Abstract
Dilated cardiomyopathy (DCM) is a fatal heart disease characterized by left ventricular dilatation and cardiac dysfunction. Recent genetic studies on DCM have identified causative mutations in over 60 genes, including RBM20, which encodes a regulator of heart-specific splicing. DCM patients with RBM20 mutations have been reported to present with more severe cardiac phenotypes, including impaired cardiac function, atrial fibrillation (AF), and ventricular arrhythmias leading to sudden cardiac death, compared to those with mutations in the other genes. An RSRSP stretch of RBM20, a hotspot of missense mutations found in patients with idiopathic DCM, functions as a crucial part of its nuclear localization signals. However, the relationship between mutations in the RSRSP stretch and cardiac phenotypes has never been assessed in an animal model. Here, we show that Rbm20 mutant mice harboring a missense mutation S637A in the RSRSP stretch, mimicking that in a DCM patient, demonstrated severe cardiac dysfunction and spontaneous AF and ventricular arrhythmias mimicking the clinical state in patients. In contrast, Rbm20 mutant mice with frame-shifting deletion demonstrated less severe phenotypes, although loss of RBM20-dependent alternative splicing was indistinguishable. RBM20S637A protein cannot be localized to the nuclear speckles, but accumulated in cytoplasmic, perinuclear granule-like structures in cardiomyocytes, which might contribute to the more severe cardiac phenotypes.
Similar content being viewed by others
Introduction
Dilated cardiomyopathy (DCM) is a fatal cardiac disease characterized by enlargement of the cardiac chambers and impaired systolic function1. A recent study reported that the prevalence of DCM is estimated to be 1 in 250–500 people2. Among the cases of idiopathic DCM, 20–35% are familial, with autosomal dominant inheritance in most cases3, and over 60 genes have been identified as causative genes for DCM4.
RBM20, encoding RNA binding motif protein 20 (RBM20), was identified as one of the causative genes for DCM5. RBM20 is a major regulator of heart-specific alternative splicing of the TTN gene, which is found to be most frequently mutated in patients with idiopathic DCM (approximately 20–25%)6,7. The TTN gene has the largest number of exons (364 in humans) and titin, a sarcomeric protein encoded by the TTN gene, is the largest known protein in vertebrates8. In an Rbm20 mutant rat strain lacking nearly all the Rbm20 exons, the shortest cardiac titin isoform N2B is not expressed. A longer isoform N2BA is predominant in heterozygotes, and an aberrantly giant isoform N2BA-G is exclusively expressed in homozygotes9. This indicates that RBM20 is a key regulator of Ttn pre-mRNA processing in the heart. Since titin-based passive tension is negatively correlated with its molecular size10,11,12, the ratio of the titin isoforms is considered to impact the passive stiffness of the myocardium and the progression of DCM13,14,15. It is therefore assumed that RBM20 mutations cause DCM phenotypes through altered splicing of the RBM20-regulated genes9.
Extensive searches for mutations in RBM20 in patients with idiopathic DCM revealed a hotspot of missense mutations in a highly conserved RSRSP stretch, within an arginine/serine (RS)-rich region and not in the RNA-binding domains5,9,16,17,18,19,20. Recently, the RSRSP stretch has been found to be critical for the nuclear localization of RBM20, and mutations in this stretch resulted in the loss of RBM20-dependent alternative splicing21.
Clinically, the features of DCM with an RBM20 mutation were reported as (1) the severely compromised cardiac systolic function leading to the need for heart transplantation at a very young age compared to that in DCM caused by mutations in other genes22, (2) higher incidence of sustained ventricular arrhythmias and sudden cardiac death than that in patients with a mutation in TTN23, and (3) higher incidence of atrial fibrillation (AF) than in patients with other idiopathic DCM16.
Cardiac functions of Rbm20 have been analyzed in mutant mice in which only an RNA-recognition motif (RRM) domain is deleted24 or exons 4 and 5 are deleted for a frame-shift23. Homozygotes of these Rbm20 mutations demonstrated severe splicing defects in Ttn and other target genes; however, the cardiac dysfunctions was mild23,24, as seen in the Rbm20 mutant rat strain9. Despite the phenotypic discrepancies between patients of DCM with a heterozygous RBM20 missense mutation and the homozygous Rbm20 mutant animals, the consequences of the mutations in the RSRSP stretch have not yet been assessed in an animal model.
We compared the cardiac phenotypes between Rbm20 knock-in and knock-out mice. We found that the Rbm20 mutant mice harboring a missense mutation in the RSRSP stretch mimicking a mutation in a patient with DCM, showed severe cardiac dysfunction with atrial and ventricular arrhythmias, mimicking clinical patients. The mutant RBM20 protein is specifically localized to perinuclear granule-like structures in the cytoplasm of the cardiomyocytes, which might lead to the more severe cardiac phenotypes.
Results
Rbm20S637A/S637A and Rbm20KO/KO mice lose RBM20-dependent alternative splicing
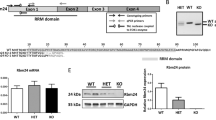
To understand the discrepancies in cardiac dysfunctions between patients with DCM and rodent models, we generated Rbm20S637A knock-in21 and Rbm20 knock-out mice by CRISPR/Cas9-mediated genome editing (Supplementary Information Fig. S1). We first analyzed the alternative splicing of known RBM20-regulated genes Ttn, Ldb3, Camk2d, and Ryr223,24 in the ventricles, by reverse transcription (RT)-polymerase chain reaction (PCR). The splicing patterns of these genes in Rbm20S637A/S637A and Rbm20KO/KO mice were drastically different from those in the wild-type (WT), and were indistinguishable between the Rbm20S637A/S637A and the Rbm20KO/KO mice (Fig. 1a,b). Consistent with the results of RT-PCR, agarose gel electrophoresis of titin proteins revealed exclusive expression of the N2BA-G isoform of titin, which harbors all exons between exons 50 and 219 in both Rbm20S637A/S637A and Rbm20KO/KO mice (Fig. 1c). RT-PCR analysis of the ventricular RNAs from Rbm20S637A/+ and Rbm20KO/+ mice revealed intermediate effects on alternative splicing of the Ttn, Ldb3 and Camk2d genes (Fig. 1a,b), and protein analysis confirmed the predominant expression of the N2BA isoform of titin (Fig. 1c). These results indicated that RBM20-dependent alternative splicing was completely dysregulated in both Rbm20S637A/S637A and Rbm20KO/KO mice, which is consistent with previous reports on Rbm20 mutant animals9,21,24.
Rbm20S637A/S637A and Rbm20KO/KO mice are defective in RBM20-dependent alternative splicing. (a) RT-PCR analysis of Ttn mRNAs with exon (E) 50 forward, E51 reverse and E219 reverse primers (top), E50 forward, E215 forward and E219 reverse primers (middle) and E115 forward, E215 forward and E219 reverse primers (bottom). Splicing patterns of the PCR products and names of corresponding titin isoforms are indicated on the right. (b) RT-PCR analysis of Ldb3 (top), Camk2d (middle), and Ryr2 (bottom) mRNAs. Gel-like images by Bioanalyzer (Agilent) are shown (a,b). (c) Vertical SDS-agarose gel electrophoresis and CBB staining of cardiac proteins from the hearts of 4-week-old Rbm20 mutant mice. Genotypes of the individual mice are indicated above. Titin isoforms (N2B, N2BA and N2BA-G) and myosin heavy chain (MHC) are indicated. n = 3 mice (a–c), each showing similar results. Full length original images are shown in Supplementary Information.
