A.T. and Dr. N.C. conceived and designed the experiments; Ms. S.G. performed the experiments; Ms. S.G. analyzed the data; Ms. S.G. contributed reagents/materials/ analysis tools; Ms. S.G. wrote the paper, which were further corrected by Dr. N.C. and Dr. U.J.; B.T.M., A.D.P. helped in simulations included molecular docking and Metropolis Monte Carlo adsorption studies.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gupta, S., Jain, U., Murti, B.T. et al. Nanohybrid-based immunosensor prepared for Helicobacter pylori BabA antigen detection through immobilized antibody assembly with @ Pdnano/rGO/PEDOT sensing platform. Sci Rep 10, 21217 (2020). https://doi.org/10.1038/s41598-020-78068-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-78068-w
This article is cited by
-
Simulation of electrical conductivity for polymer silver nanowires systems
Scientific Reports (2023)
-
Carbon-Based Nanomaterials in Wound Care Management: A New and Pristine Strategy
Biomedical Materials & Devices (2023)
-
Biosensors, modern technology for the detection of cancer-associated bacteria
Biotechnology Letters (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.