Abstract
Vascular endothelial growth factor A (VEGF-A) and its binding to VEGFRs is an important angiogenesis regulator, especially the earliest-known isoform, VEGF-A165a. Yet several additional splice variants play prominent roles in regulating angiogenesis in health and in vascular disease, including VEGF-A121 and an anti-angiogenic variant, VEGF-A165b. Few studies have attempted to distinguish these forms from their angiogenic counterparts, experimentally. Previous studies of VEGF-A:VEGFR binding have measured binding kinetics for VEGFA165 and VEGF-A121, but binding kinetics of the other two pro- and all anti-angiogenic splice variants are not known. We measured the binding kinetics for VEGF-A165, -A165b, and -A121 with VEGFR1 and VEGF-R2 using surface plasmon resonance. We validated our methods by reproducing the known affinities between VEGF-A165a:VEGFR1 and VEGF-A165a:VEGFR2, 1.0 pM and 10 pM respectively, and validated the known affinity VEGF-A121:VEGFR2 as KD = 0.66 nM. We found that VEGF-A121 also binds VEGFR1 with an affinity KD = 3.7 nM. We further demonstrated that the anti-angiogenic variant, VEGF-A165b selectively prefers VEGFR2 binding at an affinity = 0.67 pM while binding VEGFR1 with a weaker affinity—KD = 1.4 nM. These results suggest that the − A165b anti-angiogenic variant would preferentially bind VEGFR2. These discoveries offer a new paradigm for understanding VEGF-A, while further stressing the need to take care in differentiating the splice variants in all future VEGF-A studies.
Similar content being viewed by others
Introduction
The vascular endothelial growth factors have been extensively studied as signaling molecules in angiogenesis1,2, and their signaling comprises several components. The mammalian VEGF family includes five homodimeric ligands: VEGF-A, VEGF-B, VEGF-C, VEGF-D, and placental growth factor (PlGF)3. VEGF-A signal transduction unfolds following the pattern common to other tyrosine kinase receptors (RTKs), like the fibroblast growth factor (FGF) and platelet-derived growth factor (PDGF) families: (1) ligands bind a receptor monomer, promoting dimerization with another free receptor; (2) phosphorylation occurs at specific tyrosine residues depending on conformational changes allowed by the ligand—i.e. signaling is not directly coupled to binding, but dependent on ligand structure4,5; (3) adaptor proteins bind these tyrosine residues and undergo phosphorylation6, and (4) phosphorylated adaptor proteins initiate effector signaling cascades7 that can ultimately mediate cell-level responses such as cell migration, proliferation, and cell survival8.
The most well-studied ligand is VEGF-A9. VEGF-A has a wide array of isoforms produced through alternative mRNA splicing10,11, including: VEGF-A121, VEGF-A121b, VEGF-A145, VEGF-A145b, VEGF-A165, VEGF-A165b, VEGF-A183, VEGF-A189, and VEGF-A206 (Fig. 1), in addition to VEGF-A111, an abnormal splice variant induced by genotoxic stressors12. Members of the VEGF-Axxx family are pro-angiogenic, whereas those designated as VEGF-Axxxb have been described as ‘anti-angiogenic’ due to their less-effective activation of VEGFRs1,13, but are more accurately described as weak VEGFR2 agonists14. Additionally, VEGF-Axxxb splice variants cannot bind heparin-sulfate chains or the VEGFR2 coreceptor, neuropilin-115. Given this modulatory signaling role, several of these VEGF-Axxxb isoforms have been proposed as targets for treating such angiogenic-dependent diseases as diabetic retinopathy16, peripheral vascular disease17, and cancers18.
Developing tailored treatment to angiogenesis-related diseases, such as in tumors or ischemia, can be enhanced by the use computational modeling. Computational modeling offers a systemic approach for optimizing ligand, receptor, and signaling targeting. It has predicted how anti-VEGF drug efficacy may depend on VEGFR1 concentrations on tumor endothelial cells19. It has provided predictions on anti-VEGF pharmacodynamics and pharmacokinetics in the body13. It has provided systematic analysis comparing the roles of heparin-binding and non-heparin-binding VEGF-A isoforms in tumors20. The advantage of these computational approaches is that predictions are defined by protein concentrations and kinetics (e.g. protein–protein-interactions). We can similarly apply computational modeling towards investigating how best to target these VEGF-Axxxb isoforms. Such models will require both quantitative insight into the relative concentrations for each isoform and also insights into their binding characteristics with the VEGF receptors, i.e. ligand:receptor kinetic rate constants and binding affinities.
Developing computational models requires precise measurements of the protein–protein interactions involved in the systems. Computational modeling has, therefore, been limited by VEGF-A splice variant kinetic binding data availability. More broadly speaking, precise measurements of ligand:receptor binding affinities and kinetics are crucial to understanding the systems at play: determining whether pro- or anti-angiogenic splice variants dominate the signaling axis is based both on the ligand concentrations and their binding affinities. Several models have incorporated the commonly-expressed VEGF-A121a to study its role in physiological conditions, and several pathologies including breast cancers and peripheral artery disease (PAD)21,22,23,24,25,26. Current computational models have additionally evaluated anti-angiogenic VEGF-A165b in diseases like PAD27. Each model operated under the assumption that VEGF-A121a and VEGF-A165b splice variants bind VEGF receptors with the same affinity as VEGF-A165a.
