Abstract
The targeted sequencing of the 16S rRNA gene is one of the most frequently employed techniques in the field of microbial ecology, with the bacterial communities of a wide variety of niches in the human body have been characterised in this way. This is performed by targeting one or more hypervariable (V) regions within the 16S rRNA gene in order to produce an amplicon suitable in size for next generation sequencing. To date, all technical research has focused on the ability of different V regions to accurately resolve the composition of bacterial communities. We present here an underreported artefact associated with 16S rRNA gene sequencing, namely the off-target amplification of human DNA. By analysing 16S rRNA gene sequencing data from a selection of human sites we highlighted samples susceptible to this off-target amplification when using the popular primer pair targeting the V3–V4 region of the gene. The most severely affected sample type identified (breast tumour samples) were then re-analysed using the V1–V2 primer set, showing considerable reduction in off target amplification. Our data indicate that human biopsy samples should preferably be amplified using primers targeting the V1–V2 region. It is shown here that these primers result in on average 80% less human genome aligning reads, allowing for more statistically significant analysis of the bacterial communities residing in these samples.
Similar content being viewed by others
Introduction
This communication highlights off-target amplification of human DNA in 16S rRNA gene sequencing, detailing the circumstances necessary for this to occur, and the effects on ensuing research. Such artefacts are not a universal problem, and only occur in samples containing an overwhelming ratio of human to bacterial DNA. This leaves stool samples and skin samples which contain less than 10% and 90% human DNA respectively, unaffected, but can critically impact on analysis of human biopsy samples, where over 97% of the DNA present is of human origin1. Given the increased use of human biopsies from a number of body sites in microbiome research2,3,4,5, this communication serves as a timely and, to our knowledge, unique methodological warning and remedy, particularly as only one mention of this issue can currently be found in the literature6.
Currently, comparisons of primer pairs and the hypervariable regions they target in the 16S rRNA gene have focused exclusively on differing levels of taxonomic resolution and specificity7,8. The degree to which bacterial resolution is lost to the production human-derived amplicons has, so far, received no attention. This is because workflows for the analysis of 16S rRNA gene sequencing data typically remove reads falling too far from the mean or median sequence length, or if they are not classified taxonomically as originating from bacterial DNA. This is effective in ensuring that the presence of amplified human DNA does not have any impact on downstream analysis. Unaddressed is the fact that in a sequencing experiment yielding a finite amount of data (up to 15 Gb on a typical Miseq run9), a significant proportion of these can be wasted due to this off target amplification. This affects sequencing studies in two ways.
-
Prospectively: If this loss of data is anticipated, fewer samples can be sequenced on a given sequencing run, adding to the expense which is already prohibitive for smaller labs.
-
Retrospectively: If this loss if data is not anticipated, insufficient bacterial reads may be yielded to accurately characterise the samples being sequenced, particularly if attempting to identify the prevalence of rare taxa between different treatment groups.
Here, we show that the most commonly-used primer set for 16S rRNA sequencing, targeting the V3–V4 hypervariable regions, is particularly susceptible to this off-target amplification, while another commonly used primer set, targeting the V1–V2 primer region, shows almost no off-target amplification, as outlined in Fig. 1 below. While this off-target amplification does not appear to affect research using stool or skin swab samples, we would urge all groups carrying out metataxonomic analysis of low microbial biomass human biopsy samples using high throughput sequencing to use the V1–V2 primer set in future.
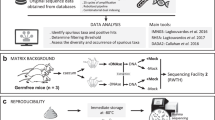
Proposed mechanism for off target amplification of mammalian DNA by V3–V4 primers, as opposed to V1–V2. (A) DNA extracted from human biopsies is known to contain large proportions of human DNA. In these circumstances V3–V4 degenerate primers, which also align to region in human mitochondrial DNA as shown can bind and amplify human DNA. There is no such alignment for V1–V2 degenerate primers. (B) Off target amplification significantly alters the 16S rRNA gene sequencing profile of a sample.
Materials/methods
Sample collection
Breast tissue was collected from women undergoing breast surgery at Cork University Hospital, Cork, Ireland. Breast tumour core-biopsies were aseptically resected using an Achieve 14G Breast Biopsy System (Iskus Health, UT, USA). The specimens were transported in sterile PBS to the lab, where they were flash-frozen and kept at − 80 °C until further processing. DNA from the specimens was purified following the protocol and reagents provided in the Ultra Deep Microbiome Prep (Molzym, GmbH & Co. KG., Bremen, Germany) and eluted in 100 µl of Tris–HCl.
DNA purification
Samples were processed and DNA purified following the procedures specified in protocols listed in Table 1. In all cases, DNA was eluted in Tris–HCl buffer and stored at − 20 °C until further analysis.