Rbm20S637A/S637A mice show more severe cardiac phenotypes
To assess the cardiac phenotypes of the Rbm20 mutant mice, we performed ultrasound echocardiography (UCG) and electrocardiography (ECG) at the age of 12–16 weeks. In the UCG analysis, Rbm20KO/KO mice showed slightly but significantly lower fractional shortening (%FS) compared to that in the WT mice (28.3 ± 1.2% vs 33.1 ± 1.4%, p = 0.002). Rbm20S637A/S637A mice had far lower %FS (16.2 ± 2.5%, p < 0.001 compared to WT) than that in others (Fig. 2a and Supplementary Information Fig. S2). Left ventricular end-diastolic diameter (LVDd) was also significantly larger, and the end-diastolic left ventricular posterior wall thickness (LVPWd) was significantly thinner in the Rbm20S637A/S637A mice (4.59 ± 0.36 mm and 0.70 ± 0.05 mm, respectively) compared to that in the Rbm20KO/KO (3.62 ± 0.30 mm, p < 0.001, and 0.84 ± 0.08 mm, p < 0.001, respectively) and WT mice (3.71 ± 0.15 mm, p < 0.001, and 0.85 ± 0.07 mm, p < 0.001, respectively) (Fig. 2a). We also performed the same assessment on Rbm20S637A/+ and Rbm20KO/+ mice. Slightly but significantly lower %FS was found in the Rbm20S637A/+ mice compared to that in the WT, whereas the Rbm20KO/+ mice showed normal cardiac function comparable to the WT (Fig. 2a and Supplementary Information Fig S2). In the ECG analysis, all six Rbm20S637A/S637A mice and one of the six Rbm20S637A/+ mice surprisingly demonstrated spontaneous AF, which was not seen in any of the Rbm20KO/KO, Rbm20KO/+, or WT mice (Fig. 2b, c). Our ECG and UCG assessments thus revealed more severe DCM-like phenotypes, including AF, in Rbm20S637A/S637A mice than in the Rbm20KO/KO mice, despite their indistinguishable defects in RBM20-dependent alternative splicing.
UCG and ECG at the age of 12–16 weeks reveal DCM-like phenotypes and AF in Rbm20S637A/S637A mice. (a) Measured value of %FS, LVDd and LVPWd by UCG. n = 6 each. p values with statistical significance after Tukey's HSD test are indicated. Error bars, SEM. (b) Proportions of mice with AF in body-surface ECG. n = 6 each. PeAF, Persistent atrial fibrillation; PAF, paroxysmal atrial fibrillation; SR, sinus rhythm. (c) Representative ECG from WT and Rbm20S637A/S637A mice. Values in the ECG indicate RR intervals. Note that the Rbm20S637A/S637A mouse demonstrated AF with irregular RR intervals.
Impairment of cardiac function is the primary pathological change in Rbm20S637A/S637A mice
Impaired cardiac function can cause and exacerbate AF and vice versa. To determine the primary change among the cardiac phenotypes, we checked the ECG of the Rbm20 mutant mice every two weeks until the age of 12 weeks, and every four weeks thereafter. Body-surface ECG revealed that Rbm20S637A/S637A mice started developing AF at the age of approximately 4 weeks and the presence of AF was evident in all the Rbm20S637A/S637A mice at 10 weeks of age (Fig. 3a). Only one out of 20 (5%) Rbm20KO/KO mice developed AF at 16 weeks of age, while the WT mice did not develop AF (Fig. 3a).
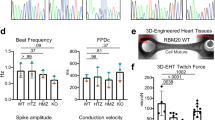
Rbm20S637A/S637A mice show high prevalence of AF and VT, and severe cardiac dysfunction precedes the AF onset. (a) AF-free survival curve. n = 20–24 each. (b) Burden of atrial tachycardia (AT) and AF in Rbm20S637A/S637A mice revealed by telemetry ECG. Three mice were analyzed. Asterisks indicate records in which VT was also observed in the telemetry ECG. (c) Spontaneous sustained VT with tachycardia cycle length of 50 ms observed in an Rbm20S637A/S637A mouse with telemetry ECG. (d) Survival curve. n = 20–40 each. (e) Measured values of %FS, LVDd and LVPWd by UCG at the age of 4 weeks. n = 6 each. (f) Gene expression of Nppa and Nppb in the heart assessed by qPCR at the age of 4 weeks. n = 3 each. p values with statistical significance after Tukey's HSD test are indicated. Error bars, SEM.
To precisely evaluate the onset of AF in Rbm20S637A/S637A mice, we performed ambulatory ECG using telemetry ECG in three Rbm20S637A/S637A mice. AF was first observed at the age of approximately 4 weeks (Fig. 3b and Supplementary Information Fig. S3), which is in line with the body-surface ECG findings. In two of the three mice, the burden of atrial arrhythmias significantly increased between 6–12 weeks of age, and AF persisted throughout the day by the age of 12 weeks. The third mouse still showed sinus rhythm (SR) and paroxysmal AF (PAF) at the age of 16 weeks. During the telemetry ECG, spontaneous, sustained, and self-terminating ventricular tachycardia (VT) and ventricular fibrillation (VF) were also observed in two of the three mice (Fig. 3b, c and Supplementary Information Fig. S4). During the VT episode, the mice suffered syncope, and VT sometimes occurred as an incessant form in these mice (Supplementary Information Fig. S4). In the survival curve analysis, sudden unexpected death was observed in approximately 30% of the Rbm20S637A/S637A mice during the 36 weeks of follow-up (Fig. 3d).
Evaluation of the cardiac function by UCG revealed significantly lower %FS, larger LVDd, and lower LVPWd in Rbm20S637A/S637A mice at the age of 4 weeks (Fig. 3e), when AF was not observed in the body-surface ECG. These results indicate that cardiac dysfunction preceded the onset of AF as the primary cardiac change. Gene expression levels of Nppa and Nppb, which are the biomarkers for heart failure, were assessed by quantitative PCR (qPCR), and both were significantly higher in the Rbm20S637A/S637A mice than in the WT and Rbm20KO/KO mice at 4 weeks of age (Fig. 3f), supporting the onset of functional phenotypes.
Morphological and histological changes in the heart of Rbm20S637A/S637A mice are consistent with DCM
Considering that RBM20S635A, a missense mutation in humans corresponding to Rbm20S637A in mice, is causative for DCM9, morphological and histological changes in the heart were evaluated in the Rbm20S637A/S637A mice. Hematoxylin and eosin (H&E) staining revealed that the hearts of the Rbm20S637A/S637A mice showed enlarged cardiac chambers and thinning of the ventricular wall at the age of 4 weeks (Fig. 4a), which was compatible with DCM cardiac phenotypes. Cardiac fibrosis was also found in Rbm20S637A/S637A mice with Masson’s trichrome staining (Fig. 4b, c). qPCR analysis revealed upregulation of the genes related to cardiac fibrosis, including Col1a2, Col3a1, and Mmp2 (Fig. 4d), which was consistent with the histological observation. At 12 weeks of age, these morphological and fibrotic LV changes were exacerbated in the Rbm20S637A/S637A mice (Fig. 4a–c). These typical DCM changes were barely observed in the hearts of Rbm20KO/KO mice (Fig. 4a–c).
