Abstract
Many emerging technologies are reliant on circulating cell-free DNA (cfDNA) and cell-free RNA (cfRNA) applications in the clinic. However, the impact of diurnal cycles or daily meals on circulating analytes are poorly understood and may be confounding factors when developing diagnostic platforms. To begin addressing this knowledge gap, we obtained plasma from four healthy donors serially sampled five times during 12 h in a single day. For all samples, we measured concentrations of cfDNA and cfRNA using both bulk measurements and gene-specific digital droplet PCR. We found no significant variation attributed to blood draw number for the cfDNA or cfRNA. This indicated that natural diurnal cycles and meal consumption do not appear to significantly affect abundance of total cfDNA, total cfRNA, or our two selected cfRNA transcripts. Conversely, we observed significant variation between individual donors for cfDNA and one of the cfRNA transcripts. The results of this work suggest that it will be important to consider patient-specific baselines when designing reliable circulating cfDNA or cfRNA clinical assays.
Similar content being viewed by others
Introduction
Liquid-biopsy based diagnostic platforms are highly desired and increasingly accepted in clinical settings1. Although many diseases could potentially benefit from liquid-biopsy technology, the need for non-invasive platforms is especially apparent in the field of cancer screening and diagnostics because of potential risks involved with invasive needle biopsy procedures (e.g.2,3) and concerns of radiation exposure during imaging tests4,5. In addition, there is substantial interest in accurate screening methods that can be performed at frequent intervals to stratify patient cancer risk and detect potentially lethal cancers at early, treatable stages6. Plasma or serum-based platforms are of particular interest because the circulatory system interacts with the entire body and therefore provides a means to sample all organs. The sensitivity of current methods to reliably characterize nucleotide sequences and subcellular particles at single-event resolution has generated substantial interest in utilizing circulating cell-free DNA (cfDNA, e.g.7,8,9,10) and cell-free RNA (cfRNA, e.g.11,12,13,14,15) as clinically-relevant biomarkers. Human plasma contains both vesicular and extravesicular RNA and DNA; these different components may have distinctive contents with potential clinical relevance16. While promising, translation of cfDNA and cfRNA to the clinic has been slow, in part, because the natural temporal and interpersonal variation of the circulating analytes remains poorly understood. It is well-known that mammalian blood cell/tissue gene expression and physiology changes drastically during the daily diurnal cycle or following meals17,18,19,20. Recent evidence also suggests that some human bodily fluid-derived micro-RNAs (miRNAs) may follow a daily cycle of fluctuation21,22. Therefore, establishing the extent to which circulating cfDNA and cfRNA is affected by normal physiology is critical for the analytes to have successful clinical implementation.
To address the potential influence of daily cycles on circulating analytes, we characterized total cfDNA and cfRNA in plasma obtained from four healthy volunteers sampled multiple times across 2 days. We measured bulk cfDNA and cfRNA concentration using fluorometric or automated electrophoresis methods, as well as sequence-specific cfDNA and cfRNA copy number concentration using digital droplet PCR (ddPCR). The droplet-counting approach of ddPCR allows for absolute quantitation of nucleic acid templates and more sensitive resolution of fold-changes compared to conventional quantitative PCR23,24,25. Using this ddPCR approach, we targeted two single-copy genomic DNA regions as proxies for cfDNA abundance: one locus containing the gene telomerase reverse transcriptase (TERT) and one locus containing the gene N-acetylglucosamine kinase (NAGK); for cfRNA we targeted two commonly used genes used for mRNA normalization: β-actin (ACTB) and glyceraldehyde-3-phosphate dehydrogenase (GAPDH). Our results suggest that while cfDNA and cfRNA are overall stably expressed diurnally, several of the analytes demonstrate significant interpersonal or daily variation. Deeper characterization of these sources of variation will likely be required before the circulating analytes gain greater acceptance as clinically practical liquid-biopsy platforms.
