Abstract
Animal tuberculosis (TB), caused by Mycobacterium bovis, is maintained in Portugal in a multi-host system, with cattle, red deer and wild boar, playing a central role. However, the ecological processes driving transmission are not understood. The main aim of this study was thus to contribute to the reconstruction of the spatiotemporal history of animal TB and to refine knowledge on M. bovis population structure in order to inform novel intervention strategies. A collection of 948 M. bovis isolates obtained during long-term surveillance (2002–2016, 15 years) of cattle (n = 384), red deer (n = 303) and wild boar (n = 261), from the main TB hotspot areas, was characterized by spoligotyping and 8 to 12-loci MIRU-VNTR. Spoligotyping identified 64 profiles and MIRU-VNTR distinguished 2 to 36 subtypes within each spoligotype, enabling differentiation of mixed or clonal populations. Common genotypic profiles within and among livestock and wildlife in the same spatiotemporal context highlighted epidemiological links across hosts and regions, as for example the SB0119-M205 genotype shared by cattle in Beja district or SB0121-M34 shared by the three hosts in Castelo Branco and Beja districts. These genomic data, together with metadata, were integrated in a Bayesian inference framework, identifying five ancestral M. bovis populations. The phylogeographic segregation of M. bovis in specific areas of Portugal where the disease persists locally is postulated. Concurrently, robust statistics indicates an association of the most probable ancient population with cattle and Beja, providing a clue on the origin of animal TB epidemics. This relationship was further confirmed through a multinomial probability model that assessed the influence of host species on spatiotemporal clustering. Two significant clusters were identified, one that persisted between 2004 and 2010, in Beja district, with Barrancos county at the centre, overlapping the central TB core area of the Iberian Peninsula, and highlighting a significant higher risk associated to cattle. The second cluster was predominant in the 2012–2016 period, holding the county Rosmaninhal at the centre, in Castelo Branco district, for which wild boar contributed the most in relative risk. These results provide novel quantitative insights beyond empirical perceptions, that may inform adaptive TB control choices in different regions.
Similar content being viewed by others
Introduction
The control and eradication of infectious diseases can be very difficult for pathogens that are able to thrive across multiple host species. This is the case of animal tuberculosis (TB), an infectious disease mainly caused by Mycobacterium bovis (M. bovis), which circulates within livestock and across several wildlife species worldwide, through direct and indirect routes1,2,3,4. Animal TB affects livestock productivity and trading, animal welfare, and also biodiversity, as it globally or locally threatens ecosystem services. The zoonotic potential of M. bovis bacteria is also liable as new infection human cases are reported every year5,6.
Cattle (Bos taurus) is described as the main host species for M. bovis, while several wildlife hosts can support pathogen maintenance, depending on the ecosystem and epidemiological scenario. Over the years several molecular reports demonstrated inter-species transmission between cattle and wildlife, and spatial interaction studies associated wildlife as a risk factor for TB infection in cattle. Badger (Meles meles) in United Kingdom and Republic of Ireland; African buffalo (Syncerus caffer) in South Africa, brushtail possums (Trichosurus vulpecula) in New Zealand, white-tailed deer (Odocoileus virginianus) in EUA, and red deer (Cervus elaphus) and wild boar (Sus scrofa) in the Iberian Peninsula (Portugal and Spain) are the best characterized wildlife reservoirs5,7,8,9,10. In Iberian ecosystems, cattle and non-bovine livestock species managed in extensive husbandry, and red deer, wild boar, fallow deer and small carnivores compose the host community that can support M. bovis maintenance11. Some spatial clusters of infection in central and south-eastern regions of Portugal, in the periphery of the Iberian TB core area, have been detected, where average TB prevalence may reach as much as 50% in wild ungulate populations alone7,10,12. Expansion from the core Iberian area is believed to be driven by high host densities, artificial management and super-shedder wild populations13,14.
In Portugal, a national eradication program for animal TB in cattle population commenced in 1987, being presently based on the detection and culling of reactors to the single intradermal comparative tuberculin (SICCT) test and the interferon-γ test of cattle aged over 6 weeks, also including routine surveillance at slaughterhouses, mandatory sanitary classification of herds and regions, compulsory slaughter of positive animals, compensation to owners of slaughtered animals and pre-movement assessments15. Bacteriological culture and histopathology are the officially accepted diagnostics to confirm animal TB infection. Portugal has made significant progress in the control of bovine TB since the nationwide implementation of the eradication program approved by the European Union in 1992 (Council Decision 92/299/CE). Currently, the program covers mainland Portugal and the autonomous regions of Azores and Madeira16. During the last decade, TB herd prevalence remained low, varying between 0.11% in 2008 and 0.29% in 2017, with a peak in 2010 (0.91%)17. However, the national epidemiological indicators reflect different contexts throughout mainland Portugal, with TB herd prevalence ranging from 0.08% in the North up to 1.24% in Alentejo (south) in 201717. Algarve, in the extreme south of Portugal, is the single region that already achieved the officially TB-free status (Decision 2012/204/EU), where a specific surveillance program applies for the maintenance of this status (Fig. 1). The eradication program has suffered changes along the years since its first implementation. While in vivo surveillance and SICCT testing used to be homogenous across cattle herds, it has recently adapted towards the evolution of the epidemiological situation. There are now regional differences in the application of the surveillance scheme, according to herd type (beef, dairy, breeding), TB history and geographical location.
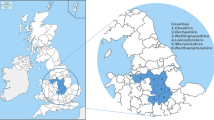
Map of mainland Portugal highlighting the high TB prevalence area for cattle and the epidemiological risk area for wild ungulates (red deer and wild boar) and evidencing the geographic distribution of M. bovis isolates under analysis. The middle plots represent the evolution of the total number of M. bovis isolated from the different host species (red for cattle (C), green for red deer (RD) and blue for wild boar (WB)) sampled per geographic region, and the plot at the right compares the global number of M. bovis isolates per geographic region.
The heterogeneity of TB herd prevalence across mainland Portugal indicates that some regional circumstances may favour intraspecific and interspecific M. bovis transmission, such as: (1) the management in extensive husbandry of livestock in the central and south regions that promote aggregation and enable the direct and indirect interaction of domestic with wild animals in communal pastures; (2) the wide host community of wildlife species potentially susceptible to TB that free-range in Portugal; and (3) the overabundance of widely distributed big game species that are artificially managed for hunting purposes, namely red deer and wild boar.
Following several scientific publications reporting M. bovis occurrence in wild populations across two regional foci10,18,19,20 and the field knowledge reported by veterinary services of some municipalities, combined with evidence for growing abundances of wild ungulates and the proliferation of big game hunting, in 2011 the national authorities defined an epidemiological risk area for big game species in south-central Portugal, along the border with Spain, where specific surveillance measures apply21. All big game specimens hunted in this risk area are mandatorily subjected to post-mortem examination in the field by a credentialed veterinarian. Samples from suspected TB lesions are collected and forwarded to the National Reference Laboratory (NRL) to confirm TB. Hunting by-products and animals with TB-compatible lesions (fully rejected) are disposed according to European Regulations (EC) no. 1069/2009 and 854/200421.
The resilience of M. bovis in wildlife hosts remains a challenge for TB eradication in the cattle population and raises cattle and wildlife management issues in addition to public health concerns. The national context in Portugal is of special complexity, since M. bovis is maintained in this multi-host system with at least three maintenance hosts. Their single or joint contribution for M. bovis transmission and spatial patterns are not yet quantified. Although reports of M. bovis occurrence in other mammal species (e.g. caprine, ovine, carnivores) have remained occasional22,23, the role of each species in these transmission processes is unknown for now.
Under these complex circumstances, appropriate epidemiological studies applied to high burden areas are required. So, in this work, we have focused on perceived hotspot areas where the livestock-wildlife interface is prominent, with widely distributed cattle herds of varying dimensions and abundant wild ungulate populations. We take advantage of a large collection of M. bovis isolates which has been gathered for almost two decades in the NRL and that can be exploited at the molecular level. A detailed molecular approach making use of spoligotyping (Spacer oligonucleotide typing) and MIRU-VNTR (Mycobacterial Interspersed Repetitive Units-Variable Number Tandem Repeats)22,23,24,25,26,27, combined with Bayesian clustering inference analyses, phylogenetic and network analyses, was envisaged with the aim to refine knowledge on the M. bovis population structure and to assess the spatiotemporal distribution of the most common genotypes, enabling the inference of the spatial risk, the most epidemiologically relevant hosts and the most probable transmission routes.
Methods
The general workflow followed in this study that involved genotypic characterization by two methods and statistical analyses is described in detail in Fig. 2.
General workflow followed in this study (general template obtained in https://tinyppt.com/).
Study area
The study area comprises the three regions regarded in Portugal as the main hotspot areas for animal TB at the livestock-cattle interface. They are enclosed in the districts (administrative level sample unit) of Castelo Branco, Portalegre and Beja, which locate on inner central and south Portugal, within the epidemiological risk area for TB in big game species that simultaneously holds chronically infected cattle herds that are managed in an extensive regimen (Fig. 1). Castelo Branco was the most represented district, with 535 M. bovis isolates, followed by Portalegre, with 217, and Beja, with 196 isolates (Fig. 1). The study area is simultaneously used for livestock production and big game hunting activities.
Bacterial isolation and identification
Nine hundred and forty-eight M. bovis isolates were recovered (n = 948), in a 15-year period (2002–2016), from three host species, namely cattle (Bos taurus) (n = 384), red deer (Cervus elaphus) (n = 303) and wild boar (Sus scrofa) (n = 261) (Fig. 1).