Histological assessment of the hearts from Rbm20S637A/S637A mice shows morphological changes compatible with DCM. (a) Representative images of H&E staining at the age of 4 weeks (top) and 12 weeks (bottom). Scale bars, 5 mm. (b) Representative images of Masson’s trichrome staining at the age of 4 weeks (top) and 12 weeks (bottom). Scale bar, 100 µm. (c) Proportions of fibrosis area measured with the images of the Masson’s trichrome staining. n = 4–6 each. p values with statistical significance after Tukey's HSD test are indicated. Error bars, SEM. (d) qPCR analysis of genes related to cardiac fibrosis at the age of 4 weeks. n = 3 each. p values with statistical significance after Tukey's HSD test are indicated. Error bars, SEM.
ECG in Rbm20S637A/S637A mice shows abnormal electrophysiological properties in both the atrium and ventricle
Body-surface ECG at the age of 4 weeks revealed significantly prolonged P wave duration, PR interval, QRS duration and heart rate-corrected QT interval (QTc), as well as significantly smaller R amplitude in the Rbm20S637A/S637A mice compared to those in the WT mice as shown in Fig. 5 (15.5 ± 2.2 vs 13.0 ± 2.3 ms, p = 0.038 for P wave duration, 39.8 ± 2.5 vs 32.3 ± 1.8 ms, p < 0.001 for PR interval, 9.8 ± 0.9 vs 7.7 ± 0.9 ms, p < 0.001 for QRS duration, 41.4 ± 7.2 vs 31.5 ± 4.2 ms, p = 0.001 for QTc). As shown in Supplementary Information Table S1, the coefficient of variation for these ECG parameters was acceptable and consistent for each measurement, indicating the reproducibility of the ECG results. These observations are compatible with those observed in patients with DCM except for the QTc interval25. In particular, the prolongation of the P wave duration and PR interval reflects conduction disturbance in the atria, which indicates that the substrate of AF was present before the development of AF.
Body-surface ECG at the age of 4 weeks reveals atrial and ventricular electrophysiological abnormalities in Rbm20S637A/S637A mice. (a) Representative images of the ECG. Genotypes are indicated at the top. (b) Measured ECG parameters. n = 7–8 each. p values with statistical significance after Tukey's HSD test are indicated. Error bars, SEM.
Rbm20KO/KO mice showed the same tendency in the ECG parameters as the Rbm20S637A/S637A mice, yet the differences were not significant except for the QTc interval (Fig. 5). These observations in the Rbm20KO/KO mice were consistent with those in a previous report23.
RNA-seq reveals upregulation of the skeletal muscle genes and downregulation of the DCM-related genes in the heart of Rbm20S637A/S637A mice
To compare the gene expression profiles in Rbm20S637A/S637A and Rbm20KO/KO mice, we performed massive RNA sequencing (RNA-seq) of poly(A)+ RNAs from the heart, at 4 weeks of age. We detected 1840 differentially expressed genes (DEGs) between the WT and Rbm20S637A/S637A mice (857 upregulated and 983 downregulated genes), whereas only 219 genes were differentially expressed between the WT and Rbm20KO/KO mice (89 upregulated and 130 downregulated genes) (Fig. 6a and Supplementary Information Fig. S5a,b). A total of 1359 transcripts were differentially expressed between the Rbm20S637A/S637A and Rbm20KO/KO mice (694 upregulated and 664 downregulated in the Rbm20S637A/S637A mice). Gene ontology analysis revealed differences in the biological processes especially in the muscle system process, muscle structure development and extracellular matrix organization between the Rbm20S637A/S637A and Rbm20KO/KO mice (Fig. 6b), whereas there was a difference with low p values in genes related to muscle contraction between the WT and Rbm20KO/KO mice (Supplementary Information Fig. S5c, d). We also found that skeletal muscle genes including Tnni1, Tnni2, Tnnt3, Myl1, Casq1, Jsrp1, Myl6b, and Mybpc2 were significantly upregulated, and DCM-related genes including Hopx, Camk2d, Myl3, Sod2, Lmod2, Mylk3, Myoz2, Ttn, Hand2, Tnnc1, Myh7b, and Ctf1 were significantly downregulated in the heart of the Rbm20S637A/S637A mice (Fig. 6c–d, Supplementary Information spreadsheet 1, 2). For some of the DEGs, we tested the mRNA and protein expression levels (Fig. 6e, Supplementary Information Fig. S6), and found them to be consistent with the results of RNA-seq. Additionally, a long non-coding RNA related to cardiac hypertrophy, Chaer1, and genes related to hereditary arrhythmic disorders, including Ank2, Gpd1l, and Cacna2d1, were downregulated in the Rbm20S637A/S637A mice (Fig. 6c–e). These results confirmed that the gene expression profile of the Rbm20S637A/S637A mice is distinct from that of the Rbm20KO/KO mice.
Gene expression profile of Rbm20S637A/S637A mice is distinct from those of WT and Rbm20KO/KO mice. Poly(A)+ RNAs extracted from ventricles of WT and Rbm20 mutant mice (n = 3 each) were subjected to RNA-seq analysis. (a) Numbers of DEGs between indicated genotypes of mice. A cut-off of a FDR < 0.05 and gene expression fold change > 1.5 or < 0.667 were applied. (b) Enrichment analysis of Rbm20S637A/S637A vs Rbm20KO/KO. Bar colors represent p values. (c) A scatter plot for Rbm20S637A/S637A vs Rbm20KO/KO. Each dot represents each gene. Skeletal muscle genes and DCM-related genes top-ranked among the DEGs are indicated. (d) Relative expression levels of DEGs between Rbm20S637A/S637A and Rbm20KO/KO mice with a GO term ‘muscle system process’ (GO:0003012). Skeletal muscle genes, DCM-related genes, and causative genes of hereditary arrhythmia are colored as indicated. (e) qPCR analysis of DEGs top-ranked in the RNA-seq analysis. n = 3 each. p values with statistical significance after Tukey's HSD test are indicated. Error bars, SEM.
RBM20S637A protein accumulates in cytoplasmic, perinuclear granule-like structures
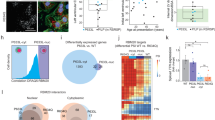
Remarkable DCM-like phenotypes of Rbm20S637A/S637A mice described above were not observed in the Rbm20KO/KO mice or other Rbm20 mutant animals reported so far, indicating that the phenotypes are caused by expression of the mutant RBM20 protein and not just by loss-of-function of the Rbm20 gene. We therefore investigated the subcellular localization of RBM20S637A protein in the cardiomyocytes of Rbm20S637A/S637A mice. In a previous study, we reported the cytoplasmic localization of FLAG-tagged RBM20S637A protein in heterologous cells21. In this study, we created a custom-made antibody against the C-terminus of mouse RBM20 protein, and detected the endogenous RBM20 proteins by immunofluorescence staining of the isolated cardiomyocytes (Fig. 7). Cytoplasmic, perinuclear granule-like RBM20 signals were detected in the Rbm20S637A/S637A cardiomyocytes (Fig. 7), whereas the RBM20 signals were localized only to the nuclear speckles in the WT cardiomyocytes (Fig. 7), as has been reported previously26,27. No specific signals were detected in the Rbm20KO/KO cardiomyocytes (Fig. 7), confirming the specific detection of endogenous RBM20 proteins on staining. In cardiomyocytes from Rbm20S637A/+ mice, RBM20 signals were present in both nuclear speckles and cytoplasmic perinuclear granule-like structures (Fig. S7). These results indicated that RBM20S637A protein cannot be localized to the nuclear speckles but is specifically accumulated in the cytoplasmic, perinuclear granule-like structures in cardiomyocytes.
