Abstract
The Heterobasidion annosum s.l species complex comprises the most damaging forest pathogens to Norway spruce. We revisited previously identified Quantitative Trait Loci (QTLs) related to Heterobasidion-resistance in Norway spruce to identify candidate genes associated with these QTLs. We identified 329 candidate genes associated with the resistance QTLs using a gene-based composite map for Pinaceae. To evaluate the transcriptional responses of these candidate genes to H. parviporum, we inoculated Norway spruce plants and sequenced the transcriptome of the interaction at 3 and 7 days post inoculation. Out of 298 expressed candidate genes 124 were differentially expressed between inoculation and wounding control treatment. Interestingly, PaNAC04 and two of its paralogs in the subgroup III-3 of the NAC family transcription factors were found to be associated with one of the QTLs and was also highly induced in response to H. parviporum. These genes are possibly involved in the regulation of biosynthesis of flavonoid compounds. Furthermore, several of the differentially expressed candidate genes were associated with the phenylpropanoid pathway including a phenylalanine ammonia-lyase, a cinnamoyl-CoA reductase, a caffeoyl-CoA O-methyltransferase and a PgMYB11-like transcription factor gene. Combining transcriptome and genetic linkage analyses can help identifying candidate genes for functional studies and molecular breeding in non-model species.
Similar content being viewed by others
Introduction
One of the most pressing challenges to the forest industry is facing the increasing damages caused by forest pests and pathogens1,2. Although good management practices can reduce the problems3,4, examining and utilizing the genetic resistance, through forest tree breeding, offers an additional powerful method to reduce damages1,3.
Norway spruce [Picea abies (L.) Karst.] is an important conifer species for the forest industry in Europe2. For instance, it constitutes 41% of the standing Swedish tree volume5,6. The major pathogen, in economic terms2, on Norway spruce is the species complex Heterobasidion annosum sensu lato (s.l.), which causes stem and root-rot in Europe7,8. Although new airborne infections of H. annosum s.l. can be reduced by good forest management and stump treatment with the biocontrol agent Phlebiopsis gigantea9, carryover between rotations in already infected sites is an important factor for the health of the new generation trees10. Thus, replantation of plant material with improved resistance would be a valuable resource in the management of H. annosum s.l. in Norway spruce.
Genetic variation for resistance to H. annosum s.l. exists in Norway spruce11,12,13,14,15,16 and there are no adverse correlations between resistance to Heterobasidion infection and growth or wood quality traits12,13,16. Hence, selection for resistance to H. annosum s.l. in breeding programs could lead to considerable gain12, without compromising other breeding achievements. Resistance to wood rotting pathogens, such as H. annosum s.l. in conifers, presents a challenge to tree breeders as the disease often manifests late in the tree’s life, which makes phenotypic selection difficult and time consuming. Marker Assisted Selection (MAS) holds promises for increasing the gain in tree breeding and specifically in resistance breeding17. Yet, MAS has not been widely implemented within tree breeding programs mainly due to the difficulty in translating Quantitative Trait Loci (QTLs) analysis into operational MAS i.e. validation of potential markers17,18. Recently, one of the marker candidates, PaLAR3 was identified in the first Quantitative Trait Locus (QTL) analysis for resistance against H. parviporum (Fr.) Niemelä & Korhonen14,15, which is a member of the H. annosum s.l. complex and lives almost exclusively in Norway spruce19. The validation of PaLAR3, a gene that encodes for a leucoanthocyanidin reductase that is involved in the flavonoid-biosynthetic pathway20, was done through the integration of information of phenotypic, transcriptional, metabolic, and genetic evidence15. Similarly, two other reported markers that are ready to be used for MAS of trees with improved resistance to pathogens and pests in conifers21,22; build on the integration of genetic information and complementing techniques.
The QTL analysis in Norway spruce for resistance to H. parviporum identified 13 QTLs linked with four traits related to host resistance14: lesion length at the inoculation site (LL), exclusion of the pathogen from the host after initial infection (E), infection prevention from establishing at all (IP) and fungal spread within the sapwood (SWG). The validated marker, PaLAR3, comprised one of the four QTLs associated with SWG14,15. In this study, we aimed to use the high degree of synteny and macrocollinearity within Pinaceae to identify novel candidate resistance genes associated with the QTLs for resistance to H. parviporum14. We used the composite map of the Pinaceae family and gene expression patterns in Norway spruce after challenging it with H. parviporum; therefore, we could identify candidate resistance genes for future validation and functional analyses.
First, we used a Pinaceae composite linkage map to identify additional Norway spruce candidate genes associated with already described resistance QTLs. Second, we predicted that the combination of genetic and transcriptional information, would enable us to identify candidate resistance genes of induced response for future analyses; allowing us to hypothesize that the genes in the QTL regions, which are likely to be important for controlling spread of H. parviporum, are also likely to be more strongly regulated by inoculation than by wounding alone. Previous studies have highlighted broad similarities in defence responses to H. parviporum infection and to wounding control compared to naïve material, although inoculated samples showed a heightened response23,24.
The composite map of the Pinaceae family25 integrated published maps of Norway spruce14, Picea glauca and Picea mariana26 and Pinus taeda27 with genetic maps from multiple crosses of Pinus pinaster. QTL markers from the Norway spruce linkage map14 were included in the Pinaceae composite map25. As the composite map is considerably denser than the Norway spruce map we chose to focus on markers in and around a significant QTLs for resistance identified by Lind et al.14 to identify candidate genes in the Pinaceae composite map25. This summed up to 329 candidate genes associated with 12 genomic regions of Norway spruce. We further determined the transcriptional responses of these candidate genes at three and seven days after inoculation with H. parviporum. We detected 124 differentially expressed candidate resistance genes, including two putative NAC family transcription factor genes in response to H. parviporum inoculation compared to the control treatment of mechanical wounding.
Materials and methods
Identification of conifer candidate genes associated with QTL regions from a composite map
The markers in the Norway spruce QTL map14 were integrated in the Pinaceae composite map25. The markers in the Norway spruce QTL map14 were used to identify the corresponding QTL regions in the Pinaceae composite map25. The markers associated with the QTL regions were identified around a significant QTL LOD-peak which contained significant markers (P value < 0.05) according to the Kruskal–Wallis test in Norway spruce linkage map14. The candidate genes between significant markers in the confidence interval, in the Pinaceae composite map were identified and selected as confidence interval candidate genes (CCGs) for future analyses (Supplementary file1). The candidate genes in between the next subsequent markers outside the confidence interval were also selected. These genes were referred as putative candidate genes (PCGs) (Supplementary file1). We chose to include PCGs because all the markers used in the Norway spruce QTL map14 were not included in the Pinaceae composite map25. For some QTL regions this discripancy made it difficult to delineate the QTL region, thus the genes in the category represent genes which are suspected to associate with the QTL region. Therefore, differentially expressed PCGs could still be interesting in this experimental set up as a part of an induced defence system.