This 15-year molecular analysis concerns three time periods: period 1 (2002–2008; 7 years) in which herd prevalence steadily decreased and surveillance in wildlife was sporadic and uneven; period 2 (2009 and 2012; 4 years) in which national herd prevalence raised more than fourfold and carcass examination of hunted big game species became mandatory in the epidemiological risk area from 2011 onwards; and, finally, period 3 (2013–2016; 4 years) that was characterized by a decrease in herd prevalence after mandatory augmented surveillance by stakeholders, both at the livestock and wildlife flanks.
Sample collection was performed within an official context according to specific national legislation or in the context of research projects held by the NRL. No animals were slaughtered or hunted for this study purpose. Cattle samples (lungs, lymph nodes, liver and/or spleen) were obtained after sanitary slaughter following previous positive intradermal testing or from evidence of macroscopic TB-like lesions at routine abattoir inspection. Wildlife samples (lungs, lymph nodes, liver and/or spleen) were collected from hunted animals presenting TB-compatible lesions that were detected during veterinarian examination in the field, which is obligatory for 19 counties (municipal level) enclosed in the hotspot areas.
Animal tissue samples were pooled and processed in a biosecurity level 3 facility at INIAV, IP (National Institute for Agrarian and Veterinary Research), the NRL for animal TB diagnosis, following the protocol guidelines recommended in the OIE Manual for Terrestrial Animals (a detailed procedure is described in Cunha et al.7); and inoculated onto Stonebrink (Biogerm, Portugal) and Löwenstein-Jensen pyruvate (Biogerm, Portugal) solid media and liquid medium (BACTEC 9000 MB system, Becton Dickinson Microbiology Systems). Cultures were incubated at 37 °C and inspected weekly for growth for a minimum period of 12 weeks.
Pelleted cultures were heat-killed to extract the DNA and species identification was performed by restriction fragment length polymorphism analysis of PCR-amplified gyrB gene (PCR-REA) as described in Cunha et al.7 or using commercially available systems [GenoType Mycobacterium (Hain diagnostics, Germany)], according to manufacturer instructions. In all PCR batch assays, DNA from two positive controls (M. tuberculosis H37RV and M. bovis BCG) and a no-template control (purified water) were included.
Spatiotemporal cluster analyses
A space–time analysis with a multinomial probability model28 was performed in SaTScan (version 9.6, https://www.satscan.org/) to assess the influence of host species on spatio-temporal risk clustering. In total, 249 TB cases in cattle, grouped in 135 livestock herds, and 474 TB cases in wildlife hosts, grouped in 91 officially-delimited hunting areas, were considered, meaning 76.4% of the global dataset, for which geographic coordinates of the location centroids of herds/hunting areas, were available. A database was created by geocoding each TB case to the geographic coordinates of the corresponding livestock herd or officially-delimited hunting area and no substantial change in geographic coordinates was assumed over the study period. The multinomial probability model applied here evaluates whether there are any clusters where the distribution of TB cases is different from the rest of the region under study. To operationalize the model, the study area is scanned using cylinders of varying sizes, with the base representing space and the height representing time, and the number of cases inside the cylinders is compared to the case expectations outside. Space–time clusters were identified at a maximum temporal and spatial window of 50.0% of the study period and case data, respectively. It was only allowed one set of geographical coordinates per location (herd/hunting area) and no overlapping clusters were permitted. Statistically significant clusters were evaluated at a p-value < 0.05, using Monte Carlo permutations with 999 replications of the dataset, under the null hypothesis of complete spatial randomness.
The map was generated using QGIS (Quantum GIS development Team 2018, version 3.10, https://www.qgis.org/en/site/) to visualize spatial‐temporal clusters and outbreak locations.
Molecular characterization of M. bovis isolates
Spoligotyping was performed with in-house made membranes following the standard published protocol29, with the modifications described by Botelho et al.30. The presence or absence of the 43 spacers contained in the Direct Repeat (DR) locus was represented by a binary code. Spoligotyping patterns (prefix SB followed by four digits) were obtained from the Spoligotype Database website (http://www.mbovis.org/index.php). DNA from M. bovis BCG and M. tuberculosis H37Rv isolates were used as positive controls and purified water as no-template control.
The relative proportion of each one of the top six most common spoligotyping profiles, over the remaining in the different study years was obtained. A global and regional trend analysis was performed by applying a linear regression.
A representative collection of isolates (n = 524) was further subjected to MIRU-VNTR analyses. Two selection criteria were applied: firstly, isolates were selected to cover all spoligotyping groups; and secondly, to cover in equal proportion different host species, geographic regions and year of isolation. Moreover, nine of the 29 orphan spoligotypes (i.e. with only one isolate) were also included.
MIRU-VNTR typing was performed using a previously established set of 8-loci panel constituted by VNTR3232, ETR-A, ETR-B, ETR-C, QUB11a, QUB11b, MIRU4 and MIRU2620. These loci were amplified by PCR, in single reaction tubes, using the primers and cycling conditions previously described by Duarte et al.20. Positive control (DNA extracted from M. bovis BCG) and a no-template control (purified water) were included in all PCR assays. PCR products were separated in 2% agarose gels, prepared in Tris Buffer EDTA 1×, at 90 V for 3 h. The number of tandem repeats in each amplified locus was attributed by correlation with the amplicon size according to previously published tables31.
Complete profiles were grouped by similarity in MIRU-VNTR types (M types). Profiles with non-amplifiable loci were excluded from this classification.
A small subset of the MIRU-VNTR results (< 10%) reported in this study have been previously published by Duarte et al.20 (n = 23, between 2002 and 2007) and by Cunha et al.7 (n = 27, between 2007 and 2009), but were included to span the analyses for a wider period.
A case was considered as polyclonal infection when two amplification products for one or more MIRU-VNTR loci were detected, as in Reis et al.32. In these cases, the presence of double alleles was confirmed by repeating the PCR and electrophoresis experiments. For the subgroup with double alleles (n = 37), four additional loci (MIRU16, MIRU40, ETR-E and QUB26) were added to the initial panel with the purpose of further discriminating the isolates, following the rational indicated by Reis et al.32. Similar to the 8-loci panel, these four additional loci were amplified by PCR, in single reaction tubes, using the primers described by Supply et al.33 for MIRU16, MIRU40 and ETR-E, and Skuce et al.34 for QUB26. The cycling and electrophoresis conditions applied were equal to the ones already described for the previous panel.
The genotypic profiles were defined by the combination of spoligotyping profile and MIRU type.
The calculation of the discriminatory power (D), that is the average probability that the typing method will assign a different type to two unrelated isolates sampled in the population, for both typing methods, and the individual allelic diversity (h) of the loci panel tested were obtained. These parameters were calculated based on Simpson's index of diversity, as described by Hunter and Gaston35, using an online tool available on the website In silico simulation of molecular biology experiments (In silico, 2018)36. Only complete profiles with single alleles were considered for the calculation of the discriminatory power of MIRU-VNTR typing.
Network analyses
To explore the relationships established within each host-geographic region pair, two networks involving spoligotyping (n = 948) or genotypic profiles (n = 487) (i.e. based on spoligotyping and MIRU) were visualized with Gephi (version 0.9.2, https://gephi.org/). Each host-geographic region combination was considered a node, and the number of shared spoligotyping or genotypic profiles were used as connection lines between nodes. The connection lines were established based on absolute values; no statistical transformation was performed.
Phylogenetic analysis
BioNumerics software (version 6.6, http://www.applied-maths.com/bionumerics) (Applied Maths, Sint-Martens-Latem, Belgium) was used to visualize evolutionary relationships among the M. bovis isolates by generating Minimum Spanning Trees (MSTs), using a combined dataset composed by spoligotyping and 8-loci MIRU-VNTR (n = 487). Profiles with double alleles were excluded from these analyses.
A composite character experiment was created with spoligotyping and MIRU-VNTR data, and an advanced cluster analysis was performed using the categorical option as a similarity coefficient. The single locus variant was used as a criterion for maximum distance between nodes.
Bayesian population analysis
To further explore the population structure of M. bovis isolates, a Bayesian clustering algorithm, implemented in the software STRUCTURE (version 2.3.4, https://web.stanford.edu/group/pritchardlab/structure.html)37 was applied to a dataset composed by MIRU-VNTR profiles (n = 487). Bayesian inference method and MST analysis were applied to the same M. bovis isolates. STRUCTURE was used to infer the most likely number of putative ancestral populations (K), and to assign individuals to these populations. It was ran using the admixture model as an ancestry prior model. The admixture model considers that the data may have been originated from the admixture of K putative ancestral populations at some time in the past, so it was selected because it is a more flexible model and, thus, suitable for dealing with the complexities of a real data population. Five independent simulations were performed using Markov Chain Monte Carlo (MCMC) with an initial 10.000 ‘burn-in’ period and 100.000 iterations, following ‘burn-in’. For each simulation, K values were tested between 2 and 15, with 10 replicated runs for each value of K.
To estimate the number of putative ancestral populations, delta K was calculated by the Evanno method38, using STRUCTURE HARVESTER39. The estimate of delta K is based on the rate of change in the log probability of data between successive K values.
The individual Q matrix corresponds to the estimated ancestry coefficients for each individual, that is the part of the genome that each individual inherited from ancestors. A threshold of Q ≥ 0.50 was applied to assign individuals to ancestral populations.