RBM20S637A protein accumulates in cytoplasmic, perinuclear granule-like structures in cardiomyocytes. Isolated cardiomyocytes from WT, Rbm20KO/KO and Rbm20S637A/S637A mice were stained with anti-RBM20 and anti-α-actinin antibodies and DAPI. Representative images are shown with pseudo colors. Dotted regions around nuclei are magnified and shown on the left as merged images. Scale bars, 25 µm. n = 3 mice, each showing similar results.
Discussion
In this study, we demonstrated for the first time that.
-
1.
Rbm20S637A/S637A mice and not Rbm20KO/KO mice showed severe cardiac dysfunction with a high prevalence of AF and VT mimicking the clinical DCM phenotypes with a mutation in RBM20.
-
2.
The hearts of the Rbm20S637A/S637A mice showed upregulation of the skeletal muscle genes and downregulation of the cardiomyopathy-related genes.
-
3.
RBM20S637A protein accumulates in the cytoplasmic, perinuclear granule-like structures in the Rbm20S637A/S637A cardiomyocytes.
One notable characteristic of the Rbm20S637A/S637A mice is the development of AF (Fig. 2b, c). It is generally very difficult to induce AF in mice, and spontaneous AF is very unusual, especially at 4 weeks of age. In previous studies, few transgenic strains have been reported to develop spontaneous AF28,29. In contrast to previous AF mouse models, Rbm20S637A/S637A is the first mouse strain to show spontaneous, persistent AF with only a single nucleotide substitution, mimicking a clinical patient. Moreover, a recent genome-wide association study revealed RBM20 as one of the AF-related genes in the general population30, although the exact RBM20 variant associated with AF is yet to be determined. Therefore, the use of Rbm20S637A/S637A mice is likely to be a great advantage in future studies on the mechanism of AF development in vivo.
A previous study using cardiomyocytes from Rbm20 null mice revealed increased spontaneous release of Ca2+ from the sarcoplasmic reticulum due to increased L-type Ca2+ current, intracellular Ca2+ overload, and increased sarcoplasmic reticulum Ca2+ content, likely caused by splicing disturbance of Camk2d23, although the Rbm20 null mice did not show VT in vivo23. Therefore, we speculated that this abnormal Ca2+ handling could contribute to the development of VT in the Rbm20S637A/S637A mice. Moreover, it is also well known that Ca2+ leaks from the sarcoplasmic reticulum, and intracellular Ca2+ overload in the atrial cardiomyocytes is crucial for the development of AF31,32. Therefore, similar to that in the ventricular cardiomyocytes, the abnormal Ca2+ handling by loss of RBM20-dependent splicing might contribute to atrial arrhythmogenicity as observed in both Rbm20S637A/S637A and Rbm20KO/KO mice (Fig. 3a). However, this mechanism cannot completely explain the higher incidence of ventricular arrhythmias in the Rbm20S637A/S637A mice compared to that in the Rbm20KO/KO mice.
Prolongation of the QTc interval in Rbm20 null mice was previously reported23, and was considered to be due to the loss of RBM20-dependent alternative splicing. In this study, the Rbm20S637A/S637A mice showed QTc prolongation similar to that in the Rbm20KO/KO mice, and we speculated that the prolongation of QTc interval in the Rbm20S637A/S637A mice might be likewise affected by splicing defects. The study also mentioned that the difference in potassium currents between humans and mice might cause a discrepancy in the QTc prolongation between RBM20 mutation carriers and the Rbm20 null mice23.
Previous in vivo studies on the function of RBM20 were conducted on Rbm20 null mice23 or rats9, or mice with a deletion of the RRM domain24. These Rbm20 mutant animals or the Rbm20KO/KO mice in this study did not show severe cardiac dysfunction or development of AF. Considering the indistinguishable splicing defects (Fig. 1) and the identical genetic background of the Rbm20S637A/S637A and Rbm20KO/KO mice, the phenotypic differences between them indicate that the Rbm20S637A mutation is not only a loss-of-function mutation in terms of the splicing defects, but also a gain-of-function mutation that leads to the severe DCM phenotypes. In particular, both a homozygous and heterozygous Rbm20S637A mutation caused impaired cardiac function and AF in adulthood (Fig. 2). These observations are consistent with the fact that all reported RBM20 missense mutations in DCM patients were heterozygous20.
A molecular feature of the Rbm20S637A mutation is the presence of RBM20S637A protein, which accumulates in the perinuclear granule-like structures of unknown characters in the cardiomyocytes. Nuclear clearance and cytoplasmic deposition of a nuclear RNA-binding protein RBM20 in non-dividing cells in the DCM model are analogous to a pathological hallmark of neurodegenerative diseases in which predominantly nuclear RNA-binding proteins such as TAR DNA-binding protein 43 kDa (TDP-43) form cytoplasmic inclusions. TDP-43 proteinopathy is an established disease concept, and is well known as the cause of amyotrophic lateral sclerosis (ALS), frontotemporal lobar degeneration (FTLD), and other neurodegenerative diseases33. Mutations in the TARDBP gene encoding TDP-43 as well as in other genes lead to the cytoplasmic aggregation of TDP-43 in the neurons. A small subset of familial ALS is associated with missense mutations in the MATR3 gene, a paralogue of RBM2034,35. Considering that the Rbm20S637A mutation specifically led to the DCM phenotypes as demonstrated in this study, and the analogy to neurodegenerative diseases, it is reasonable to assume that cytoplasmic RBM20S637A protein contributes to the development of DCM. However, the pathogenesis of the TDP-43 proteinopathy remains elusive, despite extensive studies36,37,38, partly because TDP-43 is essential for viability. Future studies on the pathogenesis of RBM20S637A would lead to a better understanding of the diseases caused by aggregation and/or altered functions of the RNA-binding proteins.
Comprehensive mRNA analysis using the hearts from Rbm20S637A/S637A mice revealed misexpression of skeletal muscle genes and downregulation of DCM-causative genes compared to those from the WT and Rbm20KO/KO mice (Fig. 6 and Supplementary Information Fig. S5). Misexpression of skeletal muscle genes has been previously reported to cause cardiac malfunction39,40,41. Hopx, Myl3, Sod2, Lmod2, Mylk3, Myoz2, Ttn, Hand2, Tnnc1, Myh7b and Ctf1 have been reported as definitive or putative DCM genes4,42,43,44,45,46,47,48,49,50,51,52, and a long noncoding RNA gene Chaer1 was also reported to be an important factor in cardiac hypertrophy53. Therefore, downregulation of these genes is consistent with the severe DCM-like phenotypes of the Rbm20S637A/S637A mice, although it is currently unclear whether this is the cause or the result of the phenotypes. The DEGs in the Rbm20S637A/S637A mice included many other genes related to cardiac dysfunction such as Slc5a154 and Mylk455 (Supplementary Information spreadsheet 1, 2). With respect to arrhythmia, expressions of Ank2, a causative gene of long QT syndrome56, and Gpd1l and Cacna2d1, the causative genes of Brugada syndrome57,58, were affected, which might contribute to the arrhythmogenicity in the heart of the Rbm20S637A/S637A mice.