Identification of Norway spruce candidate resistance genes associated with QTL regions
The FASTA sequences of the unigenes derived markers corresponding to genes in regions of the genome associated with the resistance QTLs were downloaded from the P. pinaster unigene catalogue (https://www.scbi.uma.es/sustainpinedb/unigens) and the most probable Norway spruce orthologues were identified by a blastN query (E-value cutoff:1e-3) in the Norway spruce gene catalogue (Pabies v1.0, www.congenie.org), excluding low confidence candidate genes with less than 30% coverage.
Norway spruce materials
For the RNAseq study, six 7-year-old rooted cuttings of each of the genotypes S21K0220126 and S21K0220184, originating from a well-studied full-sib family (S21H9820005) of Norway spruce11,14,23, were used. The cuttings were grown in a greenhouse with an 18 h light regime. Watering and nutrients were supplied twice a week. The Real-Time Quantitative Reverse Transcription PCR (qRT-PCR) validation experiment included cuttings of six genotypes (S21K0220263, S21K0220240, S21K0220237, S21K0220161, S21K022136 and S21K022346) from the same Norway spruce full-sib family23.
Inoculation experiment
Branches were artificially inoculated with H. parviporum (isolate Rb175) as previously described23. The same isolate was used to generate the QTL map in Norway spruce14. Briefly, branches were wounded with a five-mm cork borer and wooden plugs covered with mycelium from H. parviporum were attached to the wound with Parafilm®; control branches on the tree were also wounded and a sterile wooden plug was attached and sealed with Parafilm.
Based on the difference in necrotic lesion length (LL) extension from the inoculation site after inoculation with H. parviporum the genotypes S21K0220126 and S21K0220184 (short and long, respectively, data not shown) were selected for RNA sequencing. For the RNAseq study, bark and phloem samples were harvested at three and seven days post-inoculation (dpi). At the time of harvest, bark surrounding the wounds and inoculation sites was cut into two sections and samples were collected at the inoculation site (A) 0–0.5 cm around the wound, and distal to the inoculation site (C) 1.0–1.5 cm. We used six ramets per clone and three inoculations per twig were done. The bark samples were frozen separately in liquid nitrogen and stored at − 80˚C until further use. For the qRT-PCR study sampling is described in detail elsewhere23, briefly one bark and phloem sample was taken for each treatment and time point from six separate full-sib genotypes.
RNA extraction, transcriptome sequencing and qRT-PCR
RNA extraction
Total RNA was isolated by using a modified CTAB extraction protocol28. The samples were treated with DNase I (Sigma-Aldrich) to eliminate contamination of genomic DNA. The RNA integrity was analysed by using the Agilent RNA 6,000 Nano kit (Agilent Technologies Inc.).
Transcriptome sequencing and bioinformatics analyses
Three biological replicates of clones S21K0220126 and S21K0220184 per treatment were used for Illumina sequencing. Sequencing libraries were prepared at the SNP&SEQ Technology Platform (SciLifeLab, Uppsala) using the TruSeq stranded mRNA sample preparation kit according to the manual TruSeq stranded mRNA sample preparation guide. Sequencing was done using HiSeq 2,500, paired-end 125 bp read length, v4 sequencing chemistry. The raw sequences were submitted to the Sequence Read Archive (SRA) portal (NCBI) under BioProject accession number PRJNA522265.
RNAseq analyses were performed with the Tophat-Cufflinks pipeline as previously described29. Briefly, Nesoni clip 0.97 (https://github.com/Victorian-Bioinformatics-Consortium/nesoni) was used to filter adaptors and low-quality bases. Illumina reads were filtered based on phred-scale with a quality score cut-off of 20, minimum adapter length match of 20, with maximum errors of one in the adaptor and reads shorter than 35 were discarded. A Bowtie reference from the ‘Pabies1.0-all-cds.fna’ was constructed, downloaded from the Norway spruce genome portal (https://congenie.org/) using Bowtie2 version 2.2.4 (https://bowtie-bio.sourceforge.net/bowtie2/index.shtml) to enable alignments to a reference database. The filtered read pairs were aligned to ‘Pabies1.0-all-cds’ reference gene model with Tophat version 2.0.1330. Cufflinks version 2.2.1 was used to assemble all transcripts of each sample with the results of the alignment from TopHat. Cuffmerge included in the cufflinks package was used to merge all assemblies. Cuffquant (https://coletrapnell-lab.github.io/cufflinks/manual/) calculated transcript abundance from the single assembly of the sample, and the aligned read files produced by the Tophat output were run separately for each sample. Cuffdiff was used for differential expression analysis using default settings30,31.
qRT-PCR
One µg of total RNA was reverse transcribed to cDNA with the iScript cDNA Synthesis Kit (Bio-Rad) in a total reaction volume of 20 μl according to the manufacturer’s instructions. A ten-fold dilution of the cDNA was stored at − 20 °C. cDNA equivalent to 25 ng of total RNA worked as template for each PCR reaction, using SSoFast EVAGreen Supermix (Bio-Rad). Primers for candidate genes were designed within the exons of the predicted candidate genes in the P. abies v 1.0 release using Primer3 software32 with a melting temperature (Tm) between 60 °C and 61 °C. A final concentration of 0.15 μM of each primer (Supplementary Table 1) was used. The thermal-cycling condition parameters, ran on an iQ™5 Multicolor Real-Time PCR Detection system (Bio-rad) using the following cycling parameters: 98 °C for 2 min; 40 cycles of 98 °C for 5 s, 60 °C for 10 s. A melt-curve analysis followed the qRT-PCR reactions, to confirm that the signal was the result of a single product amplification. Primer amplification efficiency was determined by amplification of serial dilutions of cDNA from Norway spruce with PCR conditions described above. The relative expression was calculated from threshold cycle values (Ct) using the 2ΔΔCT-method33. Transcript abundance was normalized to the reference genes eukaryotic translation initiation factor 4A (elF4A)34 and elongation factor 1-α (ELF1α)23. The gene expression experiments were done with six biological and two technical replicates. Gene expression data were analysed by analysis of variance (ANOVA) using a general linear model approach implemented in R-program (https://www.r-project.org/).