The conclusions reached, therefore, are highly dependent on whether VEGF-A splice variants truly bind their receptors with identical binding affinities. Therefore, measuring their precise kinetics to calculate their affinities is crucial to ensuring their predictions are accurate. Previously, only a small subset of the pro-angiogenic isoforms, VEGF-A165a, -A121, and -A183 have had their binding affinities measured12,14, and only VEGF-A165a has binding kinetic constants (ka and kd) measured12. The binding affinities (e.g., KD = kd/ka) of the anti-angiogenic VEGF-A165b and VEGF-A165a to VEGFRs were previously compared qualitatively via a radiolabeled-ligand binding assay14, concluding that VEGF-A165b and VEGF-A165a bind with similar affinities, but also without providing quantitative measurements of the binding affinity (KD). However, these previous studies provided insufficient evidence for identical VEGFR binding affinities for two reasons: (1) Radiolabeled-ligand binding assays are unable to compare differences in ligand-binding kinetics constants (ka and kd) and only provide binding data at equilibrium conditions (e.g., KD). However, no mathematical fittings were employed to quantify binding affinity KD, rendering it difficult to interpret the qualitative binding curves, which show VEGF-A165a saturated VEGFR1 at a higher concentration than VEGF-A165b. Their results, suggest that VEGF-A165b binds VEGFR1 with a stronger affinity, contrary to their reported conclusions. (2) The concentration range was limited. The lowest dose explored was sufficient to saturate VEGFR2 for both splice variants, preventing conclusive comparisons of their relative binding affinities. If the receptors start saturated, the results cannot conclude whether one ligand binds with a stronger affinity than the other, only that both ligands have affinities sufficient to saturate at those low-dose conditions. Differentiating whether VEGF-A165b binds with identical affinity versus greater affinity than VEGF-A165a would require testing doses below the receptor-saturating concentrations for both VEGF-A165a and VEGF-A165b.
Surface plasmon resonance (SPR) is an ideal approach for addressing the unanswered questions of (a) whether VEGF-A165a, VEGF-A165b, and VEGF-A121 bind VEGFR1 and VEGFR2 with identical binding affinities and (b) the kinetic rate constants for each interaction. SPR measures binding kinetics (ka and kd) by detecting biomolecular interactions in real-time with high sensitivity28,29. Recent work has further advanced reference signal choice to minimize the detection of non-specific binding while measuring ligand:receptor kinetic constants for VEGF family receptors, specifically30. Recently, the SPR-based approach has been applied to compare pro- and anti-angiogenic binding properties, but these studies only qualitatively reported that the isoforms bound VEGFRs with binding affinities within two orders of magnitude of each other12, and these studies were performed before new reference subtraction methods were demonstrated30 . Kinetic rate constants have been reported only for VEGF-A165:VEGFR interactions12,30,31. These currently-unknown binding characteristics are needed to develop physiologically-accurate and predictive computational models.
Here we apply surface plasmon resonance (SPR) to measure the unknown VEGF-Axxxa:VEGFR1 and -Axxxb binding kinetics. We observe differential binding that should lead to differential activation of the VEGFRs. These results will enable the construction of detailed, accurate computational models to help further our understanding of angiogenic signaling in human health and pathology.
Results
Establishing VEGFR binding kinetics for VEGF-A splice variants
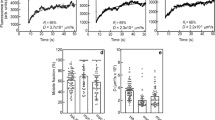
We utilized the SPR biosensor-based assay to measure the unknown kinetics for VEGF-A121 and VEGF-A165b binding to VEGFR1 and VEGFR2. We confirmed that the canonical view of VEGFR:VEGF-A splice variant binding holds: VEGF-A165a, -A165b, and -A121 bind both VEGFR1 and VEGFR232. We validated the method with VEGF-A165a binding kinetics30,31. We measured a VEGF-A165a:VEGFR1 binding affinity (Fig. 2A; and Supplementary Table S1) binding affinity of KD = 1 pM (Fig. 3A); which is consistent with a prior SPR affinity of KD = 7.5 pM31. We measured a VEGF-A165a:VEGFR2 binding affinity of KD = 9.8 pM, which is within an order of magnitude of a previous SPR measurement found by Tiedemann et al. (KD = 52 pM33). We observe that VEGFA165a binds VEGFR1 with 3-orders of magnitude stronger affinity than either the anti-angiogenic -A165b or -A121: VEGF-A165a binds VEGFR1 with KD = 1.0 pM. VEGF-A165b, however, binds VEGFR1 with a KD = 1.4 nM. VEGF-A121 binds VEGFR1 with the weakest affinity of the three isoforms with a KD = 3.7 nM. (Fig. 2A, and Supplementary Table S1). In contrast, we observe that the -A165b variant binds VEGFR2 10 × stronger than the -A165a variant with a VEGF-A165b:VEGFR2-affinity constant of KD = 0.67 pM versus a VEGF-A165a:VEGFR2 KD = 9.8 pM (Fig. 2A, Supplementary Table S1). In contrast, we observe VEGFA121:VEGFR2 binding occurred with the weakest strength, with an affinity constant KD = 660 nM (Fig. 2A). These binding affinities disagree with recent SPR studies of VEGF-A121:VEGFR1 and VEGFR234. In their studies, however, Teran et al. also report nM affinities for the well-established picomolar affinities measured for VEGF-A165a to VEGFR1 and VEGFR2. We attribute these discrepancies to their use of FGFR1 as a receptor reference protein. Structurally similar receptor proteins enable greater non-specific binding with structurally similar ligands35, thereby allowing an overestimated reference signal, underestimating the binding affinities. Alternatively, VEGF-A165a:FGFR1 may specifically bind cross-family, similar to the PDGF:VEGFR2 binding which was recently demonstrated to occur30.