There are some limitations in this study. First, we did not address the precise mechanism by which RBM20S637A protein causes DCM-like phenotypes, including AF and VT. Although this study clearly demonstrated that altered splicing of RBM20-regulated genes was not alone responsible for causing the phenotypes, it is still unclear whether altered splicing is involved in the pathogenesis. In particular, in terms of arrhythmogenicity, it is unclear whether there is a common mechanism for atrial and ventricular arrhythmias related to the RBM20S637A protein. Further studies are necessary to understand the roles of the RBM20S637A protein accumulation on the cytoplasmic structures and alternative splicing changes in the target genes in cardiac phenotypes. Second, although the development of AF is a unique feature in the Rbm20S637A/S637A mice, the roles of the RBM20S637A protein and impaired cardiac function in the development of AF are unclear. In addition to the Rbm20S637A/S637A mice, one of the Rbm20KO/KO mice developed AF spontaneously during long-term follow up (Fig. 3a), indicating that the loss of RBM20 also contributes to the development of AF. In addition, heart failure is a well-known factor for exacerbation of AF59. Thus, high AF prevalence in the Rbm20S637A/S637A mice might be the consequence of heart failure, loss of RBM20-dependent alternative splicing, presence of cytoplasmic RBM20S637A protein, and other unknown defects. To elucidate the primary effect of the Rbm20S637A mutation on the development of AF, a mouse with atrium-specific RBM20S637A expression will be required. Furthermore, the most common form of AF is not associated with cardiac dysfunction, and its molecular phenotype is different from that of AF with heart failure60. Therefore, another appropriate animal model is required for common AF. Third, the subcellular localization of RBM20 proteins has not been investigated in patients with clinical DCM, with and without a mutation in RBM20. To evaluate the relevance of the granule-like structures in Rbm20S637A/S637A mice with the DCM phenotypes of the patients, further experiments using human biopsy samples or cardiomyocytes differentiated from patient-derived induced pluripotent stem cells (iPSCs) are necessary.
In conclusion, Rbm20S637A, a missense mutation in the RSRSP stretch of RBM20, leads to severe DCM-like phenotypes as well as AF and VT. Rbm20S637A/S637A mice offer a novel path for future therapeutics as a unique mouse model for DCM and AF.
Methods
Animals
All animal experiments were in accordance with the guidelines for the Care and Use of Laboratory Animals published by National Research Council (The National Academy Presses, eighth edition, 2011), pre-approved and performed under the regulation of the Institutional Animal Care and Use Committee of Tokyo Medical and Dental University (Approval #A2019-105C5 and #A2019-178C2). Generation of the Rbm20S637A knock-in allele was previously reported21. Briefly, the Rbm20S637A allele was generated by utilizing a cloning-free CRISPR/Cas system61. The Rbm20KO allele was obtained from the identical heterozygous founder mouse of the Rbm20S637A allele as a result of non-homologous end joining that yielded a 26-bp deletion (Supplementary Information Fig. S1 ). Wild-type (WT) littermate mice were used as controls. All experiments were performed with male mice.
Anti-RBM20 antibody generation
The polyclonal antibody against mouse RBM20 was generated in rabbit by using a synthetic C-terminal peptide PERGGIGPHLERKKL (amino acids 1185–1199, accession number NP_001164318) as antigen by Cosmobio. The anti-RBM20 antibody was sequentially purified with affinity columns coupled with protein A and the synthetic C-terminal peptide.
Electrophoresis and staining of cardiac proteins
Vertical SDS-agarose gel electrophoresis of cardiac proteins from mice were performed essentially as described previously21,62. Protein samples were extracted from ventricular tissues with urea buffer (8 M urea, 2 M thiourea, 3% SDS w/v, 75 mM DTT and 0.03% bromophenol blue in 50 mM Tris–Cl, pH 6.8) and separated by using 1% Sea Kem Gold agarose (FMC bioproducts) gel. The proteins were detected by staining with CBB (Bio Craft). The images of the stained gels were captured with a scanner GT-X700 (Epson) and processed by using Photoshop CC (Adobe).
Western blot
Protein samples were extracted from murine heart ventricular tissues with 10 mM Tris-Maleate buffer (pH 7.0). Protein concentration was measured by BCA Protein Assay Reagent Kit (Thermo Fisher Scientific). Twenty μg of protein samples were loaded on a 4–15% SDS-PAGE gel (Bio-Rad) for separation and transferred to PVDF membranes with Trans-Blot Turbo system (Bio-Rad). After blocking with Blocking One (Nacalai tesque), the membranes were incubated overnight at 4 °C with the custom-made anti-RBM20 affinity-purified antibody described above (1 µg/ml), anti-Tnni1 (1:500, rabbit polyclonal, Proteintech), anti-Tnni2 (1:500, rabbit polyclonal, Proteintech), or anti-GAPDH (1:10,000, mouse monoclonal, Santa Cruz Biotechnology). HRP-labeled secondary antibodies (1:10,000, Dako) were used for detecting the specific bands. Chemiluminescense signals (GE Healthcare) were detected by using iBright CL1500 (Thermo Fisher Scientific), and analyzed using iBright Analysis Software (Thermo Fisher Scientific).
Electrocardiography and ultrasound echocardiography
Body-surface electrocardiography (ECG) and ultrasound echocardiography (UCG) were performed in a blinded manner as described previously63. ECG parameters were obtained by averaging those from three different ECGs. Coefficient of variation was calculated from them. QT interval was defined as an interval between the onset of the QRS complex and the end of the negative component of the T wave. QTc was calculated by the following formula: QTc = QT interval (ms)/√(RR interval (s) × 10). To assess the incidence of AF, 10-min ECG was performed at the age of 4, 6, 8, 10 and 12 weeks, and every 4 weeks after the age of 12 weeks. In this study, we defined AF as an irregular RR rhythm persisting longer than 5 s and not accompanying a clear P wave on ECG, paroxysmal AF (PAF) as self-terminating AF within 10 min, and persistent AF (PeAF) as AF persisting throughout 10-min ECG. Atrial tachycardia (AT) was defined as regular RR tachycardia distinguishable on ECG from sinus rhythm (SR).
Telemetry ECG
A telemetry ECG transmitter (ETA-F10, DSI) was subcutaneously implanted into the back of the mouse64. A telemetry receiver was placed under the cage and the output signal of ECG was digitized with PowerLab (ADInstruments). Ambulatory ECG recording for 24 h was conducted every 1–2 weeks after the age of 3 weeks.
RNA extraction and RT-PCR
Total RNAs were extracted from heart ventricular tissues by using RNeasy Mini Kit (QIAGEN) according to the manufacturer's protocol. cDNAs were synthesized by using High Capacity cDNA Reverse Transcription Kit (Thermo Fisher Scientific) or PrimeScript II 1st strand cDNA Synthesis Kit (Takara) from 1 μg of total RNAs. qPCR was performed with PowerSYBR Green Master Mix (Thermo Fisher Scientific) and custom-made primers. Expression levels were normalized with Gapdh. Semi-quantitative RT-PCR for analyzing splicing patterns was performed by using PrimeStarGXL (Takara) and primers previously described21. The PCR products were analyzed by utilizing Bioanalyzer 2100 Expert with DNA1000 Kit (Agilent). PCR primers used in this study were synthesized by Fasmac or Eurofins. Sequences of the primers are available in Supplementary Information Table S2.
Histological assessment
Excised hearts were immersed in 10% buffered formalin and paraffin sections were subjected to H&E staining or Masson’s trichrome staining. Stained sections were photographed (DZ-710, Keyence), and fibrotic area was quantified by using ImageJ software (NIH). Proportion of fibrotic area was calculated as ratio of the fibrotic area to the total cross-sectional area.
