Abstract
Many species change their coloration during ontogeny or even as adults. Color change hereby often serves as sexual or status signal. The cellular and subcellular changes that drive color change and how they are orchestrated have been barely understood, but a deeper knowledge of the underlying processes is important to our understanding of how such plastic changes develop and evolve. Here we studied the color change of the Malawi golden cichlid (Melanchromis auratus). Females and subordinate males of this species are yellow and white with two prominent black stripes (yellow morph; female and non-breeding male coloration), while dominant males change their color and completely invert this pattern with the yellow and white regions becoming black, and the black stripes becoming white to iridescent blue (dark morph; male breeding coloration). A comparison of the two morphs reveals that substantial changes across multiple levels of biological organization underlie this polyphenism. These include changes in pigment cell (chromatophore) number, intracellular dispersal of pigments, and tilting of reflective platelets (iridosomes) within iridophores. At the transcriptional level, we find differences in pigmentation gene expression between these two color morphs but, surprisingly, 80% of the genes overexpressed in the dark morph relate to neuronal processes including synapse formation. Nerve fiber staining confirms that scales of the dark morph are indeed innervated by 1.3 to 2 times more axonal fibers. Our results might suggest an instructive role of nervous innervation orchestrating the complex cellular and ultrastructural changes that drive the morphological color change of this cichlid species.
Similar content being viewed by others
Introduction
Coloration is an important feature that plays crucial roles in terms of natural and sexual selection. It can serve in predator avoidance, prey capture through camouflage, conspecific communication and protection from radiation1,2. Besides these ecological and evolutionary aspects, the formation of pigment patterns provides insights into the genetic basis of adaptive evolution3 as well as the formation of complex tissues4. In vertebrates the color of the integument is shaped by the multilayered arrangement and interaction of cells with different pigmentary and structural properties5. Variations in the density, shape and properties of chromatophores and the intracellular organization of pigments and reflective molecules shapes the macroscopic appearance of the integument6,7. Several types of pigment-bearing and light-reflecting chromatophores have been identified in teleosts8. The most common cell types are melanophores (containing the brown to black pigment melanin), xanthophores/erythrophores (containing yellow to red pigments including carotenoids and pteridines), leucophores (white) and iridophores (containing guanine platelet crystals that produce structural coloration)9.
Although coloration is often perceived and studied as a static trait, it can change very dynamically on different time scales during the lifetime of an organism. Rapid changes (milliseconds to hours) are referred to as physiological color change and are triggered by neuronal and hormonal signals. These signals cause changes in the intracellular distribution of pigments or reflective molecules. In contrast, changes that involve differences in pigment quantity or cell number are referred to as morphological color change and occur over longer periods of time (hours to months)8,10,11.
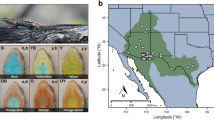
Teleost fishes lend themselves well for studying the developmental and cellular mechanisms and the molecular bases of color and color change phenotypes12,13,14,15. With their richness in pigmentation patterns, fishes in the family Cichlidae (cichlid fishes) are an attractive model for studying how genomic changes facilitate variation in molecular and developmental mechanisms and thereby the evolution of pigmentation phenotypes15,16,17,18,19,20. Cichlid fishes also offer remarkable examples for morphological color change, color polymorphism and polyphenisms as well as sexual dichromatism, traits that have received much less attention, especially from a molecular and cellular perspective. For example, in the Central American Midas cichlid Amphilophus citrinellus, 5 to 10% of the individuals lose their typical dark pigmentation and obtain a “golden” orange sometimes also white, yellow or red coloration21,22. In the haplochromine cichlid Astatotilapia burtoni, the coloration of males serves as a social signal and can change between bright blue and bright yellow23,24. This color change is regulated by the neuroendocrine melanocortin system: the yellowness of the fish body is hereby modulated by α-melanocyte-stimulating hormone (α-MSH)25. Another striking case of morphological color change occurs in the Malawi golden cichlid Melanochromis auratus (Fig. 1a,b)26,27 and was described to be the “perhaps most remarkable case” of “color change reported for any cichlid”26. M. auratus is a species endemic to the southern part of Lake Malawi. However, while the ecology28, phylogeny29, populations genetics30 and the sexual dimorphism31 of the species have been thoroughly investigated since it’s discovery in 1897 by George Albert Boulenger32, the color change and its underlying molecular mechanisms have not been studied, so far. While juveniles, females and subordinate males of M. auratus are bright yellow with two melanic horizontal stripes that is referred to as yellow morph26 (Fig. 1a,c), dominant males undergo a drastic morphological color change and become dark with two light blue horizontal stripes (dark morph; Fig. 1b,d).
