Abstract
Norway spruce has a broad natural distribution range, which results in a substantial variety of its physiological and genetic variation. There are three distinct altitudinal ecotypes described in this tree species. The physiological optimum of each ecotype may be shifted due to ongoing climate change, especially in traits associated with water demand that might be crucial for adaptation. Dehydrins are proteins that help to mitigate the adverse effects of dehydration. Dehydrin gene expression patterns appeared to be a suitable marker for plant stress assessment. Genetically determined differences in response between individuals and populations were formerly studied, however, mainly in controlled conditions. We evaluated ecotypic variation in dehydrin gene expression in a clonal bank comprised of all three ecotypes. A genetic relationship among targeted trees was uncovered utilizing GBS (Genotyping by Sequencing) platform. We sampled 4–6 trees of each ecotype throughout 15 months period. Subsequently, we assessed the RNA expression of dehydrin genes by qRT-PCR. For this study, we deliberately selected dehydrins from different categories. Our findings detected significant differences among ecotypes in dehydrin expression. The association of recorded climatic variables and individual gene expression across the study period was evaluated and revealed, for certain genes, a correlation between dehydrin gene expression and precipitation, temperature, and day-length.
Similar content being viewed by others
Introduction
Norway spruce (Picea abies [L.] Karst.) is a dominant species in mountainous, sub-alpine, and boreal forest ecosystems. Its large natural range extends from the French Alps in the west (5°E longitude) to the Ural Mountains (155°E) in the east and from 70°N latitude in northern Norway to 42°N in Macedonia1. Its altitudinal range stretches from sea level to above 2300 m in the Italian Alps2. Norway spruce is one of the essential coniferous tree species of the European continent in terms of wood and pulp production.
Owing to Norway spruce broad biogeographic range, natural selection has resulted in a substantial genetic diversity. However, high intra-population variability was conveyed to the extent surpassing the inter-population variability3.
In general, three morphological forms of Norway spruce are distinguished. Those forms also differ in their ecological requirements and are tightly connected to their altitudinal origin4. Wide crown, toothed cone scales, and comb-like branches characterize the low–elevation form acuminata (naturally growing up to 500 m a.s.l. in Central Europe). The high-elevation form (obovata; naturally growing approx. above 1100 m a.s.l.) has a typically narrow crown, round cone scales, and flat branches. It usually occurs close to the tree line. The medium-elevation form (europaea) has a rhombic, toothed cone scales with crown mostly conical of medium width. Its branches exhibit a brush type, and they are usually shorter, dense, and hanging4. Moreover, for each ecotypic form, specific adaptations for the given geographic area, climatic area, and forest vegetation zone are reported4,5,6.
With ongoing climate change, various models predict rapid shifts in microclimatic conditions along longitudinal and altitudinal gradients. Current local adaptations may not be sufficient for future forests7. Namely, traits connected to water demand are crucial for adaptation as they are strongly affected by climate change, and Norway spruce is sensitive to drought stress8. Drought-induced stress is considered a cardinal factor connected with an ongoing bark beetle outbreak and possibly leads to Norway spruce decline across the Central European region. On the contrary, high adaptive variation in drought-related traits within Norway spruce populations has been described9.
Lately, stress adaptive traits have been investigated through the gene expression analysis of dehydrins, proteins of the late embryogenesis abundant (LEA) family. Conditions of drought, low temperatures, frost, or high salinity have been associated with dehydrin gene expression10,11,12,13. It has been described that dehydrins maintain or mimic aqueous conditions in a dehydrated cell14,15. Dehydrin proteins are divided into categories differing in the presence and order of typical amino acid motifs. Conserved K-segment is present in all dehydrins; other commonly described motifs are S-segment and Y-segment16. Using the abbreviation YSK according to the quantity and sequence of segments Y, S, K, the dehydrins can be sorted into subcategories11.
