Abstract
Many chronic pain conditions show sex differences in their epidemiology. This could be attributed to sex-dependent differential expression of genes (DEGs) involved in nociceptive pathways, including sensory neurons. This study aimed to identify sex-dependent DEGs in estrous female versus male sensory neurons, which were prepared by using different approaches and ganglion types. RNA-seq on non-purified sensory neuronal preparations, such as whole dorsal root ganglion (DRG) and hindpaw tissues, revealed only a few sex-dependent DEGs. Sensory neuron purification increased numbers of sex-dependent DEGs. These DEG sets were substantially influenced by preparation approaches and ganglion types [DRG vs trigeminal ganglia (TG)]. Percoll-gradient enriched DRG and TG neuronal fractions produced distinct sex-dependent DEG groups. We next isolated a subset of sensory neurons by sorting DRG neurons back-labeled from paw and thigh muscle. These neurons have a unique sex-dependent DEG set, yet there is similarity in biological processes linked to these different groups of sex-dependent DEGs. Female-predominant DEGs in sensory neurons relate to inflammatory, synaptic transmission and extracellular matrix reorganization processes that could exacerbate neuro-inflammation severity, especially in TG. Male-selective DEGs were linked to oxidative phosphorylation and protein/molecule metabolism and production. Our findings catalog preparation-dependent sex differences in neuronal gene expressions in sensory ganglia.
Similar content being viewed by others
Introduction
Many inflammatory, neuropathic and idiopathic chronic pain conditions, such as migraine, fibromyalgia, temporomandibular disorder (TMD/TMJ), irritable bowel syndrome (IBS), rheumatoid arthritis, neuropathies, have greater prevalence and/or symptom severity in women as compared to men1,2,3,4. Additionally, clinical and preclinical data suggest that sex differences in pain prevalence and chronicity is more often encountered for conditions associated with the trigeminal ganglion (TG) system than those that affect the dorsal root ganglia (DRG)5,6,7,8. Pain symptoms in women with these chronic conditions can change in line with alterations in gonadal hormone concentrations, and some pain conditions decrease in frequency or intensity after menopause9,10,11,12. These facts and the plethora of clinical and rodent data suggest a critical role for gonadal hormone in regulation of nociceptive plasticity2,3.
Gonadal hormones mainly regulate neuronal plasticity via a classic genomic pathway13. The genomic pathway involves binding of gonadal hormones to their receptors, nuclear translocation of the gonadal hormone receptor and regulation of transcription either via gonadal hormone responsive elements, such as the estrogen response element14 or modulation of other transcription factors15. These points suggest that sex differences in gene expression could be detected in nociceptive pathways of mice and humans. For instance, sex differences in expression of genes involved in pain and inflammation, such as Ntrk1, Ccl2, Pdgfr, Traf3 and Vegfa were observed in human tibial nerve16. Neuropathic pain conditions are also associated with sex-dependent differential expression of inflammatory genes and transcription factors in human DRG tissues17.
Sex differences in gene expression can be identified in RNA sequencing (RNA-seq) experiments. A weakness of RNA-seq from whole tissue is that contributions of different cell types presented in tissue remain unsolved after analysis of RNA-seq data. Thus, when looking for sex differences in gene expression in sensory neurons, previously published conclusions often relied on RNA-seq data generated from whole ganglia. This approach is limited, since whole DRG or TG has many other cell types, and these non-neuronal ganglion cells are numerically predominant compare to sensory neurons. Therefore, the aims of the present study are (1) to examine sex differences in gene expression in enriched sensory neuron populations from both the TG and DRG of mice; and (2) to compare these results with expression in whole ganglia and hind paw of naïve mice. Recent publications indicate that sensory neuronal phenotypes could be influenced by the tissue target of innervation18. Accordingly, we also sought to define how innervation target impacts sex differences in gene expression. Overall, our work elucidates DEGs in sensory neurons of the DRG and TG in male and female mice and suggests potential pathways that may play differential roles in nociception and pain chronicity between sexes.
Results
RNA-Seq of female and male whole DRG and hind paw tissues
Since DRG neurons can interact with cells within the ganglion as well as cells in target tissues like skin to promote pain plasticity19,20, we have first evaluated potential sex differences in transcriptional profiles of whole DRG and hind paw tissues in females (all in the estrous phase) versus males (n = 3 per group/sex)21. For further analysis of all presented in this manuscript RNA-seq data, we deselected X and Y chromosome-linked genes, such as inactive X specific transcript (Xist), ubiquitously transcribed tetratricopeptide repeat gene (Uty), lysine (K)-specific demethylase 5D (Kdm5d) and eukaryotic translation initiation factor 2, subunit 3, structural gene Y-linked (Eif2s3y). Considering this deselection, transcriptomic analysis showed that there were as low as 14 sex-dependent differentially expressing genes (DEGs; RPKM > 1; FC > 2 and Pval < 0.05) in DRG tissues (Fig. 1A, Supplementary excel file). The only DEG in male was Penk (proenkephalin), an endogenous opioid, which has anti-nociceptive actions by producing opioid peptides such as enkephalin. Gene clustering according to biological processes using the PANTHER software assigned 5 female-selective DEGs to the cell population homeostasis process (Fig. 1B). This small numbers of female-predominant DRG DEGs were Lilrb3 (leukocyte immunoglobulin-like receptor subfamily B member 3); Rhag (aka CD241; rh-associated glycoprotein); IL7r (aka CD127; interleukin-7 receptor), which is critical for V(D)J recombination; Alas2 (delta-aminolevulinate synthase 2) genes, which participate in tissue repair22,23 and Rag1 (recombination activating gene 1), which is also vital for T-cell receptor V(D)J recombination. Strengthening selection criteria for sex-dependent DRG DEGs to RPKM > 1; FC > 2 and Padj < 0.05 left a solitary female-predominant DEG—myeloperoxidase (Mpo).
RNA-Seq of female and male whole DRG and hindpaw tissues. Summary plots of DEGs (RPKM > 1; FC > 2; Pval < 0.05) in female and male DRG (A) and hindpaw tissues (C); (n = 3). Data from mean values of DEGs are shown. Blue dots are male DEGs, red dots are female DEGs. (B) Female-predominant DRG DEGs linked to the cell population homeostasis process. Numbers of DEGs involved in female (D) and male selective (E) biological processes in hindpaws.
The number of sex-dependent DEGs (RPKM > 1; FC > 2 and Pval < 0.05) in the hind paw was higher at 142 (Fig. 1C; Supplementary excel file). Female-predominant DEGs in the hind paw covered several biological processes, which were mainly immune cell function related (Fig. 1D). A majority of these female DEGs encode pro inflammatory proteins (Fig. 2A), which are known to sensitize nociceptive pathways. These DEGs include IL-6 and IL-1β cytokines, which are key inflammatory signals in many pain conditions such as fibromyalgia, rheumatoid arthritis, irritable bowel syndrome, etc.24; Cxcl1, Cxcl2, Cxcl5, Ccl7 and Ccl8 chemokines, which are known to be involved in nerve injury and chemotherapy-induced neuropathic pain25,26,27. Unlike pro-inflammatory DEGs participating in pain promotion, among female-predominant DEGs was TNF-α regulated Tnfaip3 (tumor necrosis factor, alpha-induced protein 3), which can inhibit NF-kappa B activation and is critical for limiting inflammation28. Female DEGs in hind paws also contained lipid metabolism related genes—Alox12e and Olah, which increases arachidonic and linoleic acid lipid metabolites that are also known to activate the nociceptive pathway29,30. The male-specific DEGs in the hind paw included mostly genes modulating extracellular matrix and keratinocyte functions via increased expression of many members of keratin (Krt) and keratin associated protein (Krtap) family members (Figs. 1E, 2B). The contribution of these genes to pain pathways has not been studied. Strengthening selection criteria to RPKM > 1; FC > 2 and Padj < 0.05 reduced numbers of sex-dependent hind paw DEGs to 18 female and 33 male-selective DEGs. Importantly, these female DEGs are still linked to immune processes, and male DEGs are associated with keratinocyte functions.
RNA-Seq of female and male DRG and TG sensory neuronal enriched fractions
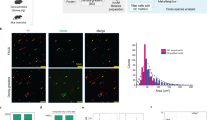
Previous reports suggest that sensory neuronal mechanisms, which are manifested at either peripheral or central terminals, could be one of the primary contributors to sexual dimorphisms in chronic pain mechanisms3. Our flow cytometry measurements of DRG single-cell suspensions from CGRPcre-ER/+/Rosa26LSL-tDTomato/+ reporter mice showed that only 2 ± 1.3% (n = 3) of total viable cells were CGRP-cre+. Since approximately 40–50% of DRG neurons are peptidergic (i.e. CGRP)31, we can estimate that the overall percentage of sensory neurons among all DRG cells is ~ 3–5%. Although, neurons contain more total RNA than non-neuronal DRG cells due to their larger size, RNA-seq data from whole ganglia likely mainly represent the transcriptome of non-neuronal DRG cells, and at the very least under-represents neurons. Hence, to gain better insight into transcriptomic sex differences at the sensory neuron level, we purified DRG and TG sensory neuronal enriched fractions using a Percoll gradient-based approach. Flow cytometry data with a purified sensory neuronal fraction from CGRPcre-ER/+/Rosa26LSL-tDTomato/+ reporter mouse DRG showed that 35 ± 6.1% (n = 3) of viable cells were CGRP-cre+. This indicates that sensory neurons constitute 85–95% of a purified neuronal fraction, since ≈ 40–50% of all DRG sensory neurons are CGRP+ as measured by immunohistochemistry31.
