Abstract
Natural coastal microbial mat communities are multi-species assemblages that experience fluctuating environmental conditions and are shaped by resource competition as well as by cooperation. Laboratory studies rarely address the natural complexity of microbial communities but are usually limited to homogeneous mono-cultures of key species grown in liquid media. The mat-forming filamentous cyanobacteria Lyngbya aestuarii and Coleofasciculus chthonoplastes were cultured under different conditions to investigate the expression of circadian clock genes and genes that are under their control. The cyanobacteria were grown in liquid medium or on a solid substrate (glass beads) as mono- or as co-cultures under a light–dark regime and subsequently transferred to continuous light. TaqMan-probe based qPCR assays were used to quantify the expression of the circadian clock genes kaiA, kaiB, and kaiC, and of four genes that are under control of the circadian clock: psbA, nifH, ftsZ, and prx. Expression of kaiABC was influenced by co-culturing the cyanobacteria and whether grown in liquid media or on a solid substrate. Free-running (i.e. under continuous light) expression cycle of the circadian clock genes was observed in L. aestuarii but not in C. chthonoplastes. In the former organism, maximum expression of psbA and nifH occurred temporally separated and independent of the light regime, although the peak shifted in time when the culture was transferred to continuous illumination. Although functionally similar, both species of cyanobacteria displayed different 24-h transcriptional patterns in response to the experimental treatments, suggesting that their circadian clocks have adapted to different life strategies adopted by these mat-forming cyanobacteria.
Similar content being viewed by others
Introduction
The study of the ecology and evolution of natural microbial assemblages is often hindered by the complexity of these systems as well as by the fluctuations of environmental conditions. Laboratory experiments avoid these drawbacks and their outcomes are sometimes extrapolated in an attempt to understand the ecology of the microbial community in nature. Such laboratory experiments are usually carried out using pure cultures growing under well-defined conditions and high nutrient concentrations and under constant temperature and light. However, the results of these experiments often do not allow to predict the natural behaviour of the microorganisms. The emerging field of synthetic microbial ecology tackles these shortcomings by combining a lower complexity with mimicking conditions of natural ecosystems. This holds the promise of a better understanding of the ecology and evolution of microorganisms in natural ecosystems1,2. Currently, applications of synthetic microbial ecology address processes such as recycling of waste products3, industrial fermentation4, dairy industry5, production of chemical compounds6, and improvement of human health through fecal transplantation7.
Coastal microbial mats are amongst the most complex and diverse ecosystems8 and are exposed to microscale gradients (among others light, oxygen, salinity, and sulfide) fluctuating with the day-night and/or tidal cycles9,10,11. The dominant primary producers in these mats are cyanobacteria that form complex assemblages and maintain tight interactions with microorganisms belonging to different functional groups9. Cyanobacteria possess a circadian clock, a well-characterized molecular mechanism that produces approximately 24-h rhythms in gene expression and protein activity12. True circadian rhythms persist even under continuous illumination (free-run), and are able to shift their phases (timing of highest and lowest expression) according to environmental variations (entrainment), while their period is more or less insensitive to temperature (temperature compensated)13,14,15. The endogenous rhythmicity enables cyanobacteria to anticipate and react to daily environmental changes thereby enhancing their fitness16,17. The core genes involved in the cyanobacterial circadian clock are kaiA, kaiB and kaiC. They encode proteins that generate circadian rhythms of KaiC phosphorylation. The phosphorylation state of KaiC is a key regulator of the transcription-translation machinery of the cell. Overexpression of KaiA increases KaiC autophosphorylation while excess of KaiB leads to KaiC dephosphorylation. Synchronization to the environment is regulated by KaiA and CikA through the indirect sensing of light. The bacteriophytochrome-like circadian input kinase CikA plays an important part in the timing of the circadian period by sensing changes in light intensity through the redox state of the plastoquinone pool18 and it also acts as a phosphatase that is regulated by the KaiB-KaiC protein complex19. Entrainment of the cyanobacterial circadian clock happens also through photosynthetic activity that produces ATP, which is—amongst others—a fundamental requirement for the phosphorylation of KaiC and the production of circadian rhythmicity20,21.
The nature of the cyanobacterial circadian clock has been vastly studied in planktonic mono-cultures of the unicellular cyanobacterium Synechococcus elongatus PCC794222. The attractiveness to study this model strain is its small genome size, simple cellular structure, accessibility to genetic modification and comprehensive standardization of high-throughput assays. However, little is known about how circadian clock-controlled gene expression is regulated in complex communities and whether the environmental complexity and presence of other species is of influence.
A potential external source of influence on rhythmic gene expression is the complexity of the environment, as was reported for Staphylococcus aureus and Vibrio cholerae in which gene regulation and metabolic activity depended on whether these bacteria were grown as a homogenized liquid culture or biofilm-grown23,24.
An additional source of influence on gene expression might be presented by another, independent circadian oscillator which is linked to the ubiquitous antioxidant enzyme peroxiredoxin. Peroxiredoxin follows a 24-h oxidation–reduction cycle and is highly conserved in all domains of life25, however, the underlying mechanisms are not well understood.
To investigate whether growth condition and/or species interaction influences circadian gene expression patterns, co-cultures of two filamentous, non-heterocystous cyanobacteria Lyngbya aestuarii and Coleofasciculus chthonoplastes were subjected to quantitative PCR-based gene expression analysis. This technique was chosen over metatranscriptomics due to the large number of variables and replicates tested that add up to 648 samples to be analysed. In addition, alternative techniques like the use of reporter genes are not yet available for the two cyanobacterial species. aTargeted genes were the circadian clock controlling genes kaiABC, and the clock dependent genes nifH (encoding the nitrogenase iron protein), psbA (encoding photosystem II protein D1), ftsZ (encoding a cell division protein) and prx (encoding the peroxiredoxin protein). The quantified expression was normalized against the expression of two housekeeping genes, rnpA (ribonuclease P protein) and ppc (phosphoenolpyruvate carboxylase). The species were grown in either liquid media or biofilm-grown on glass beads and as mono- or co-cultures. Furthermore, after a standard light–dark regime, the cultures were exposed to continuous light to study the free-running circadian clock.
Results
Total RNA and qPCR statistics
The extracted total RNA was of good quality (average RIN: 7.7 ± 1) and yielded higher concentrations from C. chthonoplastes than from L. aestuarii (Table 1). Co-cultures yielded less total RNA than mono-cultures. Similarly, biofilm-grown cultures yielded less total RNA when compared to liquid cultures (Table 1).