Dehydrin gene expression analyses are frequently reported in economically important horticultural species17,18,19. However, information on genetic variation in dehydrin gene expression is scarce in gymnosperm species. A study of dehydrin gene expression in Pinus pinaster revealed that expression patterns of several dehydrin transcripts differed significantly among various provenances. And even more contrasting results were observed between drought-sensitive and drought-tolerant genotypes20. The dehydrin gene expression of three Picea glauca genotypes detected only minor differences in single dehydrin gene expression (PgDhn10)21. Such experiments have been carried out in controlled conditions (tissue cultures, greenhouse planting, or growth chambers). In contrast, knowledge about genetically determined differences in dehydrin expression in forest trees grown in natural conditions is scarce.
Our study aimed to evaluate variation among Norway spruce ecotypic forms in dehydrin gene expression. We conducted our research in a Norway spruce clone bank, consisting of all three ecotypes. For this study, we selected three dehydrins from different categories (Kn, SKn, and KnSKn): PaDhn4.5 from SK2 subcategory, PaDhn6 from K3 subcategory, and PaCAP1.1 from subcategory K4SK2. To justify clustering into ecotypic groups objectively, we further reconstructed a genetic relationship among sampled trees utilizing Genotyping by Sequencing (GBS). Besides, the association of recorded climatic variables with the specific gene expression across the study period was evaluated.
Material and methods
Plant material and forest stand description
The original field trial corresponded to a Norway spruce clone bank, established in 1970 with clonally propagated accessions originating from several autochthonous Czech Norway spruce populations22. The plot was planted as clonal rows with ten individuals per clone in 3 m distances; distances between rows were 6 m. The whole site was subsequently pruned so that the final spacing is 6 m × 6 m. The trial is located in the central part of the Czech Republic (N 49°56.37′, E 14°20.96′), situated on a mild NW slope with a low relief of an altitude of 320–345 m a.s.l. The average precipitation (mean between years 1980–2016) in the area was 587 mm, with an average annual temperature of 8.6 °C.
Bedrock is constituted from clayey Algonkian phyllite slates with variously thick loess and sloping clay overlaps. The soils can be characterized as medium-deep cambisols in strongly skeletal bases, in places with signs of reduction processes. The upper horizons are clayey, the lower horizons heavier, silty clay. There is a lack of loess cover in places, and the soils are generally rich in the skeleton (particles > 2 mm). The average tree height on the plot was 20.6 m; sd = 2.95 m, and the average DBH was 33.1 cm; sd = 7.7 cm.
The sampling procedure consisted of randomly selecting strips from a given ecotype. Within these strips, trees were chosen randomly, ensuring that we cover different microsites within the trial. All sampled trees retained their morphological characteristics (ecotypic appearance) correspondingly to their altitudinal origin. The typical appearance of ecotypic forms is displayed in Supplementary Figure S1.
Trees (N = 4) representing a low-elevation form (acuminata) came from an altitude of 360 m a.s.l., trees (N = 6) belonging to medium-elevation form (europaea) originated from elevation of 770–775 m a.s.l., whereas trees (N = 4) representing high-elevation form (obovata) originated from stand at altitude of 1145–1175 m a.s.l.
Samples were collected monthly through April to December 2016, and in February, March, April, and June 2017. We targeted last-year needles from the middle part of the crown (8–10 m above ground) from the southern direction. Branches were cut with pole-scissors. Samples were immediately placed into liquid nitrogen in 50 mL Falcon vials, then stored in the freezer (− 80 °C) until further processing.
Climatic data
Reported climatic variables (daily temperature and its extremes, day length, sunshine duration, daily mean precipitation, and air humidity) were obtained from the nearest state meteorological station (Praha–Libuš), located approx. 10 km away from the study site.
Laboratory analysis
For the RNA isolation, needles (100 mg) were put into 2 mL Eppendorf Safe-Lock tubes containing three steel grinding balls and frozen under liquid nitrogen. Subsequently, the tissue was ground with Retsch Mixer Mill 400. Total RNA was extracted with Epicentre MasterPure RNA Purification Kit (Epicentre). RNA was quantified by a 260/280 nm absorbance ratio using the NanoDrop spectrophotometer (Thermo-Fisher Scientific). Complementary DNA (cDNA) was synthesized with the High Capacity cDNA Reverse Transcription Kit (Thermo-Fisher Scientific) according to the manufacturer's instructions. The final volume of 40 µl cDNA solution was further diluted to 160 µl volume.