Results
Markers and candidate genes associated with resistance QTLs
To identify additional Norway spruce unigene derived SNPs markers in the already described resistance QTLs, we identified 369 P. pinaster unigene derived SNPs markers in the composite map25 based on markers in the QTLs in Norway spruce14 in the composite map25 (Table 1). The P. pinaster unigene derived SNPs markers were used to query the Norway spruce gene catalogue. A total of 329 candidate genes previously not known to associate with the resistance QTLs were successfully identified, of which 83 were CCGs in between the significant markers within the confidence interval and 246 were PCGs in between the subsequent markers outside the confidence interval in Pinaceae composite map25 (Table 1). Twelve out of the original 13 QTLs were identified, and only the QTL for LL on linkage group (LG) 9 could not be identified in the composite map (Table 1).
Transcriptome analysis
The Illumina HiSeq sequencing generated 14.2–17.8 M reads per sample that passed Illumina’s quality control and between 12.6 M and 16.4 M read-pairs were kept after Nesoni filtering (Supplementary file 2). The read mapping rate from Tophat was 44.3%–34.9% (Supplementary file 3). Depending on the contrast, between 4,401 and 8,767 genes were significantly differentially expressed (Supplementary Table 2). The fractions of induced and repressed genes were similar in all comparisons. Of the 329 candidate genes selected in the Pinaceae composite map25, 298 were expressed (80 in CCG and 218 in PCG categories respectively), in at least one of the treatments in the RNAseq experiment (Supplementary Table 3). The candidate genes showed differentially expression at three and seven dpi after H. parviporum inoculation compared to wounding, of which 41 were differentially expression in CCG and 83 in PCG categories in the treatments in RNAseq experiment (Fig. 1, Supplementary Table 3).
Heat map of the 124 differentially expressed candidate genes associated with Heterobasidion-resistance QTLs in Norway spruce in response to H. parviporum. The red to yellow colours indicate the highest to lowest FPKM (fragments per kilo base of exon model per million reads mapped) values at 3 and 7 dpi at the proximal (A) and distal (C) sampling sites. The red-grey-blue scale shows the highest to lowest Fold change (FC, log2 values) in H. parviporum inoculation compared to wounding alone. Bold font indicate candidate genes corresponding to the original QTL on the Linkage groups, “E” stands for exclusion, “IP” stands for infection prevention, “LL” stands for lesion length and “SWG” stands for fungal growth in sapwood. Asterisks (*) indicates subgroup III-3 NAC TFs. cM indicates the position in centiMorgans of the marker in the Pinaceae composite map.
DEGs in QTLs associated with infection prevention
The QTLs associated with IP positioned on LG1, LG2 and LG1114 were identified in the composite map. Twelve expressed and six differentially expressed candidate genes (DEGs) associated with the QTL on LG1 (Fig. 1, Supplementary file 4). Four were upregulated adjacent to the inoculation site and two DEGs were downregulated at three dpi proximal to the inoculation site compared to wounding (Fig. 1). Most expression changes were quite moderate at LG1 with maximum log2 fold change of 1.75.
Fifteen expressed genes were associated with IP on LG2 (Supplementary file 4). Twelve of these were classified as CCGs and three as PCGs. None of PCGs were differentially expressed (Supplementary file 4). Four DEGs were associated with IP on LG2 were identified as CCGs (Fig. 1, Supplementary file 4). Two DEGs were upregulated at three dpi at the inoculation site, and one DEG was still upregulated at seven dpi. Again, most changes in expression patterns were quite moderate on LG2, except a putative glycosyltransferase gene (MA_186971g0010), an ortholog of the Arabidopsis protein UGT85A1 that encodes a UDP-Glycosyltransferase (UGT) protein. This gene has an interesting expression pattern, it is highly upregulated around the inoculation site at three and seven dpi, although its expression level was low (Fig. 1; Supplementary file 4). It should be mentioned that a second UGT like gene, MA_10436196g0010 was among the identified candidate genes. MA_10436196g0010 correspond to the original marker 0.276-A13-934 significantly associated with SWG on LG914, but this candidate gene was not differentially expressed (Table 2, Supplementary file 4).
Linkage group 11 associated with a QTL for IP contained a total of 29 expressed genes and 15 DEGs (Fig. 1; Supplementary file 4). Only five of the expressed genes were categorized as CCGs, three of these were differentially expressed. Interestingly, the PgMYB11-like (R2R3-MYB transcription factor PgMYB11-like) candidate gene MA_24271g0020/BT103501, associating with the QTL in the original study14, was highly upregulated at both three and seven dpi adjacent to the pathogen inoculation site (Fig. 1, Supplementary file 4). Likewise, the candidate gene MA_4742g0010 (PabiesFT1-1,251/BT115191 encoding spruce Mother of FT1, MFT1), was also positioned in this QTL in the original study. It was up-regulated in response to the inoculation at seven dpi even though its expression level was generally low (Fig. 1, Supplementary file 4). The candidate gene MA_6931g0010 encoding a putative caffeoyl-CoA O-methyltransferase (CCoAOMT) gene was highly expressed in all treatments and was differentially upregulated at seven dpi proximal to the inoculation site. It is noteworthy, that all three DEGs PgMYB11-like, Mother of FT1, MFT1, caffeoyl-CoA O-methyltransferase were identified as CCGs.
DEGs in QTLs associated with fungal sapwood growth
The QTLs for SWG were located on LG2, LG6 and LG9 in the original study14. Twenty five candidate genes associated with SWG QTLs were differentially expressed in our experiment. Four DEGs, associated with the marker BT105733, showed differential regulation found on LG2. Three DEGs were clearly downregulated adjacent to the inoculation site (Fig. 1; Supplementary file 4).
LG6 has two separate QTL regions for control of SWG (Fig. 1). The first region included six DEGs and comprises the marker BT105286, which corresponds to the Norway spruce candidate gene MA_10428976g0010 (PLAC8), which was moderately upregulated at three dpi at the inoculation site.