Methods
Participant plasma sample collection
All experimental protocols were reviewed and approved by the Oregon Health & Science University Institutional Review Board (protocol #8316). All methods were carried out in accordance with relevant guidelines and regulations. Informed consent was obtained from all volunteers, and volunteers were compensated for participating. Healthy donors (HDs) were consented for multiple blood draws over a 12-h period at the Oregon Health & Science University Oregon Clinical and Translational Research Institute (OCTRI) inpatient research clinic. HD age and sex distributions are given in Table 1. Each HD had 2 days, separated by 1 week, to provide five blood draws each day (Fig. 1). The HDs were advised to not engage in rigorous exercise 24 h prior to the blood draw dates. HDs were given access to recliner chairs and had the ability to walk around freely between blood draws. All individuals had an IV inserted for the multiple blood draws, but at times when the IV failed, ad hoc venipuncture was performed about 50% of the time for each participant. Blood was drawn from individuals every 2 h 45 min beginning at 8:30 am. Meals ordered from the hospital menu were consumed by individuals between draw 1–2, draw 2–3, and draw 4–5. To replicate a typical patient arriving in a clinical setting, diets were not restricted. Approximately 20 ml of blood was drawn into 10 ml EDTA tubes (Cat# 366643, BD Vacutainer) per time point. Blood was processed within 15 min of the draw and plasma was obtained as follows: EDTA tubes were centrifuged at 1000 × g for 10 min at room temperature, plasma was extracted down to ~ 500 μl from the buffy coat interface, plasma supernatant was centrifuged for a second time at 2500 × g for 10 min at room temperature, and the resulting plasma supernatant was extracted down to ~ 200 μl from the debris pellet interface. The final supernatant was distributed into 1 ml aliquots and immediately frozen at − 80 °C until analysis.
Nucleic acid extractions
For each HD and time point, cfDNA and cfRNA was extracted separately (Fig. 1). CfDNA was extracted from 1 ml plasma using the QIAamp Circulating Nucleic Acid Kit (Qiagen #55114) according to manufacturer instructions and eluted into 20 μl buffer EB (10 mM Tris–Cl, pH 8.5). CfRNA was extracted from 1 ml plasma using the Plasma/Serum Circulating and Exosomal RNA Purification Kit (Norgen #42800) and eluted into 100 μl nuclease-free deionized water. To remove genomic DNA contamination, the cfRNA samples were treated with 2 MBU of Baseline-ZERO DNase in 1X Baseline-ZERO DNAse buffer (Lucigen #DB0715K) for 20 min at 37 °C, purified using the RNA Clean & Concentrator-5 kit (Zymo #R1013), and eluted in 14 μl nuclease-free water. For no plasma controls, 1 ml nuclease-free deionized water was used in place of plasma. These purified cfDNA and cfRNA samples were used for all subsequent nucleic acid analyses.
Bulk quantitation of cfDNA and cfRNA
Purified cfDNA concentration was first determined using the Qubit dsDNA HS Assay Kit (quantification range 0.2–100 ng DNA, Thermo Fisher Scientific #Q32854) and Qubit 3 Fluorometer (Thermo Fisher Scientific). Purified cfRNA concentration was quantified using the Agilent RNA 6000 Pico Kit (quantification range 50–5000 pg/μL RNA in water, Agilent #5067–1513) and Agilent 2100 Bioanalyzer instrument (Agilent) within a window of 50–500 bp.
First-strand cDNA synthesis of cfRNA
For first-strand cDNA synthesis of cfRNA, 10 μl reverse transcription reactions were prepared using 3 μl total cfRNA in 1X SuperScript IV VILO Mastermix (Thermo Fisher Scientific #11756050). cDNA synthesis reactions were incubated at 25 °C for 10 min, followed by 50 °C for 10 min, and were terminated with incubation at 85 °C for 5 min. The cDNA reactions were used as direct template for cDNA copy number quantification by ddPCR.