New MST analyses were then performed using BioNumerics software (version 6.6), according to the criterion described in “Phylogenetic analysis” section, and identifying M. bovis by the classification provided by STRUCTURE analysis.
Allelic richness
For allelic richness analyses, M. bovis isolates were grouped according to geographic region and populations defined by STRUCTURE. The mean allelic richness of each M. bovis group was evaluated using the statistical method of rarefaction implemented in the software HP-RARE (version 1.0, https://www.montana.edu/kalinowski/software/hp-rare.html)40, which standardizes samples of different sizes, compensating for sampling disparity, so that the allelic diversity across large and small sample datasets could be compared.
Statistical analyses
The statistical significance between the associations of spoligotyping profiles proportion and geographic region or host species were tested using a chi-square test (α = 0.05). An unbiased estimate of the exact significance level was performed through Monte Carlo approximation with confidence interval of 99% and 10,000 repeated sampling. Chi-square test (α = 0.05) was also used to evaluate the statistical significance of the association between M. bovis ancestral populations defined by STRUCTURE software and geographic region or host species. Contingency tables and tests were performed in IBM SPSS Statistics (version 24.0.0, https://www.ibm.com/analytics/spss-statistics-software). Results were considered to be significant if p < 0.05.
Results
Demography analyses of M. bovis isolates
The M. bovis dataset considered in this study is well represented as far as time, space and animal hosts are concerned, however the strain set of the first period, especially from 2002 to 2006, is restricted (n = 85), corresponding to only 9% of the total dataset, which may be explained by reduced and uneven surveillance during that time period. The number of isolates has two peaks in 2007 and 2010 and values rise since 2013, coinciding with the increase in TB surveillance of the wildlife component.
Until 2006, Beja was the most represented region (cumulative number of isolates = 46), but since then and until 2016, Castelo Branco (n = 511) outnumbered the number of M. bovis isolates recovered (n = 150 for Beja and n = 202 for Portalegre) (Fig. 1).
Cattle was the most represented host species until 2011 (n = 258), then wildlife species were sampled in higher numbers (n = 437) and with similar proportions each. M. bovis became regularly isolated from wildlife hosts after 2011, which is related with the settlement of the epidemiological risk area for TB in game species (Fig. 1).
The space–time clustering behaviour of TB cases was evaluated with a multinomial probability model, in a total of 723 cases, reported between 2002 and 2016, in cattle (n = 249), red deer (n = 247) and wild boar (n = 227). Two significant clusters were identified (p-value = 0.001) and were both composed by TB cases originated from the three host species (Fig. 3). The 173.17 km-radius spatial cluster in Barrancos county, Beja district, included a total of 108 TB cases (n = 105 for cattle, n = 2 for red deer and n = 1 for wild boar) and persisted between 2004 and 2010, pointing out to a remarkable higher relative risk (RR) for cattle (RR = 4.16), when comparing with wild boar (RR = 0.025) and red deer (RR = 0.047) (Fig. 3). The other cluster, with 31.49 km-radius, holds the county Rosmaninhal at the centre, in Castelo Branco district, and embraces 270 TB cases (n = 13 for cattle, n = 131 for red deer and n = 126 for wild boar) from 2012 to 2016. In this cluster, wild boar (RR = 2.08) presents a higher relative risk compared to red deer (RR = 1.90) and cattle (RR = 0.093) (Fig. 3).
Multinomial space–time clusters (black-lined circles) of M. bovis calculated for the 2002–2016 period. Categories refer to host species. Relative Risk (RR) estimates indicate whether or not, greater than expected numbers occurred in a given category. RR > 1 represents a greater than expected number of individuals of a certain category inside the spatial-time cluster compared to outside. The radius of each space–time cluster is indicated. Map was generated with QGIS software (version 3.10, https://www.qgis.org/en/site/).
Genotypic diversity of M. bovis
Spoligotyping was used as first-line genotyping tool for the selected 948 M. bovis isolates, distinguishing 64 patterns with varying prevalence (overall discriminatory power, D = 0.92) (Supplementary Fig. S1). Three spoligotypes, SB2529, SB2530 and SB2531, were recorded for the first time, and were introduced at the M.bovis.org database. Twenty-nine patterns with lower frequency (encompassing 2 to 42 isolates) and 29 orphan patterns represented 32% and 3% of total isolates, respectively (Supplementary Fig. S1 and Supplementary Table S1).
SB1174 (n = 158; 16.7%), SB0122 (n = 123; 13.0%), SB0121 (n = 111; 11.7%), SB1264 (n = 92; 9.7%), SB0265 (n = 75; 7.9%) and SB0119 (n = 57; 6.0%) were the top 6 profiles and together account for 65% of total isolates (Supplementary Fig. S1, Table S1). These six profiles were documented in the three host species and, with the exception of SB1174 and SB1264, they were distributed across the three geographic regions under analysis. The proportion of SB0119 and SB0121 varied across the study period, exhibiting a global negative trend, while SB0122 and SB1264 showed a positive tendency (Fig. 4).
Variation in the proportion of M. bovis isolates included within the top 6 spoligotyping profiles. The solid line represents global predictions from a linear regression with 95% confidence interval, while punctuated lines represent the prediction for each geographic hotspot. Each graph is identified by the spoligotyping profile.
For a subset of 524 M. bovis isolates (selected based on host species, location, year—see “Methods” section), a panel of 8-loci MIRU-VNTR was applied, resulting in 447 complete profiles with single alleles, grouped by similarity in 157 MIRU types and following the denomination of Cunha et al.7; 37 complete profiles with double alleles in one or more loci; and 40 profiles with one systematically un-amplified locus (Supplementary Fig. S2, Table S2).
The majority of MIRU type groups (n = 95; 60.1%) were singletons, whereas a minority comprised 2 to 37 isolates (Supplementary Fig. S2). The six most common MIRU types (M34, M205, M219, M225, M251 and M261) account for 35% of sub-typified isolates (Supplementary Table S2). The contribution of singletons for global analysis was higher in the first time period, representing 26% of total MIRU types, whereas in the last considered time period singletons decreased to 21%.
The largest MIRU type groups (M261, M34 and M225), encompassing over 30 isolates each, were found in the three host species under analysis; whereas M34 was recorded in the three regions, and M225 and M261 were restricted to Castelo Branco and Portalegre (Supplementary Table S2).
The overall discriminatory power of MIRU-VNTR typing was higher than spoligotyping (D = 0.97) and the allelic diversity of individual locus ranged from 0.28 to 0.68, in decreasing order: ETR-A (h = 0.68), QUB11b (h = 0.58), ETR-B (h = 0.57), ETR-C (h = 0.53), VNTR 3232 (h = 0.42), QUB11a (h = 0.41), MIRU4 (h = 0.32) and MIRU26 (h = 0.28). All MIRU-VNTR loci were polymorphic, ranging between three (MIRU4) and 10 (VNTR3232) different alleles per locus.
MIRU-VNTR typing enabled a finer discrimination of M. bovis isolates within the same spoligotype group, with the exception of two isolates with profile SB1273 that presented the same MIRU type. Therefore, the combined dataset of results from both typing methods enabled the definition of 208 genotypic profiles (Supplementary Table S3). There were genotypic profiles common to the three host species, but never to the three geographic regions under analysis; however, common profiles across bordering regions (Castelo Branco and Portalegre or Portalegre and Beja) were found.
Polyclonal infection was suggested in 37 cases, through the identification of double alleles in one to four loci. Double alleles were registered across all loci, with the exception of QUB11a, and in the majority of profiles (n = 31; 83.8%) were restricted to one single locus (Supplementary Table S4). M. bovis genotypic variants were isolated from the three host species and geographic regions throughout the 2006–2015 period. In a small proportion of cases (n = 6), restricted to wild hosts from Castelo Branco, double alleles were recorded in two to four loci (Supplementary Table S4).
M. bovis genotypic diversity across TB hotspot regions
The most represented region, Castelo Branco, was also the region that yielded the highest number of spoligotypes, with the identification of 48 profiles (D = 0.8974), followed by Beja, with 28 (D = 0.8966), and Portalegre, with 23 (D = 0.7762) (Fig. 5). Analysis of spoligotyping diversity evolution through time, reveals an overall reduction in the three TB hotspots (Fig. 5). From time period 1 to time period 3, D values varied between 0.88 to 0.85 in Castelo Branco, 0.92 to 0.67 in Portalegre and 0.81 to 0.76 in Beja.
Spoligotyping analysis per geographic region: (a) comparison between the total number of M. bovis isolates and the spoligotyping profiles registered; (b) common and specific-spoligotyping profiles; (c) evolution of the discriminatory power (D values) of spoligotyping for the global dataset (black line) and for each geographic region. Blue represents Castelo Branco (CB), orange Portalegre (PG) and purple Beja (BJ).
Eleven spoligotypes are common to the three geographic regions, accounting for more than half of M. bovis isolates. The panel of Castelo Branco-specific spoligotypes (56%) was higher when comparing with Beja (36%) and Portalegre (13%) (Fig. 5, Supplementary Table S1).
The prevalence rates varied across geographic regions, with SB0122 (21.6%) being most commonly reported in Castelo Branco, SB1174 (43.5%) in Portalegre and SB0265 and SB1190 (15.3%) equally prevalent in Beja (Supplementary Table S1).
In Castelo Branco and Portalegre, the cumulative percentage of the six most common profiles increased by 20%, representing 78 and 81%, respectively, of the dataset in the time period 3. In Beja, profiles SB1174 and SB1264 were substituted by SB0295 and SB1190, and this cumulative evolution was subtle, from 61% in period 1 to 76% in period 3.