Samples for DNA extraction were treated similarly—100 mg plant tissue was ground under nitrogen liquid. Genomic DNA was extracted using the DNeasy Plant Mini Kit (Qiagen) following the manufacturer's instructions, and the DNA product was evaluated by spectrophotometrical measurement. After initial quality control, DNA samples were shipped to Cornell University Biotechnology Resource Center (BRC) and genotyped on Genotyping by Sequencing platform23.
Quantitative real-time PCR (qRT-PCR)
Specific primers for reference actin gene and dehydrin genes PaDhn6, PaDhn4.5, and PaCAP1 were designed according to24. Oligonucleotides were synthesized by Eurofins MWG Operon (Ebersberg, Germany). The specific dehydrin sequences were previously identified as plausible sensitive markers of drought stress in Norway spruce13.
Quantitative real-time polymerase chain reaction (qRT-PCR) was performed in a 12-μl volume containing 250 nM of reverse and forward primer, 2 µg of cDNA, 500 nM ROX (Top-Bio), and 6 μl of SYBR Green PCR Master Mix (Top-Bio) using the StepOnePlus Real-Time PCR System (Applied Biosystems) with the parameters recommended by the manufacturer. Reaction conditions were set as 2 min at 50 °C, 10 min at 95 °C, and 25 cycles of 95 °C for 15 s, 72 °C for 20 s and 63 °C for 30 s. Each PCR reaction was run in triplicates on a plate, and each plate was duplicated. The threshold cycle (Ct) values were calculated by StepOnePlus software (Applied Biosystems). All the gene expression levels were normalized to actin gene expression, chosen as endogenous control24,25. Dhn4.5 to Dhn6 ratio was calculated according to13.
Statistical analysis
For statistical analysis, the R statistical software26 was utilized. Pearson's product-moment correlation coefficients and their statistical significance were assessed using the 'cor.test' function, as implemented in R. The mean of 14 days that preceded sampling was calculated for all analyzed climatic variables and subsequently correlated to relative dehydrin gene expression. For visualization, Locally Estimated Scatterplot Smoothing (LOESS) was utilized with a span set on 0.75.
We fitted a univariate linear mixed model was fitted to evaluate dehydrin gene expression considering repeated measurements with the terms:
where Y corresponds to the data vector; μ is the overall mean effect; a is the fixed vector of individual ecotypes; b is the fixed vector of repeated events (months), ba is a fixed vector of the interaction of ecotypes with months, and e is the random vector of errors, with
R is a 13 × 13 matrix of variance–covariance components for residuals defined as a uniform correlation among measurements for the same individual but different variances per month, and In is an identity matrix identifying each tree. The letters X designate incidence matrices for all fixed effects, and 1 is a vector of ones.
Genomic data acquisition and analysis
As a prerequisite for dehydrin gene expression analysis, we evaluated genomic profiles of targeted trees. Genotyping by sequencing (GBS) is an NGS-based platform highly suitable for large genome species as the genome complexity is reduced, and the genome coverage enhanced, using restriction enzymes treatment.
Genomic data in VCF file format (approx. 9 Mio SNPs of raw data) obtained from BRC was filtered in VCF Tools software27 to keep only biallelic loci with the minor allele frequency higher than 0.01 and mean depth values between 20 to 80. From here, ~ 6 k high-quality SNPs were retained for subsequent analysis. Discriminant analysis of principal components (DAPC) implemented in R package adegenet 2.1.028 was used for genetic clustering. These methods allowed us to assess the putative ecotypes based not only on historical records but also on real genetic similarity among individuals.
Results
Genetic analysis of Norway spruce ecotypic forms
As grouping into three ecotypic units is based on phenotypic appearance (mostly on crown morphology), we evaluated these groups with a comparative genetic analysis (Fig. 1). Our results indicate that even though the sample size is insufficient for accurate cluster analysis, we can verify genetic proximity among individuals belonging to the same ecotypic groups and the distinct genetic structure among these forms, respectively. The genetic distance of medium-elevated ecotype (europaea) is, interestingly, the most delimited compared to the other two groups (Fig. 1).