SPR sensograms for VEGFA165a, -A121, and -A165b binding VEGFR1 and VEGFR2. (A) Kinetic response curve of VEGFA165a–VEGFR1 and (D) VEGFR2 binding confirmed previous high affinity measurements to both receptors. (B) Both VEGFA121 and the anti-angiogenic VEGFA165b variants bind VEGFR1 with affinities 3 orders of magnitude weaker than VEGFA165a. (E) VEGFA121 binds VEGFR2 with the weakest affinity, and (F) the anti-angiogenic VEGFA165b binds VEGFR2 with a strong affinity that could displace the angiogenic form. Receptors were immobilized on BIAcore CM5 sensors. All kinetic studies were obtained by simultaneously injecting ligand across VEGFR1, VEGFR2, and Angiopoietin-4, a non-VEGFA binding protein used for reference subtraction. The following curves were obtained by subtracting this reference signal from the raw receptor-ligand interaction curve, removing signals associated with non-specific interactions.
Association and dissociation rate constants reveal affinity trends
We can further understand the VEGF-Axxx:VEGFR interaction affinities via their association and dissociation rate constants (Fig. 2B, C) obtained from analyzing the SPR sensograms (Fig. 3A–F). Here we observe that the difference in -A165a and -A165b binding affinities with VEGFR2 is attributed to their dissociation kinetics. VEGFA165a and VEGF-A165b bind VEGFR2 with similar association rates—i.e. ka ≃ 106 M−1 s−1; however, A165a:VEGFR2 has a dissociation rate 50-fold faster than does -A165b:VEGFR2, resulting in the higher binding affinity between the latter pair (Fig. 3B, C). In contrast, -A165b:VEGFR1 dissociates ~ 30-fold faster than does VEGF-A165a:VEGFR1, but with a, -A165b:VEGFR1 association rate ~ 2 orders of magnitude slower than for VEGF-A165a:VEGFR1 ka ≃ 4.9 * 103 versus 4.0 * 105). Together this results in a 1,000-fold weaker -A165b:VEGFR1 interaction (Fig. 3A–C).
In contrast to the variation in dissociation rates, we observed minor association rate variation across VEGF-A121, -A165a, and -A165b. For both VEGFR1 and VEGFR2, the splice variants association rates varied within an order of magnitude. VEGF-A121 bound VEGFR1 with an association rate within an order of magnitude of VEGF-A165a and -A165b. Likewise, all three variants bound VEGFR2 with association rates within an order of magnitude (Fig. 3B).
Discussion
We have shown two key findings: (1) differential binding between VEGFRs and the three most common36,37,38 VEGF-A splice variants, VEGF-A165a, VEGF-A121, and VEGF-A165b; and (2) we measured the kinetics and affinities for each interaction. The differential VEGFR binding may be key to understanding VEGF-Axxx protein expression patterns in numerous pathologies, including peripheral artery disease (PAD), cancers, obesity, and other disorders like systemic sclerosis39,40,41,42,43.
Differential VEGF-Axxx:VEGFR binding kinetics in VEGFR signaling
Our measurements for VEGF-Axxx:VEGFR binding fill an important gap in our understanding of splice variant interactions with their receptors. Previous work had measured binding affinities for a subset of the “pro-angiogenic” VEGF-Axxxa isoforms—VEGF-A165a, -A121, and -A183, but precise binding kinetics were found previously only for -A165a31. Additionally, the binding affinities were only qualitatively compared between the weak-agonist VEGF-A165b and VEGF-A165a to VEGFRs via a radiolabeled-ligand binding assay14. These studies concluded that VEGF-A165b and VEGF-A165a bind with similar affinities, without providing quantitative measurements of the binding affinity (KD) nor providing binding kinetics necessary to model their dynamics computationally. Our results address this gap by measuring the binding kinetic constants, under experimental conditions that reflect the relative ligand and receptor expression levels found physiological-comparable conditions, for VEGF-A121 and VEGF-A165b, revealing stronger VEGF-A165b:VEGFR2 binding versus -A165a, and overall weaker VEGF-A121:VEGFR binding.