The second QTL region on LG6 included 10 DEGs, all categorized as CCGs (Supplementary file 4). The previously validated marker PaLAR3 (MA_176417g0010/ BT109050) is positioned in this QTL in the original linkage map14. However, the homologous marker (sp_v3.0_unigene4600) is positioned nearly 30 cM away from BT116508 in the composite map25. PaLAR3 was not expressed in this study. However, the candidate gene MA_10433955g0020, encoding the homolog of the original QTL marker BT116508 also associated with SWG in the original study, is highly but not differentially expressed in this study (Table 2, Supplementary file 4). The candidate gene MA_853405g0010 (hypothetical protein) upregulated proximal to the inoculation at seven dpi (and three dpi) in response to inoculation, but it was not expressed in the distal samples collected from the inoculation treatment at seven dpi (Fig. 1). MA_122748g0010 (a putative GRX, Glutaredoxin) also showed significant up-regulation proximal to the inoculation in our RNAseq analysis. None of the candidate genes associated with the SWG QTL were upregulated distal to the inoculation site (Fig. 1).
On LG9, we found 15 expressed candidate genes in the SWG QTL region (Supplementary file 4), five of these were also differentially expressed, including MA_19215g0010, the homolog of the original marker BT115393. All the DEGs in this region showed higher expression after wounding than after inoculation with H. parviporum (Fig. 1).
DEGs in QTLs associated with lesion length
The only original marker (02739-B22-309) we could identify with the QTL for lesion length in the phloem included in the Pinaceae composite map was located on LG8. We found 13 candidate genes that were expressed in this region, and eight of these were differentially expressed. All of the candidate genes except UDP-glucuronate 4-epimerase (MA_52380g0010), which was induced at seven dpi at site A and C respectively, were downregulated in response to inoculation (Fig. 1, Supplementary file 4). A Leucine-rich repeat protein kinase family protein gene (MA_17691g0010) was down regulated proximal to inoculation site at three and seven dpi. UDP-glucuronate 4-epimerase and Leucine-rich repeat protein kinase family protein gene were categorized as PCGs. It is noteworthy that a cinnamoyl-CoA reductase (MA_10435810g0010) was repressed at seven dpi at both proximal and distal to the inoculation. It is also the only candidate gene in CCG category (Fig. 1, Supplementary file 4).
DEGs in QTLs associated with exclusion of the H. parviporum from the Norway spruce
We could identify the QTLs associated with exclusion of the fungus from the host located on LG1, LG2, LG3 and LG614 in the composite map. On LG1 five DEGs were found and two DEGs of which were upregulated at seven dpi one at proximal and other at distal site (Fig. 1, Supplementary file 4).
On LG2 we found 16 DEGs associated with the QTL and only one of these is a CCG, which is the homolog of the original QTL marker (BT100742) MA_10431443g0010 a superoxide dismutase (Fig. 1, Table 2, Supplementary file 4). Both MA_10431443g0010 and MA_48816g0010 were downregulated proximal to the inoculation site at seven dpi. A Catalase (MA_10437148g0010) and Esterase family protein (MA_10435680g0010) were induced at seven dpi at proximal site and at distal site respectively (Fig. 1, Supplementary file 4). The candidate gene MA_28209g0010 (NAD(P)-binding Rossmann-fold superfamily protein) was upregulated proximal and distal to the inoculation site at both three and seven dpi. It was notable that candidate gene (MA_10436080g0010) dehydroquinate dehydratase, / shikimate dehydrogenase was induced in response to inoculation in all conditions tested (Fig. 1, Supplementary file 4).
There were three DEGs associated with the exclusion QTL on LG3. Interestingly, phenylalanine ammonia-lyase (PAL) (MA_15852g0010), an ortholog of the Picea glauca PgPAL3 (Genbank: BT119163)35 identified as PCG, was upregulated at both proximal and distal site; while the other two were downregulated proximal to inoculation, at both three and seven dpi (Fig. 1, Supplementary file 4).
Like with the second QTL for SWG on LG6, the exclusion QTL on LG6 had a large number of expressed CCGs (19) and 72 PCGs. Forty two of these candidate genes were also differentially expressed in at least one comparison (Fig. 1, Supplementary file 4). The orthologs (MA_14707g0010, MA_16728g0010 and MA_942991g0010) of the three markers associated with the original QTL, all showed differential expression in this study (Fig. 1). Interestingly enough, the QTL region for exclusion on LG6 showed three candidate genes encoding class III-3 NAC transcription factors: MA_264971g0010 (PaNAC04), MA_86256g0010 and MA_103386g0010. The candidate gene PaNAC04 and MA_103386g0010 were classified as CCGs. However, MA_86256g0010 was classified as a PCG. All three candidate genes were relatively highly expressed and showed clear induction, both proximal and distal, to the inoculation compared to wounding. This was especially noticeable at one week after the inoculation (Fig. 1), although, only PaNAC04 was differentially regulated in S21K0220184 distally from the inoculation site at seven dpi (Fig. 1, Supplementary file 4). This expression pattern together with the previously published phylogeny of the sub group III-3 NACs29 led us to investigate if the MA_103386g0010 candidate gene represents a different gene from the previously described PaNAC04. Additionally, an analysis of PaNAC04 (MA_264971g0010) and MA_103386g0010 expression by qRT-PCR in six well characterized Norway spruce genotypes23 showed that, on average PaNAC04 was not significantly differentially expressed. However, MA_103386g0010 was differentially expressed at seven dpi in response to inoculation (Table 3). Furthermore, expression of PaNAC04 was only detected in three of the genotypes (Fig. 2b, d) whereas MA_103386g0010 was expressed in all genotypes and treatments (Fig. 2a, c).
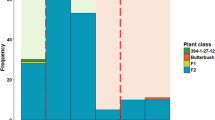
qRT-PCR of NAC candidate gene (MA_103386g0010 and PaNAC04) in bark of 7 year old genotypes of Norway spruce in response to wounded and inoculated with H. parviporum. The data have been normalized to the constitutive expressed genes Elongation-factor-1α and Elongation-factor-4 α. Different symbols with different colours represent six well characterized Norway spruce genotypes. (a) and (c) shows relative expression pattern of MA_103386g0010 at 3 and 7 dpi and (b) and (d) shows relative expression pattern of PaNAC04 at 3 and 7 dpi, in response to wounding and inoculation (N = 6).