Differences in dehydrin genes expression between ecotypic groups
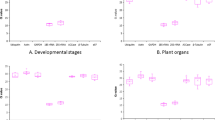
A repeated measurement assessed differences in dehydrin gene expression among Norway spruce ecotypes in a span of 15 months. Statistically non-significant differences in the expression of PaDHN4.5 and PaDHN6 genes among observed ecotypes were found, whereas statistically significant differences among ecotypes were detected in PaCAP1 gene expression (Fig. 2).
Furthermore, differences in individual months in PaCAP1 dehydrin gene expression were analyzed by Duncan's multiple range test. No differences between acuminata and europaea ecotypic forms were detected during the monitored time-span. In contrast, significant differences were found between both acuminata/obovata and europaea/obovata ecotypic forms. Diverging expression appeared most frequently between acuminata and obovata originating from the most distant locations in terms of altitude (Table 1).
Recorded climatic variables versus dehydrin expression
Mean values of PaDhn4.5 and PaDhn6 relative gene expression in given ecotypic forms were not significantly different throughout the measured period. With this in mind, the expression data of various ecotypes were pooled for posterior correlation study. PaCAP1 relative expression displayed significant differences among ecotypic forms, as reported above. For this reason, each ecotype was considered separately within the correlation study. The relation of dehydrin expression to climatic variables is outlined in Table 2, and detailed patterns of notable correlations are depicted in Fig. 3.
Relative dehydrin expression is connected to climatic variables. (a) yellow line depicts the LOESS curve of PaDHN4.5 expression related to mean daily precipitation (mm) averaged across 30 day period; (b) blue line represents the LOESS curve of PaDHN6 expression related to observed trend in temperatures throughout the 15 months, the red line follows mean daily temperature (°C) averaged across 14 days periods, the interval of corresponding max and min daily temperatures is illustrated in pink ribbon; (c) green line depicts LOESS curve of PaDHN4.5/PaDHN6 expression ratio, the yellow line represents day-length (h); black dots display the relative expression of 14 samples.
A significant negative correlation (r = −0.64) between PaDhn4.5 expression and mean daily precipitation was found in our dataset, whereas all the other remaining climatic variables did not significantly correlate with PaDhn4.5 expression.
A relatively strong negative correlation between daily temperature (mean, max, min) and PaDhn6 expression level (− 0.82 on average) was observed. However, the strongest negative correlation (− 0.86) was found with day length. Relative expression of PaDhn6 negatively correlated also with precipitation (− 0.63). On the other hand, a moderately positive correlation was observed with air humidity (0.69).
The significant negative correlation between PaCAP1 expression and precipitation was found only for obovata ecotype (− 0.61).
Discussion
We assume that Norway spruce ecotypes in the Czech Republic represent several populations sharing a similar set of adaptations. In our case, trees belonging to their respective ecotypes appear to be more alike based on SNPs data analysis. Although sampled trees have been growing under similar conditions, they retained their ecotypic membership's significant morphological features. Thus, we expected that each ecotype's divergent adaptations would lead to the different dehydrin gene expression, as the same environmental signals will be interpreted differently.
Consistently with our primitive assumption, we detected significant differences among ecotypic forms in PaCAP1 expression. The acuminata (low-elevation) form displayed the highest relative expression across the sampling period, whereas the obovata (high-elevation) form showed the lowest one. Finally, the europaea (medium-elevation) form showed the intermediate values; however, it was closer to acuminata. These differences might be attributable to a more extreme high elevation environment of obovata and its specific adaptation. These adaptations might have been crucial for pushing up the threshold of stress levels necessary for increasing the dehydrin expression.
It should be noted that PaCAP1 dehydrin expression did not appear to be driven by climatic variables—only a moderate negative correlation between precipitation and PaCAP1 in obovata ecotypic form was found. In accordance, a significant difference between early and late flushing families of Norway Spruce in PaCAP1 expression was observed12.