Primers for ddPCR analysis
Primers and probes for cfDNA ddPCR analysis to target single-copy number genes TERT (F primer 5′–3′: CCTCACATAAATGCTACCAAACGA; R primer 5′–3′: TTCCAAGAAGGAGGCCATAGTC; Probe 5′–3′: AAGAAATGAACAGACCCATCCCCCAGG; fluorescent probe: HEX; quencher: ZEN/IBFQ) or NAGK (F primer 5′–3′: TGGGCAGACACATCGTAGCA; R primer 5′–3′: CACCTTCACTCCCACCTCAAC; Probe 5′–3′: TGTTGCCCGAGATTGACCCGGT; fluorescent probe: FAM; quencher: ZEN/IBFQ) and were purchased from IDT (Integrated DNA Technologies, Coralville, IA, USA). Primer and probe sequences for cfDNA were chosen using sequences reported by Devonshire et al.26 for these two genomic loci. To quantify cDNA copy number, gene expression ddPCR assays for ACTB (assay dHsaCPE5190199; fluorescent probe: FAM) and GAPDH (assay dHsaCPE5031597; fluorescent probe: HEX) were purchased from Bio-Rad and contained both probes and primers premixed at 10X concentration.
ddPCR of cfDNA and cDNA samples
To measure cfDNA copy number by ddPCR, 22 μl ddPCR reaction mixtures were prepared using 2.2 μl purified cfDNA in 1X ddPCR Supermix for Probes (No dUTP) (Bio-Rad #1863024) and with final primer/probe concentrations of 0.9 µM/0.25 µM or 0.2 µM/0.1 µM for TERT and NAGK, respectively. Each cfDNA ddPCR reaction was multiplexed for both TERT and NAGK. To measure cDNA copy number by ddPCR, 22 μl ddPCR reaction mixtures were prepared using 1.5 μl undiluted cDNA template in 1X ddPCR Supermix for Probes (No dUTP) (Bio-Rad #1863024), 1X ACTB gene expression ddPCR assay mix, and 1X GAPDH gene expression ddPCR assay mix. For no template controls (NTCs), 1 μl nuclease-free water was used instead of cfDNA or cDNA template. Reactions were performed in semi-skirted 96-well plates (Eppendorpf #951020362). Plates for droplet generation were heat-sealed with pierceable foil (Bio-Rad #1814040), vortexed briefly, then spun down using a tabletop plate spinner. Droplet generation was performed using a QX200 AutoDG Droplet Digital PCR System (Bio-Rad) with Automated Droplet Generation Oil for Probes (Bio-Rad #1864110) and DG32 Automated Droplet Generator Cartridges (Bio-Rad #1864108). Droplets were deposited into a clean 96-well plate held in a pre-chilled cold block to prevent evaporation. Plates were then heat-sealed with pierceable foil and PCR was performed in a Bio-Rad C1000 thermocycler using the following temperature conditions: 95 °C for 10 min, 40 cycles of 94 °C for 30 s followed by 60 °C for 1 min, 98 °C for 10 min, and then cooling to 4 °C until droplets were read. Droplets were counted using the QX200 Droplet Reader (Bio-Rad) using manufacturer’s instructions. Positive and negative droplets were subsequently analyzed using QuantaSoft Analysis Pro (v1.0.596, Bio-Rad, Hercules, California, USA).
Statistics and plots
Graphs of cfDNA and cfRNA measured over time were prepared using R (v.3.6.1) and Rstudio (v.1.2.5019). Permutation tests for each analyte were performed in Rstudio using the “coin” R package and default parameters27,28. For significant permutation tests, post-hoc pairwise permutation tests were performed using the R package “rcompanion” (v. 2.3.2, https://rcompanion.org) with the “fdr” P value adjustment method. To test for a significant difference between draws performed on day 1 versus day 2, a permutation test of symmetry was performed on the draws paired by day. To test for a significant difference between draws or between individuals, the values between day 1 and day 2 for each draw were averaged and one-way permutation tests of independence were performed using either draws or individuals as factors. Summary statistics, correlation plots, Spearman nonparametric correlation coefficient, and two-tailed correlation P value analysis were prepared using GraphPad Prism (v8.3.0, GraphPad Software, San Diego, CA, USA). For all statistical tests, P values were determined to be significant at a threshold of P ≤ 0.05.