The qPCR-derived Ct values of the biological replicates of the targeted circadian clock gene cluster kaiABC, nitrogenase gene nifH, cell division protein gene ftsZ, photosynthesis D1 protein gene psbA, peroxiredoxin gene prx and two commonly used cyanobacterial housekeeping genes, rnpA and ppc ranged from 23 to 39 in L. aestuarii and from 21 to 38 in C. chthonoplastes. The average r2 and amplification efficiencies of the qPCR assays were 0.95 and 100% in L. aestuarii and 0.92 and 92% in C. chthonoplastes (Table 2). Regardless the taxon and treatment, the highest absolute transcription was observed for the photosystem II protein D1 coding gene psbA with on average 153,425 gene-copies/µl in L. aestuarii and 78,592 gene-copies/µl in C. chthonoplastes.
Validation of housekeeping genes
Each qPCR run was accompanied by the quantitation of housekeeping genes for normalization. The genes rnpA (protein component of ribonuclease P) and ppc (phosphoenolpyruvate carboxylase) were selected as potential housekeeping genes (HKG) based on a previous qPCR study of L. aestuarii26. Validation of the stability of expression of these genes under different growth conditions was tested using the housekeeping gene determination software package BestKeeper27 of which standard deviation (SD) and correlation coefficient (r2) were used to verify HKG stability. According to the BestKeeper results for all treatments and light regimes, both HKGs revealed a stable expression in L. aestuarii (rnpA: SD = ± 0.31–0.71, r2 = 0.89–0.99; ppC: SD = ± 0.30–0.71, r2 = 0.90–0.99) and C. chthonoplastes (rnpA: SD = ± 0.33–0.65, r2 = 0.83–0.96; ppc: SD = ± 0.31–0.67, r2 = 0.87–0.95) and therefore both are suitable housekeeping genes for both species (Table 3).
Effect of culturing on gene expression
A constrained correspondence analysis (CCA) was carried out in order to investigate the influence of culturing on the expression patterns of the target genes (Figs. 1 and 2 for L. aestuarii and C. chthonoplastes, respectively.). Housekeeping gene-normalized transcript levels at the different time points served as input data. The results of the CCAs were subjected to an analysis of variance like permutation test provided by the ‘vegan: Adonis’ package in R28 providing Chi-square (Χ2) and probability values (p(> F)) (p < 0.05) for each treatment (culture condition and light regime) (Table 4). For visualization purposes CCA plots were used to display data relationships (Figs. 1 and 2). For each strain, comparisons were performed relative to cells grown under “standard” laboratory conditions; i.e. liquid mono-cultures grown under LD light regime. Boxplots of the original data per gene per growth condition are presented in Figs. 3 and 4 for L. aestuarii and C. chthonoplastes, respectively. A direct comparison of the average gene expression levels between the different growth conditions is presented in supplementary figures S1A and B for L. aestuarii and C. chthonoplastes, respectively.
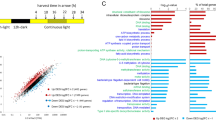
CCA plots of kaiABC, prx, psbA, nifH and ftsZ expression patterns in L. aestuarii. These genes were significantly influenced (p < 0.05, 95% confidence ellipse) by one or more of the treatments (L = liquid culture, B = biofilm-grown culture, M = mono culture, C = co-culture). Only conditions that were significantly affecting gene expression patterns are shown.
CCA plots of kaiABC, prx, psbA, nifH and ftsZ expression patterns of C. chthonoplastes. These genes were significantly influenced (p < 0.05, 95% confidence ellipse) by one or more of the treatments (L = liquid culture, B = biofilm-grown culture, M = mono culture, C = co-culture). Only conditions that were significantly affecting gene expression patterns are shown.
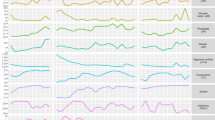
Boxplots presenting the relative normalized expression patterns in liquid mono- (ML) and co-cultures (CL) and biofilm-grown mono- (MB) and co-cultures (CB) of L. aestuarii under (a) LD and (b) LL regime. X-axes show Zeitgeber time (ZT: light–dark regime) (a) and circadian time (CT: constant light regime) (b). Red boxplots display the on average highest expression. Expressions patterns which display a blue (dotted) line display significant (p < 0.05) circadian rhythmicity as estimated by MetaCycle29. The dotted lines indicated significantly rhythmicity under LD illumination, while continuous lines symbolize significant rhythmicity under both light regimes (a, b). Boxplots display the minimum (lower error bar), maximum (upper error bar), median (horizontal line) and the first (box below the median) and third quartile (box above the median) of the expression values of the biological replicates (n = 3) per sample. A bar on top of the first row of the boxplots indicates the dark (black) and light (white) period (a). Treatments are distributed into columns and each row represents a gene.
Boxplots presenting the relative normalized gene expression patterns in liquid mono-(ML) and co-cultures (CL) and biofilm-grown mono-(MB) and co-cultures (CB) of C. chthonoplastes under a (a) LD and (b) LL regime. X-axes show Zeitgeber (ZT: light–dark regime) (a) and circadian (CT: constant light regime) (b). Red boxplots display the on average highest expression. Expressions patterns which display a blue (dotted) line display significant (p < 0.05) circadian rhythmicity as estimated by MetaCycle29. The dotted lines indicated significantly rhythmicity under LD illumination, while continuous lines symbolize significant rhythmicity under both light regimes (a, b). Boxplots indicate the minimum (lower error bar), maximum (upper error bar), median (horizontal line) and the first (box below the median) and third quartile (box above the median) of the expression values of the biological replicates (n = 3) per sample. A bar on top of the first row of the boxplots indicates the dark (black) and light (white) period (a). A bar on top of the first row of the boxplots indicates the dark (black) and light (white) period (a). Treatments are distributed into columns and each row represents a gene.