The kinetic measurements in these studies focused on the three most prevalent VEGF-A splice variants36,37,38, however, several other human splice variants still require precise kinetic and affinity quantification measurements, such as VEGF-A189, and the other -Axxxb variants besides VEGF-A165b. The systemic experimental approach here provides a framework for approaching this task. For example, this can be used to understand splice variant binding to the co-receptor neuropilin-1 and heparin sulfate (HS) chains surrounding the cell, since they are known to binddifferentially11,44,45. The VEGF-Axxxb variants specifically have a diminished capacity to bind neuropilin-1 and HS-chains, preventing their cooperative binding to VEGFR214. These phenomena may explain differences between the results reported here and studies46 by Ganta et al. This prior work reported VEGF-A165b outcompeting VEGF-A165a, for VEGFR1 binding, via qualitative competitive binding studies on endothelial cells. In contrast, we directly measure binding to isolated recombinant receptors. The difference between the recombinant VEGF-A:VEGFR binding affinities versus the in vitro binding to cell-bound receptors likely reflects these differential interactions between VEGF-A165x splice variants with co-receptors and ECM components, as these alternate binding partners will sequester splice variants to different degrees11,44,45. Measuring the ‘effective’ binding affinities in their presence could be accomplished by performing SPR experiments in the presence of neuropilins, heparins, and glycoproteins such as fibronectin, or by performing SPR directly on cells using a number of approaches recently reviewed34. The differences between the strength of these interactions, coupled with the differential ligand:receptor binding affinities, will need to be considered together to predict precisely the physiological role for these splice variants.
Towards further understanding the significance of additional binding partners, it will be key to quantitatively measure NRP1 and HS-chain abundance. Quantification of NRP1 would indicate that it could significantly sequester VEGF-ligands, with NRP1 ranges of 40,000–68,000 on the plasma membrane of a human blood or lymphatic endothelial cell, in vitro47,48 and over 100,000 membrane NRP1 per fibroblast, in vitro47. Moreover, HS chain estimates of ~ 1 per NRP1 on endothelial cells, and ~ 1–4 on basement membrane proteoglycans49, further demonstrate the significance of HS significance. Such protein measurements can be obtained using quantitative flow cytometry50, and could further compare how NRP1 and ECM expression differ across physiological and pathological states. In silico work has predicted that NRP1 can significantly sequester VEGFs in the pathological condition of cancer and in doing so, improve anti-VEGF efficacy (decreased free-VEGF). Two conditions were predicted to enable this improved therapeutic efficacy: (1) increased numbers of NRP1 on cells51 or (2) inhibition of VEGFR2–NRP1 interactions52. Additional explorations of NRP1 or HS-chain effects can be achieved via modeling, and a recent model provided an accessible guide for understanding for how concentrations and binding kinetics would alter the signaling landscapes53. However, future studies would require measurement of VEGF-NRP1 and VEGF-HS kinetics to faithfully evaluate how they alter angiogenic responses.
Additionally, our measurements of differential binding between VEGF-A splice variants stress the importance of understanding how each splice variant effects different responses downstream of ligand:receptor binding. While ligand-receptor binding is a key first-step in RTK activation, the resulting receptor conformational changes induced are critical in allowing tyrosine residue phosphorylation5. Therefore, different VEGF-A splice variants can induce receptor dimer confirmations that selectively inhibit the activity of some tyrosine residues while promoting phosphorylation at other residues. This is best exemplified by VEGF-A165b, which can promote comparable VEGFR2Y1175 phosphorylation while inhibiting VEGFR2Y1052 phosphorylation54. In contrast, VEGF-A165b:VEGFR1 binding has been shown to inhibit VEGFR1Y1333 phosphorylation46. Since, VEGFR functional responses are regulated by differential tyrosine residue phosphorylation55,56,57, it is critical that future work fully characterize the signaling elicited by each splice variant.
Our results, demonstrating that VEGF-A165b binds VEGFR2 selectively with stronger affinity than -A165a, may help explain why isoform specific up-regulation and down-regulation have either pro- or anti-angiogenic results in different pathologies. This is based on existing views that VEGF-A165b does not induce significant signaling through its binding to VEGFR2. For instance, Kawamura et al.54 demonstrated VEGF-A165a treatment, at receptor-saturation levels, resulted in reduced VEGFR2 phosphorylation levels. However, it is critical to note that recent evidence suggests that the both so-called “pro-” and “anti-angiogenic isoforms”, VEGF-A165b and -A165a, can phosphorylate VEGFR2Y1175 in peripheral artery disease46. Ganta et al.46 demonstrated that inhibiting VEGF-A165b did not result in significantly increased activity at VEGFR2Y1175, suggesting that -A165b was not acting as a full antagonist at this VEGFR2 residue under non-receptor saturating conditions. Taken together, the action of VEGF-A165b depends on the concentration of VEGF-A165a: high-affinity binding of VEGF-A165b to VEGFR2 would result in a significant decrease in VEGFR2 activity when VEGF-A165a concentrations saturate VEGFR2.
While research is currently limited for VEGF-A165b-induced VEGFR2 phosphorylation, it is possible that VEGF-A165b may play a partial agonist role at some tyrosine residues, while inhibiting other pro-angiogenic tyrosine sites. .Indeed, a recent review concludes that VEGF-A165b ultimately acts as a less efficacious agonist, and only at some residues15. From this evidence, one can speculate that VEGF-A165b acts with functional selectivity, more commonly known as biased agonism. This is a common phenomenon in G-coupled protein receptor signaling whereby different ligands, binding the same receptor, promote activation of different downstream signaling pathways58,59,60. Therefore, under biased agonism, VEGF-A165b may act as an agonist at certain tyrosine residues while acting antagonistically at others.