Our transcriptomic analysis revealed 963 sex-dependent DEGs (RPKM > 1; FC > 2 and Pval < 0.05) in TG (Fig. 3A, Supplementary excel file), and 540 genes in DRG sensory neurons (Fig. 3B, Supplementary excel file). This DEG numbers were far greater than we observed using a whole DRG approach for RNA-seq (14 DEGs). This suggests that neuronal enrichment increases the ability to discern sensory neuronal-based sex differences.
RNA-Seq of female and male DRG and TG sensory neurons. Summary plots of DEGs (RPKM > 1; FC > 2; Pval < 0.05) in female (red dots) and male (blue dots) TG (A) and DRG (B) neurons; n = 3. Data from mean values of DEGs are shown. (C) Venn diagram for RPKM > 1, FC > 2 and Pval < 0.05 female-predominantly expressing TG and DRG sensory neuronal genes. All female DEGs are divided: 555 (76.2%)—only in TG; 135 (18.5%)—only in DRG; and 38 (5.2%)—in both TG and DRG sensory neurons. (D) Venn diagram for RPKM > 1, FC > 2 and Pval < 0.05 male-predominantly expressing TG and DRG sensory neuronal genes. All male DEGs are divided: 292 (44.2%)—only in TG; 291 (44%)—only in DRG; and 78 (11.8%)—in both TG and DRG sensory neurons.
Female-selective DEGs were greater in number in TG compared to DRG, 594 to 172, respectively. Moreover, only 38 female-selective DEGs were common for DRG and TG (Fig. 3C). In contrast, DRG and TG neurons contained approximately equal numbers of male-selective DEGs, 368 and 369, respectively. Although male-selective DEGs were almost equal in DRG versus TG, only 79 of these DEGs with male-predominant expression were differentially expressed in both TG and DRG neurons (Fig. 3D). Consequently, there were many unique DEGs for both DRG and TG (Fig. 3D). Highly expressed and/or regulated female- and male-selective DEGs in TG and DRG neurons are presented in Table 1 (see also Supplementary excel files). These data suggest that a surprising number of sex differences are not consistent between ganglia, a finding that is consistent with other measures indicating important differences between the TG and DRG32,33.
Female-selective DEGs in DRG and TG sensory neuronal enriched fractions
We used the PANTHER over-representation test (https://www.pantherdb.org/) to identify distinct biological process clusters for female-selective DEGs from Percoll enriched DRG and TG sensory neuronal fractions (Figs. 4A, 4B). While the TG contains as much as threefold more female-selective DEGs (Fig. 3C), our analysis showed that biological processes linked to female-selective DEGs were similar in DRG vs TG. However, two key differences were noted. First, the TG contains a separate large female-selective DEG cluster involved in nervous system development, including synapse pruning, synaptic vesicle docking, neuron remodeling, neurogenesis, dendritic spine morphogens and regulation of neuronal apoptotic processes (Fig. 4C). Second, similar biological processes, such as extracellular matrix organization, angiogenesis and vascular development, cell adhesion, lipid metabolism, and regulation in epithelial cell, endothelial cell, bone and fibroblasts, include different sets of female-selective DEGs in TG compared to DRG neurons. We also performed a different DEG analysis using PANTHER software, which groups genes according to their functions.
Female-selective DEGs in DRG and TG sensory neurons. Biological processes for female-selective DEGs (RPKM > 1, FC > 2 and Pval < 0.05) in DRG (A) and TG (B) neurons. (C) A set of female-selective DEGs in TG sensory neurons that are involved in the nervous system development process. (D) Growth factor and their receptor female-selective DEGs in TG and DRG neurons. (E) Transcription machinery female-selective DEGs in TG and DRG sensory neurons. Genes belonging to RNA binding factors are marked as RNA-bind (green); DNA binding factors are DNA-bind (khaki); transcription factor binding proteins are TrF-bind (red); and transcription factors are TrF (blue). Yellow boxes on the panels D and E indicate female-selective DEGs, which were found in both female TG and DRG sensory neurons. Location of TG and DRG neuron female-selective DEGs are marked with vertical lines on the panels (D) and (E).
The first cluster contained growth factors and their receptor genes (Fig. 4D). These genes are involved in nervous system development, epithelial cell proliferation, bone development, reposes to lipid, immune system processes, epidermis development, cell–cell adhesion, cell-extracellular matrix adhesion, angiogenesis, etc. (www.uniprot.org). Female DEGs of this cluster were found primarily in TG sensory neurons (Fig. 4D). Among noteworthy growth factor DEGs in TG neurons were Notch, Pdgf (platelet derived growth factor), Fgfr (fibroblast growth factor receptor), Rara and Rxra (retinoic acid receptors) and Tgf (transforming growth factor) families (Fig. 4D). This functional cluster of DEGs are also found in female DRG neurons (Fig. 4D). Thus, Igf2 (insulin-like growth factor 2) and Egfr (epidermal growth factor receptor) are deferentially expressed only in female DRG neurons (Fig. 4D). Additionally, some DEGs belonging to this cluster could be found in both TG and DRG neurons (marked by yellow boxes; Fig. 4D). For instance, platelet derived growth factor, beta polypeptide (Pdgfb) and transforming growth factor, beta induced (Tgfbi) were enriched in both female TG and DRG neurons.
The second cluster contained female-selective DEGs encoding proteins for the transcriptional and translational machinery (Fig. 4E). These DEGs participate in several biological processes, such as dendritic spine morphogenesis, synapse assembly, morphogenesis of epithelium, regulation of osteoclast proliferation, responses to inflammation, angiogenesis, etc. (www.uniprot.org). Female sensory neurons had DEGs encoding RNA-binding proteins, such as Nr family (nuclear receptors); DNA-binding proteins, such as E4f1 (E4F transcription factor 1); and transcription factor-binding co-factors, such as the Med family, Crtc1 (CREB regulated transcription coactivator 1) and Tada3 (transcriptional adaptor 3). Some of these transcription factors, notably Stat5 (signal transducer and activator of transcription 5), Socs3 (suppressor of cytokine signaling 3), Sox 8 (sex determining region Y-box 8) and Hoxb4 (homeobox beta4), have been implicated in regulation of pain chronicity (Fig. 4E)34,35. It is also possible that higher expression of these DEGs in female sensory neurons could play a protective role during pain conditions35,36,37.
The third cluster of female-selective DEGs were genes involved in immune system processes (www.uniprot.org; Fig. 5A,B). Like growth factor genes, these DEGs were more prominent in TG than DRG neurons. DEGs found enriched in female TG included members of the complement system (C1q and C3), which is part of the innate immune system and is involved in nociception regulation38. Both TG and DRG neurons contained female-selective DEGs belonging to the chemokine system (Cx3cl1, Cx3cr1, Ccl3, Cxcl12 and Cmklr1), which regulate monocyte, macrophage and natural killer (NK) cell migration/trafficking39 and neuropathic pain40,41,42; the cytokine system (IL6ra and IL33); the integrin system (Itga8, Itga3 and Itgam); the tumor necrosis factor super family members (Tnfrsf1a, Tnfrsf1b, Ltbr, Ngfr and Ntrk1) and their effectors (Nfkb2, Nfkbid, Tifab and Icam1), many of which are key regulators of pain chronicity24,38,42,43,44; and cluster differential factor (CD74 and CD93) (Fig. 5A,B). Some of these DEGs, marked with yellow boxes, were found in both TG and DRG neurons (Fig. 5A).
Female-selective DEG functional clusters in DRG and TG sensory neurons. Inflammatory DEGs in female TG (A) and DRG (B) sensory neurons. Genes belonging to the complement family are marked as Comp; chemokines and their receptors are Chem; the integrins are Itg; and the tumor necrosis factor super family genes are TNFSF. Yellow boxes on the panel A indicate DEGs in both female TG and DRG sensory neurons. (C) Channel genes predominantly expressing in female TG sensory neurons. Structural protein genes expressing higher in female TG (B) than in DRG (C) sensory neurons. Structural protein DEGs belonging to extracellular matrix are marked as Extra matrix; synaptic machinery is Synp; cell adhesion is Cell-adh; and cytoskeleton are Cytosk. Yellow boxes on the panel (D) indicate DEGs identified in both male TG and DRG sensory neurons.
The fourth functional cluster of female-selective sensory neuronal DEGs consists of channel, metabotropic receptor and transmembrane protein genes (Fig. 5C). They were exclusively enriched in female TG, but not female DRG neurons, and play key roles in nervous system function/development. Almost all these genes, including Hcn (hyperpolarization-activated, cyclic nucleotide-gated K+) and Kcn families (potassium voltage-gated channels), as well as Orai1 (ORAI calcium release-activated calcium modulator 1) and Trpv4 (transient receptor potential cation channel, subfamily V, member 4), are known to modulate nociceptor excitability, transmission of nociceptive information and regulation of pain chronicity45,46,47.