Results and discussion
Over the past decade, it has become increasingly clear that detecting unpredictable disease via blood biopsy, especially at the earliest stages, will require intricate understanding of naturally occurring circulating biomarker variation. Workflow standardization for liquid-biopsy based analytes is the first step towards identifying and minimizing sources of non-biologically relevant variation29,30,31. However, confounding factors related to meals, time of day, and the intrinsic interpersonal variation could critically affect analyte abundance, normalization, and multi-omic integration. To begin addressing these concerns, here we provide the first descriptions of diurnal cfDNA and cfRNA derived from the same cohort.
Differences in plasma cfDNA abundance across donors is attributed to interpersonal variation
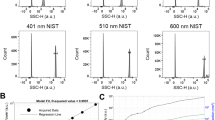
First, we characterized total donor plasma cfDNA abundance using Qubit as well as total genome copy numbers via locus-specific ddPCR assays. Following cfDNA extractions of plasma from the four HDs, we observed overall averages of 3.05 ng cfDNA / ml plasma when measured by Qubit (SD = 1.2 ng cfDNA / ml plasma, Fig. 2A,B, Supplemental table S1). When cfDNA was measured by ddPCR, we observed 758.6 copies/ml plasma (SD = 286.2) and 723.5 copies/ml plasma (SD = 299.3) using TERT and NAGK probes, respectively (Figs. 2C–F). The ddPCR workflow for cfDNA yielded an average of 14,658 accepted droplets (range = 10,426–18,130; SD = 1366) per ddPCR reaction well (Supplemental Figure S1). No plasma controls and NTCs for TERT/NAGK ranged from 0–9.3 copies/ml (Supplemental Figure S2). We observed a strong correlation between the two cfDNA extraction replicates (Spearman rs = 0.73, P < 0.0001, Figure S3A) and between the two analysis methods (Spearman rs = 0.97, P < 0.0001, Figure S3B). We found no significant difference in cfDNA abundance between the five draws when measured by either Qubit or ddPCR (Table 2, Supplemental tables S1–S3). This finding contrasts with a recent report by Madsen et al.32 which found a decrease in plasma cfDNA concentration at their final draw when five draws were performed three hours apart. However, our second centrifugation step was performed at 2500 × g, rather than 13,000 × g as done by Madsen et al.32, and therefore the composition of cell-free plasma may not be directly comparable. We observed no significant difference between the two draw days (Table 2), although we did observe a significant source of variation attributed to the individuals (P < 0.05, Table 2, Supplemental Table S1). For example, the cfDNA level of individual HD1 was consistently lower than the overall average (Fig. 2A, B, Supplemental Tables S1–S3). Post-hoc pairwise permutation tests also identified individuals with significantly different cfDNA levels (Supplemental Tables S1–S3). Plasma cfDNA was previously described by Zhong et al.33 to fluctuate 1.9–67.9 fold in healthy, nonpregnant individuals when sampled across 12-h or longer time points. Similar to Zhong et al., we observed inconsistent cfDNA fluctuation across time in the HDs and attribute the primary source of plasma cfDNA abundance differences to be from interpersonal variation.
Abundance of plasma-derived cfDNA across the five sampled time points. CfDNA abundance was measured by Qubit (A, B) and by ddPCR with TERT (C, D) or NAGK probes (E, F). For left panels, dashed lines represent the average of the four individuals and the shaded area corresponds to 95% confidence intervals. For right panels, the individual data points overlaying boxplots are color coded sequentially starting from the first blood draw of day 1.