16S rRNA gene sequencing library preparation
Genomic DNA was amplified by PCR with primers targeting the hypervariable V1–V2 region or the V3–V4 region of the 16S rRNA gene. Table 2 details the primers sequences (underlined) included for compatibility with the Illumina 16S Metagenomic Sequencing Protocol (Illumina, CA, USA).
For Breast Tumour and Normal Adjacent samples, amplification was performed in 50 µl reactions, containing 1X NEBNext High Fidelity 2X PCR Master Mix (NEB, USA), 0.5 µM of each primer, 8 µl template (5–15 ng/µl) and 12 µl nuclease free water. The thermal profile included an initial 98 °C × 30 s denaturation, followed by 25 cycles of denaturation at 98 °C × 10 s, annealing at 55 °C × 30 s for V3–V4 or 62 °C × 30 s for V1–V2 and extension at 72 °C × 30 s. Plus a final extension at 72 °C × 5 min. Amplification was confirmed by running 5 µl of PCR product on a 2% agarose gel, by visualisation of a ≈ 310 bp band for V1–V2 and ≈ 460 bp band for V3–V4.
Faecal microbial genomic DNA was amplified using Phusion High-Fidelity DNA Polymerases (Thermo Scientific, Massachusetts, USA) with the PCR thermocycler protocol as follows: Initiation step of 98 °C for 3 min followed by 25 cycles of 98 °C for 30 s, 55 °C for 60 s, and 72 °C for 20 s, and a final extension step of 72 °C for 5 min.
Oesophageal biopsies and skin swab samples microbial genomic DNA was amplified using MTP Taq DNA Polymerase (Merck KGaA, Darmstadt, Germany) with the PCR thermocycler protocol as follows: Initiation step of 94 °C for 1 min followed by 35 cycles of 94 °C for 60 s, 55 °C for 45 s, and 72 °C for 30 s, and a final extension step of 72 °C for 5 min.
An index PCR was performed to add sample specific DNA barcodes to sample amplicons in accordance with the Illumina 16S Metagenomic Sequencing Protocol (Illumina, California, USA)16. Libraries DNA concertation was quantified using a Qubit fluorometer (Invitrogen) using the ‘High Sensitivity’ assay and samples were pooled at a standardised concentration16. The pooled library was sequenced on the Illumina MiSeq platform (Illumina, California, USA) utilising 2 × 300 bp chemistry.
16S rRNA sequence analysis
The quality of the paired-end sequencing data was visualised using FastQC v (0.11.9), and trimmed using Trimmomatic v (0.39) ensuring a minimum average quality of 25. Reads were then imported into R environment v (3.6.3)17 to be resolved into Amplicon Sequence Variants by the DADA2 package v (1.12).
Contamination control
In all samples a contamination control strategy was implemented in keeping with the RIDE checklist as proposed by Eisenhofer et al.18, incorporating aseptic techniques and a variety of negative controls from different stages of the sample-to-sequence data process. Retrospective contamination assessment and removal based on sequencing data from negative controls was also performed following published guidelines19.
Retrospective bioinformatics based removal of human amplicons
Sequencing reads aligning to the human genome (GRCh38) within the fasta file generated by DADA2 were identified using bowtie220. To confirm reads mapped to the human genome were not erroneously aligned bacterial reads, all human aligning reads were classified with Mothur21, using the RDP database v (11.4) as a reference.
Statistical analysis and data visualisation
All statistical analysis was carried out in the R environment, using the following libraries: Phyloseq v (1.30), Vegan v (2.5.6), ggplot2 v (3.3.0), reshape2 v (1.4.3).
Ethical approval
All procedures in this study were performed in accordance to national ethical guidelines, following ethical approval from the University College Cork Clinical Research Committee.
Informed consent
Patients provided written informed consent for sample collection and subsequent analyses.
Results and discussion
All three sampled biopsy sites where an overwhelming ratio of host DNA was expected (breast, breast tumour and oesophageal) showed significant off target amplification of human DNA when amplified using the V3–V4 primer set (Fig. 2). This was not seen when sequencing samples with lower levels of human DNA, such as skin swabs and stool samples. An average of 34.1% of all Amplicon Sequence Variants (ASV) detected in normal breast tissue samples were shown to align to the human genome GRCh38 using bowtie2.This included the most prevalent ASV, which was identified further using BLAST as Homo sapiens haplogroup H8 mitochondrion, complete genome (Accession no. MN986463.1) with an E-value of 7e − 138 and 100% identity. In the breast tumour samples, 77.2% of all ASV’s detected aligned to the human genome, with the most prevalent ASV again being identified as Homo sapiens haplogroup H8 mitochondrion, complete genome (Accession no. MN986463.1) with an E-value of 7e − 138 and 100% identity. This situation was identical in Oesophageal biopsies, with a 55.6% of ASVs aligning to the human genome (Homo sapiens haplogroup H8 mitochondrion, complete genome (Accession no. MN986463.1) with an E-value of 7e − 138 and 100% identity). The skin swab samples showed a much lower level of amplification of human DNA, but these reads aligned to chromosomal DNA, most frequently Homo sapiens chromosome 17, clone RP11-646F1, complete sequence and were present in very low levels.