Contrastingly, these ecotypic differences were not found in PaDhn4.5 and PaDhn6 gene expression. It may be explained by the acclimation of trees to given stand conditions resulting in homogeneous stress response as described by29. An alternative explanation could be that gene expression of these dehydrins exhibits a low variation within the species, and a higher number of individuals would have to be analyzed to detect minor differences.
Dehydrin expression concerning annual climatic shifts
To protect themselves from frost's adverse effects in the winter, temperate perennial plants typically enter dormancy and cold hardiness. This period is connected with photosynthesis cession, alteration of carbon metabolism, and cryo- and osmoprotectants synthesis. LEA proteins and proline are considered the stress proteins with osmoprotective function. Dehydrins, as a part of the LEA proteins family, can attract common stress-signaling phospholipids30. The commonly mentioned protecting the binding activity of dehydrins to the proteins was not proved. However, a very similar process could be expected because many polar amino acids are part of the dehydrins.
Nevertheless, the plant usually involves many other substances that lower the plant water potential and maintain the cell's water molecules, known as osmoprotectants. These organic, highly soluble, and electrically neutral substances could be non-reducing sugars and sugar alcohols (e.g., trehalose or inositol), betaines (e.g., glycine betaine), and amino acids as proline31. Many sugar osmoprotectants play several essential roles in the plant as the growth regulator and signal molecule and can be participants in the communication network within a plant32,33.
We documented the relation of dehydrin genes expression with climatic variables across the 15 months sampling period. As reported in other studies, levels of dehydrin gene expression in forest trees were associated with winter dormancy34,35,36, bud burst12, and with both temperature and photoperiod37,38,39. Furthermore, it was documented that dehydrin gene expression is under the influence of temperature in Norway spruce25,40. We are bringing a new insight into the problem in the context of the unique common-garden experiment in a prolonged time-span. This setting allowed various environmental impacts on the studied material. In the next section, we will discuss individual dehydrin genes expression patterns related to climatic conditions.
PaDhn4.5 from SK2 subcategory
The PaDhn4.5 relative expression was reported to be regulated by water status and dormancy level12 and decreased correspondingly in the spring months from April to June. PaDhn4.5 expression was significantly downregulated in trees undergoing drought stress13. Following these findings, we observed the lowest value of the relative expression for all three ecotypes in July 2016 during the longer-lasting dry period. The opposite trend occurred within months with the highest precipitation (August, November, and December 2016), when relative expression values oscillated around their maxima (see Fig. 3b). Exception from this trend can be noted in June 2017, where both precipitation and PaDhn4.5 expression were high. However, across the evaluation period, a significant negative correlation (r = −0.64) of PaDhn4.5 expression with precipitation was present, whereas all the other remaining climatic variables did not significantly correlate with PaDhn4.5 expression. These findings may suggest that the relative expression of PaDhn4.5 responds primarily to drought stress.
PaDhn6 from K3 subcategory
Similarly, PaDhn6 was described as a drought stress-responsive marker13 with higher expression during drought periods. It was also shown that both day length and minimum temperature are regulation factors41,42. The decreasing PaDhn6 transcript level was observed from April to June12. We noticed the same trends in our results, with a strong negative correlation (r = −0.82) of PaDhn6 expression levels with both daily mean temperature and recorded extremes.
Short days were hypothesized to trigger the same dehydrin expression response as cold or drought stress in forest trees43. In accordance, we recorded a strong negative correlation of dehydrin PaDhn6 expression with day length (r = −0.79). PaDhn6 was described to be highly up-regulated after even a short period of short days12.
PaCAP1 from K4SK2 subcategory
PaCAP1 was described to be responsive only to severe drought stress13. It is related to drought resistance under high water loss. Up-regulated expression of homologous PoCAP1 was also recorded in extremely low temperatures in closely related Siberian spruce25. In our study, the recorded levels of stress probably did not reach such extreme levels, and therefore, PaCAP1 did not exhibit any significant variation across the evaluated period. Only a moderate negative correlation between precipitation and PaCAP1 in obovata ecotypic form was found in our study. In this regard, the estimated temporal correlation (r = 0.25 ± SE 0.10) among repeated measures across the study period is relatively self-explanatory because PaCAP1 expression was not responsive to temporal changes. The stable transcript level of the PaCAP1 gene during the period from April to June was also reported12.
