Abstract
The outbreak of COVID-19 has brought great threat to human health. Its causative agent is a severe acute respiratory syndrome-related coronavirus which has been officially named SARS-CoV-2. Here we report the discovery of extremely low CG abundance in its open reading frames. We found that CG reduction in SARS-CoV-2 is achieved mainly through mutating C/G into A/T, and CG is the best target for mutation. Meanwhile, 5′-untranslated region of SARS-CoV-2 has high CG content and is capable of forming an internal ribosome entry site (IRES) to recruit host ribosome for translating its RNA. These features allow SARS-CoV-2 to reproduce efficiently in host cells, because less energy is consumed in disrupting the stem-loops formed by its genomic RNA. Notably, genomes of cellular organisms also have very low CG abundance, suggesting that mutating C/G into A/T occurs universally in all life forms. Moreover, CG is the dinucleotide related to CpG island, mutational hotspot and single nucleotide polymorphism in cellular organisms. The relationship between these features is worthy of further investigations.
Similar content being viewed by others
Introduction
The outbreak of COVID-19 (coronavirus disease 2019) was listed as a public health emergency of international concern on 30 January 2020 and declared a global pandemic on 11 March 2020 by the World Health Organization. As of 15 June 2020, it has caused more than 7.8 million infection cases and over 430,000 deaths worldwide1. Its causative virus (officially named SARS-CoV-2) has a genome of single-strand positive-sense RNA with approximately 30,000 nucleotides2. Based on its genome sequence, analyses have been conducted to characterize genomic features and to trace origin of the virus3,4. Meanwhile, many reports have been focused on developing efficient methods for detection5,6 and screening effective drugs for treatment7,8 against COVID-19.
Coronaviruses have some of the largest RNA genomes among all viruses. The single-strand genomic RNA of coronavirus has a cap-like structure at 5′-UTR (untranslated region) and a poly(A)-tail at 3′-UTR, both of which allow it to assume a structure similar to mRNA of host cells. After being released into cytoplasm of a host cell, the viral RNA is translated immediately to produce viral proteins by using the translation machinery of host cells9. The viral RNA is much longer than host mRNAs. Such a long single-strand RNA will form multiple stem-loops through base-pairing between its adjacent segments. These stem-loops present an obstacle to the translation machinery, because they must be disrupted to expose coding information contained in them. The stability of a stem-loop structure is dependent on number of hydrogen bonds formed between bases in the stem part. Because C-G and T-A base-pairs are formed through three and two hydrogen bonds respectively, a viral RNA strand with high number of C and G bases will form more stable stem-loops than that with high number of T and A bases.
In order to understand whether translation of coronavirus RNA is greatly affected by base composition, we analysed dinucleotide distribution and RNA stability of twenty-four coronavirus species. We found that open reading frames (ORFs) of SARS-CoV-2 have an extremely low abundance of CG dinucleotide. Moreover, the secondary structure formed by SARS-CoV-2 genomic RNA is less stable than many other coronaviruses. Therefore, it is suggested that SARS-CoV-2 is more efficient in reproduction than other coronaviruses, because less energy is consumed in disrupting the stem-loops formed by its genomic RNA.