Yellow and dark morph of Melanochromis auratus. (a,b) Females and subordinate males of M. auratus (a) are brightly yellow colored with two black stripes (yellow morph). Dominant males (b) transform into the dark morph that has two grey to blue stripes on a black background (dark morph). (c,d) To comparatively analyze the skin of the yellow (c) and dark (d) morph we defined five homologous regions: The dorsolateral stripe (DLS, black in yellow morph, purple/blue in dark morph), the interstripe (INT, white/gray in yellow morph, black in dark morph), the midlateral stripe (MLS, black in yellow morph, blue in dark morph), the dorsal part of the ventral integument (dVEN, white in yellow morph, black in dark morph) and the ventral part of the ventral integument (vVEN, yellow in yellow morph, black in dark morph).
With a few exceptions as for example the recent investigation of seasonal camouflage in snowshoe hares33, the molecular mechanisms and genetic control of color change remain barely understood34,35. A detailed understanding of such extreme examples, where we observe complex changes in adult characteristics, will give insights into how such changes can be orchestrated, how they manifest as well as what levels of biological organization are mechanistically involved. Moreover, they might also provide a unique opportunity understand the molecular mechanism that underly the evolution of phenotypic plasticity36 and sexual dimorphisms37.
To specifically test what levels of biological organization are involved in driving the color change of M. auratus we comprehensively analyze how ultrastructural (using transmission electron microscopy), cellular (using light microscopy and immunohistochemistry) and transcriptomic (using RNA-sequencing) changes contribute to these remarkable differences in adult morphology. Hereby, our work reveals a surprising association of morphological color change with increased neural innervation. Taken together, our results provide novel insights into the cellular, and molecular underpinnings of a remarkable case of morphological color change that differentiates both females and male subordinates from dominant males.
Results
Chromatophore number, organization and properties differ between yellow and dark morph of M. auratus
Both yellow morph and dark morph of M. auratus are characterized by two longitudinal (horizontal) stripes (Fig. 1a,b). As a first step, we histologically compared the two morphs and defined five regions across dorsal–ventral axis that differ in their coloration in the two morphs (Fig. 1c,d): dorsolateral stripe (DLS), interstripe (INT), midlateral stripe (MLS), the dorsal portion of the ventral region (dVEN), and the ventral portion of the ventral region (vVEN).
To test whether and how the morphological color change in M. auratus can be explained by changes in chromatophore number, distribution and characteristics, we compared chromatophores in yellow and dark morph using light microscopy of whole-mount scale preparations. In line with previous descriptions for cichlids7,38, three types of chromatophores could be detected in both morphs: melanophores with black to dark brown pigmentation, xanthophores with yellow to orange pigmentation, and iridophores that produce iridescent/reflective colors (Supplementary Fig. S1). To describe chromatophore distributions and characteristics we measured (a) chromatophore coverage (melanophores, xanthophores and iridophores), (b) chromatophore density (melanophores and xanthophores), and (c) chromatophore size (melanophores and xanthophores) in the epidermal layer that covers the scales (Fig. 2).