While human contamination is a very common problem in amplification-free shotgun metagenomic sequencing strategies22, it is under reported as an issue for 16S rRNA gene sequencing, due to the use of bacteria/archaea specific primers. However, degenerate primers are routinely used for 16S rRNA sequencing23. This increases coverage, in terms of the number of 16S rRNA sequences matched by at least one primer, but also allows for off target amplification of non-bacterial DNA. Figure 1A shows that the V3–V4 primers align to a region within the human mitochondrial DNA. We show here that when the ratio of host:bacterial DNA is overwhelming, human mitochondrial DNA can be amplified by primers targeting the 16S rRNA gene region. To ensure the validity of the results, reads identified as aligning to the human genome using Bowtie2 were classified using the Mothur21 classifier trained on the RDP database. In all cases the reads identified as aligning to the human genome could not be classified when screened against the RDP database as shown in Table 3 below.
The most heavily affected sample type in our study (breast tumour tissue) was reanalysed by performing a pairwise comparison of samples amplified with the V3–V4 and V1–V2 primer sets (Fig. 3).
Looking initially at the rarefaction curves produced by the sequencing data corresponding to the previously mentioned paired V1–V2 and V3–V4 primer pair amplified breast tumour sample there is a clear difference between the two groups. This is done by plotting new species against number of reads per sample. Figure 3A below shows that the distribution of samples in this 2D plane appears to be stochastic prior to the removal of human reads. Figure 3B, following removal of human reads, shows clearly that samples amplified with the V1–V2 primer pair consistently yield more observable species, a greater number of reads per sample, and a plateauing of the rarefaction curve which suggests sufficient sampling depth is available for accurate characterisation.
The community structure in samples amplified with V1–V2 primers was visually similar to those amplified with V3–V4 primers (Fig. 4A) and no bacterial family was found to be significantly elevated using one primer set over the other as per Wilcoxon signed-rank test, once p-values had been corrected for multiple testing using the FDR method (Supplementary Table 1). There was also no significant difference in terms of Shannon diversity (Fig. 4B), indicating choice of primers did not have any adverse effect on the downstream results. Of considerable interest to any groups carrying out low biomass research in the future, is the huge discrepancy in the number of reads yielded once human contamination had been filtered out. As can be seen in Fig. 4C, samples amplified with primers targeting the V1–V2 region have a consistently and significantly higher number of ASVs per sample following the removal of ASV’s aligning to the human genome.
Pairwise comparison of matched samples using primers targeting the V1–V2 and V3–V4 regions of the 16S rRNA gene fragment. (A) Sample composition at the family level of paired samples. (B) Average Shannon Diversity comparison between samples amplified using V1–V2 primers (red) and V3–V4 primers (blue). (C) Percentage of total sequencing reads aligning to human genome. In both (B) and (C) statistical testing is performed using Wilcoxon signed-rank test.
Future perspectives
Third generation sequencing technologies, such as those produced by Oxford Nanopore Technologies and Pacific BioSiences are now being utilised in 16S rRNA gene sequencing experiments. The Pacific BioSciences SMRT platform has seen the greatest promise in this regard with the implementation of “Circular Consensus Sequencing” in conjunction with denoising algorithms, allowing for the production of long reads of high quality24. Earl et al. showed that this new method using degenerate primers targeting the entire 16S rRNA gene, still resulted in off target amplification of the human genome25. This study also noted that this off target amplification was related to the ratio of human to bacterial DNA. The human genome must be considered when designing or choosing primers now and in the future.
References
Pereira-Marques, J. et al. Impact of host DNA and sequencing depth on the taxonomic resolution of whole metagenome sequencing for microbiome analysis. Front. Microbiol. 10, 1277 (2019).
Deshpande, N. P. et al. Signatures within the esophageal microbiome are associated with host genetics, age, and disease. Microbiome 6(1), 227 (2018).
Riquelme, E. et al. Tumor microbiome diversity and composition influence pancreatic cancer outcomes. Cell 178(4), 795-806.e12 (2019).
Grice, E. A. & Segre, J. A. The skin microbiome. Nat. Rev. Microbiol. 9(4), 244–253 (2011).