Results
Low CG content in human SARS-CoV-2
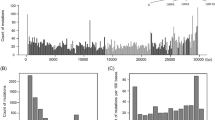
DNA or RNA sequences are composed of four nucleotides, i.e. adenylate (A), thymidylate (T), guanylate (G) and cytidylate (C). They can also be considered polymers of 16 dinucleotides. Odds ratio is a value defined to indicate relative abundance of a nucleotide, which is the ratio of observed to expected frequency of a dinucleotide10. The genome of SARS-CoV-2 (29,903 nucleotides2, sequence number NC_045512) has 29.94% of A, 32.08% of T (T is used here instead of U for simplicity), 19.61% of G and 18.37% of C. Thus, the expected frequency of CG dinucleotide in viral genome is 3.60% (i.e. 19.61% × 18.37%). However, only 439 CGs are observed, which means the observed frequency of CG dinucleotide is 1.47% (i.e. 439/29,902). Therefore, odds ratio of CG in SARS-CoV-2 is 0.41 (i.e. 1.47%/3.60%). Furthermore, odds ratio of CG in open reading frames (ORFs) of the virus is 0.39, being the lowest among 24 coronaviruses under survey (Fig. 1a and Table S1). Because a codon is composed of three nucleotides, a dinucleotide (e.g. CG) has three possible locations. Herewith, they are designated as (CG)12, (CG)23 and (CG)31 respectively. We found that the odds ratio of (CG)23 in ORFs of SARS-CoV-2 is as low as 0.25, while that of (CA)23 and (CT)23 is as high as 1.54 and 1.92 respectively (Fig. 1c). Moreover, odds ratio of (CG)31 in ORFs of SARS-CoV-2 is 0.50, while that of (AG)31 and (TG)31 is 1.52 and 2.64 respectively (Fig. 1d). These data strongly suggest that (CG)23 has been mutated into (CA)23 and (CT)23, and (CG)31 has been mutated into (AG)31 and (TG)31.
Odds ratios of dinucleotides in open reading frames of SARS-CoV-2. (a) odds ratios of dinucleotides at all codon positions. (b–d) odds ratios of dinucleotides at codon positions 1 and 2, 2 and 3, 3 and 1, respectively. Value shown in the figure is weighted average odds ratio of each dinucleotide. Odds ratio of each dinucleotide in ten ORFs (i.e. ORF1ab and ORF 2–10) of SARS-CoV-2 is calculated respectively first. Then, a weighted average odds ratio is obtained based on length of each ORF.
The above-stated mutations are possible because very few of these mutations lead to changes in amino acids. To be specific, there are four codons containing (CG)23. They are TCG, CCG, ACG and GCG which code for serine, proline, threonine and alanine, respectively. Mutation of G at codon position 3 into T, C or A in all of them does not change the amino acid they encode. As for (CG)31, there are 16 codons having C at position 3. If this C is mutated into T, all 16 codons have the same meanings. And if it is mutated into A, 9 out of 16 codons still have the same meanings. Therefore, it is concluded that SARS-CoV-2 has evolved to reduce CG in ORFs mainly through mutating its G of (CG)23 and C of (CG)31 into A and T. Among them, C-to-T (i.e. C-to-U in RNA) occurs at a very high frequency probably because it is the simplest way to change a nucleotide (C becomes U after deamination). Besides, odds ratio of (CC)23 is much lower than that of (CA)23 and (CT)23 (Fig. 1c). This does not mean that (CG)23 has not been mutated into (CC)23. In fact, low odds ratio of (CC)23 is due to high frequency of C-to-T mutation at position 3, i.e. from (CG)31 into (TG)31 (Fig. 1d). The above views are also supported by codon usage bias in SARS-CoV-2 (Fig. 2), which shows that A/T-ended codons are much more frequently used than their synonymous G/C-ended codons. Besides, all four codons containing (CG)23 have the lowest percentages of usage among their correspondent synonymous codons.
Percentages of codon usage in open reading frames of SARS-CoV-2. Usage of synonymous codons for eighteen amino acids (except methionine and tryptophan) and three stop codons are shown in the figure. Percentages of codons with A, T, G and C at codon position 3 are in yellow, brown, green and aqua blue background, respectively. Total number of codons for each amino acid is indicated at top of the percentage bar. Arrows indicate four codons that contain CG at positions 2 and 3.