Conclusion
Alternative splicing produces VEGF-A variants with different binding partners, and as we show for the first time, different binding kinetics. As recent evidence has pointed to important roles the pro- and anti-angiogenic variants can play in human pathology, it is crucial for researchers to not only have access to differences in their expression, but these differences we have identified in their binding kinetics. Cell signaling systems, like the VEGF-A-axis regulating angiogenesis, are complex: receptors activate a multitude independent and overlapping downstream pathways, with dozens of growth factors binding receptors, either activating, inhibiting, or modulating their downstream action. Deconvolving the contributions of any one molecule requires the experimental tools provided by systems biology. To this end, further work remains, in characterizing the binding, receptor activation, and downstream biological effect produced by other ‘anti-angiogenic’ VEGF-Axxxb splice variants, such as VEGF-A121b—which have hitherto received less investigation12,15,61, as well as measuring differences in splice variant binding to the ECM, heparin sulfates chains, and their associated proteoglycans, and co-receptors like neuropilins17,46. By having a more complete picture of the VEGF-A splice variants will we be able to advance more effective therapeutic interventions for angiogenesis-related pathologies.
Materials and methods
Surface plasmon resonance kinetic studies with dextran-coated gold sensors
SPR studies were performed with the BIAcore 3,000 instrument (Biacore International AB, Uppsala, Sweden) at 25 °C on dextran-coated gold sensor chips (CM5, Research grade, GE Healthcare Bio-sciences AB, Uppsala) following a previously established procedure30. The BIAcore 3,000 divides CM5 sensor chips into four separate flow cells. We immobilized a different receptor protein in each flow cell: The first cell was reserved for measuring non-specific binding by immobilizing recombinant angiopoietin-4 (Cat. #964-AN-025/CF, R&D Systems) to a flow cell: it has no known interaction with VEGF-A splice variants. VEGFR1 and VEGFR2 were immobilized in two of the remaining flow cells. Running buffer: 1 × HBS-EP pH 7.4 (10 mM HEPES, 3 mM EDTA, 150 mM NaCl, 0.005% TWEEN-20, cat. # BR100188, GE Life Sciences).
Protein immobilization
Pre-concentration studies were performed to determine optimal pH conditions for protein immobilization to ensure target levels of immobilized protein can be achieved precisely and to conserve protein. 10 mM acetate buffers were prepared ranging from pH 3.5 to 0.5–1.0 below the protein’s isoelectric point (Supplementary Table S1). Receptor solutions were prepared at 20 µg/mL with each acetate buffer. We injected 20 µL of each pH solution at a flow rate of 5 µL/min, followed by a 5-µL injection of ethanolamine-HCL (GE Healthcare AB, Uppsala, Sweden) to clear the surface. We selected the optimal acetate buffer pH for each protein based on (1) the maximum level of protein immobilization reached and (2) the rate of immobilization observed in the pre-concentration study sensograms.
Recombinant VEGFR1 (Recombinant Human VEGF R1/Flt-1 Fc Chimera Protein, CF, Cat. #321-FL-050/CF, R&D Systems) and VEGFR2 (Recombinant Human VEGF R2/KDR Fc Chimera Protein, CF, Cat. #357-KD-050/CF, R&D Systems) were immobilized irreversibly to separate flow cells via amine coupling with the dextran matrix. Additionally, we immobilized angiopoietin-4 (Recombinant Human Angiopoietin-4 Protein, CF, Cat. #964-AN-025/CF, R&D Systems), a protein with no known interaction with any VEGF-A splice variant, to a flow cell as a reference signal protein to subtract the effects of non-specific interactions. The surface was activated by injecting 35 μL of a 1:1 volumetric mixture of 0.05 M NHS (N-hydroxysuccinimide, GE Healthcare AB) and 0.2 M EDC (1-ethyl-3-(3-dimethylaminopropyl) carbodiimide hydrochloride GE Healthcare AB) at a flow rate of 5 μL/min. Each receptor was dissolved in 20 µg/mL of the 10 mM acetate buffer at its optimal pH and injected at 5 µL/min until the target level was reached (approximately 200–500 R.U. of immobilized receptor, preventing quick receptor saturation at injected ligand concentrations.) After sufficient protein was coupled, the surface was de-activated by injecting 35 μL ethanolamine (ethanolamine hydrochloride-NaOH pH 8.5, GE Healthcare AB).
Ligand-receptor kinetics measurements
Fresh ligand solutions were prepared across a range of ligand concentrations in HBS-EP running buffer—10 nM, 20 nM, and 40 nM—each experimental day, including: human recombinant VEGFA (R&D Systems, Cat. #293-VE-010), VEGFA121, and VEGFA165b. These concentration ranges were selected such that the injected concentrations exceeded the immobilized receptor concentrations, which enables these experiments to best reflect the physiological reality, and follow the configuration of recent computational models27. 120 μL of each ligand solution was injected into flow cells containing immobilized receptor and Ang-4—a reference for non-specific binding—at 30 μL/min (association). This was followed by a 10 min running-buffer injection (dissociation). Between each sample, we injected, in series, 5 μL of 5 mM HCl and 5 μL of 10 mM NaOH at 5 μL/min to remove any remaining bound ligand. We repeated this cycle for each concentration tested (40, 20, and 10 nM). Each concentration series were performed in triplicate. All recombinant human cytokines were obtained from R&D Systems.