Discussion
Throughout evolution, species in the Pinaceae have maintained a high degree of synteny and collinearity in their genomes25,26,36,37,38. In this study we have capitalized on this feature to expand the array of potential candidate genes associated with the resistance QTLs originally reported by Lind and co-workers14. As the composite map25 is considerably more dense than the original map, it may allow for identification of additional candidate genes associated with the reported QTL regions. Using shared markers associated with the QTL regions, identified around a significant LOD peak inside the confidence interval and subsequent markers outside the LOD peak in the Pinaceae composite map25, we found 329 potential Norway spruce candidates associated with 12 out of 13 original QTLs. Two hundred and ninety eight of the candidate genes were expressed, of which 80 were within confidence interval candidate genes (CCGs) and 218 putative candidate genes (PCGs). Out of these 41 CCGs and 83 PCGs were differentially expressed in the RNAseq study.
The products of the candidate genes could affect the resistance trait either as part of the constitutive or the induced defence39,40. Candidate genes involved in constitutive defence response are important for the host, but were not included in our current study. Compared to induced candidate genes associated with induced defences, whose involvement can be detected by transcriptional, protein or metabolite accumulation the contribution of constitutive defences are very difficult to quantify in short-term experiments. Therefore, we delibrately chose to focus on identifying candidate genes with a potential role in the induced defences in Norway spruce. It is reasonable to hypothesize that candidate genes associated with the resistance QTLs and also showing differential regulation in response to H. parviporum inoculation may be connected to the resistance phenotype15,24. In this study we used progenies from one Norway spruce cross and one H. parviporum strain to create expression profiles of the candidate genes associated with the resistance QTLs. Plant materials used in this study was represented by a set of closely-related individuals drawn from the original QTL population11,14. This approach may have limitations as the use of more host and pathogen genotypes could have given a broader picture. However, in this study we wanted to study the expression pattern of the candidate genes mapped in the original QTL mapping study14 in response to same fungal isolate H. parviporum Rb175 that was used to detect the QTLs. Using more H. parviporum isolates in the study would have perhaps generated information on how broad the transcriptional responses of the candidate genes associated with the QTLs were. Even though the choice to use full-sib plants and H. parviporum Rb175 might have the drawback of not being able to identify some of the broader defence responses in Norway spruce, it can help removing “signal noise” as resistance against H. parviporum in Norway spruce is a quantitative trait with multiple genes having small effect where highly resistant individuals might rely on a different set of genetic factors. The approach in this study allowed us to work with well-studied plant materials and to use a narrow genetic base to generate initial data41,42 and identify potential resistance candidates for further work, which was one of the primary objectives of this study. Clearly we missed out on potential candidate genes which are not associated with the QTL regions in the genetic linkage map and also candidates associated with QTLs which are not present in the original study43,44. Nevertheless, combining genomic and transcriptomic analysis we identified 124 candidate resistance genes which could be considered as candidates for induced resistance.
Fungal sapwood growth (SWG) is a trait that reflects the trees capacity to restrict the spread of the pathogen in its sapwood. The general assumption is that the trees with shorter SWG after inoculation with H. annosum s.l. in the sapwood would also display shorter decay columns after natural infections45. We could locate all of the original QTLs for SWG in the composite map14,25. Interestingly, the inspection of the QTL for SWG2 on LG6 including the previously validated marker PaLAR315 in the composite map, positioned PaLAR3 nearly 30 cM away from the other markers in this QTL region. This could suggest that the markers found around 154 cM on LG6, including BT116508/ MA_10433955g0020 encoding magnesium-protoporphyrin, may not be linked to the original SWG QTL or the original QTL14 comprises several independent QTLs which could not be separated due to low density of markers in this QTL region. Despite this we chose to include these candidate genes in the subsequent analyses, and 39 expressed candidate genes associated with the SWG QTL were identified. In fact, 16 of the candidate genes differentially expressed (10 CCGs and 6 PCGs) in response to inoculation were found at this position in the map. Among the upregulated candidate genes associated with the SWG QTLs MA_853405g0010 (a hypothetical protein containing a domain of unknown function, DUF4228), and the MA_122748g0010 (a putative GRX, Glutaredoxin) showed significant up-regulation during inoculation compared to wounding at both three and seven dpi in our RNAseq analysis. Both of these DEGs were positioned at 154 cM on LG6. The expression patterns of the candidate genes in this region could possibly indicate that they are involved in the expression of the resistance trait. Clearly, fine mapping the region between BT116508 and BT109050/PaLAR3 would be very helpful to resolve the locus structure.
One of the two QTLs for lesion length (LL) could be identified in the composite map14,25. We found two DEGs associated with cell wall modifications and specialized metabolism in this QTL region. The moderately upregulated DEG MA_52380g0010 identified as PCG (log2 Fold Change of 0.75) with similarity to UDP-glucuronate 4-epimerase that presumably catalyzes the formation of the key building block of pectins, UDP-d-galacturonic acid46,47. The second identified as CCG in this QTL that is putatively associated with cell wall modifications encodes a CCR-like protein (cinnamoyl-CoA reductase, MA_10435810g0010) and was repressed both proximally and distally at seven dpi. CCR is the first committed enzyme in the lignin-specific pathway48,49, and it is possible that the downregulation of this gene is part of a redirection of resources away from the lignin biosynthesis pathway to other branches of the phenolics metabolism (see also discussion on NAC transcription factors below). The repressed CCR gene would thus be in line with Norway spruce allocating more resources to potential antifungal low molecular weight phenolics following challenges with H. annosum24. It has been shown that suppression of CCR gene expression in Norway spruce decreases lignin content50. Suppression of CCR gene expression in tobacco has been accompanied by accumulation of phenolic substances51,52. Further analyses, eg. with RNAi- or overexpression constructs would shed light on cinnamoyl-CoA reductase’s (MA_10435810g0010) role in the interaction with H. parviporum.
There are two expression hotspots associated with the QTL regions for IP and E. The traits IP and E are measures of the ability the host have to stop the fungus from entering the wound upon inoculation or the ability to hem in and exclude an invading pathogen14. Thus, these traits could potentially reflect the “true” resistance to infection, and consequently the candidate genes associated with these QTLs were of special interest to us. Two thirds of the expressed genes as well as differentially expressed genes were associated with these QTLs. It is difficult to decide when to best capture gene expression patterns associated with IP and E in the artificial inoculation system we employ, as the read out of the traits takes place after several weeks of interaction14,53. Consequently, the expression data gathered for these traits should be seen as a probe and genes not differentially expressed in the current study could still be highly relevant to induced defence responses in H. parviporum resistance.