Low CG content in other coronaviruses
Odds ratios of CG in ORFs of other coronaviruses are also very low (mean value = 0.50, Fig. 3 and Table S1). This could have profound effect on viral replication, because ORFs of coronaviruses are immediately translated by host ribosomes after being released into the cytoplasm of host cells9. The translation of viral RNA is affected by two factors. One is that host ribosomes must be recruited to the 5′-UTR (untranslated region) of viral RNA for initiation of translation. The other is that stem-loops formed by ORFs of viral RNA must be disrupted to expose coding information during translation. In contrast to ORFs, 5′-UTR of coronaviruses have quite high odds ratios of CG (mean value = 0.84, Table S2). This would facilitate formation of stable secondary structure that could serve as the internal ribosome entry site (IRES)11,12,13 for host ribosome (Fig. 4). Meanwhile, the viral RNA beginning at the translation start site (TSS) forms relatively unstable secondary structure, because its stem-loops are maintained by less hydrogen bonds (A-T and C-G base pairs have two and three hydrogen bonds respectively).
Odds ratios of dinucleotides in open reading frames of coronaviruses and cellular organisms. (a) odds ratios of dinucleotides at all codon positions. (b–d) odds ratios of dinucleotides at codon positions 1 and 2, 2 and 3, 3 and 1, respectively. Data of coronaviruses are from Table S1, which are shown in blue background. Those of cellular organisms are from our previous work15. Filled triangle or filled inverter triangle indicates that odds ratio of a dinucleotide in coronavirus is significantly higher or lower than that in cellular organisms at p = 0.05 level. Open triangle or open inverter triangle indicates that odds ratio of a dinucleotide in coronavirus is insignificantly higher or lower than that in cellular organisms.
Secondary structure formed by 5′-UTR of poliovirus (a) and SARS-CoV-2 (b). The secondary structure is based on 200 nucleotides immediately upstream of the translation start site. Sequence number of poliovirus is MG212486. That of SARS-CoV-2 is NC_045512. Both structures and their free energy (indicated in centre of the structure) are drawn/calculated using RNAstructure (version 5.7)27.
Stability variations of viral genomes at 5′-UTR and TSS-to-end regions could probably determine virulence of different viruses, because high stability of IRES structure means high efficiency in initiating translation, and high stability of TSS-to-end region means high energy consumption during translation. After high, medium and low stability of both 5′-UTR and TSS-to-end regions is given 3, 2 and 1 points respectively, virulence of coronaviruses can be classified into five grades, i.e. very high, high, medium, low and very low (Table 1). For example, human MERS (Middle East respiratory syndrome) coronavirus has very high virulence, because both its 5′-UTR and TSS-to-end regions are highly stable. High stability of 5′-UTR means that host ribosomes can be recruited to translate viral RNA at high rate. And, high stability of ORFs means that more energy is consumed to disrupt stem-loops in viral RNA during translation. Thus, normal translation of host cell mRNAs is greatly affected, suggesting that MERS coronavirus is highly virulent. SARS (severe acute respiratory syndrome) coronavirus has high virulence, because its 5′-UTRs is less stable than MERS coronavirus. SARS-CoV-2 has medium virulence, because it has medium stability in both 5′-UTR and TSS-to-end regions. This classification is consistent with estimations on case fatality ratio of MERS, SARS and COVID-19, which is 35%, 9% and 2.4% respectively14 and with our observations on odds ratio of CG in their ORFS, which is 0.56, 0.44 and 0.39 respectively (Table S1). Moreover, compared to SARS coronavirus, SARS-CoV-2 could infect and replicate more efficiently in human lung tissues but induce expression of less inflammatory cytokines/chemokines and mediators15. In our opinion, it is the lower C/G content in genomic RNA that allows SARS-CoV-2 to reproduce higher number of virus particles before triggering the immunoreaction of host cells, because less energy is consumed in replicating each virus particle.
Two other human coronaviruses have medium virulence as well. Among them, NL63 has medium stability in both 5′-UTR and TSS-to-end regions, whereas 229E has low stability in 5′-UTR but high stability in TSS-to-end region. Another human coronavirus (i.e. HKU1) has very low virulence, because it has low stability in both 5′-UTR and TSS-to-end regions (Table 1). The worldwide transmission of SARS-CoV-2 probably means that a coronavirus with medium virulence is more likely to spread rapidly. In comparison, a coronavirus with high or very high virulence could kill its host before causing severe epidemic, whereas a coronavirus with low or very low virulence is not able to replicate itself efficiently for further transmission.