The fifth and final cluster of female-predominantly expressing genes in TG and to a lesser extent in DRG sensory neurons were structural proteins (Fig. 5D,E). These DEGs represent extracellular matrix proteins, such as Col (collagen), Lmn (lamin), Lam (laminin), Thbs (thrombospondin) and Fbn (fibrillin) families; synaptic machinery proteins, such as Cplx2 (complexin 2), Nrxn2 (neurexin II) and Nlgn2 (neuroligin 2); and a few cell adhesion and cytoskeletal proteins, including Smtn (smoothelin), Stab1 (stabilin 1), Svil (supervillin), Vtn (vitronectin) and Cdh5 (cadherin 5).
Male-selective DEGs in DRG and TG sensory neuronal enriched fractions
Although there were many unique male-selective DEGs for TG or DRG neurons (Fig. 3D), the PANTHER over-representation test (https://www.pantherdb.org/) grouped these DEGs into similar biological process clusters for TG and DRG neurons (Fig. 6A, B). These biological processes, including oxidation–reduction, cellular respiration, macromolecule and protein metabolism, translation and gene expression, were represented by a set of distinct male-selective DEGs in TG versus DRG neurons. Similar to how we organized female-selective DEGs, male-selective DEGs were clustered by enriched functional families using the PANTHER classification system (https://www.pantherdb.org/). This analysis highlighted four main DEG clusters in male TG and DRG neurons.
Male-selective DEGs in DRG and TG sensory neurons. Biological processes for male-selective DEGs (RPKM > 1, FC > 2 and Pval < 0.05) in DRG (A) and TG (B) neurons. (C) Proteasome subunit encoding DEGs are male-enriched in TG sensory neurons. (D) Male-predominant mitochondrial genes in TG sensory neurons. (E) Male-predominant mitochondrial genes in DRG sensory neurons. Yellow boxes on the panels C and D indicate DEGs in both male TG and DRG sensory neurons. Genes belonging to oxidative phosphorylation complexes I–V are marked as CI–CV; mitochondrial ribosomal proteins are MRib; the anti-oxidative glutathione pathway are Glut; and the cytochrome pathway are Cyto.
The first cluster consisted of DEGs encoding proteasome subunits and assembly chaperones (Fig. 6C). These DEGs were found in both TG and DRG neurons (yellow boxes; Fig. 6C); and are closely involved in a vast array of biological functions related to protein metabolism and complex assembly (www.uniprot.org; Fig. 6A,B). Proteasomes could play key roles in pain processing, since they contribute to recovery from tissue injury48, from nerve damage49 and resolution of inflammation50.
The second cluster consisted of mitochondrial genes, which had higher levels of expression both in the male TG (Fig. 6D) and DRG sensory neurons (Fig. 6E). These mitochondrial genes can be divided into several sub-groups according to their involvement in different biological processes, such as oxidation–reduction, electron transport chain, cellular respiration, and co-factor and small molecule metabolism (www.uniprot.org; Fig. 6D,E). One of the DEG sub-groups in male TG and DRG sensory neurons represents the oxidative phosphorylation proteins, which are also critical for cellular respiration and recovery from tissue and nerve injury51,52. Genes belonging to all five main complexes of the oxidative phosphorylation pathway—reduced nicotinamide adenine dinucleotide (NADH) dehydrogenase (complex I), succinate dehydrogenase (complex II), cytochrome c reductase (complex III), cytochrome c oxidase (complex IV) and adenosine triphosphate (ATP) synthase (complex V)—had male-predominant expression in both TG and DRG sensory neurons (Fig. 6D,E). Another male sensory neuronal DEG sub-group contained coenzymes and cofactors, such as Cycs (cytochrome c, somatic), Cyb5r4 (cytochrome b5 reductase 4), Cmc1 (cyclooxygenase (COX) assembly mitochondrial protein 1), Coq3 (coenzyme Q3 methyltransferase), Coq10a (coenzyme Q10A), Moab (monoamine oxidase B), Sucl family (succinate-CoA ligase) and the Acot family (acyl-coenzyme A (CoA) thioesterase), supporting the oxidative phosphorylation pathway. Genes involved in protection from oxidative damage, Gpx (glutathione peroxidase), Glrx (glutaredoxin) and the Gst families (glutathione S-transferase) were also higher in male sensory neurons (Fig. 6D,E). Finally, mitochondrial protein anabolism genes also showed higher expression in males including Mterfd (Mitochondrial Transcription Termination Factors), Rpp30 (ribonuclease P/MRP 30 subunit), Tfb2m (transcription factor B2, mitochondrial) Mtg1 (mitochondrial ribosome-associated GTPase 1), and the Mrpl and Mrps families.
The third and fourth clusters of DEGs in male sensory neurons related to non-mitochondrial transcription and translation machinery (Fig. 7A,B). Interestingly, these male-selective DEGs did not overlap with female-selective DEGs representing transcription machinery (Fig. 4E). Transcription machinery genes with higher expression in males could be separated into several sub-groups: genes of RNA-binding proteins, such as Snhg6 (small nucleolar RNA host gene 6) and Snrpb2 (U2 small nuclear ribonucleoprotein B); DNA-binding proteins, such as Polr2d (polymerase (RNA) II (DNA directed) polypeptide D), Tcea2 (transcription elongation factor A (SII), 2) and Orc (origin recognition complex); transcription factor-binding co-factors, such as C1d (C1D nuclear receptor co-repressor), Tifa (TRAF-interacting protein with forkhead-associated domain) and Med (mediator complex subunit); histone-binding proteins, such as Hat1 (histone aminotransferase 1), Asf1a (anti-silencing function 1A histone chaperone) and L3mbtl4 (L3MBTL4 histone methyl-lysine binding protein); ligases and helicases, such as Rtcd1 (RNA 3′-Terminal Phosphate Cyclase) and Nae1 (NEDD8 activating enzyme E1 subunit 1); and, finally, general transcription factors, such as the Zfp family (zinc finger proteins), Gtf2b (general transcription factor IIB) and the Taf family (TATA-box binding protein associated factors). In contrast to female DEGs in this class of genes (Fig. 4E), male DEGs include histone-binding proteins and ligases/helicases (Fig. 7A,B). Male-enriched mRNAs in the translation machinery family were mainly composed of genes encoding Eif family (eukaryotic translation initiation factors), Naa20 (N(alpha)-acetyltransferase 20, NatB catalytic subunit), and Rpl and Rps families (ribosomal proteins).
Male-selective DEG functional clusters in DRG and TG sensory neurons. Male-predominant transcription and translation machinery genes in TG (A) and DRG (B) sensory neurons. Yellow boxes on the panel A indicate DEGs in both male TG and DRG sensory neurons. Genes belonging to RNA binding factors are marked as RNA-bind; DNA binding factors are DNA-bind; transcription factor binding proteins are TrF-bind; transcription factors are TrF; histone binding proteins are His-bind; ligases and helicases are Ligase Helicase or Lig-Hel; and translation machinery proteins are Translation.
RNA-Seq of female and male sensory neurons innervating hindpaw skin and thigh muscle
Recent reports indicate that peripheral tissue innervation influences sensory neuron phenotype and potentially sensory neuron plasticity18. We sought to clarify whether the transcriptome of sensory neurons innervating hindpaw skin or thigh muscle in males or females would be distinct from data generated on Percoll enriched fraction of DRG neurons. To do this, we injected WGA-488 simultaneously into left and right hindpaws as well as left and right thighs of mice. The rational for combine injection into skin and muscle was to increase a number of WGA-488+ neurons. These injections were performed for females and males (n = 4 each group). Two days post-injection L3-L5 DRG were dissected and generated single-cell suspensions were generated. A mixture of skin and thigh innervating neurons were isolated by double sorting WGA-488+ neurons using a 130 μm nozzle. The use of the 130 μm nozzle on a FACS sorter is essential to avoid losing medium-to-large (> 35 μm) neurons during the sorting procedure. Each group (female or male) had 4 samples and 6 DRG (L3-L5 levels from left and right sides) within sample; and this sorting procedure generated 1,500–2,500 WGA-488+ neurons for each sample. cDNA library from extracted RNA was synthesized using PCR-based SmartSeq II protocol.
Our transcriptomic analysis showed that there are 88 female-selective DEGs (RPKM > 1; FC > 2 and Pval < 0.05) in WGA+ neurons. Six DEGs, Jub (LIM domain-containing protein ajuba), Cd74 (CD74 antigen), Abi3bp (ABI gene family, member 3 (NESH) binding protein), Bgn (biglycan), Mxra8 (matrix-remodeling associated 8) and Islr (immunoglobulin superfamily containing leucine-rich repeat), are identified in WGA+ and Percoll purified DRG neurons (Fig. 8A). Moreover, Cd74 and Islr were female-selective DEGs in Percoll purified TG neurons as well (Table 1). Cd74 and Jub play a critical role in pro-inflammatory processes (www.uniprot.org). Abi3bp, Islr, Bgn and Mxra8 contribute to cell–cell adhesion and extracellular matrix organization processes (www.uniprot.org). The PANTHER over-representation test (https://www.pantherdb.org/) groups these 88 female-selective DEGs into antigen processing and nervous system development biological processes. Analysis of 285 female-predominant DEGs selected according to RPKM > 1; FC > 1.5 and Pval < 0.05 revealed additional biological processes, such as gliogenesis, cell adhesion, epithelium and vasculature development (Fig. 8B).