Kinetic analysis
The raw ligand:receptor sensograms were aligned and the background, non-specific binding was subtracted using the sensogram trace from the ligand:Ang-4 flow cell. Both the raw ligand:receptor and ligand:Ang-4 sensogram curves were obtained within the BIAcore 3,000’s detection window (10–70,000 R.U.) for all interactions62 to ensure detected interactions did not represent system noise. BIA evaluation removes momentary signal spikes resulting from transient air bubbles.
Global analysis is considered to produce more accurate results than fitting of a single response curve, so global fitting was performed with BIAevaluation software (Version 4.1.1, GE Healthcare) following a 1:1 Langmuir adsorption isotherm (Eq. 1)63. The software applies nonlinear least squares analysis to determine association (ka) and dissociation (kd) rates fitting best to multiple response curves simultaneously. Additionally, the software provides the goodness-of-fit parameter χ2 and the peak magnitude of the signal response, Rmax.
Classifying binding
Both the instrument manufacturer (BIAcore) and previous researchers have suggested that when fitting kinetic rate constants using global analysis, a χ2-to-Rmax value (a measure of noise-to-signal) < 0.2 is ideal for confidence in the kinetic parameters obtained when studying known interactions64,65,66. A low noise-to-signal indicates that the sensogram signal includes minimal contributions from the following three noise-factors: (1) overall instrument noise, (2) heterogeneities in immobilized receptor or ligand, and (3) non-specific interactions. We applied a cut-off value we established previously30 of χ2-to-Rmax < 1.0 that differentiates real, 1:1 Langmuir interactions, from predominantly non-specific interactions, where χ2-to-Rmax > 1.0 to confirm that detected interactions represented true receptor binding. (χ2-to-Rmax values obtained are summarized in Supplementary Table S2).
References
Koch, S., Tugues, S., Li, X., Gualandi, L. & Claesson-Welsh, L. Signal transduction by vascular endothelial growth factor receptors. Biochem. J. 437, 169–183 (2011).
Woolard, J., Bevan, H. S., Harper, S. J. & Bates, D. O. Molecular diversity of VEGF-A as a regulator of its biological activity. Microcirculation 16, 572–592 (2009).
Holmes, D. I. R. & Zachary, I. The vascular endothelial growth factor (VEGF) family: angiogenic factors in health and disease. Genome Biol. 6, 209 (2005).
Li, E. & Hristova, K. Receptor tyrosine kinase transmembrane domains: function, dimer structure and dimerization energetics. Cell Adhes. Migr. 4, 249–254 (2010).
Sarabipour, S., Ballmer-Hofer, K. & Hristova, K. VEGFR-2 conformational switch in response to ligand binding. Elife 5, e13876 (2016).
Roskoski, R. & Roskoski, R. Jr. VEGF receptor protein-tyrosine kinases: structure and regulation. Biochem. Biophys. Res. Commun. 375, 287–291 (2008).
Simons, M., Gordon, E. & Claesson-Welsh, L. Mechanisms and regulation of endothelial VEGF receptor signalling. Nat. Rev. Mol. Cell Biol. 10, 611 (2016).
Schlessinger, J. Receptor tyrosine kinases: legacy of the first two decades. Cold Spring Harb. Perspect. Biol. 6, a008912 (2014).
Mac Gabhann, F. & Popel, A. S. Systems biology of vascular endothelial growth factors. Microcirculation 15, 715–738 (2008).
Robinson, C. J. & Stringer, S. E. The splice variants of vascular endothelial growth factor (VEGF) and their receptors. J. Cell Sci. 114, 853–865 (2001).
Woolard, J. et al. VEGF 165b, an inhibitory vascular endothelial growth factor splice variant: mechanism of action, in vivo effect on angiogenesis and endogenous protein expression. Cancer Res. 64, 7822–7835 (2004).
Delcombel, R. et al. New prospects in the roles of the C-terminal domains of VEGF-A and their cooperation for ligand binding, cellular signaling and vessels formation. Angiogenesis 16, 353–371 (2013).
Finley, S. D. & Popel, A. S. Predicting the effects of anti-angiogenic agents targeting specific VEGF isoforms. AAPS J. 14, 500 (2012).
Cébe Suarez, S. et al. A VEGF-A splice variant defective for heparan sulfate and neuropilin-1 binding shows attenuated signaling through VEGFR-2. Cell. Mol. Life Sci. 63, 2067–2077 (2006).
Peach, C. J. et al. Molecular pharmacology of VEGF-A isoforms: binding and signalling at VEGFR2. Int. J. Mol. Sci. 19, 1264 (2018).
Boucher, J. M. & Bautch, V. L. Antiangiogenic VEGF-A in peripheral artery disease. Nat. Med. 20, 1383–1385 (2014).
Chu, L.-H. et al. A multiscale computational model predicts distribution of anti-angiogenic isoform VEGF165b in peripheral arterial disease in human and mouse. Sci. Rep. 6, 37030 (2016).
Harper, S. J. & Bates, D. O. VEGF-A splicing: the key to anti-angiogenic therapeutics?. Nat. Rev. Cancer 8, 880–887 (2008).
Weddell, J. C. & Imoukhuede, P. I. Quantitative characterization of cellular membrane-receptor heterogeneity through statistical and computational modeling. PLoS ONE 9, e97271 (2014).