Chromatophore measurements in scales of the yellow and dark morph of M. auratus. (a–f) Photographs of individual scales from yellow morph (a–c) and dark morph (d–f). Scale bars correspond to 500 μm. (g–i) Measurements for melanophores in the five homologous regions (DLS, INT, MLS, dVEN and vVEN) of yellow and dark morph including coverage (g), cell density (h) and dispersal (i). (j–l) Same measurements for xanthophores including coverage (j), cell density (k) and dispersal (l). (m) Measurement of iridophore coverage. Differences between morphs in the same homologous regions were evaluated by two-tailed t test, n = 5 (individual points). Each point represents one individual (mean value of five scales). Error bars indicate means + SD. Significant sign: *** P < 0.001, **P < 0.01, *P < 0.05, n.s. non-significant. Full data for (g–m) see Supplementary Tables S1, S2. (n) Principal component analysis (PCA) using all measurements, demonstrating that similarly colored regions cluster together and that the morphological changes of the stripe regions (MLS, DLS) and other regions (INT, dVEN, vVEN) between yellow (white circles) and dark morph (dark circles) occur in opposite directions. PC1 mainly correlates with melanophore properties (negatively) and xanthophore properties (positively), PC2 correlates negatively with differences in iridophore coverage.
Chromatophore coverage was calculated by measuring the percentage of the scale that is covered by dark pigment (melanophores), yellow to orange pigment (xanthophores—detected by autofluorescence) and iridescent substances (iridophores—detected using polarized light). Chromatophore coverage is influenced both by number and size of the cells (as well as intracellular dispersal/aggregation of pigments) that we also quantified individually. Size of the pigmented area of melanophores and xanthophores (that is influenced both by cell size and melanosome/xanthosome dispersal within the cell) was estimated by measuring the diameter of the minimally sized circle that encloses all pigmented parts. To obtain reliable cell number estimates we treated scales with adrenaline to aggregate the melanosomes, thereby easing both quantifications of xanthophore and melanophore number (Supplementary Fig. S1). As cell delimitations could be only visualized for xanthophores and melanophores, for iridophores solely the chromatophore coverage was measured.
The most pronounced differences (see a more detailed report in the supplementary text) were found for melanophores and are consistent with the strong phenotypic differences in pigmentation (Figs. 1 and 2a–f), melanophore coverage, melanophore cell density, and average melanosome dispersal diameter differed significantly between dark and yellow morph (all P < 0.01, two-tailed t test; Fig. 2g–i, Supplementary Tables S1, S2). Differences in xanthophore coverage, xanthophore cell density and xanthophore size/dispersal were restricted to the ventral regions (vVEN and dVEN) (Fig. 2j–l, Supplementary Tables S1, S2). Although we could identify iridophores by polarized light illumination (Supplementary Fig. S1), we were not able to demarcate individual cells. Therefore, we only measured iridophore coverage but not density and diameter of iridophores. Iridophore coverage increased significantly in the two regions with iridescent white/blue coloration in the dark morph (DLS and MLS) (Fig. 2m, Supplementary Tables S1, S2). When all data were analyzed by a principal component analysis, we observe that the five homologous regions of the two morphs largely cluster by color (Fig. 2n).
Three-dimensional arrangement of chromatophores and their properties shape the phenotypic differences between morphs
Differences in coloration between yellow and dark morph might likely not only be driven by chromatophore differences in scales but also in the underlying skin. While scales can be easily investigated using light microscopy as there is only one cell layer of chromatophores, in the underlying integument such investigation is more challenging due to the three-dimensional arrangement of chromatophores39,40. To address this problem, we used transmission electron microscopy (TEM) to examine the multilayered arrangement of chromatophores as well as the ultrastructure of the pigment-bearing and structural organelles.
All chromatophores could be classified using TEM. Melanophores were identified by the presence of melanin-containing, black melanosomes (Supplementary Fig. S1)41,42. Xanthophores could be recognized by their round pigment organelles that contain carotenoid and/or pteridine and appeared grey (in contrast to the dark melanosomes) on the TEM images (Supplementary Fig. S1)40,41,42. Iridophores are characterized by stacks of white layers (Supplementary Fig. S1)40,41,42,43,44. These areas indicate the position of iridosomes (also referred to as reflecting platelets) that are composed of guanine crystals (guanine crystals are lost during the sample preparation and therefore appear as white areas on TEM images)43,44. Iridosomes reflect light and can therefore, if orientated horizontally, intensify the colors of chromatophores on top of the platelets (i.e. the yellow to orange color of xanthophores). Tilted iridosome arrangements causing light interference can result in structural coloration8,43,44. We therefore hypothesized that the main differences in dark and yellow coloration will be explained by differences in melanophore and xanthophore numbers, respectively—or at a subcellular level by differences in melanosomes and xanthosomes. The shiny appearance of yellow and white regions might be caused by horizontally (or fully randomly) oriented guanine-platelets, while the iridescent blue coloration of the MLS of the dark morph might be due to a tilted, parallelly oriented and lamellar-like organized iridosomes.