PaDhn6/PaDhn4.5—Expression ratio
In addition to various single dehydrin gene expression profiles, PaDhn6 to PaDhn4.5 expression ratio was suggested as a sensitive marker of plant drought stress in Norway spruce13. This relative expression ratio follows temperature with a significant positive correlation (Table 2).
A previously published study recorded the lowest values of the respective ratio during the relatively moist periods13. In our case, the highest precipitation was recorded in November 2016. This month was also connected with the low value in the respective ratio. We found a strong negative correlation with air humidity, but we did not detect a significant association with precipitation across the whole period (Table 2).
In contrast to all single dehydrin genes, the PaDhn6/PaDhn4.5 significantly positively correlated with day length and sunshine duration in our study, which could explain the strong negative correlation with air humidity.
Conclusions
In this study, we analyzed dehydrin gene expression concerning Norway spruce variation based on ecotypic elevation forms planted ex-situ on a clone bank plot. We believe this unique setting makes our results interesting for a broader scientific community. Methodically similar research has been primarily performed under controlled conditions in greenhouses or growth chambers13,21, while such studies under natural conditions are scarce12. It is worth noting that all three ecotypic forms growing at the same site retained the specific morphological characteristics according to their origin based on particular altitudes. We confirmed the underlying genetic background of these morphologic features observing inherent grouping patterns based on SNPs data.
Dehydrin gene expression profiles were analyzed by a linear mixed model to evaluate dehydrin gene expression over the 15 months. Significant variation among Norway spruce ecotypic forms in dehydrin gene expression was found. The ecotypic forms differ in PaCAP1, where acuminata and obovata are the most different forms in terms of dehydrin gene expression. The other observed dehydrin genes do not exhibit significant variation in ecotypic forms. These differences might be connected to relatively stronger adaptations that the obovata ecotype acquired in its more extreme habitat.
PaDhn4.5 and PaDhn6 dehydrin genes correlate rather strongly with a range of climatic variables such as temperature and precipitation. In contrast, PaCAP1 expression did not appear to be driven by climatic variables: only a moderate negative correlation between precipitation and PaCAP1 in obovata ecotypic form was found.
References
Oleksyn, J., Modrzýnski, J., Tjoelker, M. G., Reich, P. B. & Karolewski, P. Growth and physiology of Picea abies populations from elevational transects: Common garden evidence for altitudinal ecotypes and cold adaptation. Funct. ecol. 12(4), 573–590 (1998).
Jansson, G. et al. Norway spruce (Picea abies (L.) H. Karst.) Pâques L. (ed.) forest tree breeding in Europe. Manag. Ecosyst. 25, 123–176 (2013).
Müller-Starck, G., Baradat, Ph. & Bergmann, F. Genetic variation within European tree species. New For. 6(1–4), 23–47 (1992).
Morgenstern, E. K. of tree ecotypes in Geographic Variation in Forest Trees: Genetic Basis and Application of Knowledge in Silviculture 109–115 (Vancouver, Amsterdam, 1996).
Androsiuk, P. et al. Genetic status of Norway spruce (Picea abies) breeding populations for northern Sweden. Silvae Genet. 62(1–6), 127–136 (2013).
Farjon, A. & Filer, D. Specific Adaptations in An atlas of the world’s conifers: An Analysis of Their Distribution, Biogeography, Diversity and Conservation Status (Springer, The Netherlands, 2013).
Chakraborty, D. et al. Selecting populations for non-analogous climate conditions using universal response functions: The case of Douglas-fir in central Europe. PLoS ONE 10(8), e0136357 (2015).
van der Maaten-Theunissen, M., Kahle, H. P., & van der Maaten, E. Drought sensitivity of Norway spruce is higher than that of silver fir along an altitudinal gradient in southwestern Germany. Ann. Sci. 70(2), 185–193 (2013).