Vempati, P., Popel, A. S. & Mac Gabhann, F. Formation of VEGF isoform-specific spatial distributions governing angiogenesis: computational analysis. BMC Syst. Biol. 5, 59 (2011).
Finley, S. D. et al. Effect of tumor microenvironment on tumor VEGF during anti-VEGF treatment: systems biology predictions. J. Natl. Cancer Inst. 105, 802–811 (2013).
Finley, S. D., Engel-Stefanini, M. O., Imoukhuede, P. I. & Popel, A. S. Pharmacokinetics and pharmacodynamics of VEGF-neutralizing antibodies. BMC Syst. Biol. 5, 193 (2011).
Finley, S. D., Angelikopoulos, P., Koumoutsakos, P. & Popel, A. S. Pharmacokinetics of anti-VEGF agent aflibercept in cancer predicted by data-driven, molecular-detailed model. CPT Pharmacomet. Syst. Pharmacol. 4, 641–649 (2015).
Stefanini, M. O. et al. The presence of VEGF receptors on the luminal surface of endothelial cells affects VEGF distribution and VEGF signaling. PLoS Comput. Biol. 5, e1000622 (2009).
Mac Gabhann, F., Ji, J. W. & Popel, A. S. Multi-scale computational models of pro-angiogenic treatments in peripheral arterial disease. Ann. Biomed. Eng. 35, 982–994 (2007).
Mac Gabhann, F. & Popel, A. S. Dimerization of VEGF receptors and implications for signal transduction: a computational study. Biophys. Chem. 128, 125–139 (2007).
Clegg, L. E., Ganta, V. C., Annex, B. H. & Mac Gabhann, F. Systems pharmacology of VEGF165b in peripheral artery disease. CPT Pharmacomet. Syst. Pharmacol. 6, 833–844 (2017).
Salamon, Z. & Tollin, G. Surface plasmon resonance. Theory 3, 2311–2319 (1999).
Homola, J., Yee, S. S. & Gauglitz, G. Surface plasmon resonance sensors: review. Sens. Actuators B Chem. 54, 3–15 (1999).
Mamer, S. B. et al. Discovery of high-affinity PDGF-VEGFR interactions: redefining RTK dynamics. Sci. Rep. https://doi.org/10.1038/s41598-017-16610-z (2017).
Von Tiedemann, B. & Bilitewski, U. Characterization of the vascular endothelial growth factor-receptor interaction and determination of the recombinant protein by an optical receptor sensor. Biosens. Bioelectron. 17, 983–991 (2002).
Ladomery, M. R., Harper, S. J. & Bates, D. O. Alternative splicing in angiogenesis: the vascular endothelial growth factor paradigm. Cancer Lett. 249, 133–142 (2007).
Cunningham, S. A., Tran, T. M., Arrate, M. P. & Brock, T. A. Characterization of vascular endothelial cell growth factor interactions with the kinase insert domain-containing. J. Biol. Chem. 274, 18421–18427 (2000).
Teran, M. & Nugent, M. A. Characterization of receptor binding kinetics for vascular endothelial growth factor-A using SPR. Anal. Biochem. 564–565, 21–31 (2019).
Haseley, S. R., Talaga, P., Kamerling, J. P. & Vliegenthart, J. F. Characterization of the carbohydrate binding specificity and kinetic parameters of lectins by using surface plasmon resonance. Anal. Biochem. 274, 203–210 (1999).
Bates, D. O. et al. VEGF165b, an inhibitory splice variant of vascular endothelial growth factor, is down-regulated in renal cell carcinoma. Cancer Res. 62, 4123–4131 (2002).
Dokun, A. O. & Annex, B. H. The VEGF165b ‘ICE-o-form’ puts a chill on the VEGF story. Circ. Res. 109, 246–247 (2011).
Finley, S. D., Dhar, M. & Popel, A. S. Compartment model predicts VEGF secretion and investigates the effects of VEGF trap in tumor-bearing mice. Front. Oncol. 3, 196 (2013).
Carmeliet, P. Angiogenesis in life, disease and medicine. Nature 438, 932–936 (2005).
Qutub, A., Gabhann, F., Karagiannis, E., Vempati, P. & Popel, A. Multiscale models of angiogenesis. IEEE Eng. Med. Biol. Mag. 28, 14–31 (2009).
Krishna, S. M., Moxon, J. V. & Golledge, J. A review of the pathophysiology and potential biomarkers for peripheral artery disease. Int. J. Mol. Sci. 16, 11294–11322 (2015).
Manetti, M. et al. Overexpression of VEGF165b, an inhibitory splice variant of vascular endothelial growth factor, leads to insufficient angiogenesis in patients with systemic sclerosis. Circ. Res. 109, e14–e26 (2011).
Ngo, D. T. M. et al. Antiangiogenic actions of vascular endothelial growth factor-A165b, an inhibitory isoform of vascular endothelial growth factor-A, in human obesity. Circulation 130, 1072–1080 (2014).
Guyot, M. & Pagès, G. VEGF splicing and the role of VEGF splice variants: from physiological-pathological conditions to specific pre-mRNA splicing. Signal. Pathw. Liver Dis. 1332, 3–23 (2015).
Ikeda, S. et al. Novel role of ARF6 in vascular endothelial growth factor-induced signaling and angiogenesis. Circ. Res. 96, 467–475 (2005).