Statistically significant associations were established within the spoligotyping profiles and the three locations. Beja was the district with more positive associations established (p-value < 0.01), wherein seven profiles were highlighted (SB0119, SB0120, SB0140, SB0265, SB0295, SB1172 and SB1190); Castelo Branco with four (SB0122, SB1195, SB1264 and SB 1266); and, finally, Portalegre with three (SB1167, SB1174 and SB1483) (p-value < 0.01) (Supplementary Table S5).
MIRU-VNTR typing grouped the 254 isolates from Castelo Branco into over 100 distinctive MIRU types, while Portalegre and Beja isolates were split into 48 and 35 different MIRU types, respectively (Supplementary Fig. S3). M34 (15.9%) was the most frequent in Beja; M225 (13.4%) in Castelo Branco; and M261 (21.4%) in Portalegre (Supplementary Table S2).
The discriminatory power of MIRU-VNTR was higher than spoligotyping (D = 0.97) and the stratified analysis also disclosed similar values. ETR-A and MIRU26 were the loci with highest and lowest allelic diversity, respectively, both in Castelo Branco and Portalegre, while in Beja, QUB11a was the most discriminatory and QUB11b the least.
Three MIRU types (M34, M109 and M199), encompassing 42 isolates, were common to the three geographic regions (Supplementary Fig. S3, Table S2). However, when combining spoligotyping and MIRU type, only adjacent geographic regions presented common genotypic profiles, in a total of 19 common to Castelo Branco and Portalegre and 2 common to Portalegre and Beja (Supplementary Table S3).
M. bovis genotypic diversity across host species
Red deer yielded the highest number of spoligotyping profiles, with the 303 M. bovis isolates being grouped into 39 profiles, when comparing with the 384 cattle isolates that grouped into 38 profiles and 261 wild boar isolates that collapsed into 30 profiles, with 17 profiles common to the three host species (Fig. 6). The discriminatory power presents a differential evolution through time periods, and the underlying overall discriminatory power ranged from 0.92 in cattle isolates to 0.88 in red deer (Fig. 6).
Spoligotyping analysis per host species: (a) comparison between the total number of M. bovis isolates and the spoligotyping profiles registered; (b) common and specific-spoligotyping profiles; (c) evolution of the discriminatory power (D values) of spoligotyping, for the global dataset (black line) and for each host species. Red represents cattle (C), green red deer (RD) and blue wild boar (WB).
Frequencies of spoligotyping profiles vary across host species, SB0121 (17.7%) is the most frequent in cattle, SB0122 (21.8%) in red deer and SB1174 (21.8%) in wild boar. The spoligotypes SB0122, SB1174 and SB1264 are the top three most common in the wildlife hosts, only with variable frequencies. When considering the percentages of the six most common profiles, per host species, the increase of the cumulative effect, through time, only occurred in wildlife datasets. The contribution for cattle dataset is stable, around 55%; while for wild boar it ranges from 61 to 76% and for red deer from 52 to 85%, from period 1 to 3, respectively.
Positive and significant relationships were established with all host species, being one profile (SB1264) associated to both wild boar and red deer (p-value < 0.01) (Supplementary Table S6). Red deer established positive associations with SB0122, SB1195 and SB1264; wild boar was associated with SB1174, SB1257, SB1264 and SB1265; and cattle with SB0119, SB0121, SB0140, SB0295, SB1090 and SB1172 (p-value < 0.01).
Complete MIRU-VNTR profiles included 194 isolates from cattle grouped into 100 MIRU types, 138 from red deer clustered into 55 MIRU types and 115 from wild boar into 58 MIRU types (Supplementary Fig. S4). Seventeen MIRU types were common to three host species, representing 45% of the global dataset (Supplementary Fig. S4). The M34, M225 and M261 types were most commonly reported in specific host species datasets, M34 (11.8%) in cattle, M225 (13.9%) in wild boar and M225 and M261 (12.3%) in red deer.
With an overall discriminatory power of 0.96, the wildlife datasets were approximately as diverse as cattle’ (D = 0.97). ETR-A was the locus registering the highest allelic diversity for all host species, while MIRU4 was the least diverse locus for cattle, MIRU26 for red deer and QUB11a for wild boar.
Fifteen genotypic profiles were shared by the three host species, whereas the number of shared genotypic profiles between pairs of host species is equal or inferior to 10.
Network analysis
A network analysis was performed to get a visual image of the strength of relationships established within each host-geographic region pair, using as connection lines the number of shared spoligotyping or genotypic profiles (Fig. 7a,b). Overall, results highlighted a strong regional spoligotyping network, linking the three host species within Castelo Branco and between Castelo Branco and Portalegre, with approximately half of the connections representing more than 12 shared spoligotyping profiles. Second, the high number and strength of epidemiological connections involving cattle, across and within all regions, is particularly impressive when considering spoligotyping profiles as linking criteria. In fact, in the spoligotyping network, all connections, with only one exception, represent over ten shared profiles. However, when considering genotypic profiles, the higher connection strengths (> 10 shared genotypic profiles) are exclusive of Castelo Branco and the cattle connections are established with an inferior number of profiles (< 5 genotypic profiles).
Global spoligotyping (a) and genotypic profiles (b) networks. The nodes represent the different host-geographic region pairs (RD, red deer; WB, wild boar; C, cattle—BJ, Beja; PG, Portalegre; CB, Castelo Branco) and the width of lines between them is proportional to the number of shared spoligotyping/genotypic profiles. For the spoligotyping-based network, the shared profiles varied between one and 16, while for the genotypic profile network the variation is from one to 19. The networks were generated with Gephi (version 0.9.2, https://gephi.org/).
Epidemiological tracing within livestock herds and officially-delimited hunting areas
Information concerning the herd of origin was available for 93% (n = 356) of cattle specimens, while the identity of officially-delimited hunting areas was obtainable for 86% (n = 479) of wildlife animals.
Criteria were established to discriminate two epidemiologically distinctive situations, small outbreaks and chronically infected scenarios. A chronically infected herd/hunting area was defined by the recovery of three or more M. bovis isolates in different years (consecutive or not). According to this, 48 cases, 19 in herds and 29 in hunting areas were identified. In the case of small outbreaks, the cases were confined to a single year, with indication of 14 events, being the majority (n = 12) in herds. The livestock herd with largest outbreaks account for 19 M. bovis and five spoligotyping profiles, as well as five genotypic profiles, while the hunting area with more isolates recovered registers 70 M. bovis isolates distributed across 15 spoligotypes and 23 genotypic profiles (Supplementary Fig. S5).
Globally, M. bovis isolates recovery was higher in hunting areas, when considering chronically infected scenarios, while the reverse situation was registered if small outbreaks are considered.
A detailed analysis of livestock herds (n = 4) and officially-delimited hunting areas (n = 12) with more than 10 M. bovis isolates recovered was performed (Supplementary Fig. S5). The outbreaks in livestock herds were all heterogeneous, with the existence of spoligotyping profiles common to two or more years, consecutive or not, and also with common genotypic profiles, but only in consecutive years. However, when considering officially-delimited hunting areas, the outbreaks are more complex, extending over larger time periods and with M. bovis recovered from the two wild host species. So, as expected, even when comparing outbreaks with the same number of isolates, the total of spoligotyping/genotypic profiles generated is higher in hunting areas, and two different situations can be described: specific genotypic profiles regularly detected throughout the outbreaks and particular genotypic profiles occasionally detected in a specific time period.
Phylogenetic inferences of M. bovis isolates by MST
The majority of isolates (n = 428; 87.9%) were grouped into one of 14 clonal complexes (CC), with the largest complex encompassing 45% of isolates (n = 220) (Fig. 8). Some clonal complexes were specific to a single host species, while others were geographically clustered. The clonal complexes 4, 5 and 8 were exclusive of cattle, while CC 3 only encompasses red deer (Fig. 8). Geographic segregation is evident for clonal complexes CC 3, CC 6 and CC 12, only present in Castelo Branco; and for CC 5, exclusive for Beja isolates (Supplementary Fig. S6).
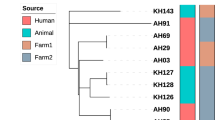
MST illustrating the evolutionary relationships among M. bovis isolates (n = 487) based on combined spoligotyping and 8-loci MIRU-VNTR data, using single locus variant analysis. Circle size is proportional to the number of isolates within each group; nodes are colored by host species (red for cattle (C), green for red deer (RD) and blue for wild boar (WB)); and the different clonal complexes are identified by different colors. The complexity of the lines denotes the number of differences in spoligo-MIRU type profile between two nodes: solid lines (1, 2 or 3 differences), gray dashed lines (4 differences) and gray dotted lines (5 or more differences). Profiles with double alleles were not included in this analysis. MST was generated with Bionumerics (version 6.6, http://www.applied-maths.com/bionumerics).
Clonal complexes 2 and 10, and two largest branches of CC 1, displayed a star-like shape typical of expanding populations41, with high-frequency central nodes surrounded by diffusing layers of variants (Fig. 8).
Downsizing the analysis to geographic region, identified 11 CC in Castelo Branco, eight in Portalegre and four in Beja (Supplementary Fig. S7). Central nodes of the largest three complexes in Castelo Branco are shared by two or three host species and display a structure coincident with an expanding population (Supplementary Fig. S7). In Portalegre, this is also evident for one clonal complex, while in Beja none of the defined complexes exhibit this architectural arrangement (Supplementary Fig. S7).