GAPDH counts, but not ACTB counts or total cfRNA, varied significantly by donor
Next, we measured total plasma cfRNA abundance by Bioanalyzer and mRNA-specific abundance characterization using ddPCR. Across the four HDs, we observed an overall average of 3.05 ng/ml plasma cfRNA when measured by Bioanalyzer (SD = 1.2 ng/ml plasma, Fig. 3A,B, Supplemental table S4). We did not find significant cfRNA variation by day, draw, or individual when total cfRNA abundance was measured using this method (Table 2). When cfRNA was measured by ddPCR, we observed 25022 copies/ml plasma (SD = 5932) and 5983 copies/ml plasma (SD = 1703) using ACTB and GAPDH probes, respectively (Fig. 3C–F). The ddPCR workflow for cfRNA yielded an average of 16,216 droplets per well (range = 11,912–19,618; SD = 1554) for ACTB and GAPDH cDNA templates (Supplemental Figure S2). No plasma controls and NTCs for ACTB/GAPDH ranged from 0 to 4.3 copies/ml (Supplemental Figure S5). Similar to our bulk measurement of cfRNA by Bioanalyzer, we did not observe significant variation due to day of draw or draw number for either ACTB or GAPDH (Fig. 3C–F; Table 2, Supplemental Tables S5 and S6). However, we observed significant variation attributed to the individuals for GAPDH counts (P < 0.05, Table 2). Post-hoc pairwise permutation tests for GAPDH counts did not reveal specific significant differences between individuals (Supplemental Tables S6).
Abundance of plasma-derived cfRNA across the five sampled time points. CfRNA abundance was measured by Bioanalyzer (A, B) and by ddPCR with probes targetting ACTB (C, D) or GAPDH (E, F). For left panels, dashed lines represent the average of the four individuals and the shaded area corresponds to 95% confidence intervals. For right panels, the individual data points overlaying boxplots are color coded sequentially starting from the first blood draw of day 1.
ACTB and GAPDH are commonly used mRNA normalization genes for liquid biopsy applications34,35 despite accumulating evidence suggesting their expression can be highly variable and require situation-specific considerations36,37,38. Currently, there are few reports describing the long-term stability of plasma mRNA expression in individuals. Recent work by Max et al.31 found no significant variation miRNA plasma/serum profiles following meals and also described interpersonal differences in miRNA abundance that could be stable for up to a year. While we similarly did not find an effect of meals on bulk cfRNA abundance or ACTB/GAPDH expression, we found significant variation attributed to individual donors for GAPDH transcripts. Like our finding with plasma cfDNA abundance, our results suggest that at least some plasma-derived cfRNA transcripts may have baseline expression levels that are specific to the donor; therefore, a thorough understanding of cfRNA normalization transcripts across time and between individuals may be necessary for future cfRNA diagnostic applications.
Conclusions
Liquid-biopsy technology for patient risk-stratification, diagnosis, or disease progression monitoring will likely require biomarker thresholds tailored specifically for each patient. In our pilot cohort, we observed significant interpersonal variation for each of the two analytes examined. Remarkably, distinct baseline levels of cfDNA and cfRNA of each individual were persistent through the draws over time. Future multi-omic studies such as the work presented here, but with larger and more inclusive cohorts, will be essential for determining the full extent of interpersonal and sample collection variation that may be present across populations.
References
Sheridan, C. Investors keep the faith in cancer liquid biopsies. Nat. Biotechnol. 37(9), 972 (2019).
Loeb, S., Carter, H. B., Berndt, S. I., Ricker, W. & Schaeffer, E. M. Complications after prostate biopsy: data from SEER-medicare. J. Urol. 186(5), 1830–1834 (2011).
Berger-Richardson, D. & Swallow, C. J. Needle tract seeding after percutaneous biopsy of sarcoma: risk/benefit considerations. Cancer 123(4), 560–567 (2017).
Brenner, D. J. Radiation risks potentially associated with low-dose CT screening of adult smokers for lung cancer. Radiology 231(2), 440–445 (2004).