Trujillo-Moya, C. et al. Drought sensitivity of norway spruce at the species’ warmest fringe: Quantitative and molecular analysis reveals high genetic variation among and within provenances. G3 Genes Genom. Genet. g3, 300524 (2018).
Close, T. J. Dehydrins: Emergence of a biochemical role of a family of plant dehydration proteins. Physiol. Plantarum. 97(4), 795–803 (1996).
Campbell, S. A. & Close, T. J. Dehydrins: Genes, proteins, and associations with phenotypic traits. New Phytol. 137(1), 61–74 (1997).
Yakovlev, I. A. et al. Dehydrins expression related to timing of bud burst in Norway spruce. Planta 228(3), 459–472 (2008).
Eldhuset, T. D. et al. Drought affects tracheid structure, dehydrin expression, and above-and below ground growth in 5-year-old Norway spruce. Plant Soil 366(1–2), 305–320 (2013).
Hara, M. The multifunctionality of dehydrins: An overview. Plant Signal. Behav. 5(5), 503–508 (2010).
Graether, S. P. & Boddington, K. F. Disorder and function: A review of the dehydrin protein family. Front. Plant Sci. 5, 576 (2014).
Hanin, M. et al. Plant dehydrins and stress tolerance: Versatile proteins for complex mechanisms. Plant Signal. Behav. 6(10), 1503–1509 (2011).
Kosová, K. et al. Expression of dehydrin 5 during the development of frost tolerance in barley (Hordeum vulgare). J. Plant Physiol. 165(11), 1142–1151 (2008).
Yamasaki, Y., Koehler, G., Blacklock, B. J. & Randall, S. K. Dehydrin expression in soybean. Plant Physiol. Biochem. 70, 213–220 (2013).
Liu, H. et al. Overexpression of ShDHN, a dehydrin gene from Solanum habrochaites enhances tolerance to multiple abiotic stresses in tomato. Plant Sci. 231, 198–211 (2015).
Velasco-Conde, T., Yakovlev, I., Majada, J. P., Aranda, I. & Johnsen, Ø. Dehydrins in maritime pine (Pinus pinaster) and their expression related to drought stress response. Tree Genet. Genomes. 8(5), 957–973 (2012).
Stival Sena, J., Giguère, I., Rigault, P., Bousquet, J. & Mackay, J. Expansion of the dehydrin gene family in the Pinaceae is associated with considerable structural diversity and drought-responsive expression. Tree Physiol. 38(3), 442–456 (2018).
Šindelář J. of experimental plot in Klonové Archivy Smrku Ztepilého Picea abies Karst. na PLO Zbraslav-Strnady—Polesí Jíloviště (VÚLHM, 1975).
Elshire, R. J. et al. A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PLoS ONE 6(5), e19379 (2011).
Yakovlev, I. A., Fossdal, C. G., Johnsen, O., Junttila, O. & Skrøppa, T. Analysis of gene expression during bud burst initiation in Norway spruce via ESTs from subtracted cDNA libraries. Tree Genet. Genomes. 2(1), 39–52 (2006).
Kjellsen, T. D., Yakovlev, I. A., Fossdal, C. G. & Strimbeck, G. R. Dehydrin accumulation and extreme low-temperature tolerance in Siberian spruce (Picea obovata). Tree Physiol. 33(12), 1354–1366 (2013).
R Core Team. R. A language and environment for statistical computing. Preprint at https://www.R-project.org/ (2018).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27(15), 2156–2158 (2011).
Jombart, T. & Ahmed, I. New tools for the analysis of genome-wide SNP data. Bioinformatics 27(21), 3070–3071 (2011).
Gömöry, D., Foffová, E., Kmeť, J., Longauer, R. & Romšáková, I. Norway spruce (Picea abies [L.] Karst.) provenance variation in autumn cold hardiness: Adaptation or acclimation?. Acta Biol. Cracov. Bot. 52(2), 42–49 (2010).