kaiABC expression in L. aestuarii
Comparison of the 24-h expression patterns of kaiA, kaiB and kaiC growing under different conditions (Fig. 1) revealed that their expression was significantly influenced by the way of culturing (liquid versus biofilm-grown growth; Χ2 = 0.033–0.074, p = 0.001–0.002) as well as whether co-cultured or as mono-culture (Χ2 = 0.019–0.212, p = 0.001–0.003) (Table 4) and explained ~ 81% of the observed variance in their expression. Unexpectedly, only kaiA was significantly influenced by the light regime (Χ2 = 0.017, p = 0.02) (Table 4). Potential rhythmic expression was deduced from comparing and fitting the expression patterns with a sine curve using the circadian cycle predictor program MetaCycle29 in accordance with a previous study in natural microbial mats30 (Supplementary Figure S2). Expression of kaiA followed a significant sinusoidal curve only in co-cultures under all growth conditions (p = 0.0084–0.01) and in biofilm-grown mono-culture under continuous illumination (p = 0.005–0.007) (Fig. 3a, b, Supplementary Figure S2). Highest expression of kaiA was observed in the light period, at point ZT9 in liquid mono- and co-cultures (ML & CL) and at ZT0 in biofilm-grown cultures (MB & CB) (Supplementary Figure S1A). A sinusoidal pattern for kaiB was only evident when biofilm-grown (LD-CB, LL-MB and LL-CB) (p = 0.006–0.02) (Fig. 3a, b, Supplementary Figure S2). Peak expression of kaiB varied with whether the organisms were co-cultured or not and ranged from ZT9 (ML, CL and MB) to ZT4.5 (CB). The highest peak expression measured for kaiB was observed in biofilm-grown co-cultures (Fig. 3a, Supplementary Figure S1A). Sinusoidal expression patterns for kaiC were mainly observed under continuous illumination (LL-ML, LL-CL and LL-MB) (p = 0.005–0.03) and in biofilm-grown mono-cultures under LD conditions (LD-MB) (p = 0.01) (Fig. 3a, b, Supplementary Figure S2). The time point of maximum kaiC expression (ZT4.5) was largely unaffected whether co-cultured or not but changed from ZT4.5 in liquid cultures to ZT9/ZT14.5 in biofilm-grown cultures. Under continuous illumination, a similar pattern was observed for kaiC and differences in peak expression were also only observed between liquid (peak at CT4.5) and biofilm-grown (peak at CT9) cultures (Fig. 3b, Supplementary Figure S1A).
prx expression in L. aestuarii
Transcription of prx was overall low and was significantly affected when grown as liquid and biofilm-grown culture (Χ2 = 0.187, p = 0.005) (Table 4). A sinusoidal pattern was observed for biofilm-grown mono-cultures (MB) under LD illumination (p = 0.01) and for CL and MB under continuous illumination (p = 0.004–0.01) (Fig. 3a, b, Supplementary Figure S2). Highest expression of prx was observed in the late dark/early light period (ZT21, ZT0) in liquid mono-cultures and in biofilm-grown mono- and co-cultures. These expression peaks shifted in liquid cultures, grown under continuous light, to CT14.5 (ML) and CT17.45 (CL) and to CT4.5 (MB) and CT0 (CB) in biofilm-grown cultures (Supplementary Figure S1A).
psbA expression in L. aestuarii
The variation of expression of psbA in L. aestuarii during a 24-h period differs significantly between different light regimes (LD versus-LL; Χ2 = 0.307, p = 0.001) and the way of cultivation (biofilm-grown versus liquid; Χ2 = 0.094, p = 0.003) (Fig. 1, Table 4). Cultures growing under a light–dark regime revealed a sinusoidal pattern of psbA gene expression (p = 0.007–0.02) (Fig. 3a, b, Supplementary Figure S2) with the exception of CB cultures. Highest psbA expression was found during the beginning of the light period (ZT4.5-ZT9) followed by a sharp drop in expression in liquid cultures at ZT14.5 (Fig. 3a, b, Supplementary Figure S1A). In biofilm-grown cultures the overall expression level of psbA was lower than in the liquid cultures, while in CB cultures the highest expression was at the end of the light, beginning of the dark period (ZT14.5-ZT17.45). Under continuous illumination the overall expression levels in liquid cultures were slightly lower and a shift in expression pattern was observed with highest expression at ZT14.5-ZT17.45 (Fig. 3b, Supplementary Figure S2). The sinusoidal pattern was maintained under continuous illumination for CL and MB conditions biofilm-grown (p = 0.01–0.04) (Fig. 3a, b, Supplementary Figure S2).
nifH expression in L. aestuarii
Under the standard dark–light regime, nifH expression was low during the light and was highest during the dark (ZT17.45-ZT21) (Fig. 3a, Supplementary Figure S1A). Moreover, nifH expression was significantly higher in liquid cultures compared to the biofilm-grown cultures. A significant sinusoidal curve under LD illumination was observed under all conditions (p = 0.005–0.01) except for the co-cultured biofilm-grown cultures (CB) (Fig. 3a, Supplementary Figure S2). The imposed light regime significantly affected the daily transcription pattern of nifH (Χ2 = 0.296, p = 0.001) (Fig. 1, Table 4). Under continuous light, the pattern of expression in the liquid cultures peaked more often than in biofilm-grown cultures with high expression at CT0, CT4.5 and CT21. (Fig. 3b, Supplementary Figure S1A). In biofilm-grown mono-cultures nifH expression peaked at CT14.5 and CT21 and in co-cultures at CT17.45. No significant sinusoidal nifH gene expression was found under continuous illumination.
ftsZ expression in L. aestuarii
The variation in expression of ftsZ in L. aestuarii during a 24-h period was large and only in co-cultured biofilm-grown cultures at LD illumination a sinusoidal like curve was observed with a maximum at ZT0 (p = 0.03) (Fig. 3a, Supplementary Figure S2). In contrast, under continuous illumination an opposite pattern is found with all the three other growth conditions providing significant sinusoidal ftsZ expression (p = 0.008–0.01) (Fig. 3b, Supplementary Figure S1A). Due to other more variable patterns, each of the imposed cultivation conditions appeared to contribute significantly to the gene expression pattern (liquid vs. biofilm-grown: Χ2 = 0.038, p = 0.001; LD/LL: Χ2 = 0.020, p = 0.004 and mono-/co-culture: Χ2 = 0.013, p = 0.012) (Fig. 1, Table 4).
kaiABC expression in C. chthonoplastes
The expression pattern of kaiA, kaiB and kaiC of C. chthonoplastes grown under a LD regime changed significantly when transferred to continuous light (Χ2 = 0.049–0.058, p = 0.001–0.007) (Fig. 2, Table 4). Furthermore, the expression of kaiA and kaiC differed when the organism was grown as a mono-culture or co-cultured with L. aestuarii (mono- vs. co-culture: Χ2 = 0.053–0.077, p = 0.001), while the way of culturing significantly impacted on the expression of kaiB and kaiC during a 24-h period (liquid- vs. biofilm-grown-culture: Χ2 = 0.040–0.833, p = 0.001–0.009) (Fig. 2, Table 4).
The expression of kaiA and kaiB did not seem to follow a particular pattern and only in liquid co-cultures a potential sinusoidal curve was discerned for kaiA expression (p = 0.005) and in biofilm-grown mono-cultures for kaiB (p = 0.03) (Fig. 4a, Supplementary Figure S1). Peak expression for kaiA in liquid co-cultures and biofilm-grown cultures was at the beginning of the light period (ZT0) and at the end of the dark period (ZT21) and peaked at ZT0 and ZT9 in liquid mono-cultures (Fig. 4a). A sinusoidal distribution was also observed for kaiC expression of cells grown in ML and CB conditions with respective highest expression at ZT0 and ZT4.5 (p = 0.01) (Fig. 4a, Supplementary Figure S2).