Cardiomyocyte isolation and immunofluorescence staining
Ventricular myocytes were isolated by using a published protocol65 with minor modification. Animals were pretreated with heparin (1000 units/kg body weight) and anesthetized with intraperitoneal infusion of 100 mg/kg of pentobarbital sodium. Hearts were excised, and cannulated aorta was fixed to a Langendorff apparatus. The hearts were perfused with Ca2+-free Tyrode’s solution (120 mM NaCl, 5.4 mM KCl, 1.2 mM MgSO4, 1.2 mM NaH2PO4 and 5.6 mM glucose in 20 mM NaHCO3, pH 7.4) for 5 min at 2 ml/min, followed by perfusion with Tyrode’s solution supplemented with 0.4 mg/ml Collagenase type II (GIBCO), 0.06 mg/ml Protease type XIV (Sigma-Aldrich), and 1 mg/ml fatty acid free BSA (Sigma-Aldrich) for 18 min. The perfusion solution was bubbled with 95% O2/5% CO2 and maintained at 37 °C. The ventricles were cut into several pieces and subjected to gentle agitation through a cell strainer (70 µm, Falcon) to separate the cardiomyocytes.
Isolated cardiomyocytes were fixed with 2% paraformaldehyde in PBS for 10 min. The cells were permeabilized with PBS containing 0.1% Triton X-100 for 30 min. After blocking, the cells were incubated with 1 µg/ml anti-RBM20 antibody (described above) and anti-α-actinin antibody (1:200, mouse monoclonal, Sigma-Aldrich) for 1 h at 37 °C, followed by incubation with Alexa488-conjugated goat anti-rabbit IgG (1:200, Invitrogen), Alexa647-conjugated goat anti-mouse IgG (1:200, Invitrogen) and DAPI for 30 min. Fluorescence images were captured by utilizing optimal sectioning with a digital microscope (BZ-X710, Keyence) equipped with a 100 × oil-immersion objective (NA = 1.45, Nikon).
Statistical analysis
All data except RNA-seq are shown in mean ± standard error of the mean (SEM). Tukey's HSD test was used to compare more than 2 groups. Statistical analyses were performed with JMP10 (SAS Institute Inc.). A p value < 0.05 was considered as statistically significant.
RNA-seq
After RNA extraction, NanoDrop (Thermo Fisher Scientific), and Bioanalyzer (Agilent) were used to check the purity, concentration, and RIN values. After poly(A)+ RNA selection with NEBNext Poly(A) mRNA Magnetic Isolation Module (New England Biolabs), sequencing libraries were constructed by using NEBNext Ultra Directional RNA Library Prep Kit for Illumina (New England Biolabs). The libraries were sequenced on Illumina Novaseq 6000 by Novogene. Paired-end, 150-nt reads were generated. The sequence data files have been submitted to DRA (Accession number: DRA010298).
Analysis of RNA-seq data
The quality of obtained sequence data was evaluated using FastQC program (https://www.bioinformatics.babraham.ac.uk/projects/fastqc/) and trimmed by using the Trim Galore tool (https://www.bioinformatics.babraham.ac.uk/projects/trim_galore/). Then, the reads were mapped to the mouse reference genome (mm10) utilizing HISAT266. Sequence alignment/map (.SAM) format files were converted into binary alignment/map (.BAM) format using SAMtools67. StringTie was used for transcript assembling68. Finally, the processed data was analyzed with integrated differential expression and pathway analysis (iDEP.91) (https://bioinformatics.sdstate.edu/idep/)69. In iDEP package, read count data were normalized by counts per million function in edgeR with a pseudo-count of 4. Normalized count data were transformed using the logarithm function, and transformed data was used for gene expression analysis based on DESeq2 package. A cut-off of a false discovery rate (FDR) < 0.05 and gene expression fold change > 1.5 or < 0.667 were applied. Metascape (https://metascape.org/) was used for the gene set enrichment analysis. A gene list for Metascape analysis was generated by iDEP.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding authors on reasonable request.
References
Kirk, R. et al. Outcome of pediatric patients with dilated cardiomyopathy listed for transplant: A multi-institutional study. J. Heart Lung Transpl. 28, 1322–1328. https://doi.org/10.1016/j.healun.2009.05.027 (2009).
McKenna, W. J., Maron, B. J. & Thiene, G. Classification, epidemiology, and global burden of cardiomyopathies. Circ. Res. 121, 722–730. https://doi.org/10.1161/CIRCRESAHA.117.309711 (2017).
Kimura, A. Molecular genetics and pathogenesis of cardiomyopathy. J. Hum. Genet. 61, 41–50. https://doi.org/10.1038/jhg.2015.83 (2016).
McNally, E. M. & Mestroni, L. Dilated cardiomyopathy: genetic determinants and mechanisms. Circ. Res. 121, 731–748. https://doi.org/10.1161/CIRCRESAHA.116.309396 (2017).
Brauch, K. M. et al. Mutations in ribonucleic acid binding protein gene cause familial dilated cardiomyopathy. J. Am. Coll. Cardiol. 54, 930–941. https://doi.org/10.1016/j.jacc.2009.05.038 (2009).
Herman, D. S. et al. Truncations of titin causing dilated cardiomyopathy. N. Engl. J. Med. 366, 619–628. https://doi.org/10.1056/NEJMoa1110186 (2012).
Franaszczyk, M. et al. Titin truncating variants in dilated cardiomyopathy—prevalence and genotype-phenotype correlations. PLoS ONE 12, e0169007. https://doi.org/10.1371/journal.pone.0169007 (2017).
Chauveau, C., Rowell, J. & Ferreiro, A. A rising titan: TTN review and mutation update. Hum. Mutat. 35, 1046–1059. https://doi.org/10.1002/humu.22611 (2014).
Guo, W. et al. RBM20, a gene for hereditary cardiomyopathy, regulates titin splicing. Nat. Med. 18, 766–773. https://doi.org/10.1038/nm.2693 (2012).
Cazorla, O. et al. Differential expression of cardiac titin isoforms and modulation of cellular stiffness. Circ. Res. 86, 59–67 (2000).
Anderson, B. R. & Granzier, H. L. Titin-based tension in the cardiac sarcomere: molecular origin and physiological adaptations. Prog. Biophys. Mol. Biol. 110, 204–217. https://doi.org/10.1016/j.pbiomolbio.2012.08.003 (2012).
Opitz, C. A., Leake, M. C., Makarenko, I., Benes, V. & Linke, W. A. Developmentally regulated switching of titin size alters myofibrillar stiffness in the perinatal heart. Circ. Res. 94, 967–975. https://doi.org/10.1161/01.RES.0000124301.48193.E1 (2004).
LeWinter, M. M. & Granzier, H. L. Cardiac titin and heart disease. J. Cardiovasc. Pharmacol. 63, 207–212. https://doi.org/10.1097/FJC.0000000000000007 (2014).
Hutchinson, K. R., Saripalli, C., Chung, C. S. & Granzier, H. Increased myocardial stiffness due to cardiac titin isoform switching in a mouse model of volume overload limits eccentric remodeling. J. Mol. Cell. Cardiol. 79, 104–114. https://doi.org/10.1016/j.yjmcc.2014.10.020 (2015).
Makarenko, I. et al. Passive stiffness changes caused by upregulation of compliant titin isoforms in human dilated cardiomyopathy hearts. Circ. Res. 95, 708–716. https://doi.org/10.1161/01.RES.0000143901.37063.2f (2004).
Refaat, M. M. et al. Genetic variation in the alternative splicing regulator RBM20 is associated with dilated cardiomyopathy. Heart Rhythm 9, 390–396. https://doi.org/10.1016/j.hrthm.2011.10.016 (2012).