Bayesian population structure
The Bayesian clustering approach suggested the existence of K = 5 ancestral M. bovis (AM) populations (Supplementary Fig. S8), that were named ancestral M. bovis 1 to M. bovis 5 ancestral populations (AM1 to AM5) (Figs. 9a, 10a).
The spatial analysis of these five Bayesian-inferred ancestral populations revealed a contrasted geographical distribution (Fig. 9b, Supplementary Table S7): populations AM1, AM2, AM4 and AM5 are predominant in Castelo Branco, while AM3 is more evenly distributed between Castelo Branco and Portalegre. Moreover, population AM2 and AM3 are underrepresented in Portalegre and Beja, respectively (Fig. 9, Supplementary Table S7).
Host analysis (Fig. 10b, Supplementary Table S7) highlighted a tropism of AM4 population for cattle, while the other ancestral populations were more evenly distributed across different host species. This structure was further confirmed by performing a new MST analysis of M. bovis isolates, pre-labelled as populations AM1 to AM5 (Supplementary Fig. S9). Results were very congruent between both approaches, notably for populations AM2 and AM3, that revealed to be very well structured.
The geographic adaptation of ancestral populations was marked, with each geographic region evidencing positive and negative associations (p-value < 0.01) (Supplementary Table S8). Castelo Branco evidenced a positive association (p-value < 0.01) with population AM2, Portalegre with AM3 and Beja with AM5 (Fig. 9b). The negative associations were established at a p-value < 0.01 between population AM3 and Castelo Branco and Beja, and populations AM2 and AM5 with Portalegre (p-value < 0.01) (Fig. 9b).
Concerning host species, only cattle revealed a positive association with population AM4, while AM2 was negatively linked with cattle and AM4 with red deer (Fig. 10b).
Genetic characteristics of AM1 to AM5 populations
When focusing on markers driving the structure of M. bovis populations, the locus with a higher individual allelic diversity (h) could be defined as QUB11a for populations AM2 and AM4, ETR-A for AM1, VNTR3232 for AM3 and ETR-C for AM5. MIRU4 was the least diverse locus in all populations, with the exception of AM1, for which QUB11b was the least diverse.
The mean allelic richness analysis revealed that AM4 holds the highest mean allelic richness, while AM5 reaches the lowest value.
Discussion
Understanding the transmission dynamics of M. bovis circulating in an ecosystem and the role that each host species exerts in the spread and maintenance of the pathogen is necessary to improve existing control measures. The molecular methods implemented in this work thus granted an accurate monitoring of M. bovis genotypes over a 15-year period and a better understanding of the regional and local dynamics, such as small outbreaks and chronically infected scenarios both in herds and hunting estates. Spoligotyping generated a total of 64 profiles from three hotspot areas and a D value (0.92) similar to the global mean assessed a few years ago in a nationwide survey, in mainland Portugal (D = 0.93) by previous work7. SB0121 was previously reported as the most common in Portugal7,42, as well as in Spain43,44,45, and the third most frequent in France24. In this work, SB0121 remains the most common in cattle and the third for the entire dataset. About half of the identified spoligotypes, including the most common, were also previously reported in bordering Spain43,45,46,47,48. In fact, a recent work by García-Jiménez et al.45, in Extremadura region (that borders Castelo Branco and Portalegre districts), evidenced shared spoligotypes among livestock and wildlife species. The high agreement of spoligotyping profiles reported by Portugal and Spain could be related with historical backgrounds, that include long trade relationships, in addition to wild animal migrations throughout the border and disease expansion from the core Iberian cluster of disease.
Down-sizing this analysis to the administrative regions and to the host species level, Beja (the region where the number of cattle and wildlife isolates is more unbalanced) and cattle, subsist as the scenarios in which the reduction in genetic variability is less evident. In Beja and cattle sub-datasets, the evolution of the cumulative influence of the most prevalent spoligotyping profiles (that is the cumulative proportion of most common profiles), across temporal periods, reaches around 50% in 2016. The decrease of genetic variability over time is in agreement with other long-term studies performed in livestock-wildlife epidemiological settings49, and with the scenario concerning M. tuberculosis complex members’ population structure and evolution. Clonality reflects evolution processes dominated by reductions in diversity caused by population bottlenecks and selective sweeps50. The implementation of test and cull eradication programs in cattle enables continuous surveillance over these populations, turning disease outbreaks smaller and geographically stratified, so that decrease in genotypic diversity is smaller, when compared with wildlife populations with sustained genotype flow in their vital areas.
Globally, 26 spoligotyping profiles, representing 94.7% of total M. bovis isolates, were found in two or three host species. Diversity calculations within each species offered insight into their relative contribution to the epidemiology of animal TB. The occurrence of host exclusive spoligotyping profiles was suggested as an indicator to assess the importance of maintenance hosts51 and specific host profiles were registered in all three species. The greater commonality of genotypes shared between red deer and cattle give an emphasis to the role of red deer at the domestic-wildlife interface. Previous studies on social behaviour and spatio-temporal interactions at the cattle-wildlife interface, in a similar ecosystem in south-central Spain, reported that red deer daily activity patterns slightly overlap with cattle11,52, making direct transmission events more likely to occur, when comparing with wild boar that present a different daily activity pattern as a crepuscular or nocturnal species, turning direct transmission unlikely. However, a recent spatial interaction study performed in Spain revealed that environmental conditions during summer and autumn, when water and food supplies are reduced, can promote encounters of wild boar and cattle53.
The spatial–temporal analysis applied to our dataset reinforces the notion that the cattle-wildlife interaction is important, since the two significant clusters obtained were formed by the three host species, however in different associated proportions and relative risk, pointing out for two remarkably different epidemiological scenarios. A first epidemiological setting located in Beja, occurring between the 2004 and 2010 period that highlighted a higher associated risk for cattle; and a second, posterior scenario, positioned in Castelo Branco, underlined in the 2012–2016 period, in which wildlife species, particularly wild boar, present higher relative risk. This temporal and spatial stratified situation cannot be dissociated from the characteristics of our dataset, with wildlife sampling only occurring in a regular way from 2011 onward, being underrepresented up until that year and being strongly dependent upon hunting actions. Not all municipalities did regular, systematic work examining hunter-harvested big game. Also, the hunting bags vary across regions and year. Therefore, these results need to be analysed with caution and a regular update should be performed in the coming years to understand the clustering behaviour.
Molecular data reinforce and support livestock-wildlife and wildlife-wildlife transmission, either by direct or indirect ways, highlighting wildlife as keystone hosts in animal TB dynamics and supporting the general notion that wildlife may negatively contribute to the efficiency of TB eradication programs in cattle. Knowing that homoplasy of spoligotyping and MIRU typing54,55 can impact on transmission inference, a combined approach (i.e. with spoligotyping and MIRU-VNTR panel) was applied to minimize those risks and to confer more discriminatory power and accuracy to our conclusions. A panel of 8 loci was selected, not only because it was previously applied to Portuguese isolates with high discriminatory capacity, but also because a higher number of loci decrease homoplasy risks55. Indeed, the application of this typing technique enabled a more refined discrimination of isolates.
On a regional level, the specific discriminatory capacity obtained for each locus is not in agreement with previous data from Portugal7,20, and is uneven across geographic regions. The variation in allelic diversity between geographic regions and the similarities observed across adjacent geographic regions suggest a contrasting evolutionary history of these loci across regions, as well as the biogeographic segregation of M. bovis subpopulations and local processes of evolution. In fact, the network built upon the genotypic profiles revealed strong links established within geographic regions and not in between. Work performed in Northern Ireland also demonstrated the geographical structure of MIRU profiles obtained from cattle isolates, pinpointing the importance of local dynamics56.
The uniformity of genotypic profiles found in cattle and wildlife, within the same spatiotemporal context, reinforces epidemiological connections and suggests intra- and interspecies transmission, as already described in Iberian multi-host systems (including, not only red deer and wild boar48, but also badgers57); in Italy58 and Canada59. Some genotypic profiles are restricted in time and space, such as SB0121-M219 that is shared by cattle and red deer in Castelo Branco, during 2008; or SB1174-M260 that is common to three host species in Castelo Branco, during 2016. In contrast, others are actively in circulation during long time periods, for example SB1190-M34 that was detected within the three hosts species between 2009 to 2014, both in Beja and Portalegre (see supplementary information). Interestingly, the genotypic profiles common to more than one host species are the ones that were maintained in circulation for longer periods of time, reinforcing the importance of this multi-host system for M. bovis persistence and transmission, and possibly reflecting a better strain adaptation to the environment and/or to the host. Similar results were observed in France, where the local spread of specific genotypes was registered in regions with wildlife TB cases24. When downsizing to local level of livestock herds and hunting areas, the epidemiological scenario could be evaluated in closer detail, with the identification of different genotypic profiles over the years and within the same year, supporting different sources of infection. The presence of wildlife, interaction between cattle-wildlife11,60,61 and environmental contamination4,62 could exert a strong influence in herd TB dynamics, as suggested by the genotypic diversity encountered, even in small outbreaks, with the identification of up to six genotypic profiles within the same year. The presence of wildlife, namely spatial distribution, densities and artificial management conditions, have been associated as risk factors for TB in cattle. The relationship between densities of badgers, brushtail possums and wild boar were already associated with cattle TB in the UK, New Zealand and Iberian Peninsula, respectively13,63,64. These parameters are starting to be included in modelling approaches to simulate the efficiency of different TB surveillance and control systems65,66.