Discussion
Our present study provides a novel insight into the evolution of human SARS-CoV-2. It is evident that this virus has evolved to reduce CG intensely in its ORFs. Such reduction is achieved mainly through mutating G of (CG)23 and C of (CG)31 into A or T (Fig. 1). Meanwhile, C or G not of CG may also be mutated. For example, TCA in SARS-CoV-2 of S-type has been mutated into TTA16. GTC and GGT in SARS-CoV-2 isolated from France have been mutated into TTC and GTT respectively17. Although the mutated C or G is not of CG and not at codon position 3, they do reduce C or G in viral RNA. C/G reduction is favourable for increasing efficiency of viral RNA translation, because stem-loops formed by less C/G-containing segments can be disrupted more easily. In fact, genomic RNA stability is closely related to nucleotide composition in coronaviruses (Fig. 5). First, RNA stability is positively correlated to content of C, G and C + G but negatively correlated to content of T, A and T + A (Fig. 5a). Second, RNA stability is also positively correlated to content of GC, GG, CG and CC but negatively correlated to content of AT, AA, TA and TT (Fig. 5b). Third, RNA stability is only positively correlated with odds ratio of dinucleotide GC and CG (Fig. 5c). As odds ratio measures the relative abundance of a specific dinucleotide, the extremely significant correlation between CG odds ratio and RNA stability strongly suggests that CG has been selected as the major target for mutation in coronaviruses.
Correlation between RNA stability and nucleotide composition in viral genome. Shown here are correlation coefficients of RNA stability with (a) content of nucleotide(s), (b) content of dinucleotide and (c) odds ratio of dinucleotide in genomes of 24 coronaviruses. Only TSS-to-end region of viral genome is included for analysis (TSS: translation-start-site). * and **above data bar indicate that the correlation reaches significant (0.01 < p < 0.05) and extremely significant (p < 0.01) level, respectively. Detailed data for correlation analysis are listed in rows 67 to 103 of Table S2.
Then, if reducing hydrogen bonds is the goal of base mutation, why is CG but not GC, GG or CC taken as the target for mutation? An examination on number of silent mutations of each dinucleotide at various codon positions reveals that CG has the highest number (47) of silent mutations among these four dinucleotides (Table 2 and Table S3). This explains why CG is the best target for mutation. Although CT has the same highest number like CG, it is not taken as the target for mutation because a T-to-C or T-to-G mutation would increase number of hydrogen bonds between potential base pairs, which is contradictory to the goal of mutation.
It seems that the strategy of “reducing CG content to increase gene expression efficiency” has also been adopted by cellular organisms. As we have observed, CG in both ORFs and inter-genic regions of bacteria, archaea, fungi, plants and animals has an average odds ratio of 0.81, and that in introns of fungi, plants and animals is as low as 0.69. At time of our previous report18, we did not know why CG has such a low odds ratio in surveyed organisms. Now, after analysing cases in coronaviruses, we realize that low CG content in cellular organisms should also be the evolutionary consequence of increasing gene expression efficiency, because lowered CG content means reduced number of hydrogen bonds between DNA double strands (of the same length). Expression of a gene with low CG content saves energy not only in separating DNA double strands during transcription but also in disrupting stem-loops formed by mRNA during translation. Coincidently, CG is the very dinucleotide related to existence of CpG island, mutational hotspot, and single nucleotide polymorphism (SNP) in DNA sequences of cellular organisms. A CpG island is defined as a region of DNA with less methylated C, and this region generally contains actively expressed genes19,20,21. A mutational hotspot is defined as CG with methylated C, in which the methylated C is frequently mutated into T through deamination22,23,24. SNP refers to single nucleotide difference in genome sequences among individual organisms, which is observed most frequently at CG dinucleotide25,26. The relationship between CG reduction and these three important features of cellular DNA sequences is worthy of further investigations.