Li, D. et al. Identification of novel mutations in RBM20 in patients with dilated cardiomyopathy. Clin. Transl. Sci. 3, 90–97. https://doi.org/10.1111/j.1752-8062.2010.00198.x (2010).
Millat, G. et al. Clinical and mutational spectrum in a cohort of 105 unrelated patients with dilated cardiomyopathy. Eur. J. Med. Genet. 54, e570-575. https://doi.org/10.1016/j.ejmg.2011.07.005 (2011).
Rampersaud, E. et al. Rare variant mutations identified in pediatric patients with dilated cardiomyopathy. Prog. Pediatr. Cardiol. 31, 39–47. https://doi.org/10.1016/j.ppedcard.2010.11.008 (2011).
Watanabe, T., Kimura, A. & Kuroyanagi, H. Alternative splicing regulator RBM20 and cardiomyopathy. Front. Mol. Biosci. 5, 105. https://doi.org/10.3389/fmolb.2018.00105 (2018).
Murayama, R. et al. Phosphorylation of the RSRSP stretch is critical for splicing regulation by RNA-Binding Motif Protein 20 (RBM20) through nuclear localization. Sci. Rep. 8, 8970. https://doi.org/10.1038/s41598-018-26624-w (2018).
Kayvanpour, E. et al. Genotype-phenotype associations in dilated cardiomyopathy: Meta-analysis on more than 8000 individuals. Clin. Res. Cardiol. 106, 127–139. https://doi.org/10.1007/s00392-016-1033-6 (2017).
van den Hoogenhof, M. M. G. et al. RBM20 mutations induce an arrhythmogenic dilated cardiomyopathy related to disturbed calcium handling. Circulation 138, 1330–1342. https://doi.org/10.1161/CIRCULATIONAHA.117.031947 (2018).
Methawasin, M. et al. Experimentally increasing titin compliance in a novel mouse model attenuates the Frank–Starling mechanism but has a beneficial effect on diastole. Circulation 129, 1924–1936. https://doi.org/10.1161/CIRCULATIONAHA.113.005610 (2014).
Finocchiaro, G. et al. The electrocardiogram in the diagnosis and management of patients with dilated cardiomyopathy. Eur. J. Heart Fail. https://doi.org/10.1002/ejhf.1815 (2020).
Bertero, A. et al. Dynamics of genome reorganization during human cardiogenesis reveal an RBM20-dependent splicing factory. Nat. Commun. 10, 1538. https://doi.org/10.1038/s41467-019-09483-5 (2019).
Guo, W. et al. Splicing factor RBM20 regulates transcriptional network of titin associated and calcium handling genes in the heart. Int. J. Biol. Sci. 14, 369–380. https://doi.org/10.7150/ijbs.24117 (2018).
Ihara, K. et al. Electrophysiological assessment of murine atria with high-resolution optical mapping. J. Visual. Exp.: JoVE https://doi.org/10.3791/56478 (2018).
Riley, G., Syeda, F., Kirchhof, P. & Fabritz, L. An introduction to murine models of atrial fibrillation. Front. Physiol. 3, 296. https://doi.org/10.3389/fphys.2012.00296 (2012).
Nielsen, J. B. et al. Biobank-driven genomic discovery yields new insight into atrial fibrillation biology. Nat. Genet. 50, 1234–1239. https://doi.org/10.1038/s41588-018-0171-3 (2018).
Neef, S. et al. CaMKII-dependent diastolic SR Ca2+ leak and elevated diastolic Ca2+ levels in right atrial myocardium of patients with atrial fibrillation. Circ. Res. 106, 1134–1144. https://doi.org/10.1161/CIRCRESAHA.109.203836 (2010).
Voigt, N. et al. Enhanced sarcoplasmic reticulum Ca2+ leak and increased Na+-Ca2+ exchanger function underlie delayed afterdepolarizations in patients with chronic atrial fibrillation. Circulation 125, 2059–2070. https://doi.org/10.1161/CIRCULATIONAHA.111.067306 (2012).
Gao, J., Wang, L., Huntley, M. L., Perry, G. & Wang, X. Pathomechanisms of TDP-43 in neurodegeneration. J. Neurochem. https://doi.org/10.1111/jnc.14327 (2018).
Johnson, J. O. et al. Mutations in the Matrin 3 gene cause familial amyotrophic lateral sclerosis. Nat. Neurosci. 17, 664–666. https://doi.org/10.1038/nn.3688 (2014).
Coelho, M. B., Attig, J., Ule, J. & Smith, C. W. Matrin3: connecting gene expression with the nuclear matrix. Wiley Interdiscipl Rev. RNA 7, 303–315. https://doi.org/10.1002/wrna.1336 (2016).
Bolognesi, B. et al. The mutational landscape of a prion-like domain. Nat. Commun. 10, 4162. https://doi.org/10.1038/s41467-019-12101-z (2019).
Wolozin, B. The evolution of phase-separated TDP-43 in stress. Neuron 102, 265–267. https://doi.org/10.1016/j.neuron.2019.03.041 (2019).
Gasset-Rosa, F. et al. Cytoplasmic TDP-43 De-mixing independent of stress granules drives inhibition of nuclear import, loss of nuclear TDP-43, and cell death. Neuron 102, 339-357 e337. https://doi.org/10.1016/j.neuron.2019.02.038 (2019).
Huang, Q. Q. et al. Co-expression of skeletal and cardiac troponin T decreases mouse cardiac function. Am. J. Physiol. Cell Physiol. 294, C213-222. https://doi.org/10.1152/ajpcell.00146.2007 (2008).
van Rooij, E. et al. Control of stress-dependent cardiac growth and gene expression by a microRNA. Science 316, 575–579. https://doi.org/10.1126/science.1139089 (2007).
Rao, P. K. et al. Loss of cardiac microRNA-mediated regulation leads to dilated cardiomyopathy and heart failure. Circ. Res. 105, 585–594. https://doi.org/10.1161/CIRCRESAHA.109.200451 (2009).
Pennica, D. et al. Expression cloning of cardiotrophin 1, a cytokine that induces cardiac myocyte hypertrophy. Proc. Natl. Acad. Sci. USA 92, 1142–1146. https://doi.org/10.1073/pnas.92.4.1142 (1995).
Warkman, A. S. et al. Developmental expression and cardiac transcriptional regulation of Myh7b, a third myosin heavy chain in the vertebrate heart. Cytoskeleton (Hoboken) 69, 324–335. https://doi.org/10.1002/cm.21029 (2012).
Esposito, T. et al. Digenic mutational inheritance of the integrin alpha 7 and the myosin heavy chain 7B genes causes congenital myopathy with left ventricular non-compact cardiomyopathy. Orphanet J. Rare Dis. 8, 91. https://doi.org/10.1186/1750-1172-8-91 (2013).
Liu, F. et al. Histone-deacetylase inhibition reverses atrial arrhythmia inducibility and fibrosis in cardiac hypertrophy independent of angiotensin. J. Mol. Cell. Cardiol. 45, 715–723. https://doi.org/10.1016/j.yjmcc.2008.08.015 (2008).
Pappas, C. T. et al. Knockout of Lmod2 results in shorter thin filaments followed by dilated cardiomyopathy and juvenile lethality. Proc. Natl. Acad. Sci. USA 112, 13573–13578. https://doi.org/10.1073/pnas.1508273112 (2015).