Ganta, V. C., Choi, M., Kutateladze, A. & Annex, B. H. VEGF165b modulates endothelial VEGFR1-STAT3 signaling pathway and angiogenesis in human and experimental peripheral arterial disease. Circ. Res. 120, 282–295 (2017).
Chen, S., Guo, X., Imarenezor, O. & Imoukhuede, P. I. Quantitation of VEGFRs, NRP1, and PDGFRs on endothelial cells and fibroblasts reveals serum, intra-family ligand, and cross-family ligand regulation. Cell Mol. Bioeng. 8, 383–403 (2015).
Imoukhuede, P. I. & Popel, A. S. A. S. Quantification and cell-to-cell variation of vascular endothelial growth factor receptors. Exp. Cell Res. 317, 955–965 (2011).
Sarrazin, S., Lamanna, W. C. & Esko, J. D. Heparan sulfate proteoglycans. Cold Spring Harb. Perspect. Biol. 3, 1–33 (2011).
Chen, S. et al. qFlow cytometry-based receptoromic screening: a high-throughput quantification approach informing biomarker selection and nanosensor development. In Biomedical Nanotechnology: Methods and Protocols (eds Petrosko, S. H. & Day, E. S.) 117–138 (Springer, New York, 2017). https://doi.org/10.1007/978-1-4939-6840-4_8.
Imoukhuede, P. I. et al. Pharmacokinetics and pharmacodynamics of VEGF-neutralizing antibodies. Am. J. Physiol. Heart Circ. Physiol. 5, H1085–H1093 (2013).
Mac Gabhann, F. & Popel, A. S. Targeting neuropilin-1 to inhibit VEGF signaling in cancer: comparison of therapeutic approaches. PLoS Comput. Biol. 2, 1649–1662 (2006).
Mamer, S. B., Palasz, A. A. & Imoukhuede, P. I. Mapping tyrosine kinase receptor dimerization to receptor expression and ligand affinities. Processes 7, 288 (2019).
Kawamura, H., Li, X., Harper, S. J., Bates, D. O. & Claesson-Welsh, L. Vascular endothelial growth factor (VEGF)-A165b is a weak in vitro agonist for VEGF receptor-2 due to lack of coreceptor binding and deficient regulation of kinase activity. Cancer Res. 68, 4683–4692 (2008).
Clegg, L. W. & Mac Gabhann, F. Site-specific phosphorylation of VEGFR2 is mediated by receptor trafficking: insights from a computational model. PLoS Comput. Biol. 11, e1004158 (2015).
Evans, I. VEGF Signaling. Methods in Molecular Biology Vol. 1332 (Springer, New York, 2015).
Weddell, J. C., Chen, S. & Imoukhuede, P. I. VEGFR1 promotes cell migration and proliferation through PLCγ and PI3K pathways. NPJ Syst. Biol. Appl. 4, 1 (2018).
Rajagopal, S. et al. Quantifying ligand bias at seven-transmembrane receptors. Mol. Pharmacol. 80, 367–377 (2011).
Kenakin, T., Watson, C., Muniz-Medina, V., Christopoulos, A. & Novick, S. A simple method for quantifying functional selectivity and agonist bias. ACS Chem. Neurosci. 3, 193–203 (2012).
Yamazaki, Y. & Morita, T. Molecular and functional diversity of vascular endothelial growth factors. Mol. Divers. 10, 515–527 (2006).
Dehghanian, F., Hojati, Z. & Kay, M. New insights into VEGF-A alternative splicing: key regulatory switching in the pathological process. Avicenna J. Med. Biotechnol. 6, 192–199 (2014).
GE Healthcare Life Sciences. Biacore Assay Handbook. GE Healthc. Bio-Sciences AB 1–78 (2012).
Roden, L. D. & Myszka, D. G. Global analysis of a macromolecular interaction measured on BIAcore. Biochem. Biophys. Res. Commun. 225, 1073–1077 (1996).
Karlsson, R. & Fält, A. Experimental design for kinetic analysis of protein-protein interactions with surface plasmon resonance biosensors. J. Immunol. Methods 200, 121–133 (1997).
Murphy, M., Jason-Moller, L. & Bruno, J. Using Biacore to measure the binding kinetics of an antibody-antigen interaction. Curr. Protoc. Protein Sci. Editor. Board John E Coligan al Chapter 19, Unit 19.14–Unit 19.14 (2006).
Biacore, A. B., July, E. & Biacore, A. B. BIA Evaluation Software Handbook. System (Biacore AB, 1997).
Acknowledgements
We thank Dr. Brian Imai for his immeasurably-valuable guidance towards operating the BIAcore 3000, and for allowing us to utilize it so extensively.
Author information
Authors and Affiliations
Contributions
P.I., S.M., and A.W. conceived of the studies. S.M. and A.W. designed the experiments. S.M. and A.W. collected the data and performed the analyses. S.M. and P.I. drafted the manuscript, and all authors contributed to its revision. All authors have read and have approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mamer, S.B., Wittenkeller, A. & Imoukhuede, P.I. VEGF-A splice variants bind VEGFRs with differential affinities. Sci Rep 10, 14413 (2020). https://doi.org/10.1038/s41598-020-71484-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-71484-y
This article is cited by
-
How VEGF-A and its splice variants affect breast cancer development – clinical implications
Cellular Oncology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.