Chromatophores of M. auratus are organized in three layers in the integument (Fig. 3): (a) In the basal membrane layer (BM), the uppermost layer that is directly beneath the basal membrane (Fig. 3e–h), (b) in the stratum spongiosum layer (SP) that is the middle layer within the loose connective tissue of the upper dermis (Fig. 3i–l) and (c) in the stratum compactum layer (SC), the bottommost layer between the dense, collagenous layer of the stratum compactum and the vascular layer that delimits the dermis from the muscle (Fig. 3m–p).
Ultrastructural differences between skin regions of the yellow and dark morph. (a-d) Macroscopic appearance of the two homologous regions MLS (a,b) and vVEN (c,d) that were comparatively analyzed between yellow (a,c) and dark (b,d) morph. (e–p) The ultrastructure of the integument was analyzed using transmission electron microscopy (TEM). Chromatophores were mainly found in three layers: Directly beneath the basal membrane (BM; e–h), within the loose connective tissue of the stratum spongiosum (SP; i–l) and in the deepest layer below the stratum compactum (SC; m–p). The layers were compared between the midlateral stripe (MLS) of the yellow (e,i,m) and dark morph (f,j,n) as well as the ventral part of the ventral integument (vVEN) of the yellow (g,k,o) and dark morph (h,l,p). Scale bars correspond to 2 μm. (q–t) Polar chart of the angle of iridosomes in the stratum compactum of the MLS of yellow (q) and dark morph (r) as well as the vVEN of the yellow (s) and dark morph (t). Quantifications of all layers in dark morph MLS can be found in Supplementary Fig. S3.
Two main observations emerge: (1) Clear differences between the similarly dark-pigmented regions of both morphs [MLS of the yellow morph (Fig. 3a) and vVEN of the dark morph (Fig. 3d)] and (2) striking morphological differences between homologous, but differently colored regions [i.e. MLS, dark in the yellow morph (Fig. 3a) vs. iridescently blue in the dark morph (Fig. 3b); vVEN, brilliantly yellow in the yellow morph (Fig. 3c) vs. dark in the dark morph (Fig. 3d)].
Similarly colored skin regions are caused by different cellular compositions
In dark regions of both morphs, e.g. MLS of the yellow morph (Fig. 3e,i,m) and vVEN of the dark morph (Fig. 3h,l,p), we found melanophores in all three chromatophore layers. The melanophores in dark-pigmented regions of both morphs were mostly in a dispersed state. Melanosome density is higher in MLS of the yellow morph (Supplementary Fig. S2) than in vVEN of the dark morph (Supplementary Fig. S2)—compatible with the stronger black appearance in MLS of the yellow morph (Fig. 3a). The melanophores in the bottommost layer of the yellow morph MLS and the dark morph vVEN were underlaid by iridophores with few iridosomes (Fig. 3q,t). Iridosomes are almost arranged parallel to the surface in the yellow morph MLS (mean angle to surface 11.47°) (Fig. 3q). In contrast, platelets are more tilted in the dark morph vVEN (mean angle to surface 21.53°) (Fig. 3t). In the two upper layers of the yellow morph MLS, melanophores were more scattered, thereby often overlaying small-sized xanthophores (Fig. 3e,i). In the SP of dark morph vVEN, small numbers of iridophores were found beneath the melanophores (Fig. 3l).