It is notable that several of the candidate genes associating with the QTL regions for IP are genes or orthologs to genes, which have been shown to respond to biotic stress29,54 or control other adaptive traits55,56 in the genus Picea. MA_24271g0020 (PgMYB11-like), which corresponds to the original marker BT103501 in the first QTL for IP on LG1114, is one example. This marker has been found to associate with a QTL for bud set in black spruce55. The observation that variation in BT103501 associate with two apparently different traits in spruce, indicates pleiotropic effect or tightly linked loci55,57. Another of the original markers in this QTL MA_4742g0010 (PabiesFT1-1,251/BT115191) encoding a spruce Mother of FT1- like protein MFT1, suggested to control the formation of resin ducts in male buds56. MA_4742g0010 was weakly expressed but significantly upregulated at seven dpi both proximal and distal to the inoculation. Neither PgMYB11-like nor Norway spruce MFT1, have been connected to host defence previously. In contrast, a DEG associated with this QTL that has a recognized role in the induced defences in conifers is MA_6931g0010. This candidate gene encodes a putative caffeoyl-CoA O-methyltransferase (CCoAOMT) and it is highly expressed in all treatments and was weakly upregulated at seven dpi proximal to the inoculation site. CCoAOMT1, an enzyme in the lignin biosynthesis pathway54 has been implicated in budworm and white pine weevil resistance in white spruce.
The candidate gene Phenylalanine ammonia-lyase (PaPAL3) identified as a PCG (MA_15852g0010), an ortholog of the Picea glauca PgPAL3 (Genbank: BT119163)35 at the QTL for exclusion on the LG3, was upregulated both proximal and distal to the inoculation site at three and seven dpi. Two other PaPAL1 and PaPAL2 is reported to be induced upon wounding and H. annosum s.l.24,58,59. PAL is the first enzyme committed in the phenylpropanoid biosynthesis pathway60. The activation of phenylpropanoid biosynthetic pathway which leads to the production of polyphenolics. Flavonoids and stilbene monomers plays a central role in the induced defence towards wounding and fungal infection in conifers61,62,63,64. Stilbene astringin was negatively correlated with the depth of the hyphal penetration of Heterobasidion annosum in Norway spruce bark65. Flavonoids have an antimicrobial effect on H. annosum s.l.24 and E. polonica20,61,62,63 in Norway spruce.
Just like the IP QTL on LG11, the QTL region for fungal exclusion from the sapwood on LG6 also involved a number of candidate genes that had been studied previously in conifers. However, the most interesting feature of the QTL region for exclusion on LG6 is that it harbours three of the previously identified seven Norway spruce candidate genes with similarity to subgroup III-3 NAC transcription factors29; PaNAC04, MA_86256g0010 and MA_103386g0010. All three candidate genes were relatively highly expressed and showed clear upregulation in response to inoculation with H. parviporum both proximal and distal to the inoculation (Fig. 1). Their upregulation could possibly be associated with a shifted balance from cell wall use to defense active phenylpropanoids29. Naturally, the question arose if these three predicted candidate genes indeed represent distinct genes. The difficulties in assembling the large and repetitive conifer genomes into scaffolds will lead to errors in the assembly66,67. Thus, it would not be unlikely that the three highly similar candidate genes29, correspond to one single gene located in the Exclusion QTL on LG6. However, based on the expression patterns detected by qRT-PCR, which agrees with the previous phylogenetic analysis29, placing PaNAC04 and MA_86256g0010 together on a supported branch separate from MA_103386p0010 in subgroup III-3 of the NAC transcription factor family, we argue that there are at least two NAC genes associated with this QTL. Albeit, tightly linked and not much diverged, but this must be confirmed by resequencing of the region.
This study gives an insight into Norway spruce genome organization with information of position of the candidate genes associated with resistant trait in the genome e.g. the previously described PaNAC04. PaNAC04 was not only upregulated in response to H. parviporum but it was also located in the region important for controlling resistance determined by QTL mapping14. Therefore, allelic variation in PaNAC04 needs to be further studied in future experiment to understand the role of PaNAC04 in controlling induce defence response.
In conclusion; this study, combining map-based information and expression analyses have associated previously identified candidate genes, such as PaNAC04 and MA_103386g0010, with genomic regions in Norway spruce harboring resistance QTL, strengthening their predicted role in control of H. annosum s.l. infection. This approach has also allowed us to identify a set of novel candidate genes for future analyses, the most prominent being genes associated with the phenylpropanoid pathway CCR (MA_10435810g0010), the PgMYB11-like gene and PAL (MA_15852g0010).
References
Bulman, L. et al. A worldwide perspective on the management and control of Dothistroma needle blight. For. Pathol. 46, 472–488 (2016).
Woodward, S. Heterobasidion annosum: Biology, ecology, impact and control (CABI, 1998).
Schmidt, R. A. Fusiform rust of southern pines: a major success for forest disease management. Phytopathology 93, 1048–1051 (2003).
Thor, M., Arlinger, J. D. & Stenlid, J. Heterobasidion annosum root rot in Picea abies: Modelling economic outcomes of stump treatment in Scandinavian coniferous forests. Scand. J. For. Res. 21, 414–423 (2006).
Fridman, J. & Wulff, S. Skogsdata 2018. Report No. 0280–0543 (2018).
Loman, J. Statistical Yearbook of Forestry Official Statistics of Sweden. In Loman J, editor. Jönköping: Swedish Forestry Agency (2007).
Asiegbu, F. O., Adomas, A. & Stenlid, J. Conifer root and butt rot caused by Heterobasidion annosum (Fr.) Bref. Sl.. Mol. Plant Pathol. 6, 395–409 (2005).
Garbelotto, M. & Gonthier, P. Biology, epidemiology, and control of Heterobasidion species worldwide. Annu. Rev. Phytopathol. 51, 39–59 (2013).
Holdenrieder, O. & Greig, B. Biological Control of Heterobasidion annosum. In: Heterobasidion annosum 235–258 (CAB International, Wallingford, 1998).
Oliva, J., Thor, M. & Stenlid, J. Long-term effects of mechanized stump treatment against Heterobasidion annosum root rot in Picea abies. Can. J. For. Res. 40, 1020–1033 (2010).
Arnerup, J., Swedjemark, G., Elfstrand, M., Karlsson, B. & Stenlid, J. Variation in growth of Heterobasidion parviporum in a full-sib family of Picea abies. Scand. J. For. Res. 25, 106–110 (2010).