Plus, cattle movement between herds due to trading, breeding activities, extensive production systems and proximity to hunting zones can also have an important impact in M. bovis transmission, as indicated in other works67,68,69,70. Indeed, cattle trading is often studied as a driving force of TB introduction into cattle herds71,72,73. Chronically infected officially-delimited hunting areas also evidenced a huge genotypic diversity, being the most problematic epidemiological situations located in Castelo Branco. The identification of officially-delimited hunting areas with up to 23 genotypic profiles suggests that translocations of wild ungulates for hunting purposes may change the distribution map of TB strains and highlights the disease risks that occur if appropriate veterinary control measures are not implemented. Recrudescence of specific genotypes in a hunting area were also evident by longitudinal analysis, suggesting that some genotypes may be better adapted for multi-host infection or better detected by conventional testing methods. Moreover, the presence of super-shedders in badger populations from the UK74, and red deer and wild boar from Portugal14 have been reported possibly contributing to the persistence of specific genotypes. Concomitantly with this intricate transmission scenario, concomitant infection of individual hosts with two M. bovis genotypic variants identified by the presence of double alleles. This situation suggests two possible scenarios, co-infection with different strains acquired by the host in a single occasion or on separated time points, which is a very plausible scenario if one considers the epidemiological situation just described up above, with different genotypic profiles circulating even in small outbreaks; or the occurrence of an in vivo microevolutionary event on the founder strain. In the majority of cases, double alleles were detected only in one loci, with special prevalence in VNTR3232, supporting the hypothesis of microevolution phenomena, once the infection of one host by two very similar genotypic strains is not that likely in a scenario with high genotypic diversity. However, in a smaller group of strains, double alleles were detected in two to four loci of the same genome, which would tend to be the result of a co-infection by different strains. The smaller group of cases in which this occurred was restricted to wildlife hosts, so it might be related with their increased mobility for food and water supplies that promote contact with different M. bovis contamination sources44,75,76. Recent work conducted by our research group with M. caprae isolates also revealed the presence of double alleles, exclusively in MIRU432, which contrasts with data from this work, that evidenced a higher prevalence in VNTR3232 but not exclusive to this locus. Either way, this is a subject that deserves further investigation in the future. Furthermore, MIRU-VNTR just focuses on a small fraction of the genome and has limited resolution to capture microevolution events in other genomic regions. The identification of particularities in this work regarding chronically infected scenarios at the livestock-wildlife interface in Portugal will inform and guide a more cost-effective, risk-based, TB prevention and control strategy in these geographic regions. Given that the consistency of genotypic profiles identified in the same herd/officially-delimited hunting area in different time periods suggests reactivation of the disease, the failure to abolish the infection source is underlined, requiring more stringent surveillance. In contrast, epidemiological scenarios with multiple profiles could be related with disease re-introduction by artificial management and translocations/movements, once again calling for action.
The M. bovis population structure defined through minimum spanning trees and the Bayesian clustering algorithm implemented in STRUCTURE reflect the strain diversity across the three districts. Forty-five percent of the M. bovis dataset is contained in a single clonal complex. The fact that a substantial part of genotypes was developed from single or double locus variations upon basal types highlights a plausible clonal structure that is consistent with the expansion of a founder strain. The geographic stratification analysis of genotypic composition performed per district demark a central node in Beja exclusively composed by cattle genotypes (SB0121-derived profiles) that then irradiate to deer’ and wild boar’ strains; cattle and red deer strains at the expanding central node of Portalegre; and a mixed central node in Castelo Branco that is constituted by strains from cattle and wildlife hosts that suggests an ancestral common origin. Herewith, Beja was pointed out as the most ancient population, while Castelo Branco and Portalegre appear to be expanding, a very interesting finding that correlates with the central node composition of stratified MST, considering different host species in their composition.
Currently, five M. bovis clonal complexes have been described in the literature, European 1 (Eu1), 2 (Eu2) and 3 (Eu3), and African 1 (Af1) and 2 (Af2)77,78,79,80,81. Forty-two spoligotyping profiles accounting with 64.5% of isolates within this dataset present a deletion in spacer 21 (specific of European 2), while a smaller proportion of profiles exhibit deletion in spacer 11 (specific of European 1); in spacers 3 to 7 (specific of African 2) and present SB0120 (representative profile of European 3). The analysis of specific gene profiling (e.g. SNP analysis of guaA for Eu2) was not performed, so that grouping of M. bovis isolates in the defined clonal complexes is not possible, but the majority are candidates to belong to clonal complex European 2. This indication is in agreement with the known geographic specificities of clonal complexes distribution, that suggest that the majority of M. bovis from the Iberian Peninsula are included in clonal complex European 277. Bayesian clustering inferences provided evidence for geographic specificity of defined M. bovis ancestral populations and statistically significant correlations with cattle. Although there are commonalities across the three regions, M. bovis adapted to each hotspot area present specific genotypic characteristics, which is in agreement with differential prevalence rates of the circulating strains in a region and also with the distinctive discriminatory capacity of each MIRU-VNTR locus across regions. These results suggest that genotypic differences are more dependent on location than host species, an information that can be useful when planning control measures for each epidemiological scenario.
Intersecting genotypic data with information from the studies published so far, concerning characteristics of infection, the way animals use the environment and the interactions they establish can be key in determining the local risk for disease transmission and maintenance. Previously published data concerning TB lesions revealed a high frequency of red deer presenting multiple non-calcified lesions at the abdomen and thorax, whereas most lesions in wild boar are single and calcified10,13. Multiple lesions are suggestive of higher susceptibility carrying a higher potential for excretion of the pathogen, in parallel with lesion location and structure. In one of the studied areas, Castelo Branco district, previous work revealed that density of red deer was consistently the most relevant and significant factor influencing M. bovis infection in red deer and wild boar13.
The population structure we came across with was further confirmed by performing new MST analyses with isolates classified as population AM1 to AM5. A good agreement between Bayesian clustering inference and MST analysis was established, pronouncing the value of microsatellite genetic structure that is almost as robust as combined Spoligo/MIRU-VNTR phylogeny. The agreement among our different phylogenetic approaches supports the clonal evolution model of M. tuberculosis complex members82. Using allelic richness as a criterion of diversification, population AM4 seems to be more ancient. Interestingly, this ancestral population is statistically associated with cattle, a host species described to be M. bovis main host species, and Beja, providing clues on the who infected whom, cattle or wildlife, question.
While ongoing surveillance has provided valuable information on the prevalence and spatial occurrence of TB in areas where wild boar and red deer are sympatric, the possibility of massive whole genome sequencing (WGS) at low cost could revolutionize our current view of TB epidemiology in Iberia. The ability to observe point mutations in M. bovis genome could allow tracking transmission down to the individual or farm level, and to infer population level contact structure characteristics. Specifically, WGS could identify genetic clustering of isolates indicative of intra-species transmission, and so potentially estimate how much inter-species transmission there is amongst sampled wild and cattle populations, as well contribution of environmental reservoirs. Whole genome sequencing, contrary to spoligotyping and MIRU-VNTR, would enable the identification of sequence variations across the entire genome and single nucleotide polymorphisms could be used to study the relationships among isolates, with the advantage of presenting low reverse mutation rates and homoplasy indexes, supplanting several of the typing techniques limitations.
Concluding remarks
In this work, spoligotyping and MIRU-VNTR were applied for the molecular characterization of M. bovis longitudinally obtained during a 15-year period across TB hotspot areas in Portugal. Together, these techniques provided solid information concerning genotyping diversity and updated the characteristics of M. bovis isolates circulating across host species, within populations and across geographic regions. Geographic specificities revealed by MST and Bayesian clustering inference analyses assign Beja as the more ancient population and epidemiologically link contiguous districts, Castelo Branco and Portalegre, with locally segregated but expanding M. bovis populations that seem to be better adapted for multi-host infection.
Results support the hypothesis of a good TB maintenance community across the three hosts species and locations under analysis, and the confirmation of intraspecific and interspecific transmission, namely at the livestock-wildlife interface, pointing out an important reservoir role for cattle, probable direct transmission to red deer, but also a prominent role for wild boar as a reservoir, recommending the need to harsh control on cattle and to further consider wildlife behaviour and abundance in the adjustment of the TB eradication program in Portugal.
Change history
24 March 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41598-021-86047-y
References
Gortázar, C. et al. Bovine tuberculosis in Doñana biosphere reserve: the role of wild ungulates as disease reservoirs in the last Iberian lynx strongholds. PLoS ONE 3, e2776 (2008).
Naranjo, V., Gortázar, C., Vicentea, J. & de la Fuente, J. Evidence of the role of European wild boar as a reservoir of Mycobacterium tuberculosis complex. Vet. Microbiol. 127, 1–9 (2008).
Barasona, J. A., Torres, M. J., Aznar, J., Gortázar, C. & Vicente, J. DNA detection reveals Mycobacterium tuberculosis complex shedding routes in its wildlife reservoir the Eurasian wild boar. Transbound. Emerg. Dis. 64, 906–915 (2017).
Cano-Terriza, D. et al. Management of hunting waste as control measure for tuberculosis in wild ungulates in south-central Spain. Transbound. Emerg. Dis. 65, 1190–1196 (2018).
Palmer, M. V., Thacker, T. C., Waters, W. R., Gortázar, C. & Corner, L. A. L. Mycobacterium bovis: a model pathogen at the interface of livestock, wildlife, and humans. Vet. Med. Int. 2012, 236205. https://doi.org/10.1155/2012/236205 (2012).