Liu, H. et al. HAND2 loss-of-function mutation causes familial dilated cardiomyopathy. Eur. J. Med. Genet. 62, 103540. https://doi.org/10.1016/j.ejmg.2018.09.007 (2019).
Zhang, T. et al. The deltaC isoform of CaMKII is activated in cardiac hypertrophy and induces dilated cardiomyopathy and heart failure. Circ. Res. 92, 912–919. https://doi.org/10.1161/01.RES.0000069686.31472.C5 (2003).
Kang, P. T. et al. Overexpressing superoxide dismutase 2 induces a supernormal cardiac function by enhancing redox-dependent mitochondrial function and metabolic dilation. J. Mol. Cell. Cardiol. 88, 14–28. https://doi.org/10.1016/j.yjmcc.2015.09.001 (2015).
Tougas, C. L., Grindrod, T., Cai, L. X., Alkassis, F. F. & Kasahara, H. Heterozygous Mylk3 knockout mice partially recapitulate human DCM with heterozygous MYLK3 mutations. Front. Physiol. 10, 696. https://doi.org/10.3389/fphys.2019.00696 (2019).
Frey, N. et al. Mice lacking calsarcin-1 are sensitized to calcineurin signaling and show accelerated cardiomyopathy in response to pathological biomechanical stress. Nat. Med. 10, 1336–1343. https://doi.org/10.1038/nm1132 (2004).
Ho, Y. L., Lin, Y. H., Tsai, W. Y., Hsieh, F. J. & Tsai, H. J. Conditional antisense-knockdown of zebrafish cardiac troponin C as a new animal model for dilated cardiomyopathy. Circ. J. 73, 1691–1697. https://doi.org/10.1253/circj.cj-09-0210 (2009).
Wang, Z. et al. The long noncoding RNA Chaer defines an epigenetic checkpoint in cardiac hypertrophy. Nat. Med. 22, 1131–1139. https://doi.org/10.1038/nm.4179 (2016).
Ramratnam, M. et al. Transgenic knockdown of cardiac sodium/glucose cotransporter 1 (SGLT1) attenuates PRKAG2 cardiomyopathy, whereas transgenic overexpression of cardiac SGLT1 causes pathologic hypertrophy and dysfunction in mice. J. Am. Heart Assoc. https://doi.org/10.1161/JAHA.114.000899 (2014).
Chang, A. N. et al. Cardiac myosin light chain is phosphorylated by Ca2+/calmodulin-dependent and -independent kinase activities. Proc. Natl. Acad. Sci. USA 113, E3824-3833. https://doi.org/10.1073/pnas.1600633113 (2016).
Mohler, P. J. et al. Ankyrin-B mutation causes type 4 long-QT cardiac arrhythmia and sudden cardiac death. Nature 421, 634–639. https://doi.org/10.1038/nature01335 (2003).
London, B. et al. Mutation in glycerol-3-phosphate dehydrogenase 1 like gene (GPD1-L) decreases cardiac Na+ current and causes inherited arrhythmias. Circulation 116, 2260–2268. https://doi.org/10.1161/CIRCULATIONAHA.107.703330 (2007).
Burashnikov, E. et al. Mutations in the cardiac L-type calcium channel associated with inherited J-wave syndromes and sudden cardiac death. Heart rhythm : the official journal of the Heart Rhythm Society 7, 1872–1882. https://doi.org/10.1016/j.hrthm.2010.08.026 (2010).
Li, D., Fareh, S., Leung, T. K. & Nattel, S. Promotion of atrial fibrillation by heart failure in dogs: atrial remodeling of a different sort. Circulation 100, 87–95. https://doi.org/10.1161/01.cir.100.1.87 (1999).
Schmidt, C. et al. Inverse remodelling of K2P3.1 K+ channel expression and action potential duration in left ventricular dysfunction and atrial fibrillation: implications for patient-specific antiarrhythmic drug therapy. Eur. Heart J 38, 1764–1774. https://doi.org/10.1093/eurheartj/ehw559 (2017).
Aida, T. et al. Cloning-free CRISPR/Cas system facilitates functional cassette knock-in in mice. Genome Biol. 16, 87. https://doi.org/10.1186/s13059-015-0653-x (2015).
Warren, C. M., Krzesinski, P. R. & Greaser, M. L. Vertical agarose gel electrophoresis and electroblotting of high-molecular-weight proteins. Electrophoresis 24, 1695–1702. https://doi.org/10.1002/elps.200305392 (2003).
Sugiyama, K. et al. Oxidative stress induced ventricular arrhythmia and impairment of cardiac function in Nos1ap deleted mice. Int. Heart J. 57, 341–349. https://doi.org/10.1536/ihj.15-471 (2016).
Mitchell, G. F., Jeron, A. & Koren, G. Measurement of heart rate and Q-T interval in the conscious mouse. Am. J. Physiol. 274, H747-751. https://doi.org/10.1152/ajpheart.1998.274.3.H747 (1998).
Kirchhefer, U. et al. Cardiac hypertrophy and impaired relaxation in transgenic mice overexpressing triadin 1. J. Biol. Chem. 276, 4142–4149. https://doi.org/10.1074/jbc.M006443200 (2001).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915. https://doi.org/10.1038/s41587-019-0201-4 (2019).
Li, H. et al. The sequence alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079. https://doi.org/10.1093/bioinformatics/btp352 (2009).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295. https://doi.org/10.1038/nbt.3122 (2015).
Ge, S. X., Son, E. W. & Yao, R. iDEP: an integrated web application for differential expression and pathway analysis of RNA-Seq data. BMC Bioinform. 19, 534. https://doi.org/10.1186/s12859-018-2486-6 (2018).
Acknowledgements
We thank Dr. Jun K. Takeuchi and Dr. Yasuhiro Maejima for their helpful discussions and Eichi Watabe for technical assistance. We thank Wei Guo of University of Wyoming for anti-RBM20 antibody used in our preliminary experiment. This study was supported by Japan Heart Foundation Dr. Hiroshi Irisawa & Dr. Aya Irisawa Memorial Research Grant (to K.I.), Grant-in-Aid for Scientific Research (JP20K08467 to K.I. and JP20K21385 to H.K.), and Nanken-Kyoten, TMDU (to H.K., K.I. and Y.H.). We would like to thank Editage (www.editage.com) for English language editing.
Author information
Authors and Affiliations
Contributions
K.I., T.S., M.T.-O., T.F. and H.K. contributed to animal experiments. Y.H. contributed to generation and analysis of knock-in and knock-out mice. M.T. and H.O. contributed to antibody generation. K.I., Y.S. and M.S. contributed to histological assessment. K.I. and H.K. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ihara, K., Sasano, T., Hiraoka, Y. et al. A missense mutation in the RSRSP stretch of Rbm20 causes dilated cardiomyopathy and atrial fibrillation in mice. Sci Rep 10, 17894 (2020). https://doi.org/10.1038/s41598-020-74800-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-74800-8
This article is cited by
-
Mislocalization of pathogenic RBM20 variants in dilated cardiomyopathy is caused by loss-of-interaction with Transportin-3
Nature Communications (2023)
-
Striated muscle-specific base editing enables correction of mutations causing dilated cardiomyopathy
Nature Communications (2023)
-
Modulating biomolecular condensates: a novel approach to drug discovery
Nature Reviews Drug Discovery (2022)
-
I536T variant of RBM20 affects splicing of cardiac structural proteins that are causative for developing dilated cardiomyopathy
Journal of Molecular Medicine (2022)
-
Rbm24 displays dynamic functions required for myogenic differentiation during muscle regeneration
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.