Sex-dependent DEGs in DRG neurons innervating hindpaw skin and thing muscle. Venn diagram for RPKM > 1, FC > 2 and Pval < 0.05 female (A) and male-predominantly (C) expressing WGA-488-labeled DRG neurons innervating hindpaw skin and thigh (WGA+) and Percoll purified DRG neurons (Percoll). All female DEGs are divided: 82 (32.2%)—only in WGA+; 167 (65.5%)—only in Percoll purified; and 6 (2.4%)—in both sensory neuron preparations. All male DEGs are divided: 22 (5.6%)—only in WGA+; 367 (93.9%)—only in Percoll purified; and 2 (0.5%)—in both sensory neuron preparations. Biological processes for female (B) and male-selective (D) DEGs (RPKM > 1, FC > 2 and Pval < 0.05) in DRG neurons innervating hindpaw skin and thing muscles.
RNA-seq analysis indicates that only 24 DEGs (RPKM > 1; FC > 2 and Pval < 0.05) in WGA+ neurons could be classified as male-selective. Among these 24 male-predominant DEGs, Pkib (protein kinase inhibitor beta, cAMP dependent, testis specific) and Arxes2 (adipocyte-related X-chromosome expressed sequence 2) were common for both WGA+ and Percoll purified DRG neurons (Fig. 8C). Pkib is a potent competitive inhibitor of cAMP-dependent protein kinase activity (www.uniprot.org), and Pkib up-regulation could potentially lead to anti-pain effect in inflammatory pain conditions. However, although Pkib is expressed in DRG neurons at high levels53, there is no literature its involvement in pain. Arxes2 is also highly expressed in DRG neurons53, but its function in neurons is unknown. The overlap between male-selective DEGs in WGA+ DRG neurons and Percoll purified TG neurons was wider than between DEGs in WGA+ DRG neurons and Percoll purified DRG neurons; and includes Ptbp2 (polypyrimidine tract binding protein 2), Dcun1d1 (DCN1, defective in cullin neddylation 1, domain containing 1), Pcdhb20 (protocadherin beta 20), Pkib, Tifa (TRAF-interacting protein with forkhead-associated domain), Stk32a (serine/threonine kinase 32A), Lipo1 (lipase, member O1) and Serpina3i (serine (or cysteine) peptidase inhibitor, clade A, member 3I). A low number of male-selective DEGs (24) was not enough for linking them to statistically significant biological process using the PANTHER over-representation test (https://www.pantherdb.org/). To overcome this, we decreased strength of selection criteria to RPKM > 1; FC > 1.5 and Pval < 0.05. This revealed 122 male-selective DEGs that can be linked to biological processes (Fig. 8D). Two biological processes – reactive oxygen and protein metabolism—were common for RNA-seq analysis of DEGs identified in Percoll purified neurons and WGA+ neurons (Figs. 6A,B, 8D).
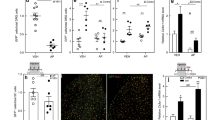
Finally, we used quantitative RT-PCR for validation of a set of DEGs that have been identified in neurons prepared by all procedures: Percoll from TG, Percoll from DRG as well as WGA-labeled DRG neurons. Accordingly, we have selected Cd74, Islr, Bgn, Mxra8, Pkib and Arxes2. RNA was prepared from female and male TG neurons enriched by Percoll. Another RNA preparation was from DRG neurons labeled by back-tracing WGA-488 from both hindpaws and thigh muscles. Figure 9A shows that Cd74, Islr, Bgn and Mxra8 have female-predominant expression, while Pkib and Arxes2 are higher expression levels in male sensory neurons in both Percoll enriched TG neurons and WGA-488+ DRG neurons.
Quantitative RT-PCR validation of RNA-seq identified sex-dependent DEGs in sensory neurons. (A) Quantitative RT-PCR for 6 sex-dependent sensory neuronal DEGs. Female (F DRG) and male (M DRG) DRG sensory neurons are identified by back-tracing from hindpaw and thigh. Female (F TG) and male (M TG) TG sensory neurons are purified using Percoll gradient. Statistic is un-paired t-test (NS p > 0.05; *p < 0.05; **p < 0.01; # p < 0.0001; n = 4). Name of DEGs are indicated. Schematic of female (B) and male (C) sensory neuronal DEGs clusters that could explain difference in pain promoters in female versus male mice.
Discussion
There is now a strong body of clinical and basic science studies demonstrating sex differences in the mechanisms contributing to the development and/or maintenance of pain in males and females. There is epidemiological evidence that males and females may be prone to different types of chronic pain disorders1,2,3,4. Sex differences in gene expression in nociceptive pathways, including sensory neurons, of humans and animals could potentially elucidate mechanistic reasons for why these differences exist. Accordingly, we aimed to identify sex differences in gene expression in sensory neurons. These differences could also be influenced by several factors, aside from biological sex. First, DRG and TG have very different physiological functions and gene expression profiles33. Second, the preparation method of sensory neurons can impact the RNA-seq outcome. Third, the innervated tissue can contribute to sensory neuronal phenotype, or different tissues can be innervated with distinct subsets of sensory neurons18. We confirm that only a small proportion of cells of DRG are sensory neurons, and the remaining cells are non-neuronal. Hence, RNA-seq data analysis of whole DRG (or TG) will reflect a strong influence of non-neuronal cells on the transcriptome. Analysis of our data showed that hindpaw and especially whole DRG of naïve mice have relatively few sex differences at the transcriptomic level (Figs. 1, 2). Consequently, we examined transcriptomic differences on isolated and purified TG and DRG sensory neuronal fractions from naïve females (estrous phase) and males where we found prominent sex differences that may have important implications for understanding how pain mechanisms differ in men and women (Figs. 3–7). Furthermore, we isolated female and male mouse sensory neurons innervating hindpaw skin and thigh muscle; and used RNA from those cells for RNA-seq and analysis (Figs. 8, 9). These experiments showed that certain sets of sex-dependent DEGs are influenced by tissue innervation. There could be two explanations for this phenomenon: first, tissue innervation could influence sensory neuron transcriptomics18; or second, the neuronal subsets innervating different tissues could be distinct to begin with and therefore, are guided to different targets. The latter notion is supported by single cell sequencing of back labelled neurons from the colon54.
Groups of female-predominant DEGs generated from Percoll purified DRG and TG neurons, and skin or thigh innervating L3–L5 DRG neurons showed low overlap. The highest numbers (593) of such DEGs (RPKM > 1; FC > 2; Pval < 0.05) were identified in Percoll purified TG neurons, and the lowest (88) were in DRG neurons innervating skin and thigh (Figs. 3C,8A). Nevertheless, there is substantial overlap in biological processes linked to these DEGs (Figs. 4A,B, 8B). We observed broadly higher expression of DEGs involved in immune processes in Percoll purified female DRG and TG neurons as well as DRG neurons innervating the skin and thigh muscle. These female-selective DEGs showed preferential expression in female DRG and TG, with notably larger differences in the TG neurons. In this cluster, the complement system38; certain chemokines regulating monocyte, macrophage and NK cell migration/trafficking40,41,42; IL-1, IL-6 and IL-33 cytokines; and the tumor necrosis factor super family members, such as Tnfrsf1a and Ntrk1 and their effector Nfkb224,38,42,43 are well documented contributors to nociception and pain development (Fig. 9B). One of the immune DEGs—CD74—was revealed in every sensory neuronal preparation (Fig. 9A, Table 1). CD74 is responsible for transport of MHC class II proteins55, which promote inflammatory and neuropathic pain56.
Other biological processes linked to female-selective sensory neuronal DEGs identified from different preparations are nervous system, vasculature and epithelium development (Figs. 4A,B, 8B). These DEG clusters contain many genes encoding growth factors and their receptors (Mxra8, Arxes2, Notch3, etc.), cell–cell and cell-extracellular matrix adhesion (Islr, Bgn, etc.), and channels and their modulators (Pkib, etc.; Figs. 4, 5, 9A). Growth factors and their receptors are modulators of nociceptive pathways and pain development. Pdgf regulates inflammatory pain57. Fgf and Tgfb signaling play a role in pain development during osteoarthritis, but they may be protective rather than pain promoting58. The Egfr signaling pathway has been implicated in nociception and pain based on human genetics as well as preclinical models where EGFR ligands promote pain via the EGFR and the mTOR pathway59. Notch signaling is involved in the development of pain via communication between sensory neurons and non-neuronal cells60. Channels and their modulators are likewise involved in regulation of nociceptive pathways. The Hcn family which plays a key role in underlying spontaneous activity and mechanical hypersensitivity after nerve injury61. Antagonism of Hcn with ZD7288 dose-dependently reduces mechanical hypersensitivity in a model of cervical radiculopathy pain61, and this antagonist is also effective for inflammatory pain62. Kcnq channels encode K+ channels that represent the M-current that regulates nociceptor signaling, especially at peripheral terminals46,63, and also plays an important role in generation of spontaneous activity after nerve injury64. The M current is also critical in mediating hyperalgesic actions of many Gq-protein receptor activators such as bradykinin65 and protease activated receptors66. Orai1 as a key component of store-operated calcium channels that regulate nociception via modulation of neuronal excitability and A-type potassium channels67. Delayed rectified potassium voltage-gated channels, Kcnc2 and Kcng2, play key roles in action potential shaping and consequently in nociception transmission and pain plasticity45. Kcnk18, a TWIK-related two-pore potassium channel is involved in regulating the resting membrane potential68,69 and has been linked to migraine with aura pathogenesis70,71. Finally, we identified higher expression of Trpv4 in female TG sensory neuronal fractions and this ion channel plays an important role in pain sensitization that has been thoroughly studied47,72. Collectively these female-selective DEGs in sensory neurons obtained from a variety of preparations suggest a potential role of enhanced interactions between inflammatory cells and sensory neurons via specific signaling pathways that are involved in the development of chronic pain (Fig. 9B). An intriguing hypothesis that emerges from these findings is the suggestion of DEGs in female sensory neurons that could shift the balance between protective neuro-immunity and neuro-immunopathology that would be predicted to promote the transition of acute to chronic pain73,74,75.