Under continuous illumination, kaiA expression was sinusoidal at LL- MB (p = 0.01) and kaiB at LL-CB (p = 0.003) with peak expression at CT17.5 and CT14.5 (Fig. 4b, Supplementary Figure S2). Potential rhythmic expression of kaiC under LL conditions was not observed.
prx expression in C. chthonoplastes
The expression pattern of prx in C. chthonoplastes differed significantly between a mono-culture and co-culture with L. aestuarii (Χ2 = 0.263, p = 0.001), and between a liquid medium and biofilm-grown culture (Χ2 = 0.053, p = 0.002), as well as upon transfer to continuous light (Χ2 = 0.022, p = 0.007) (Fig. 2, Table 4). Overall, the expression of prx was 5- (LL) to tenfold (LD) higher in biofilm-grown than in liquid culture (Fig. 4a, b, Supplementary Figure S1B). Expression of prx under standard conditions (ML-LD) gradually increased from the beginning of light to the dark period while an almost opposite trend was observed in MB cultures with highest expression at time points ZT0 and ZT21. A sinusoidal pattern was observed in the co-cultures and in biofilm-grown mono-cultures independent of the light regimes tested (p = 0.006–0.04) (Fig. 4a, b, Supplementary Figure S2).
psbA expression in C. chthonoplastes
Expression of psbA in C. chthonoplastes was significantly influenced by two of the growth conditions (mono-/co-culture: Χ2 = 0.029, p = 0.029, liquid vs. biofilm-grown: Χ2 = 0.028, p = 0.031) (Fig. 2, Table 4). A sinusoidal pattern was observed for psbA under CB conditions at LD illumination (p = 0.01) and under CL and MB conditions under continuous illumination (p = 0.003–0.01) (Fig. 4a, b, Supplementary Figure S2). The expression of psbA in LD-grown cultures peaked during the early dark to early light period (ZT17.45-ZT0) (Fig. 4a, Supplementary Figure S1B). Under continuous illumination, co-cultures and MB cultures revealed peaks in the expression of psbA between CT14.5 and CT21, while the ML culture was characterized by high psbA expression at CT0 (Fig. 4b, Supplementary Figure S1B).
nifH expression in C. chthonoplastes
The expression of nifH was significantly affected by the way of culturing (liquid vs. biofilm-grown: Χ2 = 0.266, p = 0.001, mono-/co-culture: Χ2 = 0.099, p = 0.029) and by transfer to constant light (Χ2 = 0.212, p = 0.001) (Fig. 2, Table 4). Relative nifH expression in biofilm-grown, co-culture-grown and LL-grown cultures displayed less fluctuation than displayed in liquid-grown, mono-culture-grown and LD-grown cultures. Expression of nifH in C. chthonoplastes was overall low with a single peak at the late dark/early light period in liquid cultures (ZT21 (ML), ZT0 (CL)) and during the light period in biofilm-grown cultures (ZT9 (MB), ZT4.5 (CB)) (Fig. 4a, Supplementary Figure S1B). In the liquid mono-culture at LL a single peak in the expression of nifH was observed at CT0 (Fig. 4b, Supplementary Figure S1B). No significant sinusoidal patterns were found for C. chthonoplastes nifH expression under any of the tested conditions.
ftsZ expression in C. chthonoplastes
Expression of the cell division protein coding gene ftsZ was significantly affected whether co-cultured or not (Χ2 = 0.122, p = 0.001) and liquid- or biofilm-grown culture (Χ2 = 0.025, p = 0.016) (Fig. 2, Table 4). The patterns of expression of ftsZ during a 24-h period varied. A sinusoidal curve was only observed in mono-cultures (p = 0.006–0.01) and at LL-CL and LL-MB (p = 0.008–0.04) (Fig. 4a, b, Supplementary Figure S2) with peak expression during the late dark and early light period in LD-grown cultures and at same time points in LL-grown cultures. (Fig. 4a, b, Supplementary Figure S1B).
Discussion
The circadian clock of L. aestuarii and C. chthonoplastes
Rhythmic gene expression in cyanobacteria involves the following three steps: (1) the external cues, mostly light, temperature and redox potential that (2) entrain the “clock” proteins that control the rhythmic expression of (3) the functional protein coding genes involved in processes that are separated in time.
The only controlled rhythmic cue in our experimental set-up was the illumination period while temperature was kept the same in all experiments. Given the large number of variables tested within one cycle and the need for at least 6 time points per cycle, we were not able to assay over a 48 h or longer period that would be needed to determine truly circadian control. However, the observed significant differences in expression patterns are sufficient to infer that circadian control may also be affected by the greater complexity of natural conditions.
Both the way of culturing (liquid or biofilm-grown) as well as whether the cyanobacteria were co- or mono-cultured affected the expression of the circadian clock genes kaiA, kaiB, and kaiC in both species, albeit in different ways.
The maximum expression of the circadian clock genes kaiABC of L. aestuarii shifted 4–8 h between liquid and biofilm-grown cultures. Shifting the cultures to continuous illumination for 2 days, from which samples were taken on the second day, had little influence on the variation of expression of the three kai genes over a 24-h period, which is indicative of the free-running cyanobacterial clock as has been previously shown for unicellular cyanobacteria31,32. In contrast, the cultures of C. chthonoplastes did not display a free-running clock. Under continuous light, maximum gene expression of kaiABC in this organism shifted in regard to what has been found under a dark–light regime when grown in biofilm-grown cultures and co-culture.
Based on the aforementioned results we suggest that, despite being both members of the order of Oscillatoriales, the two species display major differences in how stringent the control over their circadian clocks is. These differences may be attributed to the life strategies of the two cyanobacteria regarding their response to high light. For example, C. chthonoplastes avoids light stress by migration33,34, which may trigger several transcriptional responses in line with a metabolism that is not optimally adapted to continuous high light intensities. In contrast, Lyngbya only moves when differentiating into hormogonia: short, sheet-less motile trichomes35, and therefore cannot rapidly avoid the imposed light stress. However, Lyngbya species evolved strategies to endure high light, desiccation and UV-stress. To avoid the latter, Lyngbya species synthesize UV-screening compounds such as mycosporine amino acid-like substances (MAAs) and carotenoids, in addition to coiling of their trichomes36. These strategies may not require additional transcriptional regulation in response to continuous illumination and might explain why the expression of kaiABC in Lyngbya sp. does not differ between the two illumination set ups. The observed expression patterns in C. chthonoplastes are less well understood and perhaps appear more similar to an ‘hourglass’ clock, which has been described for kaiA-lacking species such as Prochlorococcus marinus37. However, since C. chthonoplastes does have, and expresses the kaiA gene, the differences in expression with L. aestuarii may reflect specific adaptation to their natural habitat. In support of this assumption is the low identity between kaiA nucleotide and protein sequences (< 60% identity) of L. aestuarii and C. chthonoplastes (and also compared to the kaiA sequence of S. elongatus), which is also the lowest in comparison to kaiB and kaiC (> 75% identity), suggesting that kaiA is the least conserved gene and may experience the lowest selective pressure among the kai genes (Supplementary Table S1, Supplementary Figure S3). Even though the two species co-occur in many marine microbial mats, L. aestuarii and C. chthonoplastes may occupy different ecological niches that allow them to share light and micronutrients using differently tuned circadian clocks.