Hendrick, R. E. Radiation doses and cancer risks from breast imaging studies. Radiology 257(1), 246–253 (2010).
Etzioni, R. et al. Early detection: the case for early detection. Nat. Rev. Cancer 3(4), 243 (2003).
Crowley, E., Di Nicolantonio, F., Loupakis, F. & Bardelli, A. Liquid biopsy: monitoring cancer-genetics in the blood. Nat. Rev. Clin. Oncol. 10(8), 472 (2013).
Zemmour, H. et al. Non-invasive detection of human cardiomyocyte death using methylation patterns of circulating DNA. Nat. Commun. 9(1), 1443 (2018).
Butler, T. M. et al. Exome sequencing of cell-free DNA from metastatic cancer patients identifies clinically actionable mutations distinct from primary disease. PLoS ONE 10(8), e0136407 (2015).
Newman, A. M. et al. An ultrasensitive method for quantitating circulating tumor DNA with broad patient coverage. Nat. Med. 20(5), 548 (2014).
Ngo, T. T. et al. Noninvasive blood tests for fetal development predict gestational age and preterm delivery. Science 360(6393), 1133–1136 (2018).
Arroyo, J. D. et al. Argonaute2 complexes carry a population of circulating microRNAs independent of vesicles in human plasma. Proc. Natl. Acad. Sci. 108(12), 5003–5008 (2011).
Williams, Z. et al. Comprehensive profiling of circulating microRNA via small RNA sequencing of cDNA libraries reveals biomarker potential and limitations. Proc. Natl. Acad. Sci. 110(11), 4255–4260 (2013).
Tsang, J. C. et al. Integrative single-cell and cell-free plasma RNA transcriptomics elucidates placental cellular dynamics. Proc. Natl. Acad. Sci. 114(37), E7786–E7795 (2017).
Koh, W. et al. Noninvasive in vivo monitoring of tissue-specific global gene expression in humans. Proc. Natl. Acad. Sci. 111(20), 7361–7366 (2014).
Lázaro-Ibáñez, E. et al. Different gDNA content in the subpopulations of prostate cancer extracellular vesicles: apoptotic bodies, microvesicles, and exosomes. Prostate 74(14), 1379–1390 (2014).
Scheer, F. A. et al. Impact of the human circadian system, exercise, and their interaction on cardiovascular function. Proc. Natl. Acad. Sci. 107(47), 20541–20546 (2010).
Lange, T., Dimitrov, S. & Born, J. Effects of sleep and circadian rhythm on the human immune system. Ann. N. Y. Acad. Sci. 1193(1), 48–59 (2010).
Vollmers, C. et al. Time of feeding and the intrinsic circadian clock drive rhythms in hepatic gene expression. Proc. Natl. Acad. Sci. 106(50), 21453–21458 (2009).
Buhr, E. D. & Takahashi, J. S. Molecular Components of the Mammalian Circadian Clock 3–27 (Springer, Circadian clocks, 2013).
Floris, I. et al. MiRNA analysis by quantitative PCR in preterm human breast milk reveals daily fluctuations of hsa-miR-16-5p. PLoS ONE 10(10), e0140488 (2015).
Heegaard, N. H. et al. Diurnal variations of human circulating cell-free micro-RNA. PLoS ONE 11(8), e0160577 (2016).
Hindson, B. J. et al. High-throughput droplet digital PCR system for absolute quantitation of DNA copy number. Anal. Chem. 83(22), 8604–8610 (2011).
Taylor, S. C., Laperriere, G. & Germain, H. Droplet digital PCR versus qPCR for gene expression analysis with low abundant targets: from variable nonsense to publication quality data. Sci. Rep. 7(1), 2409 (2017).
Olmedillas-López, S., García-Arranz, M. & García-Olmo, D. Current and emerging applications of droplet digital PCR in oncology. Mol. Diagn. Ther. 21(5), 493–510 (2017).