As for genes with female-predominant expression, sets of male-selective DEGs depend on sensory neuron purification approaches and selection of ganglia. The highest numbers (369–370) of such DEGs (RPKM > 1; FC > 2; Pval < 0.05) were identified in Percoll purified DRG and TG neurons, and the lowest (24) were in DRG neurons innervating skin and thigh (Figs. 3D, 8C). Again, like for female-selective DEGs, there was substantial overlap in biological processes linked to sets of male DEGs (Figs. 6A,B, 8D). Two main biological processes related to male-selective DEGs are reactive oxygen species and protein metabolism, including proteasomal machinery and mitochondrial protein production (Figs. 6A,B, 8D). Protein metabolism is involved in recovery from nerve and tissue injuries76. An example of the important role of this pathway in neuropathy and neuropathic pain is inhibition of the proteasome with the chemotherapy agent bortezomib which causes distal axonopathy49, nerve degeneration49 and painful neuropathy77,78. In contrast, activation of the proteasome can lead to effective removal of misfolded proteins and changes in transcriptomic profiles79, and contributes to recovery from tissue injury48, nerve damage49 and inflammation50. The important roles of mitochondrial proteins and reactive oxygen species (ROS) metabolism, especially oxidative phosphorylation, in nerve and tissue degeneration/regeneration is highlighted by a robust literature of hundreds of papers80,81. Enhancing the mitochondrial oxidative phosphorylation process can be an effective approach for recovery from nerve damage and degeneration82,83 as well as painful neuropathies84. Excessive ROS is a common feature for many pain conditions including a variety of neuropathy conditions76. Additionally, some male-selective DEGs from Percoll purified sensory neurons could be linked to translational machinery processes (Figs. 6A,B), while DEGs from neurons innervating skin and muscle can cover cell–cell communication and peptide secretion processes (Fig. 8D). The translation machinery is one of key players in regulation of nociception pathways and development of pain conditions85,86.
Overall, our approaches identify a number of candidate biological processes that are associated with female-selective sensory neuronal DEGs, in particular in the TG, (Fig. 9B) and male-selective DEGs (Fig. 9C). These genes include very prominent players in nociception that could be involved in differential occurrences of pain disorders in men and women (Fig. 9B). In contrast, DEGs predominantly expressed in male sensory neurons may protect male sensory neurons from some of the consequences of nerve injury that are known to promote neuropathic pain (Fig. 9C). Based on our findings, we favor the hypothesis that sex differences in nociception may be partially based on DEGs in female and male sensory neurons. Our results advance our understanding of sex dimorphisms in nociceptive pathways and provide an informatic basis for further studies on regulation of sex-dependent gene expression plasticity in sensory neurons regulated by injury and/or by gonadal hormones44,87,88.
Materials and methods
Mouse lines and tissues
All animal experiments conformed to APS's Guiding Principles in the Care and Use of Vertebrate Animals in Research and Training, and to protocols approved by the University Texas Health Science Center at San Antonio (UTHSCSA) Animal Care and Use Committee (IACUC). We followed guidelines issued by the National Institutes of Health (NIH) and the Society for Neuroscience (SfN) to minimize the number of animals used and their suffering.
Eight-to-twelve-week-old naïve C57BL/6J (The Jackson Laboratory, Bar Harbor, ME) female in estrous phase and male mice were used for all described experiments. Estrous phase was determine by vaginal gavage as described previously89. In some fluorescent activated cell sorting (FACS) experiments, we used CGRPcre/± ER/Rosa26LSL-tDTomato/+ reporter mice31.
Left and right whole L3-L5 DRG and TG tissue biopsies were dissected after perfusion of mice with phosphate buffer pH 7.3 (PB). DRG and TG tissues were used either for total RNA isolation or for single-cell suspension generation. After perfusion of mice with PB, a 3 mm biopsy-puncher was set to take epidermal-dermal layers and with partial collection of fat cells located underneath of dermal layer of hindpaws. Hindpaw biopsies do not touch muscles. Paw tissues were used for total RNA.
Sensory neuronal fraction isolation with Percoll gradient
To generate single-cell suspensions, whole L3-L5 DRG or TG were treated with 125 μg/ml liberase (Millipore-Sigma, St. Louis, MO) and 200 μg/ml dispase II (Millipore-Sigma, St. Louis, MO) in Hank’s solution for 60 min. Tissues was washed twice with DMEM/L-glutamate/5% fetal bovine serum (FBS) media and dispersed to single-cell conditions by pipette. Single-cell media solutions were filtered through 70 μm strainer and loaded on a 60%/30% Percoll (Millipore-Sigma, St. Louis, MO) gradient prepared in the same media. Gradient preparations were spun for 15 min at 1,000×g at 4 °C. The non-neuronal fraction that was accumulated between 0 and 30% Percoll was discarded, and neuronal fraction at the border between 30 and 60% Percoll was collected. The neuronal fraction was diluted eightfold and washed with media. Sensory neurons were collected by centrifugation for 5 min at 1,200×g at 4 °C.
Isolation of DRG sensory neurons innervating hind paws and thigh muscles
Left and right male or female naïve mouse hind paws and thigh muscles were injected with 10 μl (each injection) of 1% WGA-Alexa Fluor™ 488 conjugate (WGA-488) dissolved in 1% DMSO. Two days post-injection, L3–L5 DRG were isolated and single-cell suspension was generated with enzymatic treatment as described above.
Flow cytometry was used to isolate labeled back-traced from skin and muscle DRG neurons. LIVE/DEAD viability staining was done with 7-AAD Viability Staining Solution diluted at 1:50. Labeled cells were sorted using 5 laser FACS Aria-IIIu cell sorter equipped with 130 μm nozzle.
Cell sorting was also used to assess the percentage and viability of CGRP-cre+ cells in DRG of CGRPcre-ER/+/Rosa26LSL-tDTomato/+ reporter male mice.
RNA isolation and cDNA synthesis
RNA was isolated from male or female mouse whole DRG, whole TG and paw tissues, as well as single-cell sensory neuronal fractions. Fresh tissues were homogenized in Rn-easy solution using a Bead Mill Homogenizer (Omni International, Kennesaw, GA), while RNA isolation from single-cell sensory neuron suspension did not require homogenization. Sensory neuronal and tissue total RNA was extracted using Qiagen Rn-easy (Universal Mini Kit) as was previously described90. If the preparation generated > 10–20 ng RNA, then RNA quality and integrity were checked using Fragment Analyzer Agilent 2,100 Bioanalyer RNA 6,000 Nano chip (Agilent Technologies, Santa Clara, CA). For small amounts of RNA (< 10 ng), quality of RNA was accessed after cDNA preparation.
Preparation of cDNA library and RNA-seq from tissue RNA
Approximately 500 ng total RNA was used for cDNA library preparation with oligo-dT primers by following the Illumina TruSeq stranded mRNA sample preparation guide (Illumina, San Diego, CA).
Preparation of cDNA library and RNA-seq from Percoll enriched sensory neuronal fraction
The first step in the workflow involved the depletion of rRNA by hybridization of complementary DNA oligonucleotides, followed by treatment with RNase H and DNase to remove rRNA duplexed to DNA and original DNA oligonucleotides, respectively. Following rRNA removal, the mRNA was fragmented into small pieces using divalent cautions under elevated temperature and magnesium. The cleaved RNA fragments were copied into first strand cDNA using reverse transcriptase and random primers. This was followed by second strand cDNA synthesis using DNA Polymerase I and RNase H. Strand specificity was achieved by replacing dTTP with dUTP in the Second Strand Marking Mix (SMM). The incorporation of dUTP in second strand synthesis effectively quenches the second strand during amplification, since the polymerase used in the assay will not incorporate past this nucleotide. These cDNA fragments then went through an end repair process, the addition of a single ‘A’ base, and then ligation of the adapters. The products were then purified and enriched with PCR to create the final RNA-seq library.
Preparation of cDNA library and RNA-seq from Percoll enriched sensory neuronal fraction
Sensory neurons isolated by sorting DRG neurons back-traced and labeled with WGA-488 generated < 10 ng RNA. Hence, RNA-seq cDNA libraries were prepared using oligo dT according to SMART-seq-2 protocol91,92, with the following modifications: 1: All primers were designed with 5′ biotin modifications to prevent primer concatenation; 2: PCR preamplification to 10–12 cycles; 3: Two rounds beads cleanup with 1:1 ratio after cDNA synthesis, and 4: 0.6–0.8 dual beads cleanup for Nextera XT DNA-seq library purification.
All RNA-seq cDNA libraries were subjected to quantification, pooled for cBot amplification and subsequent 50 bp single read sequencing run with Illumina HiSeq 3,000 platform (Illumina, San Diego, CA). Each group have n = 3–5 samples. Depth of reads was 30–50 × 106 bp for each sample.