A kaiABC-independent circadian oscillator has been linked to the periodicity of peroxiredoxin which follows the generation of stress-induced reactive oxygen species25,38. Overall, the expression patterns of prx reveal a significant sinusoidal curve over the 24-h sampling period for most conditions except the standard liquid mono-cultures in C. chthonoplastes and under some conditions in L. aestuarii. However, the expression levels of prx shifted significantly between mono- and co-cultures of liquid (L. aestuarii) and biofilm-grown (C. chthonoplastes) cultures. Highest prx expression was found in mono-cultures in the dark to early light period. This was also found in Crocosphaera watsonii strain WH8501 and it was attributed to the accumulation of reactive oxygen species (ROS) towards the end of the light period39. In contrast to mono-cultures, prx expression in co-cultures of L. aestuarii and C. chthonoplastes displayed maxima towards the late light period. This would suggest that the accumulation of ROS occurs faster in co-cultures as the result of a higher raise of oxygen concentration due to the combined photosynthetic activity. Robust free-running prx expression25 without a peak shift was only found in co-cultures of liquid (C. chthonoplastes) and biofilm-grown (L. aestuarii) cultures under continuous illumination. Regardless of the illumination regime and treatment, average prx levels of C. chthonoplastes were more than a 100-fold higher than in cultures of L. aestuarii and showed a tenfold increase in biofilm-grown cultures in comparison to liquid cultures. The large transcriptional discrepancy between the species’ prx levels and between C. chthonoplastes’ liquid and biofilm-grown cultures suggests a higher sensitivity of C. chthonoplastes and especially of its biofilm-grown cultures, to oxidative stress and explains its migratory behaviour to avoid high light intensities.
The expression of circadian clock regulated genes in mono- and co-cultures grown in free-living or biofilm-grown mode
Depending on growth conditions, expression of psbA in C. chthonoplastes and L. aestuarii, and nifH in L. aestuarii followed a sinusoidal pattern during the 24-h period. In the unicellular cyanobacteria Cyanothece sp., expression of psbA and nifH is under control of the kaiABC regulatory network40 similar as for psbA expression in S. elongatus41. The peak in expression of psbA in the early light period and of nifH in the dark is typical for non-heterocystous, circadian clock controlled cyanobacteria in which nitrogenase activity is confined to the low light periods where photosynthetically produced oxygen is low42,43. Remarkably, we did not find such distinction between the peaks of psbA and nifH expression in L. aestuarii grown as a biofilm-grown co-cultured with C. chthonoplastes. Instead, both psbA and nifH peaked during the dark. It is unclear why the expression of psbA shifted to the dark in biofilm-grown co-cultures. Moreover, under LD conditions psbA and nifH peak transcription levels in L. aestuarii were up to 11 times lower in biofilm-grown cultures compared to liquid cultures. Albeit less distinct, the same was seen in C. chthonoplastes. The lower expression of psbA in biofilm-grown cultures may have been caused by the lower exposure to light of the biofilm-grown trichomes or by a decreased rate of growth as has been observed in aggregated cells in biofilms of Escherichia coli44. However, the latter explanation was not supported by lower transcription levels of the cell division gene ftsZ in biofilm-grown cultures. Alternatively, as is the case in S. elongatus, reactive oxygen species may target the de novo synthesis of psbA mRNA45,46 decreasing the rate of photosynthesis47. This explanation is supported by the increase of prx expression in biofilm-grown cultures that is possibly a response to an increase in ROS.
When L. aestuarii was transferred to continuous illumination the expression peak of psbA shifted 4–8 h later during the 24-h cycle. In contrast, C. chthonoplastes, which does not display a free-running circadian rhythm, the peak of the expression of psbA did not shift when the culture was transferred to constant illumination. These results challenge the degree of control of the circadian clock on the expression of psbA in these cyanobacteria and call for another controlling mechanism.
The measurement of expression of nifH in C. chthonoplastes was hindered by technical difficulties in the qPCR reaction yielding in some runs false positive reactions above the threshold level in the non-template controls, which was attributed to a known reagent contamination problem48. However, this problem is mostly neglectable when large amounts of species specific nifH containing DNA is added as template. With some caution we conclude that nifH expression in C. chthonoplastes revealed low transcript numbers and lack of temporal separation from psbA expression. This is unusual for non-heterocystous cyanobacteria. Moreover, it is still uncertain whether C. chthonoplastes actually fixes atmospheric dinitrogen since it has not yet been possible to grow the species diazotrophically in culture49. Although this cyanobacterium possesses the nifHDK gene cluster50, it lacks the accessory genes nifOTWXZ, which are present in, for instance, L. aestuarii51 . The function of these accessory nif genes is not well understood and it is therefore not certain whether they are indispensable for N2 fixation. Daily patterns of nifH expression in biofilm-grown C. chthonoplastes cultures are similar to those observed in a coastal microbial mat52 as well as metatranscriptomes from the same mat30. These studies report that nifH expression is high in the beginning and/ or middle of the light period.
In conclusion, comparison of gene expression patterns between laboratory cultures and field samples are rare and may be contradictory. The daily variations of gene expression obtained from the metatranscriptome of oceanic surface water containing abundant picocyanobacteria (Synechococcus) were similar to those obtained from liquid cultures53. The laboratory conditions for liquid cultures may better represent the natural conditions of these planktonic species than would be the case for benthic microorganisms. The transcriptomes of Salinibacter ruber and Haloquadratum walsbyi reveal large differences in relative expression levels in a subset of genes while relative expression levels of other genes are well conserved in the metatranscriptomes from natural samples in which these organisms thrive (Bolhuis, unpublished data).