Color changes of homologous regions are driven by structural and cellular modifications
For the vVEN of the yellow morph, which is brightly yellow colored, we found stacked and almost parallelly oriented reflecting platelets (mean angle to surface 9.12°) in the bottommost SC layer (Fig. 3o,s). The two upper layers, BM and SP, had numerous xanthophores (Fig. 3g,k). Melanophores were almost absent from all layers (Fig. 3g,k,o). The ventral area of the yellow morph therefore had more xanthophores, less melanophores and more and more parallel oriented iridosomes within iridophores in the SC. Usually iridosomes are quite thin (e.g. in the neon tetra, Paracheirodon innesi, that has been intensively studied in this respect, thickness ranges between 5 and 60 nm45,46). In M. auratus iridosomes were substantially thicker in all regions and both morphs (mean thickness between 94 and 144 nm; distance between iridosomes between 76 and 147 nm; Supplementary Fig. S4). This is in accordance with the hypothesis that such structural characteristics—that merely lead to a reflection of light in a mirror-like fashion—would result in the bright yellow coloration of the vVEN region of the yellow morph.
The most noticeable characteristic in the integument of the distinct bluish iridescent dark morph MLS (Fig. 1b) is that it is composed of several sheets of highly organized iridophores (Fig. 3f,j,n). In the upper layers, guanine platelets are arranged in parallel but strongly angled (mean angle to surface in BM: 26.10°; mean angle to surface in SP: 38.63°; Fig. 3f,I; Supplementary Fig. S3). In the bottom layer (SC) the orientation appears more random and essentially ranges from 0 to 90° (mean angle to surface: 50.13°; Fig. 3n,r). Some aggregated melanophores (Fig. 3n) were also found in the in the iridophore layers, especially in the SC layer. The highly organized ultrastructure of the iridophores in the dark morph MLS confirms the hypothesis that the iridescent blue coloration is caused by iridosome orientation in the integument.
Within the other dorso-ventral regions we found comparable patterns that are comprehensively summarized in Supplementary Fig. S5. The differences between the INT region of the yellow and dark morph resembled those of the vVEN region with the main difference being the much higher number of xanthophores in the vVEN of the yellow morph compared to the INT region. This is compatible with the whitish appearance of the INT region (Supplementary Fig. S5). The DLS region resembles the MLS region, with the main difference being that the DLS of the dark morph has less organized iridosomes likely resulting in the more greyish to whitish appearance of the DLS compared to the bluish iridescent MLS (Supplementary Fig. S5).
Comparative transcriptomics of M. auratus color morphs
To identify genes that are associated with the coloration differences between yellow and dark morph, we screened for differentially expressed genes using RNA-sequencing (RNA-seq) of the integument of yellow (n = 4) and dark morph (n = 4) individuals. Our comparative transcriptomic approach identified 42 differentially expressed genes: 13 genes were expressed at a significantly higher level in the integument of the yellow morph and 29 genes had significantly higher expression in integument of the dark morph (P < 0.01, lfc > log2(2); Fig. 4; Tables 1, 2). Surprisingly, among those were few genes that had been previously implicated in teleost pigmentation (6 of 42), and all of them were among the genes that were higher expressed in the integument of the yellow morph (Fig. 4; Table 2). Those six pigmentation genes were hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 1 (hsd3b1)47, pteridine biosynthesis enzyme GTP cyclohydrolase 2 (gch2)48, carotenoid droplets disperser perilipin 6 (plin6)49, melanophore-linage cell marker microphthalmia-associated transcription factor a (mitfa)50,51,52, tetratricopeptide repeat protein 39B (ttc39b)53,54,55 and oncogenic transcription factor forkhead box Q1 (foxq1a)56.
Differential gene expression analysis. Differential expression between yellow morph (n = 4) versus dark morph skin (n = 4). 42 genes show significant differential expression [P < 0.01, lfc > log2(2)]. Light points represent genes overexpressed in yellow morph (13 genes), dark points those genes overexpressed in the dark morph (29 genes). Grey points indicate genes that do not show significant differential expression. Red circles indicate genes with a reported link to pigmentation, Blue circles indicate genes involved in nervous system function (see Tables 1 and 2 for references).