RNA seq transcriptomic data generation, analyses and statistics
Sequencing data from all samples were processed in the same way. After the sequencing run, de-multiplexing with CASAVA was employed to generate the FastQ file for each sample. The combined raw reads were aligned to mouse genome build mm9/UCSC hg19 using TopHat2 default settings93,94. The BAM files obtained after alignment were processed using HTSeq-count to obtain the counts per gene, and then converted to RPKM (Read Per Kilobase of gene length per Million reads of the library)95. Then DESeq was used to identify differentially expressing genes (DEGs) after performing median normalization96. Quality control statistical analysis of outliers, intergroup variability, distribution levels, PCA and hierarchical clustering analysis was performed to statistically validate the experimental data. Multiple test controlling was performed with Benjamini–Hochberg procedure and adjusted p-value (Padj) was generated.
Three criteria have been used for selecting DEGs for the further analysis: expression levels (i.e. RPKM), fold-change (FC) and statistically significant DEGs (i.e. Pval or Padj). Selection criteria vary depending on cDNA library preparation. These selection criteria and their justifications and rationales are in the text. Venn diagrams were generated using https://bioinfogp.cnb.csic.es/tools/venny/. Genes were clustered according to biological processes using the PANTHER software (https://www.pantherdb.org/). Pathway analysis was performed with Ingenuity Pathway Analysis (IPA, Qiagen, CA).
Quantitative real-time PCR
Quantitative real-time PCR (RT-PCR) was performed on independently isolated sensory neuronal fraction from mouse male and female TG and DRG using Percoll purification protocol (see above). Quantitative RT-PCR procedure has essentially been performed as described90. Briefly, cDNA from RNA was prepared using the Superscript III First Strand Synthesis kit (Invitrogen). Amplification of target sequences was detected on ABI 7,500 Fast RTPCR system (Applied Biosystems, Foster City, CA) using TaqMan Fast Universal PCR Master Mix. The reactions were run in triplicates of 25 μl. The endogenous control for each individual gene expression assay was mouse GAPDH with Mm03302249_g1 primers (ThermoFisher Sci.), which express equally in female and male DRG and TG neurons. Specific primers were Mm00658576_m1 for Cd74, Mm01700423_m1 for Islr, Mm01191753_m1 for Bgn, Mm00470429_g1 for Mxra8, Mm00435846_m1 for Pkib and Mm00547130_s1 for Arxes2. For quantitative analysis, comparative delta-delta Ct was used to normalize the data based on the endogenous reference, and to express it as the relative fold change, after the exclusion criteria were verified by comparing primer efficiencies90.
Ethical approval and informed consent
All experimental protocols were approved by the UTHSCSA IACUC committee. Protocol numbers are 20150100AR and 20180001AR.
Data availability
RNA-seq data has been deposited to GSE121645, GSE135909 and GSE153269. Supplementary excel files for the raw gene counts per gene in all of our sequencing experiments. These supplementary files are Whole DRG (Female Vs Male) for sequencing from whole DRG; Hindpaw skin (Female vs Male) for sequencing from epidermis and dermis layer of hindpaw skin; DRG neurons by Percoll (Female vs Male) for sequencing from Percoll purified DRG neurons; TG neurons by Percoll (Female vs Male) for sequencing from Percoll purified TG neurons; and WGA-488+ (skin-thigh innerv) DRG neurons (Female vs Male) for sequencing from a mix of DRG neurons innervating hindpaw skin and thigh muscle.
References
Berkley, K. J. Sex differences in pain. Behav. Brain Sci. 20, 371–380 (1997) ((Discussion 435–513)).
Fillingim, R. B., King, C. D., Ribeiro-Dasilva, M. C., Rahim-Williams, B. & Riley, J. L. Sex, gender, and pain: a review of recent clinical and experimental findings. J Pain 10, 447–485. https://doi.org/10.1016/j.jpain.2008.12.001 (2009).
Traub, R. J. & Ji, Y. Sex differences and hormonal modulation of deep tissue pain. Front Neuroendocrinol 34, 350–366. https://doi.org/10.1016/j.yfrne.2013.07.002 (2013).
Unruh, A. M. Gender variations in clinical pain experience. Pain 65, 123–167 (1996).
Peretz, A. M., Woldeamanuel, Y. W., Rapoport, A. M. & Cowan, R. P. Spontaneous extracranial hemorrhagic phenomena in primary headache disorders: a systematic review of published cases. Cephalalgia 36, 1257–1267. https://doi.org/10.1177/0333102415618951 (2016).
Westerlund, U., Linderoth, B. & Mathiesen, T. Trigeminal complications arising after surgery of cranial base meningiomas. Neurosurg. Rev. 35, 203–209. https://doi.org/10.1007/s10143-011-0355-0 (2012) (Discussion 209–210).
Cairns, B. E. The influence of gender and sex steroids on craniofacial nociception. Headache 47, 319–324. https://doi.org/10.1111/j.1526-4610.2006.00708.x (2007).
Moulin, D. E., Foley, K. M. & Ebers, G. C. Pain syndromes in multiple sclerosis. Neurology 38, 1830–1834 (1988).
Houghton, L. A., Lea, R., Jackson, N. & Whorwell, P. J. The menstrual cycle affects rectal sensitivity in patients with irritable bowel syndrome but not healthy volunteers. Gut 50, 471–474. https://doi.org/10.1136/Gut.50.4.471 (2002).
Mathew, P. G., Dun, E. C. & Luo, J. J. A cyclic pain: the pathophysiology and treatment of menstrual migraine. Obstet. Gynecol. Surv. 68, 130–140. https://doi.org/10.1097/OGX.0b013e31827f2496 (2013).
LeResche, L., Mancl, L., Sherman, J. J., Gandara, B. & Dworkin, S. F. Changes in temporomandibular pain and other symptoms across the menstrual cycle. Pain 106, 253–261 (2003).
Slade, G. D. et al. Study methods, recruitment, sociodemographic findings, and demographic representativeness in the OPPERA study. J. Pain 12, T12-26. https://doi.org/10.1016/j.jpain.2011.08.001 (2011).
Amandusson, A. & Blomqvist, A. Estrogenic influences in pain processing. Front. Neuroendocr. 34, 329–349. https://doi.org/10.1016/j.yfrne.2013.06.001 (2013).
Klein-Hitpass, L., Schorpp, M., Wagner, U. & Ryffel, G. U. An estrogen-responsive element derived from the 5’ flanking region of the Xenopus vitellogenin A2 gene functions in transfected human cells. Cell 46, 1053–1061 (1986).
Gruber, C. J., Gruber, D. M., Gruber, I. M., Wieser, F. & Huber, J. C. Anatomy of the estrogen response element. Trends Endocrinol. Metab. 15, 73–78. https://doi.org/10.1016/j.tem.2004.01.008 (2004).
Ray, P. R. et al. Transcriptome analysis of the human tibial nerve identifies sexually dimorphic expression of genes involved in pain, inflammation, and neuro-immunity. Front. Mol. Neurosci. 12, 37. https://doi.org/10.3389/fnmol.2019.00037 (2019).
North, R. Y. et al. Electrophysiological and transcriptomic correlates of neuropathic pain in human dorsal root ganglion neurons. Brain https://doi.org/10.1093/brain/awz063 (2019).
Sharma, N. et al. The emergence of transcriptional identity in somatosensory neurons. Nature 577, 392–398. https://doi.org/10.1038/s41586-019-1900-1 (2020).
Baral, P., Udit, S. & Chiu, I. M. Pain and immunity: implications for host defence. Nat. Rev. Immunol. https://doi.org/10.1038/s41577-019-0147-2 (2019).
Fan, W. et al. The role of satellite glial cells in orofacial pain. J. Neurosci. Res. 97, 393–401. https://doi.org/10.1002/jnr.24341 (2019).
LaPaglia, D. M. et al. RNA-Seq investigations of human post-mortem trigeminal ganglia. Cephalalgia 38, 912–932. https://doi.org/10.1177/0333102417720216 (2018).
Castor, C. W., Furlong, A. M. & Carter-Su, C. Connective tissue activation: stimulation of glucose transport by connective tissue activating peptide III. Biochemistry 24, 1762–1767 (1985).
Angel, N. Z. et al. Transgenic mice overexpressing tartrate-resistant acid phosphatase exhibit an increased rate of bone turnover. J. Bone Miner. Res. 15, 103–110. https://doi.org/10.1359/jbmr.2000.15.1.103 (2000).
Huh, Y., Ji, R. R. & Chen, G. Neuroinflammation, bone marrow stem cells, and chronic pain. Front. Immunol. 8, 1014. https://doi.org/10.3389/fimmu.2017.01014 (2017).
Kwiatkowski, K. & Mika, J. The importance of chemokines in neuropathic pain development and opioid analgesic potency. Pharmacol. Rep. 70, 821–830. https://doi.org/10.1016/j.pharep.2018.01.006 (2018).
Littlejohn, G. & Guymer, E. Neurogenic inflammation in fibromyalgia. Semin Immunopathol. 40, 291–300. https://doi.org/10.1007/s00281-018-0672-2 (2018).
Kiguchi, N., Kobayashi, D., Saika, F., Matsuzaki, S. & Kishioka, S. Pharmacological regulation of neuropathic pain driven by inflammatory macrophages. Int. J. Mol. Sci. https://doi.org/10.3390/ijms18112296 (2017).