Chen, Z.-Q. et al. Early selection for resistance to Heterobasidion parviporum in Norway spruce is not likely to adversely affect growth and wood quality traits in late-age performance. Eur. J. For. Res. 1–9 (2018).
Karlsson, B. & Swedjemark, G. Genotypic variation in natural infection frequency of Heterobasidion spp. in a Picea abies clone trial in southern Sweden. Scand. J. For. Res. 21, 108–114 (2006).
Lind, M. et al. A Picea abies linkage map based on SNP markers identifies QTLs for four aspects of resistance to Heterobasidion parviporum infection. PLoS ONE 9, e101049 (2014).
Nemesio-Gorriz, M. et al. Different alleles of a gene encoding leucoanthocyanidin reductase (PaLAR3) influence resistance against the fungus Heterobasidion parviporum in Picea abies. Plant Physiol. 171, 2671–2681 (2016).
Steffenrem, A., Solheim, H. & Skrøppa, T. Genetic parameters for wood quality traits and resistance to the pathogens Heterobasidion parviporum and Endoconidiophora polonica in a Norway spruce breeding population. Eur. J. Forest Res. 135, 815–825 (2016).
Sniezko, R. A. & Koch, J. Breeding trees resistant to insects and diseases: putting theory into application. Biol. Invasions 19, 3377–3400 (2017).
Neale, D. B. & Kremer, A. Forest tree genomics: growing resources and applications. Nat. Rev. Genet. 12, 111 (2011).
Dalman, K., Olson, Å & Stenlid, J. Evolutionary history of the conifer root rot fungus Heterobasidion annosum sensu lato. Mol. Ecol. 19, 4979–4993 (2010).
Hammerbacher, A. et al. Flavan-3-ols in Norway spruce: biosynthesis, accumulation, and function in response to attack by the bark beetle-associated fungus Ceratocystis polonica. Plant Physiol. 164, 2107–2122 (2014).
Liu, J. J. et al. Saturated genic SNP mapping identified functional candidates and selection tools for the Pinus monticola Cr2 locus controlling resistance to white pine blister rust. Plant Biotechnol. J. 15, 1149–1162 (2017).
Mageroy, M. H. et al. Expression of the β-glucosidase gene Pgβglu-1 underpins natural resistance of white spruce against spruce budworm. Plant J. 81, 68–80 (2015).
Arnerup, J., Lind, M., Olson, Å, Stenlid, J. & Elfstrand, M. The pathogenic white-rot fungus Heterobasidion parviporum triggers non-specific defence responses in the bark of Norway spruce. Tree Physiol. 31, 1262–1272 (2011).
Danielsson, M. et al. Chemical and transcriptional responses of Norway spruce genotypes with different susceptibility to Heterobasidion spp. infection. BMC Plant Biol. 11, 154 (2011).
de Miguel, M. et al. Evidence of intense chromosomal shuffling during conifer evolution. Genom. Biol. Evolut. 7, 2799–2809 (2015).
Pavy, N. et al. A spruce gene map infers ancient plant genome reshuffling and subsequent slow evolution in the gymnosperm lineage leading to extant conifers. BMC Biol. 10, 84 (2012).
Eckert, A. J. et al. Patterns of population structure and environmental associations to aridity across the range of loblolly pine (Pinus taeda L., Pinaceae). Genet 185, 969–982 (2010).
Chang, S., Puryear, J. & Cairney, J. A simple and efficient method for isolating RNA from pine trees. Plant Mol. Biol. Rep. 11, 113–116 (1993).
Dalman, K. et al. Overexpression of PaNAC03, a stress induced NAC gene family transcription factor in Norway spruce leads to reduced flavonol biosynthesis and aberrant embryo development. BMC Plant Biol. 17, 6 (2017).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562 (2012).
Trapnell, C. et al. Differential analysis of gene regulation at transcript resolution with RNA-seq. Nat. Biotechnol. 31, 46 (2013).
Rozen, S. & Skaletsky, H. Primer3 on the WWW for general users and for biologists programmers. In: Bioinformatics methods and protocols. 365–386 (Springer, Berlin, 2000).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2− ΔΔCT method. Methods 25, 402–408 (2001).
Palovaara, J. & Hakman, I. Conifer WOX-related homeodomain transcription factors, developmental consideration and expression dynamic of WOX2 during Picea abies somatic embryogenesis. Plant Mol. Biol. 66, 533–549 (2008).
Raherison, E. S. M. et al. Modular organization of the white spruce (Picea glauca) transcriptome reveals functional organization and evolutionary signatures. New Phytol. 207 (2015).
Krutovsky, K. V., Troggio, M., Brown, G. R., Jermstad, K. D. & Neale, D. B. Comparative mapping in the Pinaceae. Genetics 168, 447–461 (2004).
Pavy, N. et al. Enhancing genetic mapping of complex genomes through the design of highly-multiplexed SNP arrays: application to the large and unsequenced genomes of white spruce and black spruce. BMC Genom. 9, 21 (2008).
Pelgas, B. et al. Comparative genome mapping among Picea glauca, P. mariana× P. rubens and P. abies, and correspondence with other Pinaceae. Theoret. Appl. Genet. 113, 1371 (2006).
Franceschi, V. R., Krokene, P., Christiansen, E. & Krekling, T. Anatomical and chemical defenses of conifer bark against bark beetles and other pests. New Phytol. 167, 353–376 (2005).
Keeling, C. I. & Bohlmann, J. Genes, enzymes and chemicals of terpenoid diversity in the constitutive and induced defence of conifers against insects and pathogens. New Phytol. 170, 657–675 (2006).
Petersen, B., Chen, S., Hansen, C., Olsen, C. & Halkier, B. Composition and content of glucosinolates in developing Arabidopsis thaliana. Planta 214, 562–571 (2002).
Bowerman, P. A., Ramirez, M. V., Price, M. B., Helm, R. F. & Winkel, B. S. Analysis of T-DNA alleles of flavonoid biosynthesis genes in Arabidopsis ecotype Columbia. BMC Res. Notes 5, 485 (2012).
Elfstrand, M. et al. Association genetics identifies a specifically regulated Norway spruce laccase gene, PaLAC5, linked to Heterobasidion parviporum-resistance. PCE https://doi.org/10.1111/pce.13768 (2020).
Mukrimin, M. et al. Genome-wide exon-capture approach identifies genetic variants of Norway spruce genes associated with susceptibility to Heterobasidion parviporum infection. Front. Plant Sci. 9, 793 (2018).