Our study shows that gene expression in two filamentous non-heterocystous cyanobacteria respond differently to external stimuli and potential zeitgebers. Phase shifts as well as changes in expression levels occurred as a function of the mode of growth (liquid medium versus biofilm-grown culture) and to the presence of the other competitor species. The observed species-specific expression patterns most likely reflect different life strategies, hinted at by differences in expression of prx, psbA, and nifH. In this study, neither co- or mono-cultivation nor liquid or biofilm-grown growth could be held as main responsible effector for the observed differences in gene expression patterns. However, in combination these effectors are able to alter gene expression not only in the laboratory but also in nature. As a consequence, most laboratory derived cyanobacterial circadian clock (controlled) expression patterns are insufficient predictors for expression patterns in the field. This emphasizes the need to mimic the natural environment when testing bacteria in the laboratory.
Material and methods
Experimental setup
Stock cultures of the filamentous, non-heterocystous cyanobacteria Lyngbya aestuarii PCC8106 (synonym = Lyngbya sp. CCY9616) and Coleofasciculus (Microcoleus) chthonoplastes PCC7420 (CCY9604) served as inoculum for the experiments. These cyanobacteria were isolated from similar coastal microbial mats but at different times and at a large geographical distance (Mellum, Germany (L. aestuarii) and Woods Hole, USA (C. chthonoplastes)). Although the two species, both members of the order Oscillatoriales, are naturally co-occurring in coastal microbial mats and in many other similar habitats54,55,56,57, the isolates tested here did not have any previous interaction other than the three weeks of co-culturing for the experiments described here. The cyanobacteria were grown at 23 °C under continuous fluorescent light (L: photon density: 40 µmol m−2 s−1) in 40 ml culture flasks (TPP, Switzerland) containing 25 ml of BA + medium (1:1 mix of BG11 and ASN 3) (www.dsmz.de) with added nitrate ( +) (13.29 mM). The culturing was done in triplicate. Liquid cultures (L) were grown in the 40 ml TP culture flasks. Biofilm-grown cultures (B) were grown in 6-well culture plates equipped with filter trays (0.4 µm pore size) (NUNC™, Denmark), which were filled with sterile glass beads (3.5 g/well) (ø 0.1 mm) and 4 ml/well of BA + medium (which did not submerse the beads). Sterile tweezers and disposable sterile Pasteur pipettes (VWR, USA) were used to distribute equally sized pieces of cyanobacterial trichomes on the glass beads in order to obtain an even distribution. Co-culturing (C) was done by inoculating similar sized filaments of both species (1:1 ratio) in culture flasks or by placing their trichomes on the glass bead surface. The cultures were grown for three weeks under a 16 h light and 8 h dark cycle (LD) to entrain the circadian clock. Two days prior to sampling, half of the LD cultures of each treatment was exposed to constant light (LL) in order to test the free-running characteristics of the circadian clock. During the three-week culturing period, the cyanobacteria did not show signs of degradation or bleaching. Evaporation of medium during culturing was compensated by refilling wells and culture flasks with autoclaved MilliQ water.
Sampling and RNA extraction
RNA was extracted from L. aestuarii and C. chthonoplastes cultures grown in liquid medium (L) or biofilm-grown on glass beads (B) as mono- (M) or as co-culture (C). Sampling of cell material occurred at 6 time points during a light–dark regime (LD) indicated with the Zeitgeber Time (ZT) ZT0 (05:00 h), ZT4.5 (09:30 h), ZT9 (14:00 h), ZT14.5 (19:30 h), ZT17.45 (22:45 h) and ZT21 (02:00 h). Cultures transferred to continuous light (LL) were sampled at the corresponding time points of the LD series but were labelled as Circadian Time (CT). Samples were taken in biological triplicates and immediately submerged in liquid nitrogen in a sterile mortar and ground with a sterile pestle. From the liquid cultures an equal number of subsamples were taken and from the biofilm-grown samples a complete well with cell material was used for RNA extraction. After grinding, the material was placed in bead tubes provided by the ZR-Fungal/ Bacterial RNA MiniPrep isolation kit (ZYMO research, USA) and quickly frozen in liquid nitrogen prior to short-term storage at − 80 °C. RNA extraction was done according to the manufacturers’ protocol. Quantity, quality, and efficiency of DNA removal was measured after DNAse treatment of each RNA sample (TurboDNAse, LifeTechnology, USA) by using a Bioanalyzer 2,100 (Agilent Technologies, USA). Three µl of each of the DNAse treated samples containing 2 ng/µl RNA (RIN > 5) were reverse transcribed into cDNA using SuperScript III reverse transcriptase (200U/µl) and random hexamer primers (100 ng/μl) followed by an RNase treatment to remove residual RNA according to the manufacturers’ protocol (Life Technologies, USA).
Gene quantification
Based on the genome sequences of L. aestuarii and C. chthonoplastes, primers and TaqMan probes of the targeted circadian clock gene cluster kaiABC, the circadian clock reporter gene cikA, nitrogenase gene nifH, cell division protein gene ftsZ, photosynthesis D1 protein gene psbA, peroxiredoxin gene prx and two commonly used cyanobacterial housekeeping genes, rnpA and ppc26, were designed using the genetic analysis software Geneious R 8.1.758 (Table 2). The proper annealing temperatures of the primers were established by gradient PCR on a thermocycler (BioMetra) and specificity was confirmed by Sanger sequencing of the amplicons (BaseClear, Leiden, The Netherlands). Standard curves for RT-qPCR were prepared by using dilution series of PCR products and the primers were checked for species specificity by means of crossover PCRs and gel electrophoreses. Samples, standard curves and non-template controls were run in technical triplicates on a Rotor-Gene 6000™ (Qiagen, USA).
RT-qPCR amplification of the targeted and housekeeping genes was performed in 15 µl volumes containing 2.25 µl MilliQ, 7.5 µl 2 × Multiplex qPCR Perfecta Supermix (Quanta Biosciences), 0.38 µl (0.5 µM) of each target and housekeeping gene primer, 0.19 µl of each probe (0.25 µM) and 1.5 µl of template. RT-qPCR cycling was performed at an initial activation step of 95 °C for 2 min followed by 40 cycles at 95 °C for 10 s and at 64 °C for 60 s. Gene amplification efficiencies (%), r2, Ct values and transcript abundances (copies/µl) were retrieved from the program Rotor-Gen Q 2.1.0 (Qiagen) (Supplementary Table S2). Where needed, missing Ct values of biological replicates were surrogated by the median of the remaining biological replicates59.