Luciano, S. A. & Roess, A. Human zoonotic tuberculosis and livestock exposure in low- and middle-income countries: a systematic review identifying challenges in laboratory diagnosis. Zoonoses Public Health 67, 97–111 (2020).
Cunha, M. V. et al. Implications and challenges of tuberculosis in wildlife ungulates in Portugal: a molecular epidemiology perspective. Res. Vet. Sci. 92, 225–235 (2012).
Corner, L. A. L., Murphy, D. & Gormley, E. Mycobacterium bovis Infection in the Eurasian badger (Meles meles): the disease, pathogenesis, epidemiology and control. J. Comp. Pathol. 144, 1–24 (2011).
Fitzgerald, S. D. & Kaneene, J. B. Wildlife reservoirs of bovine tuberculosis worldwide: hosts, pathology, surveillance, and control. Vet. Pathol. 50, 488–499 (2012).
Vieira-Pinto, M. et al. Combined evaluation of bovine tuberculosis in wild boar (Sus scrofa) and red deer (Cervus elaphus) from central-east Portugal. Eur. J. Wildl. Res. 57, 1189–1201 (2011).
Cowie, C. E. et al. Interactions between four species in a complex wildlife: livestock disease community: implications for Mycobacterium bovis maintenance and transmission. Eur. J. Wildl. Res. 62, 51–64 (2016).
Santos, N. et al. Spatial analysis of wildlife tuberculosis based on a serologic survey using dried. Emerg. Infect. Dis. 24, 2169–2175 (2018).
Madeira, S. et al. Factors that influence Mycobacterium bovis infection in red deer and wild boar in an epidemiological risk area for tuberculosis of game species in Portugal. Transbound. Emerg. Dis. 64, 793–804 (2015).
Santos, N., Almeida, V., Gortázar, C. & Correia-Neves, M. Patterns of Mycobacterium tuberculosis-complex excretion and characterization of super-shedders in naturally-infected wild boar and red deer. Vet. Res. 46, 129–139 (2015).
DGAV. Programme for the eradication of bovine tuberculosis, bovine brucellosis or sheep and goat brucellosis (submitted for obtaining EU cofinancing-submission number 1483698737655-9828). https://ec.europa.eu/food/funding/animal-health/national-veterinary-programmes_en Accessed Mar 2020 (2017).
DGAV. Programme for the eradication of bovine tuberculosis, bovine brucellosis or sheep and goat brucellosis (B. melitensis) (submitted for obtaining EU cofinancing). https://ec.europa.eu/food/funding/animal-health/national-veterinary-programmes_en Accessed Mar 2020 (2019).
DGAV. Relatório técnico de sanidade animal—tuberculose Bovina. http://srvbamid.dgv.min-agricultura.pt/portal/page/portal/DGV/genericos?generico=24303689&cboui=24303689 Accessed Mar 2020 (2017).
Santos, N. et al. Epidemiology of Mycobacterium bovis infection in wild boar (Sus scrofa) from Portugal. J. Wildl. Dis. 45, 1048–1061 (2009).
Cunha, M. V. et al. Multihost tuberculosis: insights from the portuguese control program. Vet. Med. Int. 2011, 795165. https://doi.org/10.4061/2011/795165 (2011).
Duarte, E. L., Domingos, M., Amado, A., Cunha, M. V. & Botelho, A. MIRU-VNTR typing adds discriminatory value to groups of Mycobacterium bovis and Mycobacterium caprae strains defined by spoligotyping. Vet. Microbiol. 143, 299–306 (2010).
DGAV. Plano de Controlo e Erradicação de Tuberculose em Caça Maior (2011).
Matos, F. et al. Snapshot of Mycobacterium bovis and Mycobacterium caprae infections in livestock in an area with a low incidence of bovine tuberculosis. J. Clin. Microbiol. 48, 4337–4339 (2010).
Matos, A. C. et al. New Insights into Mycobacterium bovis prevalence in wild mammals in Portugal. Transbound. Emerg. Dis. 63, 313–322 (2016).
Hauer, A. et al. Genetic evolution of Mycobacterium bovis causing tuberculosis in livestock and wildlife in France since 1978. PLoS ONE 10, e0117103. https://doi.org/10.1371/journal.pone.0117103 (2015).
Boniotti, M. B. et al. Molecular typing of Mycobacterium bovis strains isolated in Italy from 2000 to 2006 and evaluation of variable-number tandem repeats for geographically optimized genotyping. J. Clin. Microbiol. 47, 636–644 (2009).
Allix, C. et al. Evaluation of the epidemiological relevance of variable-number tandem-repeat genotyping of Mycobacterium bovis and comparison of the method with IS6110 restriction fragment length polymorphism analysis and spoligotyping. J. Clin. Microbiol. 44, 1951–1962 (2006).
Navarro, Y. et al. Multiple sampling and discriminatory fingerprinting reveals clonally complex and compartmentalized infections by M. bovis in cattle. Vet. Microbiol. 175, 99–104 (2015).
Jung, I., Kulldorff, M. & Otukei John, R. A spatial scan statistic for multinomial data. Stat. Med. 29, 1910–1918 (2010).
Kamerbeek, J. et al. Simultaneous detection and strain differentiation of Mycobacterium tuberculosis for diagnosis and epidemiology. J. Clin. Microbiol. 35, 907–914 (1997).
Botelho, A., Canto, A., Leão, C. & Cunha, M. V. Clustered regularly interspaced short palindromic repeats (CRISPRs) analysis of members of the Mycobacterium tuberculosis complex. In Veterinary Infection Biology: Molecular Diagnostics and High-Throughput Strategies (eds Cunha, M. V. & Inácio, J.) 373–389 (Springer, Berlin, 2015).
Supply, P. et al. Proposal for standardization of optimized mycobacterial interspersed repetitive unit-variable-number tandem repeat typing of Mycobacterium tuberculosis. J. Clin. Microbiol. 44, 4498–4510 (2006).
Reis, A. C., Albuquerque, T., Botelho, A. & Cunha, M. V. Polyclonal infection as a new scenario in Mycobacterium caprae epidemiology. Vet. Microbiol. 240, 108533. https://doi.org/10.1016/j.vetmic.2019.108533 (2020).
Supply, P., Lesjean, S., Savine, E., Kremer, K. & Locht, C. Automated high-throughput genotyping for study of global epidemiology of Society. J. Clin. Microbiol. 39, 3563–3571 (2001).
Skuce, R. A. et al. Discrimination of Mycobacterium tuberculosis complex bacteria using novel VNTR-PCR targets. Microbiology 148, 519–528 (2002).
Hunter, P. R. & Gaston, M. A. Numerical index of the discriminatory ability of typing systems: an application of simpson’s index of diversity. J. Clin. Microbiol. 26, 2465–2466 (1988).
In silico simulation of molecular biology experiments. http://insilico.ehu.es. Accessed Nov 2018.
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Evanno, G., Regnaut, S. & Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol. Ecol. 14, 2611–2620 (2005).
Earl, D. A. & von Holdt, B. M. Structure harvester: a website and program for visualizing Structure output and implementing the evanno method. Conserv. Genet. Resour. 4, 359–361 (2012).
Kalinowski, S. T. HP-RARE 1.0: a computer program for performing rarefaction on measures of allelic richness. Mol. Ecol. Notes 5, 187–189 (2005).
Merker, M. et al. Evolutionary history and global spread of the Mycobacterium tuberculosis Beijing lineage. Nat. Genet. 47, 242–249 (2015).
Duarte, E. L., Domingos, M., Amado, A. & Botelho, A. Spoligotype diversity of Mycobacterium bovis and Mycobacterium caprae animal isolates. Vet. Microbiol. 130, 415–421 (2008).
Rodríguez, S. et al. High spoligotype diversity within a Mycobacterium bovis population: clues to understanding the demography of the pathogen in Europe. Vet. Microbiol. 141, 89–95 (2010).
Rodriguez-Campos, S. et al. Splitting of a prevalent Mycobacterium bovis spoligotype by variable-number tandem-repeat typing reveals high heterogeneity in an evolving clonal group. J. Clin. Microbiol. 51, 3658–3665 (2013).
García-jiménez, W. L. et al. Spoligotype diversity and 5-year trends of bovine tuberculosis in Extremadura, southern Spain. Trop. Anim. Health Prod. 48, 1533–1540 (2016).
Romero, B. et al. Persistence and molecular evolution of Mycobacterium bovis population from cattle and wildlife in Doñana national park revealed by genotype variation. Vet. Microbiol. 132, 87–95 (2008).
Parra, A., Larrasa, J., García, A., Alonso, J. M. & de Mendoza, J. H. Molecular epidemiology of bovine tuberculosis in wild animals in Spain: a first approach to risk factor analysis. Vet. Microbiol. 110, 293–300 (2005).
de Mendoza, J. H. et al. Bovine tuberculosis in wild boar (Sus scrofa), red deer (Cervus elaphus) and cattle (Bos taurus) in a mediterranean ecosystem (1992–2004). Prev. Vet. Med. 74, 239–247 (2006).
Hauer, A. et al. Accurate phylogenetic relationships among Mycobacterium bovis strains circulating in France based on whole genome sequencing and single nucleotide polymorphism analysis. Front. Microbiol. 10, 955. https://doi.org/10.3389/fmicb.2019.00955 (2019).
Smith, N. H., Gordon, S. V., de la Rua-Domenech, R., Clifton-Hadley, R. S. & Hewinson, R. G. Bottlenecks and broomsticks: the molecular evolution of Mycobacterium bovis. Nat. Rev. Microbiol. 4, 670–681 (2006).