Devonshire, A. S. et al. Towards standardisation of cell-free DNA measurement in plasma: controls for extraction efficiency, fragment size bias and quantification. Anal. Bioanal. Chem. 406(26), 6499–6512 (2014).
Hothorn, T., Hornik, K., Van De Wiel, M. A. & Zeileis, A. A lego system for conditional inference. Am. Stat. 60(3), 257–263 (2006).
Trivedi, D. K. et al. Discovery of volatile biomarkers of Parkinson’s disease from sebum. ACS Cent. Sci. 5(4), 599–606 (2019).
van Ginkel, J. H. et al. Preanalytical blood sample workup for cell-free DNA analysis using droplet digital PCR for future molecular cancer diagnostics. Cancer Med. 6(10), 2297–2307 (2017).
Welsh, J. A., Holloway, J. A., Wilkinson, J. S. & Englyst, N. A. Extracellular vesicle flow cytometry analysis and standardization. Front. Cell Dev. Biol. 5, 78 (2017).
Max, K. E. et al. Human plasma and serum extracellular small RNA reference profiles and their clinical utility. Proc. Natl. Acad. Sci. 115(23), E5334–E5343 (2018).
Madsen, A. T., Hojbjerg, J. A., Sorensen, B. S. & Winther-Larsen, A. Day-to-day and within-day biological variation of cell-free DNA. EBioMedicine. 49, 284–290 (2019).
Zhong, X. Y. et al. Fluctuation of maternal and fetal free extracellular circulatory DNA in maternal plasma. Obstet. Gynecol. 96(6), 991–996 (2000).
Chen, M. et al. Utility of circulating cell-free rna analysis for the characterization of global transcriptome profiles of multiple myeloma patients. Cancers. 11(6), 887 (2019).
Li, Y. et al. Serum circulating human mRNA profiling and its utility for oral cancer detection. J Clin Oncol. 24(11), 1754–1760 (2006).
Dheda, K. et al. Validation of housekeeping genes for normalizing RNA expression in real-time PCR. Biotechniques 37(1), 112–119 (2004).
De Jonge, H. J. et al. Evidence based selection of housekeeping genes. PLoS ONE 2(9), e898 (2007).
Yang, Q. et al. Evaluation and validation of the suitable control genes for quantitative PCR studies in plasma DNA for non-invasive prenatal diagnosis. Int. J. Mol. Med. 34(6), 1681–1687 (2014).
Acknowledgements
We thank OCTRI for assistance with obtaining samples and the Knight Cancer Institute Biostatistics Shared Resource for statistics advisement.
Funding
This work was supported by funding from the Cancer Early Detection Advanced Research (CEDAR) center at Oregon Health & Science University’s Knight Cancer Institute. LN is supported by the National Institute of Child Health and Human Development (1K23 HD091369-01).
Author information
Authors and Affiliations
Contributions
Conceptualization: P.S., T.N., and K.J. Data Curation: J.W. Formal Analysis: J.W. and H.K. Funding Acquisition: P.S. and K.J. Investigation: J.W., H.K., K.J., L.N. and T.K. Methodology: P.S., T.N., J.W., H.K., and K.J. Project Administration: P.S. and K.J. Resources: P.S. and K.J. Software: J.W. Supervision: T.N. and P.S. Validation: J.W. and H.K. Visualization: J.W. Writing—Original Draft Preparation: J.W. and H.K. Writing – Review & Editing: all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wagner, J.T., Kim, H.J., Johnson-Camacho, K.C. et al. Diurnal stability of cell-free DNA and cell-free RNA in human plasma samples. Sci Rep 10, 16456 (2020). https://doi.org/10.1038/s41598-020-73350-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-73350-3
This article is cited by
-
Prediction of preeclampsia using maternal circulating mRNAs in early pregnancy
Archives of Gynecology and Obstetrics (2024)
-
Current challenges and best practices for cell-free long RNA biomarker discovery
Biomarker Research (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.