Statistics
To evaluate the validity of the housekeeping genes across treatments, Ct values of genes of interest and housekeeping genes were used in the program BestKeeper27. T-tests were applied to examine the sample-wise expression within each genes’ expression profile and the influence of treatments on transcription was assessed using R-based CCA scripts and ANOVA (Χ2, F- and p values (< 0.05)). In these tests normalized transcript abundances ((\(\frac{GOI }{geomean(HKGs)}\))) of each gene served as input data. CCAs were plotted and supplemented with confidence ellipses (95%) of the standard expression of the different genes. Potential significant rhythmic gene expression following a sinusoidal curve was predicted by the R based script Metacycle29 using the transcription levels at different time points as input matrix. The cycle analysis was performed with the meta2D function that integrates multiple cycle prediction methods and combines their p-values. Training of the cycling algorhythm was performed by combining the datasets, thereby mimicking a 72 h sampling pattern and a p-value cut-off of 0.05 was used to determine potential sinusoidal patterns.
References
De Roy, K., Marzorati, M., Van den Abbeele, P., Van de Wiele, T. & Boon, N. Synthetic microbial ecosystems: an exciting tool to understand and apply microbial communities. Environ. Microbiol. 16, 1472–1481 (2014).
Mee, M. T. & Wang, H. H. Engineering ecosystems and synthetic ecologies. Mol. Biosyst. 8, 2470 (2012).
Hendrickx, L. et al. Microbial ecology of the closed artificial ecosystem MELiSSA (micro-ecological life support system alternative): reinventing and compartmentalizing the Earth’s food and oxygen regeneration system for long-haul space exploration missions. Res. Microbiol. 157, 77–86 (2006).
Chen, Y. Development and application of co-culture for ethanol production by co-fermentation of glucose and xylose: a systematic review. J. Ind. Microbiol. Biotechnol. 38, 581–597 (2011).
Spus, M. et al. Strain diversity and phage resistance in complex dairy starter cultures. J. Dairy Sci. 98, 5173–5182 (2015).
Ma, Q. et al. Integrated proteomic and metabolomic analysis of an artificial microbial community for two-step production of vitamin C. PLoS ONE 6, e26108 (2011).
Petrof, E. O. et al. Stool substitute transplant therapy for the eradication of Clostridium difficile infection: ‘RePOOPulating’ the gut. Microbiome 1, 3 (2013).
Bolhuis, H. & Stal, L. J. Analysis of bacterial and archaeal diversity in coastal microbial mats using massive parallel 16S rRNA gene tag sequencing. ISME J. 5, 1701–1712 (2011).
van Gemerden, H. Microbial mats: a joint venture. Mar. Geol. 113, 3–25 (1993).
Tolker-Nielsen, T. & Molin, S. Spatial organization of microbial biofilm communities. Microb. Ecol. 40, 75–84 (2000).
Bolhuis, H., Cretoiu, M. S. & Stal, L. J. Molecular ecology of microbial mats. FEMS Microbiol. Ecol. 90, 335–350 (2014).
Cohen, S. E. & Golden, S. S. Circadian rhythms in cyanobacteria. Microbiol. Mol. Biol. Rev. 79, 373–385 (2015).
Nakajima, M. et al. Reconstitution of circadian oscillation of cyanobacterial KaiC phosphorylation in vitro. Science 308, 414–415 (2005).
Murayama, Y. et al. Low temperature nullifies the circadian clock in cyanobacteria through Hopf bifurcation. Proc. Natl. Acad. Sci. USA. 114, 5641–5646 (2017).
Johnson, C. H., Zhao, C., Xu, Y. & Mori, T. Timing the day: what makes bacterial clocks tick?. Nat. Rev. Microbiol. 15, 232–242 (2017).
Woelfle, M. A., Ouyang, Y., Phanvijhitsiri, K. & Johnson, C. H. The adaptive value of circadian clocks: an experimental assessment in cyanobacteria. Curr. Biol. 14, 1481–1486 (2004).
Welkie, D. G. et al. A hard day’s night: cyanobacteria in diel cycles. Trends Microbiol. 27, 231–242 (2019).
Ivleva, N. B., Gao, T., LiWang, A. C. & Golden, S. S. Quinone sensing by the circadian input kinase of the cyanobacterial circadian clock. Proc. Natl. Acad. Sci. USA. 103, 17468–17473 (2006).
Gutu, A. & O’Shea, E. K. Two antagonistic clock-regulated histidine kinases time the activation of circadian gene expression. Mol. Cell 50, 288–294 (2013).
Rust, M. J., Golden, S. S. & O’Shea, E. K. Light-driven changes in energy metabolism directly entrain the cyanobacterial circadian oscillator. Science 331, 220–223 (2011).
Pattanayak, G. & Rust, M. J. The cyanobacterial clock and metabolism. Curr. Opin. Microbiol. 18, 90–95 (2014).
Mackey, S. R., Golden, S. S. & Ditty, J. L. The itty-bitty time machine genetics of the cyanobacterial circadian clock. Adv. Genet. 74, 13–53 (2011).
Resch, A., Rosenstein, R., Nerz, C. & Götz, F. Differential gene expression profiling of Staphylococcus aureus cultivated under biofilm and planktonic conditions. Appl. Environ. Microbiol. 71, 2663–2676 (2005).
Moorthy, S. & Watnick, P. I. Identification of novel stage-specific genetic requirements through whole genome transcription profiling of Vibrio cholerae biofilm development. Mol. Microbiol. 57, 1623–1635 (2005).
Edgar, R. S. et al. Peroxiredoxins are conserved markers of circadian rhythms. Nature 485, 459–464 (2012).
Pinto, F., Pacheco, C. C., Ferreira, D., Moradas-Ferreira, P. & Tamagnini, P. Selection of suitable reference genes for RT-qPCR analyses in cyanobacteria. PLoS ONE 7, 1–9 (2012).
Pfaffl, M. W., Tichopad, A., Prgomet, C. & Neuvians, T. P. Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: Bestkeeper–Excel-based tool using pair-wise correlations. Biotechnol. Lett. 26, 509–515 (2004).
Oksanen, J. et al. Package ‘vegan’ Title Community Ecology Package. Community Ecol. Packag. 2, (2019).
Wu, G., Anafi, R. C., Hughes, M. E., Kornacker, K. & Hogenesch, J. B. MetaCycle: an integrated R package to evaluate periodicity in large scale data. Bioinformatics 32, 3351–3353 (2016).
Hörnlein, C., Confurius-Guns, V., Stal, L. J. & Bolhuis, H. Daily rhythmicity in coastal microbial mats. npj Biofilms Microbiomes 4, 11 (2018).
Kondo, T. et al. Circadian rhythms in prokaryotes: luciferase as a reporter of circadian gene expression in cyanobacteria. Proc. Natl. Acad. Sci. USA. 90, 5672–5676 (1993).
Tomita, J. No transcription-translation feedback in circadian rhythm of KaiC phosphorylation. Science 307, 251–254 (2005).
Whale, G. F. & Walsby, A. E. Motility of the cyanobacterium Microcoleus chthonoplastes in mud. Br. Phycol. J. 19, 117–123 (1984).