Urbaniak, C. et al. The microbiota of breast tissue and its association with breast cancer. Appl. Environ. Microbiol. 82(16), 5039 (2016).
Davis, N. M. et al. Simple statistical identification and removal of contaminant sequences in marker-gene and metagenomics data. Microbiome 6(1), 226 (2018).
Pinna, N. K. et al. Can targeting non-contiguous V-regions with paired-end sequencing improve 16S rRNA-based taxonomic resolution of microbiomes? An in silico evaluation. Front. Genet. 10, 653 (2019).
Johnson, J. S. et al. Evaluation of 16S rRNA gene sequencing for species and strain-level microbiome analysis. Nat. Commun. 10(1), 5029 (2019).
Ravi, R. K. et al. Miseq: A next generation sequencing platform for genomic analysis. Methods Mol Biol. 1706, 223–232 (2018).
Flemer, B. et al. Tumour-associated and non-tumour-associated microbiota in colorectal cancer. Gut 66(4), 633–643 (2017).
Yu, Z. & Morrison, M. Improved extraction of PCR-quality community DNA from digesta and fecal samples. Biotechniques 36(5), 808–812 (2004).
Costea, P. I. et al. Towards standards for human fecal sample processing in metagenomic studies. Nat. Biotechnol. 35(11), 1069–1076 (2017).
Browne, H. P. et al. Culturing of ‘unculturable’ human microbiota reveals novel taxa and extensive sporulation. Nature 533(7604), 543–546 (2016).
Elliott, D. R. F. et al. A non-endoscopic device to sample the oesophageal microbiota: A case–control study. Lancet Gastroenterol. Hepatol. 2(1), 32–42 (2017).
Klindworth, A. et al. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 41(1), e1 (2013).
Illumina. Amplicon, P. C. R., Clean-Up, P. C. R. & Index, P. C. R. 16S Metagenomic Sequencing Library Preparation. https://www.illumina.com/content/dam/illumina-support/documents/documentation/chemistry_documentation/16s/16s-metagenomic-library-prep-guide-15044223-b.pdf (2013).
Team, R.C. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, Vienna, 2019).
Eisenhofer, R. et al. Contamination in low microbial biomass microbiome studies: Issues and recommendations. Trends Microbiol. 27(2), 105–117 (2019).
Walker, S.P., Tangney, M., & Claesson, M.J. Sequence-based characterization of intratumoral bacteria—A guide to best practice.Front. Oncol.10(179), (2020).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9(4), 357–359 (2012).
Schloss, P. D. et al. Introducing mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 75(23), 7537–7541 (2009).
Marotz, C. A. et al. Improving saliva shotgun metagenomics by chemical host DNA depletion. Microbiome 6(1), 42 (2018).
Sambo, F. et al. Optimizing PCR primers targeting the bacterial 16S ribosomal RNA gene. BMC Bioinform. 19(1), 343–343 (2018).
Callahan, B. J. et al. High-throughput amplicon sequencing of the full-length 16S rRNA gene with single-nucleotide resolution. Nucleic Acids Res. 47(18), e103–e103 (2019).
Earl, J. P. et al. Species-level bacterial community profiling of the healthy sinonasal microbiome using Pacific Biosciences sequencing of full-length 16S rRNA genes. Microbiome 6(1), 190 (2018).
Acknowledgements
The authors acknowledge the contribution of Prof. Paul O’Toole, Microbiology Department, UCC, in providing access to sequencing data used in this study.
Author information
Authors and Affiliations
Contributions
S.W., M.B., M.J.C. and M.T. conceived and wrote the manuscript, S.W. and M.B. performed sequence and statistical analysis, M.B., G.H. and Y.F.B. generated sequence data. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Walker, S.P., Barrett, M., Hogan, G. et al. Non-specific amplification of human DNA is a major challenge for 16S rRNA gene sequence analysis. Sci Rep 10, 16356 (2020). https://doi.org/10.1038/s41598-020-73403-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-73403-7
This article is cited by
-
Using formalin fixed paraffin embedded tissue to characterize the microbiota in p16-positive and p16-negative tongue squamous cell carcinoma: a pilot study
BMC Oral Health (2024)
-
16S rRNA gene primer choice impacts off-target amplification in human gastrointestinal tract biopsies and microbiome profiling
Scientific Reports (2023)
-
Questioning the fetal microbiome illustrates pitfalls of low-biomass microbial studies
Nature (2023)
-
Higher off-target amplicon detection rate in MiSeq v3 compared to v2 reagent kits in the context of 16S-rRNA-sequencing
Scientific Reports (2022)
-
Biopsy bacterial signature can predict patient tissue malignancy
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.