Tiruppathi, C. et al. The transcription factor DREAM represses the deubiquitinase A20 and mediates inflammation. Nat. Immunol. 15, 239–247. https://doi.org/10.1038/ni.2823 (2014).
Hargreaves, K. M. & Ruparel, S. Role of oxidized lipids and TRP channels in orofacial pain and inflammation. J. Dent. Res. 95, 1117–1123. https://doi.org/10.1177/0022034516653751 (2016).
Ruparel, S. et al. Oxidized linoleic acid metabolite-cytochrome P450 system (OLAM-CYP) is active in biopsy samples from patients with inflammatory dental pain. Pain 154, 2363–2371. https://doi.org/10.1016/j.pain.2013.07.011 (2013).
Patil, M. J., Hovhannisyan, A. H. & Akopian, A. N. Characteristics of sensory neuronal groups in CGRP-cre-ER reporter mice: comparison to Nav1.8-cre, TRPV1-cre and TRPV1-GFP mouse lines. PLoS ONE 13, e0198601. https://doi.org/10.1371/journal.pone.0198601 (2018).
Rodriguez, E. et al. A craniofacial-specific monosynaptic circuit enables heightened affective pain. Nat. Neurosci. 20, 1734–1743. https://doi.org/10.1038/s41593-017-0012-1 (2017).
Megat, S. et al. Differences between dorsal root and trigeminal ganglion nociceptors in mice revealed by translational profiling. J. Neurosci. https://doi.org/10.1523/JNEUROSCI.2663-18.2019 (2019).
Tan, W., Song, Y., Mo, C., Jiang, S. & Wang, Z. Analysis of gene expression profile microarray data in complex regional pain syndrome. Mol. Med. Rep. 16, 3371–3378. https://doi.org/10.3892/mmr.2017.6950 (2017).
Gallaher, Z. R. & Steward, O. Modest enhancement of sensory axon regeneration in the sciatic nerve with conditional co-deletion of PTEN and SOCS3 in the dorsal root ganglia of adult mice. Exp. Neurol. 303, 120–133. https://doi.org/10.1016/j.expneurol.2018.02.012 (2018).
Wei, J. et al. Overexpression of suppressor of cytokine signaling 3 in dorsal root ganglion attenuates cancer-induced pain in rats. Mol. Pain. 13, 1744806916688901. https://doi.org/10.1177/1744806916688901 (2017).
Dominguez, E., Mauborgne, A., Mallet, J., Desclaux, M. & Pohl, M. SOCS3-mediated blockade of JAK/STAT3 signaling pathway reveals its major contribution to spinal cord neuroinflammation and mechanical allodynia after peripheral nerve injury. J. Neurosci. 30, 5754–5766. https://doi.org/10.1523/JNEUROSCI.5007-09.2010 (2010).
Shutov, L. P. et al. The complement system component C5a produces thermal hyperalgesia via macrophage-to-nociceptor signaling that requires NGF and TRPV1. J. Neurosci. 36, 5055–5070. https://doi.org/10.1523/Jneurosci.3249-15.2016 (2016).
Stuart, M. J., Singhal, G. & Baune, B. T. Systematic review of the neurobiological relevance of chemokines to psychiatric disorders. Front. Cell Neurosci. 9, 357. https://doi.org/10.3389/fncel.2015.00357 (2015).
Li, Y. Y. et al. Activation of STAT3-mediated CXCL12 up-regulation in the dorsal root ganglion contributes to oxaliplatin-induced chronic pain. Mol Pain. https://doi.org/10.1177/1744806917747425 (2017).
Zhang, J. et al. TGF-beta 1 suppresses CCL3/4 expression through the ERK signaling pathway and inhibits intervertebral disc degeneration and inflammation-related pain in a rat model. Exp. Mol. Med. 49, e397 (2017).
Wang, J. et al. Upregulation of CX3CL1 mediated by NF-kappaB activation in dorsal root ganglion contributes to peripheral sensitization and chronic pain induced by oxaliplatin administration. Mol. Pain. 13, 1744806917726256. https://doi.org/10.1177/1744806917726256 (2017).
Miller, R. E., Miller, R. J. & Malfait, A. M. Osteoarthritis joint pain: the cytokine connection. Cytokine 70, 185–193. https://doi.org/10.1016/j.cyto.2014.06.019 (2014).
Rosen, S., Ham, B. & Mogil, J. S. Sex differences in neuroimmunity and pain. J. Neurosci. Res. 95, 500–508. https://doi.org/10.1002/jnr.23831 (2017).
Busserolles, J., Tsantoulas, C., Eschalier, A. & Lopez Garcia, J. A. Potassium channels in neuropathic pain: advances, challenges, and emerging ideas. Pain 157(Suppl 1), S7-14. https://doi.org/10.1097/j.pain.0000000000000368 (2016).
Du, X., Gao, H., Jaffe, D., Zhang, H. & Gamper, N. M-type K(+) channels in peripheral nociceptive pathways. Br. J. Pharmacol. 175, 2158–2172. https://doi.org/10.1111/bph.13978 (2018).
Moore, C., Gupta, R., Jordt, S. E., Chen, Y. & Liedtke, W. B. Regulation of pain and itch by TRP channels. Neurosci. Bull. 34, 120–142. https://doi.org/10.1007/s12264-017-0200-8 (2018).
Magnani, N. D., Dada, L. A. & Sznajder, J. I. Ubiquitin-proteasome signaling in lung injury. Transl. Res. 198, 29–39. https://doi.org/10.1016/j.trsl.2018.04.003 (2018).
Gong, B., Radulovic, M., Figueiredo-Pereira, M. E. & Cardozo, C. The ubiquitin-proteasome system: potential therapeutic targets for alzheimer’s disease and spinal cord injury. Front. Mol. Neurosci. 9, 4. https://doi.org/10.3389/fnmol.2016.00004 (2016).
Draganidis, D. et al. Protein ingestion preserves proteasome activity during intense aseptic inflammation and facilitates skeletal muscle recovery in humans. Br. J. Nutr. 118, 189–200. https://doi.org/10.1017/S0007114517001829 (2017).
Hamed, S. A. Brain injury with diabetes mellitus: evidence, mechanisms and treatment implications. Expert Rev. Clin. Pharmacol. 10, 409–428. https://doi.org/10.1080/17512433.2017.1293521 (2017).
Figueroa-Romero, C., Sadidi, M. & Feldman, E. L. Mechanisms of disease: the oxidative stress theory of diabetic neuropathy. Rev. Endocr. Metab. Disord. 9, 301–314. https://doi.org/10.1007/s11154-008-9104-2 (2008).
Zeisel, A. et al. Molecular architecture of the mouse nervous system. Cell 174, 999–1014. https://doi.org/10.1016/j.cell.2018.06.021 (2018).
Hockley, J. R. F. et al. Single-cell RNAseq reveals seven classes of colonic sensory neuron. Gut 68, 633–644. https://doi.org/10.1136/gutjnl-2017-315631 (2019).
Denzin, L. K., Robbins, N. F., Carboy-Newcomb, C. & Cresswell, P. Assembly and intracellular transport of HLA-DM and correction of the class II antigen-processing defect in T2 cells. Immunity 1, 595–606. https://doi.org/10.1016/1074-7613(94)90049-3 (1994).
Hartlehnert, M. et al. Schwann cells promote post-traumatic nerve inflammation and neuropathic pain through MHC class II. Sci. Rep. 7, 12518. https://doi.org/10.1038/s41598-017-12744-2 (2017).
Barkai, O. et al. Platelet-derived growth factor activates nociceptive neurons by inhibiting M-current and contributes to inflammatory pain. Pain https://doi.org/10.1097/j.pain.0000000000001523 (2019).
Sun, M. M., Beier, F. & Pest, M. A. Recent developments in emerging therapeutic targets of osteoarthritis. Curr. Opin. Rheumatol. 29, 96–102. https://doi.org/10.1097/BOR.0000000000000351 (2017).
Martin, L. J. et al. Epiregulin and EGFR interactions are involved in pain processing. J. Clin. Invest. 127, 3353–3366. https://doi.org/10.1172/JCI87406 (2017).
Rajaei, S., Fatahi, Y. & Dabbagh, A. Meeting between rumi and shams in notch signaling; implications for pain management: a narrative review. Anesth. Pain Med. 9, e85279. https://doi.org/10.5812/aapm.85279 (2019).
Liu, D. L. et al. Upregulation of Ih expressed in IB4-negative Adelta nociceptive DRG neurons contributes to mechanical hypersensitivity associated with cervical radiculopathic pain. Sci. Rep. 5, 16713. https://doi.org/10.1038/srep16713 (2015).
Weng, X., Smith, T., Sathish, J. & Djouhri, L. Chronic inflammatory pain is associated with increased excitability and hyperpolarization-activated current (Ih) in C-but not Adelta-nociceptors. Pain 153, 900–914. https://doi.org/10.1016/j.pain.2012.01.019 (2012).
Zhang, X. et al. Ezogabine (KCNQ2/3 channel opener) prevents delayed activation of meningeal nociceptors if given before but not after the occurrence of cortical spreading depression. Epilepsy Behav. 28, 243–248. https://doi.org/10.1016/j.yebeh.2013.02.029 (2013).
Barkai, O. et al. The role of Kv7/M potassium channels in controlling ectopic firing in nociceptors. Front. Mol. Neurosci. 10, 181. https://doi.org/10.3389/fnmol.2017.00181 (2017).