Urmeneta, J., Navarrete, A., Huete, J. & Guerrero, R. Isolation and characterization of cyanobacteria from microbial mats of the Ebro Delta Spain. Curr. Microbiol. 46, 199–204 (2003).
Kothari, A., Vaughn, M. & Garcia-Pichel, F. Comparative genomic analyses of the cyanobacterium, Lyngbya aestuarii BL J, a powerful hydrogen producer. Front. Microbiol. 4, 363 (2013).
Rath, J. & Adhikary, S. P. Response of the estuarine cyanobacterium Lyngbya aestuarii to UV-B radiation. J. Appl. Phycol. 19, 529–536 (2007).
Holtzendorff, J. et al. Genome streamlining results in loss of robustness of the circadian clock in the marine cyanobacterium Prochlorococcus marinus PCC 9511. J. Biol. Rhythms 23, 187–199 (2008).
O’Neill, J. & Reddy, A. Circadian clocks in human red blood cells. Nature 469, 498–503 (2011).
Shi, T., Ilikchyan, I., Rabouille, S. & Zehr, J. P. Genome-wide analysis of diel gene expression in the unicellular N 2-fixing cyanobacterium Crocosphaera watsonii WH 8501. ISME J. 4, 621–632 (2010).
Červený, J., Sinetova, M. A., Valledor, L., Sherman, L. A. & Nedbal, L. Ultradian metabolic rhythm in the diazotrophic cyanobacterium Cyanothece sp. ATCC 51142. Proc. Natl. Acad. Sci. U. S. A. 110, 13210–5 (2013).
Johnson, C. H., Stewart, P. L. & Egli, M. The cyanobacterial circadian system: from biophysics to bioevolution. Annu. Rev. Biophys. 40, 143–167 (2011).
Mitsui, A. et al. Strategy by which nitrogen-fixing unicellular cyanobacteria grow photoautotrophically. Nature 323, 720–722 (1986).
Berman-Frank, I. et al. Segregation of nitrogen fixation and oxygenic photosynthesis in the marine cyanobacterium Trichodesmium. Science 294, 1534–1537 (2001).
Besharova, O., Suchanek, V. M., Hartmann, R., Drescher, K. & Sourjik, V. Diversification of gene expression during formation of static submerged biofilms by Escherichia coli. Front. Microbiol. 7, 1568 (2016).
Nishiyama, Y. et al. Oxidative stress inhibits the repair of photodamage to the photosynthetic machinery. EMBO J. 20, 5587–5594 (2001).
Nishiyama, Y., Allakhverdiev, S. I., Yamamoto, H., Hayashi, H. & Murata, N. Singlet oxygen inhibits the repair of photosystem II by suppressing the translation elongation of the D1 protein in Synechocystis sp. PCC 6803. Biochemistry 43, 11321–11330 (2004).
Los, D. A. & Zinchenko, V. V. in Lipids Photosynth. Essent. Regul. Funct. (eds. Wada, H. & Murata, N.) 329–348 (Springer Netherlands, 2010). doi:10.1007/978–90–481–2863–1_15
Zehr, J. P., Crumbliss, L. L., Church, M. J., Omoregie, E. O. & Jenkins, B. D. Nitrogenase genes in PCR and RT-PCR reagents: implications for studies of diversity of functional genes. Biotechniques 35, 996–1005 (2003).
Stal, L. J., Bolhuis, H. & Cretoiu, M. S. Phototrophic marine benthic microbiomes: the ecophysiology of these biological entities. Environ. Microbiol. 21, 1529-1551/////// (2019).
Bolhuis, H., Severin, I., Confurius-Guns, V., Wollenzien, U. I. a & Stal, L. J. Horizontal transfer of the nitrogen fixation gene cluster in the cyanobacterium Microcoleus chthonoplastes. ISME J. 4, 121–30 (2010).
Lee, S. H., Pulakat, L., Parker, K. C. & Gavini, N. Genetic analysis on the NifW by utilizing the yeast two-hybrid system revealed that the NifW of Azotobacter vinelandii interacts with the NifZ to form higher-order complexes. Biochem. Biophys. Res. Commun. 244, 498–504 (1998).
Severin, I. & Stal, L. J. NifH expression by five groups of phototrophs compared with nitrogenase activity in coastal microbial mats. FEMS Microbiol. Ecol. 73, 55–67 (2010).
Ottesen, E. a et al. Pattern and synchrony of gene expression among sympatric marine microbial populations. Proc. Natl. Acad. Sci. U. S. A. 110, E488–97 (2013).
Stal, L. J., Gemerden, H. & Krumbein, W. E. Structure and development of a benthic marine microbial mat. FEMS Microbiol. Lett. 31, 111–125 (1985).
Villbrandt, M. & Stal, L. J. The effect of sulfide on nitrogen fixation in heterocystous and non-heterocystous cyanobacterial mat communities. Algol. Stud. für Hydrobiol. Suppl. 83, 549–563 (1996).
Paerl, H. W., Pinckney, J. L. & Steppe, T. F. Cyanobacterial-bacterial mat consortia: Examining the functional unit of microbial survival and growth in extreme environments. Environ. Microbiol. 2, 11–26 (2000).
Fourçans, A. et al. Characterization of functional bacterial groups in a hypersaline microbial mat community (Salins-de-Giraud, Camargue, France). FEMS Microbiol. Ecol. 51, 55–70 (2004).
Kearse, M. et al. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28, 1647–1649 (2012).
Pabinger, S., Rödiger, S., Kriegner, A., Vierlinger, K. & Weinhäusel, A. A survey of tools for the analysis of quantitative PCR (qPCR) data. Biomol. Detect. Quantif. 1, 23–33 (2014).
Acknowledgements
We thank Rick Bouvé, Jetta Vlaming, Jeroen Atteveld and Diana Vasquez-Cardenas for their help in lab and sampling activities. This publication was financed under Project Number: 821.01.013 by the Earth and Life Sciences program (ALW) of the Netherlands Organization of Scientific Research (NWO) and the MaCuMBA Project 311975 of the European Commission FP7.
Author information
Authors and Affiliations
Contributions
Culturing was done by C.H. and M.G. followed by sampling which was executed by C.H., V.C.-G. and H.B. Molecular lab work was done by C.H. and V.C.-G. Gene expression data was analyzed by C.H. and the manuscript written by C.H., L.J.S. and H.B.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hörnlein, C., Confurius-Guns, V., Grego, M. et al. Circadian clock-controlled gene expression in co-cultured, mat-forming cyanobacteria. Sci Rep 10, 14095 (2020). https://doi.org/10.1038/s41598-020-69294-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-69294-3
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.