Methods
Genome sequences of coronaviruses were retrieved from GenBank (www.ncbi.nlm.nih.gov). Odds ratios of dinucleotides were calculated using formulae developed by Karlin and Mrázek10 and by Wang et al.18 with self-compiled computer programs (C++ scripts are available upon request). Secondary structure and free energy of viral RNA is predicted using RNAstructure (version 5.7)27. SPSS software (version 17.0) was used to conduct independent-sample t-test for comparing difference in odds ratio of nucleotide between coronaviruses and cellular organisms, and to conduct correlation analysis between RNA stability and nucleotide composition in viral genomes.
Change history
03 June 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41598-021-91385-y
References
WHO Director-General’s opening remarks at the media briefing on COVID-19, 15 June 2020, https://www.who.int/dg/speeches/detail/
Wu, F. et al. A new coronavirus associated with human respiratory disease in China. Nature 579(7798), 265–269 (2020).
Zhou, P. et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579(7798), 270–273 (2020).
Wassenaar, T. M. & Zou, Y. 2019_nCoV/SARS-CoV-2: rapid classification of betacoronaviruses and identification of Traditional Chinese Medicine as potential origin of zoonotic coronaviruses. Lett. Appl. Microbiol. 70(5), 342–348 (2020).
Jin, Y. H. et al. A rapid advice guideline for the diagnosis and treatment of 2019 novel coronavirus (2019-nCoV) infected pneumonia (standard version). Mil. Med. Res. 7(1), 4. https://doi.org/10.1186/s40779-020-0233-6 (2020).
Li, Z. et al. Development and clinical application of a rapid IgM-IgG combined antibody test for SARS-CoV-2 infection diagnosis. J. Med. Virol. https://doi.org/10.1002/jmv.25727 (2020).
Gao, J., Tian, Z. & Yang, X. Breakthrough: Chloroquine phosphate has shown apparent efficacy in treatment of COVID-19 associated pneumonia in clinical studies. Biosci. Trends 14(1), 72–73 (2020).
Lim, J. et al. Case of the index patient who caused tertiary transmission of COVID-19 infection in Korea: the application of Lopinavir/Ritonavir for the treatment of COVID-19 infected pneumonia monitored by quantitative RT-PCR. J. Korean Med. Sci. 35(6), e79 (2020).
Fehr, A. R. & Perlman, S. Coronaviruses: an overview of their replication and pathogenesis. Methods Mol. Biol. 1282, 1–23 (2015).
Karlin, S. & Mrázek, J. Compositional differences within and between eukaryotic genomes. Proc. Natl. Acad. Sci. USA 94(19), 10227–10232 (1997).
Sonenberg, N. & Pelletier, J. Poliovirus translation: a paradigm for a novel initiation mechanism. BioEssays 11(5), 128–132 (1989).
Ren, Q. et al. Alternative reading frame selection mediated by a tRNA-like domain of an internal ribosome entry site. Proc Natl Acad Sci USA 109, E630–E639 (2012).
Renaud-Gabardos, E. et al. Internal ribosome entry site-based vectors for combined gene therapy. World J. Exp. Med. 5(1), 11–20 (2015).
Peeri, N. C. et al. The SARS, MERS and novel coronavirus (COVID-19) epidemics, the newest and biggest global health threats: what lessons have we learned?. Int. J. Epidemiol. https://doi.org/10.1093/ije/dyaa033 (2020).
Chu, H. et al. Comparative replication and immune activation profiles of SARS-CoV-2 and SARS-CoV in human lungs: an ex vivo study with implications for the pathogenesis of COVID-19. Clin. Infect. Dis. https://doi.org/10.1093/cid/ciaa410 (2020).
Tang, X. L. et al. On the origin and continuing evolution of SARS-CoV-2. Natl. Sci. Rev. https://doi.org/10.1093/nsr/nwaa036 (2020).