Liu, B. et al. The acute nociceptive signals induced by bradykinin in rat sensory neurons are mediated by inhibition of M-type K+ channels and activation of Ca2+-activated Cl-channels. J. Clin. Invest. 120, 1240–1252. https://doi.org/10.1172/JCI41084 (2010).
Linley, J. E. et al. Inhibition of M current in sensory neurons by exogenous proteases: a signaling pathway mediating inflammatory nociception. J. Neurosci. 28, 11240–11249. https://doi.org/10.1523/JNEUROSCI.2297-08.2008 (2008).
Dou, Y. et al. Orai1 plays a crucial role in central sensitization by modulating neuronal excitability. J. Neurosci. 38, 887–900. https://doi.org/10.1523/JNEUROSCI.3007-17.2017 (2018).
Lamas, J. A. & Fernandez-Fernandez, D. Tandem pore TWIK-related potassium channels and neuroprotection. Neural. Regen. Res. 14, 1293–1308. https://doi.org/10.4103/1673-5374.253506 (2019).
Guo, Z. et al. TRESK K(+) channel activity regulates trigeminal nociception and headache. eNeuro https://doi.org/10.1523/ENEURO.0236-19.2019 (2019).
Lafreniere, R. G. et al. A dominant-negative mutation in the TRESK potassium channel is linked to familial migraine with aura. Nat. Med. 16, 1157–1160. https://doi.org/10.1038/nm.2216 (2010).
Yan, J. & Dussor, G. Ion channels and migraine. Headache 54, 619–639. https://doi.org/10.1111/head.12323 (2014).
Wei, X., Edelmayer, R. M., Yan, J. & Dussor, G. Activation of TRPV4 on dural afferents produces headache-related behavior in a preclinical rat model. Cephalalgia 31, 1595–1600. https://doi.org/10.1177/0333102411427600 (2011).
Talbot, S., Foster, S. L. & Woolf, C. J. Neuroimmunity: physiology and pathology. Annu. Rev. Immunol. 34, 421–447. https://doi.org/10.1146/annurev-immunol-041015-055340 (2016).
Austin, P. J. & Moalem-Taylor, G. The neuro-immune balance in neuropathic pain: involvement of inflammatory immune cells, immune-like glial cells and cytokines. J. Neuroimmunol. 229, 26–50. https://doi.org/10.1016/j.jneuroim.2010.08.013 (2010).
Huygen, F. J., de Bruijn, A. G., Klein, J. & Zijlstra, F. J. Neuroimmune alterations in the complex regional pain syndrome. Eur. J. Pharmacol. 429, 101–113 (2001).
Waseem, M., Kaushik, P., Tabassum, H. & Parvez, S. Role of mitochondrial mechanism in chemotherapy-induced peripheral neuropathy. Curr. Drug Metab. 19, 47–54. https://doi.org/10.2174/1389200219666171207121313 (2018).
Hopkins, H. L., Duggett, N. A. & Flatters, S. J. L. Chemotherapy-induced painful neuropathy: pain-like behaviours in rodent models and their response to commonly used analgesics. Curr. Opin. Support 10, 119–128. https://doi.org/10.1097/Spc.0000000000000204 (2016).
Xie, J. D., Chen, S. R., Chen, H. & Pan, H. L. Bortezomib induces neuropathic pain through protein kinase C-mediated activation of presynaptic NMDA receptors in the spinal cord. Neuropharmacology 123, 477–487. https://doi.org/10.1016/j.neuropharm.2017.06.027 (2017).
Hohn, A., Konig, J. & Jung, T. Metabolic syndrome, redox state, and the proteasomal system. Antioxid. Redox Signal 25, 902–917. https://doi.org/10.1089/ars.2016.6815 (2016).
Kiryu-Seo, S. & Kiyama, H. Mitochondrial behavior during axon regeneration/degeneration in vivo. Neurosci. Res. 139, 42–47. https://doi.org/10.1016/j.neures.2018.08.014 (2019).
Quansah, E. et al. Targeting energy metabolism via the mitochondrial pyruvate carrier as a novel approach to attenuate neurodegeneration. Mol. Neurodegener. 13, 28. https://doi.org/10.1186/s13024-018-0260-x (2018).
Hill, C. S., Menon, D. K. & Coleman, M. P. P7C3-A20 neuroprotection is independent of Wallerian degeneration in primary neuronal culture. NeuroReport 29, 1544–1549. https://doi.org/10.1097/WNR.0000000000001146 (2018).
Pieper, A. A. et al. Discovery of a proneurogenic, neuroprotective chemical. Cell 142, 39–51. https://doi.org/10.1016/j.cell.2010.06.018 (2010).
LoCoco, P. M. et al. Pharmacological augmentation of nicotinamide phosphoribosyltransferase (NAMPT) protects against paclitaxel-induced peripheral neuropathy. Elife https://doi.org/10.7554/eLife.29626 (2017).
Tansley, S. N., Wong, C., Uttam, S., Mogil, J. S. & Khoutorsky, A. Translation regulation in the spinal dorsal horn: a key mechanism for development of chronic pain. Neurobiol. Pain 4, 20–26. https://doi.org/10.1016/j.ynpai.2018.03.003 (2018).
Khoutorsky, A. & Price, T. J. Translational control mechanisms in persistent pain. Trends Neurosci. 41, 100–114. https://doi.org/10.1016/j.tins.2017.11.006 (2018).
Sorge, R. E. et al. Different immune cells mediate mechanical pain hypersensitivity in male and female mice. Nat. Neurosci. 18, 1081–1083. https://doi.org/10.1038/nn.4053 (2015).
Martin, L. J. et al. Male-specific conditioned pain hypersensitivity in mice and humans. Curr. Biol. 29, 192–201. https://doi.org/10.1016/j.cub.2018.11.030 (2019).
Caligioni, C. S. Assessing reproductive status/stages in mice. Curr. Protoc. Neurosci. https://doi.org/10.1002/0471142301.nsa04is48 (2009).
Patil, M. J., Ruparel, S. B., Henry, M. A. & Akopian, A. N. Prolactin regulates TRPV1, TRPA1, and TRPM8 in sensory neurons in a sex-dependent manner: contribution of prolactin receptor to inflammatory pain. Am. J. Physiol. Endocrinol. Metab. 305, E1154-1164. https://doi.org/10.1152/ajpendo.00187.2013 (2013).
Picelli, S. et al. Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat. Methods 10, 1096–1098. https://doi.org/10.1038/nmeth.2639 (2013).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181. https://doi.org/10.1038/nprot.2014.006 (2014).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562–578. https://doi.org/10.1038/nprot.2012.016 (2012).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111. https://doi.org/10.1093/bioinformatics/btp120 (2009).
Anders, S., Pyl, P. T. & Huber, W. HTSeq: a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169. https://doi.org/10.1093/bioinformatics/btu638 (2015).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106. https://doi.org/10.1186/gb-2010-11-10-r106 (2010).
Acknowledgements
We would like to thank Dr. Siarhei Bialuhin for advice on the experimental strategy; and Dr. Pao-Tien Chuang (UC San Francisco, San Francisco, CA) for kindly providing the Calcacre/+-ER mouse line. RNA-seq studies were conducted in the Genome Sequencing Facility (GSF) in the Greehey Children’s Cancer Research Institute (GCCRI) of UTHSCSA. The GSF facility has been constructed in part with the support from UT Health San Antonio, NIH-NCI P30 CA054174 (Cancer Center at UT Health San Antonio), S10 Shared Instrumentation Grant Program (SIG)/NIH (S10OD021805-01 to Z.L.), and Cancer Prevention Research Institute of Texas (CPRIT) Core Facility Award (RP160732). This work was supported by NINDS/NIH grants NS102161 (to T.J.P and A.N.A.); NIH/NIGMS GM112747 (to A.N.A.); NINDS/NIH NS065926 (to T.J.P.), and UT BRAIN Pilot Program ID: 1503083 (to G.D. and A.N.A.).
Author information
Authors and Affiliations
Contributions
J.M. conducted most experiments; A.V.T. and A.N.A. conducted a set of experiments; Y.Z. and A.W. analyzed RNA-seq data and prepared corresponding figures and graphical depictions; D.G. generated RNA-seq/transcriptomic data; Z.L. designed and directed RNA-seq experiments; A.V.T., G.D. and T.J.P. helped with experimental design, and contributed to the manuscript preparation; A.N.A. and T.J.P. prepared final version of the manuscript; and A.N.A designed and directed the project, wrote the first draft of the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mecklenburg, J., Zou, Y., Wangzhou, A. et al. Transcriptomic sex differences in sensory neuronal populations of mice. Sci Rep 10, 15278 (2020). https://doi.org/10.1038/s41598-020-72285-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-72285-z
This article is cited by
-
Caldendrin represses neurite regeneration and growth in dorsal root ganglion neurons
Scientific Reports (2023)
-
Associations of tissue damage induced inflammatory plasticity in masseter muscle with the resolution of chronic myalgia
Scientific Reports (2023)
-
Nociceptors regulate osteoimmune transcriptomic response to infection
Scientific Reports (2023)
-
Machine learning analysis predicts a person’s sex based on mechanical but not thermal pain thresholds
Scientific Reports (2023)
-
Sex-dependent differences in the genomic profile of lingual sensory neurons in naïve and tongue-tumor bearing mice
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.