Cortleven, A. et al. Cytokinin action in response to abiotic and biotic stresses in plants. Plant Cell Environ. 42, 998–1018 (2019).
Szabados, L. & Savoure, A. Proline: A multifunctional amino acid. Trends Plant Sci. 15, 89–97 (2010).
Zulfiqar, F., Akram, N. A. & Ashraf, M. Osmoprotection in plants under abiotic stresses: New insights into a classical phenomenon. Planta 251, 3 (2020).
Ciereszko, I. Regulatory roles of sugars in plant growth and development. Acta Soc. Bot. Pol. 87(2), 66 (2018).
Rowland, L. J. & Arora, R. Proteins related to endodormancy (rest) in woody perennials. Plant Sci. 126(2), 119–144 (1997).
Erez, A., Faust, M. & Line, M. J. Changes in water status in peach buds on induction, development and release from dormancy. Sci. Hortic. 73(2–3), 111–123 (1998).
Kalberer, S. R., Wisniewski, M. & Arora, R. Deacclimation and reacclimation of cold-hardy plants: Current understanding and emerging concepts. Plant Sci. 171(1), 3–16 (2006).
Welling, A., Moritz, T., Palva, E. T. & Junttila, O. Independent activation of cold acclimation by low temperature and short photoperiod in hybrid aspen. Plant Physiol. 129(4), 1633–1641 (2002).
Welling, A. et al. Photoperiod and temperature differentially regulate the expression of two dehydrin genes during overwintering of birch (Betula pubescens Ehrh.). J. Exp. Bot. 55(396), 507–516 (2004).
Karlson, D. T., Zeng, Y., Stirm, V. E., Joly, R. J. & Ashworth, E. N. Photoperiodic regulation of a 24-kD dehydrin-like protein in red-osier dogwood (Cornus sericea L.) in relation to freeze-tolerance. Plant Cell Physiol. 44(1), 25–34 (2003).
Carneros, E., Yakovlev, I., Viejo, M., Olsen, J. E. & Fossdal, C. G. The epigenetic memory of temperature during embryogenesis modifies the expression of bud burst-related genes in Norway spruce epitypes. Planta 246(3), 553–566 (2017).
Asante, D. K. et al. Gene expression changes during short day induced terminal bud formation in Norway spruce. Plant Cell Environ. 34(2), 332–346 (2011).
Asante, D. K. et al. Effect of bud burst forcing on transcript expression of selected genes in needles of Norway spruce during autumn. Plant Physiol. Bioch. 47(8), 681–689 (2009).
Ruttink, T. et al. A molecular timetable for apical bud formation and dormancy induction in poplar. Plant Cell 19(8), 2370–2390 (2007).
Acknowledgements
We thank Dr. Ivana Tomášková for her valuable advice in plant physiology issues. This work was funded by the grant, "Building up an excellent scientific team and its spatiotechnical background focused on mitigation of the impact of climatic changes to forests from the level of a gene to the level of a landscape (EXTEMIT-K) at the FFWS CULS Prague", No. CZ.02.1.01/0.0/0.0/15_003/0000433 financed by OP RDE; The Ministry of Education, Youth, and Sports program INTER-EXCELLENCE, subprogramme INTER-ACTION, grant ID LTAUSA19113; The project of National Agency of Agriculture research, Czech Republic (NAZV, No. QK1910480); Internal Grant Agency of the Czech University of Life Sciences Prague CIGA, Grant No. 20164306
Author information
Authors and Affiliations
Contributions
JC and JK coordinated research activities; JC, JH, and JK were collecting material in the field; JH, JC, and ZF done laboratory work; JS, SG, JC, and ML built statistical models; JK and ZF done the genomic analyses; JS, JC, and JH wrote the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Čepl, J., Stejskal, J., Korecký, J. et al. The dehydrins gene expression differs across ecotypes in Norway spruce and relates to weather fluctuations. Sci Rep 10, 20789 (2020). https://doi.org/10.1038/s41598-020-76900-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-76900-x
This article is cited by
-
Genetic diversity of Norway spruce ecotypes assessed by GBS-derived SNPs
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.