Corner, L. A. L. The role of wild animal populations in the epidemiology of tuberculosis in domestic animals: how to assess the risk. Vet. Microbiol. 112, 303–312 (2006).
Kukielka, E. et al. Spatial and temporal interactions between livestock and wildlife in south central Spain assessed by camera traps. Prev. Vet. Med. 112, 213–221 (2013).
Barasona, J. A. et al. Spatiotemporal interactions between wild boar and cattle: implications for cross-species disease transmission. Vet. Res. 45, 122. https://doi.org/10.1186/s13567-014-0122-7 (2014).
Warren, R. M. et al. Microevolution of the direct repeat region of Mycobacterium tuberculosis: implications for interpretation of spoligotyping data. J. Clin. Microbiol. 40, 4457–4465 (2002).
Reyes, J. F., Chan, C. H. S. & Tanaka, M. M. Impact of homoplasy on variable numbers of tandem repeats and spoligotypes in Mycobacterium tuberculosis. Infect. Genet. Evol. 12, 811–818 (2012).
Skuce, R. A. et al. Mycobacterium bovis genotypes in Northern Ireland: herd-level surveillance (2003 to 2008). Vet. Res. 167, 684–689 (2010).
Balseiro, A. et al. Spatial relationships between Eurasian badgers (Meles meles) and cattle infected with Mycobacterium bovis in northern Spain. Vet. J. 197, 739–745 (2013).
Serraino, A. et al. Monitoring of transmission of tuberculosis between wild boars and cattle: genotypical analysis of strains by molecular epidemiology techniques. J. Clin. Microbiol. 37, 2766–2771 (1999).
Andrievskaia, O., Turcotte, C., Berlie-surujballi, G., Battaion, H. & Lloyd, D. Genotypes of Mycobacterium bovis strains isolated from domestic animals and wildlife in Canada in 1985–2015. Vet. Microbiol. 214, 44–50 (2018).
Glaser, L. et al. Descriptive epidemiology and whole Genome sequencing analysis for an outbreak of bovine tuberculosis in beef cattle and white-tailed deer in northwestern minnesota. PLoS ONE 11, e0145735. https://doi.org/10.1371/journal.pone.0145735 (2016).
Biek, R. et al. Whole genome sequencing reveals local transmission patterns of Mycobacterium bovis in sympatric cattle and badger populations. PLoS Pathog. 8, 1003008. https://doi.org/10.1371/journal.ppat.1003008 (2012).
Barasona, J. A. et al. Environmental presence of Mycobacterium tuberculosis complex in aggregation points at the wildlife/ livestock interface. Transbound. Emerg. Dis. 64, 1148–1158 (2016).
Caley, P., Hickling, G. J., Cowan, P. E. & Pfeiffer, D. U. Effects of sustained control of brushtail possums on levels of Mycobacterium bovis infection in cattle and brushtail possum populations from Hohotaka, New Zealand Mycobacterium bovis infection in cattle and brushtail possum. N. Z. Vet. J. 47, 133–142 (1999).
Reilly, L. A. & Courtenay, O. Husbandry practices, badger sett density and habitat composition as risk factors for transient and persistent bovine tuberculosis on UK cattle farms. Prev. Vet. Med. 80, 129–142 (2007).
Cosgrove, M. K., Brien, D. J. O. & Ramsey, D. S. L. Baiting and feeding revisited: modeling factors influencing transmission of tuberculosis among deer and to cattle. Front. Vet. Sci. 5, 306. https://doi.org/10.3389/fvets.2018.00306 (2018).
Birch, C. P. D., Goddard, A. & Tearne, O. A new bovine tuberculosis model for England and Wales (BoTMEW) to simulate epidemiology, surveillance and control. BMC Vet. Res. 14, 273. https://doi.org/10.1186/s12917-018-1595-9 (2018).
Lahue, N. P., Banos, J. V., Acevedo, P., Gortázar, C. & Martínez-López, B. Spatially explicit modeling of animal tuberculosis at the wildlife-livestock interface in ciudad real province, Spain. Prev. Vet. Med. 128, 101–111 (2016).
Martínez-López, B. et al. Farm-level risk factors for the occurrence, new infection or persistence of tuberculosis in cattle herds from south-central Spain. Prev. Vet. Med. 116, 268–278 (2014).
Bouchez-zacria, M., Courcoul, A. & Durand, B. The distribution of bovine tuberculosis in cattle farms is linked to cattle trade and badger-mediated contact networks in south-western France, 2007–2015. Front. Vet. Sci. 5, 173. https://doi.org/10.3389/fvets.2018.00173 (2018).
Green, D. M., Kiss, I. Z., Mitchell, A. P. & Kao, R. R. Estimates for local and movement-based transmission of bovine tuberculosis in British cattle. Proc. R. Soc. 275, 1001–1005 (2008).
Goodchild, A. V., Watkins, G. H., Sayers, A. R., Jones, J. R. & Clifton-Hadley, R. S. Geographical association between the genotype of bovine tuberculosis in found dead badgers and in cattle herds. Vet. Res. 170, 259. https://doi.org/10.1136/vr.100193 (2012).
Gopal, R., Goodchild, A., Hewinson, G. & Domenech, R. D. R. Introduction of bovine tuberculosis to north-east England by bought-in cattle. Vet. Rec. 159, 265–271 (2006).
Grear, D. A., Kaneene, J. B., Averill, J. J. & Webb, C. T. Local cattle movements in response to ongoing bovine tuberculosis zonation and regulations in michigan, USA. Prev. Vet. Med. 114, 201–212 (2014).
Delahay, R., Langton, S., Smith, G., Clifton-Hadley, R. & Cheeseman, C. The spatio-temporal distribution of Mycobacterium bovis (bovine tuberculosis) infection in a high-density badger population. J. Anim. Ecol. 69, 428–441 (2000).
Navarro, Y. et al. Detailed chronological analysis of microevolution events in herds infected persistently by Mycobacterium bovis. Vet. Microbiol. 183, 97–102 (2016).
Gormley, E., Corner, L. A. L., Costello, E. & Rodriguez-Campos, S. Bacteriological diagnosis and molecular strain typing of Mycobacterium bovis and Mycobacterium caprae. Res. Vet. Sci. 97, 30–43 (2014).
Rodriguez-Campos, S. et al. European 2—a clonal complex of Mycobacterium bovis dominant in the Iberian Peninsula. Infect. Genet. Evol. 12, 866–872 (2012).
Berg, S. et al. African 2, a clonal complex of Mycobacterium bovis epidemiologically important in east Africa. J. Bacteriol. 193, 670–678 (2011).
Muller, B. et al. African 1, an epidemiologically important clonal complex of Mycobacterium bovis dominant in Mali, Nigeria, Cameroon, and Chad. J. Bacteriol. 191, 1951–1960 (2009).
Smith, N. H. et al. European 1: a globally important clonal complex of Mycobacterium bovis. Infect. Genet. Evol. 11, 1340–1351 (2011).
Branger, M. et al. The complete genome sequence of Mycobacterium bovis Mb3601, a SB0120 spoligotype strain representative of a new clonal group. Infect. Genet. Evol. 82, 104309. https://doi.org/10.1016/j.meegid.2020 (2020).
Smith, N. H. et al. Ecotypes of the Mycobacterium tuberculosis complex. J. Theor. Biol. 239, 220–225 (2006).
Acknowledgements
MVC thanks the national authority for nature conservation and forests, ICNF (Gonçalo Lopes), for providing georeferenced location of the hunting areas, and the national veterinary authority, DGAV (DSSPA, Ana Caria), for kindly providing georeferenced data from cattle herds. MVC also acknowledges DVM João Serejo for fruitful discussion on TB along the years and excellent field work, the generous collaboration of hunting associations and the laboratorial assistance by Celeste Matos, Alice Batalha and Ana Canto at the NRL for animal TB. This work was funded by Programa Operacional de Competitividade e Internacionalização (POCI) (FEDER component), Programa Operacional Regional de Lisboa, and Fundação para a Ciência e a Tecnologia (FCT/MEC), Portugal, in the scope of the project “Colossus: Control Of tubercuLOsiS at the wildlife/livestock interface uSing innovative natUre-based Solutions” (ref. POCI-01-0145- FEDER-029783) awarded to MVC as PI. This work was also supported by strategic funding to cE3c and BioISI Research Units (UID/BIA/00329/2020 and UID/Multi/04046/2020). ACR is the recipient of a PhD fellowship by FCT/MEC (PD/BD/128031/2016).
Author information
Authors and Affiliations
Contributions
M.V.C. conceived the study and directed all experiments. A.B. and T.A. provided the bacterial isolates from the NRL. A.C.R. performed the experimental molecular work under the supervision of M.V.C. and R.T. Formal data analyses were performed by A.C.R. and M.V.C. A.C.R. and M.V.C. wrote and revised all drafts of the manuscript. R.T. gave critical feedback on final draft. All authors approved the final version for submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Reis, A.C., Tenreiro, R., Albuquerque, T. et al. Long-term molecular surveillance provides clues on a cattle origin for Mycobacterium bovis in Portugal. Sci Rep 10, 20856 (2020). https://doi.org/10.1038/s41598-020-77713-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-77713-8
This article is cited by
-
Wild boar (Sus scrofa) as a potential reservoir of infectious agents in Portugal: a review of two decades (2001–2021)
European Journal of Wildlife Research (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.