Cleemput, S. et al. Genome Detective Coronavirus Typing Tool for rapid identification and characterization of novel coronavirus genomes. Bioinformatics 36(11), 3552–3555 (2020).
Wang, Y. et al. TA, GT and AC are significantly under-represented in open reading frames of prokaryotic and eukaryotic protein-coding genes. Mol. Genet. Genomics 294(3), 637–647 (2019).
Cooper, D. N., Mort, M., Stenson, P. D., Ball, E. V. & Chuzhanova, N. A. Methylation-mediated deamination of 5-methylcytosine appears to give rise to mutations causing human inherited disease in CpNpG trinucleotides, as well as in CpG dinucleotides. Hum. Genomics 4(6), 406–410 (2010).
Gardiner-Garden, M. & Frommer, M. CpG islands in vertebrate genomes. J. Mol. Biol. 196(2), 261–282 (1987).
Deaton, A. M. & Bird, A. CpG islands and the regulation of transcription. Genes Dev. 25(10), 1010–1022 (2011).
Shen, J. C., Rideout, W. M. & Jones, P. A. The rate of hydrolytic deamination of 5-methylcytosine in double-stranded DNA. Nucleic Acids. Res. 22(6), 972–976 (1994).
Krawczak, M., Ball, E. V. & Cooper, D. N. Neighboring-nucleotide effects on the rates of germ-line single-base-pair substitution in human genes. Am. J. Hum. Genet. 63(2), 474–488 (1988).
Hartl, D. et al. CG dinucleotides enhance promoter activity independent of DNA methylation. Genome Res. 29(4), 554–563 (2019).
Thompson, R. F., Suzuki, M., Lau, K. W. & Greally, J. M. A pipeline for the quantitative analysis of CG dinucleotide methylation using mass spectrometry. Bioinformatics 25(17), 2164–2170 (2009).
Bani-Fatemi, A. et al. Analysis of CpG SNPs in 34 genes: association test with suicide attempt in schizophrenia. Schizophr Res. 147(2–3), 262–268 (2013).
Reuter, J. S. & Mathews, D. H. RNAstructure: software for RNA secondary structure prediction and analysis. BMC Bioinform. 11, 129 (2010).
Acknowledgements
This study was supported by the National Natural Science Foundation of China (No. 31872425 and No. 31861143051).
Author information
Authors and Affiliations
Contributions
Y.W., Q.Y. and K.P.C. conceived the study and wrote the manuscript. Y.W. and J.M.M. compiled computer programs. Y.W., G.D.W., Z.P.L and L.Y. performed surveys and analyses. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this Article was revised: The original version of this Article incorrectly stated that coronaviruses have the largest of all reported RNA genomes. This statement has now been corrected for accuracy. In the Introduction, "Coronaviruses have the largest RNA genomes among all viruses." now reads: "Coronaviruses have some of the largest RNA genomes among all viruses."
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, Y., Mao, JM., Wang, GD. et al. Human SARS-CoV-2 has evolved to reduce CG dinucleotide in its open reading frames. Sci Rep 10, 12331 (2020). https://doi.org/10.1038/s41598-020-69342-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-69342-y
This article is cited by
-
Ribosome biogenesis in disease: new players and therapeutic targets
Signal Transduction and Targeted Therapy (2023)
-
The DNA glycosylase NEIL2 is protective during SARS-CoV-2 infection
Nature Communications (2023)
-
A comparison of five Illumina, Ion Torrent, and nanopore sequencing technology-based approaches for whole genome sequencing of SARS-CoV-2
European Journal of Clinical Microbiology & Infectious Diseases (2023)
-
Comparative genomic analysis of the human genome and six bat genomes using unsupervised machine learning: Mb-level CpG and TFBS islands
BMC Genomics (2022)
-
The low abundance of CpG in the SARS-CoV-2 genome is not an evolutionarily signature of ZAP
Scientific Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.