Delatour, C., von Weissenberg, K. & Dimitri, L. Host Resistance w: Heterobasidion annosum, Biology Ecology and Control (Cab International, Wallingford, 1998).
Mølhøj, M., Verma, R. & Reiter, W. D. The biosynthesis of D-galacturonate in plants. Functional cloning and characterization of a membrane-anchored UDP-D-glucuronate 4-epimerase from Arabidopsis. Plant Physiol. 135, 1221–1230 (2004).
Stevanic, J. S. & Salmén, L. Characterizing wood polymers in the primary cell wall of Norway spruce (Picea abies (L.) Karst.) using dynamic FT-IR spectroscopy. Cellulose 15, 285. https://doi.org/10.1007/s10570-007-9169-1 (2007).
Lacombe, E. et al. Cinnamoyl CoA reductase, the first committed enzyme of the lignin branch biosynthetic pathway: cloning, expression and phylogenetic relationships. Plant J. 11, 429–441 (1997).
Li, L. et al. Clarification of cinnamoyl co-enzyme a reductase catalysis in monolignol biosynthesis of aspen. Plant Cell Physiol. 46, 1073–1082 (2005).
Wadenbäck, J. et al. Lignin biosynthesis in transgenic Norway spruce plants harboring an antisense construct for cinnamoyl CoA reductase (CCR). Transgenic Res. 17, 379–392 (2008).
Pinçon, G. et al. Simultaneous down-regulation of caffeic/5-hydroxy ferulic acid-O-methyltransferase I and cinnamoyl-coenzyme A reductase in the progeny from a cross between tobacco lines homozygous for each transgene. Consequences for plant development and lignin synthesis. Plant Physiol. 126, 145–155 (2001).
Ralph, J. et al. NMR characterization of altered lignins extracted from tobacco plants down-regulated for lignification enzymes cinnamylalcohol dehydrogenase and cinnamoyl-CoA reductase. Proc. Natl. Acad. Sci. 95, 12803–12808 (1998).
Swedjemark, G., Karlsson, B. & Stenlid, J. Exclusion of Heterobasidion parviporum from inoculated clones of Picea abies and evidence of systemic induced resistance. Scand. J. For. Res. 22, 110–117 (2007).
Ralph, S. G. et al. Conifer defence against insects: microarray gene expression profiling of Sitka spruce (Picea sitchensis) induced by mechanical wounding or feeding by spruce budworms (Choristoneura occidentalis) or white pine weevils (Pissodes strobi) reveals large-scale changes of the host transcriptome. Plant Cell Environ. 29, 1545–1570 (2006).
Prunier, J. et al. The genomic architecture and association genetics of adaptive characters using a candidate SNP approach in boreal black spruce. BMC Genom. 14, 368 (2013).
Karlgren, A. et al. Evolution of the PEBP gene family in plants: functional diversification in seed plant evolution. Plant Physiol., pp. 111.176206 (2011).
Pelgas, B., Bousquet, J., Meirmans, P. G., Ritland, K. & Isabel, N. QTL mapping in white spruce: gene maps and genomic regions underlying adaptive traits across pedigrees, years and environments. BMC Genom. 12, 145 (2011).
Koutaniemi, S. et al. Expression profiling of the lignin biosynthetic pathway in Norway spruce using EST sequencing and real-time RT-PCR. Plant Mol. Biol. 65, 311–328 (2007).
Yaqoob, N. et al. Defence-related gene expression in bark and sapwood of Norway spruce in response to Heterobasidion parviporum and methyl jasmonate. Physiol. Mol. Plant Pathol. 77, 10–16 (2012).
Bagal, U. R. The phenylalanine ammonia lyase (PAL) gene family shows a gymnosperm-specific lineage. BMC Genomics 13(Suppl 3), S1 (2012).
Brignolas, F. et al. Induced responses in phenolic metabolism in two Norway spruce clones after wounding and inoculations with Ophiostoma polonicum, a bark beetle-associated fungus. Plant Physiol. 109(3), 821–827 (1995).
Brignolas, F., Lieutier, F., Sauvard, D., Christiansen, E. & Berryman, A. A. Phenolic predictors for Norway spruce resistance to the bark beetle Ips typographus (Coleoptera: Scolytidae) and an associated fungus, Ceratocystis polonica. Can. J. For. Res. 28(5), 720–728 (1998).
Lieutier, F. et al. Intra- and inter-provenance variability in phloem phenols of Picea abies and relationship to a bark beetle-associated fungus. Tree Physiol. 23(4), 247–256 (2003).
Schmidt, A. et al. Induced chemical defences in conifers: Biochemical and molecular approaches to studying their function. Recent Adv. Phytochem. 39. (2005).
Lindberg, M., Lundgren, L., Gref, R. & Johansson, M. Stilbenes and resin acids in relation to the penetration of Heterobasidion annosum through the bark of Picea abies. Eur. J. For. Pathol. 22, 95–106 (1992).
Nystedt, B. et al. The Norway spruce genome sequence and conifer genome evolution. Nature 497, 579 (2013).
Bernhardsson, C. et al. An ultra-dense haploid genetic map for evaluating the highly fragmented genome assembly of Norway spruce (Picea abies). bioRxiv, 292151 (2018).
Acknowledgements
The Swedish Research Council for Environment, Agricultural Sciences and Spatial Planning (FORMAS), Grant Nrs. 2012-1276, 2013-00965 and 2015-00081 provided financial support to the study. The authors would like to thank Dr. Mårten Lind and Dr. Jenny Arnerup for assistance in the laboratory and helpful discussions on the results and Louis Mielke for language revision. Open access funding provided by Swedish University of Agricultural Sciences.
Author information
Authors and Affiliations
Contributions
M.E. and J.S. conceived and designed the experiments. B.K. provided study material for the study. K.D. and M.N.G. performed the inoculation experiments. K.D. extracted RNA and sent it for sequencing. R.C. performed the RNAseq and statistical analysis. R.C. and M.D. performed qPCR analysis. R.C., J.S., K.L. and M.E. wrote the manuscript. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chaudhary, R., Lundén, K., Dalman, K. et al. Combining transcriptomics and genetic linkage based information to identify candidate genes associated with Heterobasidion-resistance in Norway spruce. Sci Rep 10, 12711 (2020). https://doi.org/10.1038/s41598-020-69386-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-69386-0
This article is cited by
-
Applying molecular and genetic methods to trees and their fungal communities
Applied Microbiology and Biotechnology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.