In the integument of the dark morph we found no pigmentation genes with significantly higher expression, instead a high number (80% of the genes) of neuronal genes—23 of 29 based on a literature search (Fig. 4; Table 1). Two GO-terms were significantly enriched (padj < 0.05), syntaxin-1 binding (GO:0017075) and SNARE binding (GO:0000149), both are important protein interactions involved in the regulation of synaptic vesicles at the presynaptic zone of neuronal synapses. Among the most significantly differentially expressed genes are the synaptosomal-associated protein 25 (snap25a and snap25b) that is required for the fusion of synaptic vesicles with the presynaptic membrane57, the neuronal membrane glycoprotein M6a (gpm6ab) that is involved in neurite outgrowth and synaptogenesis58,59 and the calcium sensor protein visinin-like protein 1 (vsnl1a) that is important for neuronal signaling60. The up-regulation of neural and synapse-related genes in the integument suggests differences in neural innervation of the integument that—besides the difference in pigmentation—distinguish the yellow and dark morph of M. auratus. We will elaborate in the discussion how this unexpected link between coloration phenotype and neural innervation might be explained.
Axonal innervation of the scale epithelium is increased in the dark morph of M. auratus
To find further support for the hypothesis that neural innervation is indeed increased in the dark morph, we performed immunofluorescence staining of axonal fibers on scales of both morphs61. We dissected scales from three homologous dorsal–ventral positions and counted the number of nerve fibers (Fig. 5, Supplementary Figs. S6, S7, see details in the methods section). Axons were visualized using Anti-Acetylated Tubulin antibody (AcTub). In accordance with the transcriptome data, we indeed found more AcTub+ axons on scales of the dark morph compared to scales of the yellow morph, independent of dorso-ventral level and position on the scale (all P < 0.01, two-tailed t test, 1.3 to 2-fold change; Fig. 5e, Supplementary Figs. S6, S7, Supplementary Tables S3, S4). Yet, we were concerned that the lower number of axon counts might be also linked to the increased number of melanophores that could mask the fluorescence and thereby result in an underestimation of axons in the dark morph (Supplementary Fig. S8). Therefore, we used a previously described (spontaneous) amelanistic mutant line that lacks all melanic pigmentation due to a loss of the second exon of oculocutaneous albinism II (oca2) gene18. Still, despite this loss-of-function mutation, melanophores seem to be present in both, the yellow and dark morph of the amelanistic line as previously described18 (as the morph names are rather confusing descriptions in the amelanistic strain we use quotations marks, i.e. ‘yellow morph’ and ‘dark morph’). Comparably to the wildtype-like strain of M. auratus, ‘dark morph’ individuals of the amelanistic M. auratus had significantly higher axon density than ‘yellow morph’ individuals (all P < 0.01, two-tailed t test, 1.3 to 1.8 fold change) (Fig. 5a–d, Supplementary Figs. S6, S7, Tables S3, S4). Lastly, as mentioned in the introduction and discussed below it is dominant males that undergo the color transition and transition into the dark morph, while this has not been reported (or only very rarely) for female individuals. It is therefore possible that the differences in axon density are linked to sex and not to morph. To test this possibility, we performed the same quantifications on scales of a closely related species without color change, Pseudotropheus cyaneorhabdos (previously Melanochromis cyaneorhabdos), and compared males and females. No significant differences in axon density could be detected (all P > 0.05, two-tailed t test) (Supplementary Figure S6, S7, Supplementary Tables S3, S4) suggesting that the difference in neural innervation is linked to color morph, but not sex.
Increased axon innervation in the epidermis of the dark morph. (a–d) Immunohistochemistry with Anti-Acetylated Tubulin antibody labeling axonal fibers shows increased innervation of scales of the dark morph/‘dark morph’ (b,d) compared to the yellow morph/‘yellow morph’ (a,c). Scale bars are 500 μm in (a,b) and 50 μm in (c,d). The complete analysis can be found in Supplementary Fig. S8 (e) Axon density on scales is quantified based on Acetylated Tubulin staining. Difference between morphs was evaluated by two-tailed t test, n = 5 (individual points). Each individual point represents the mean value of 15 measurement for one fish. Green bars indicate the means. Significant sign: ***P < 0.001.
Discussion
In this study, we investigated the cellular and transcriptional differences between the yellow and dark morph of M. auratus, a cichlid fish species that is endemic to Lake Malawi62,63. Specifically, we asked how the morphological differences can be explained (a) by changes in chromatophore number (b) chromatophore characteristics, (c) multi-dimensional arrangement of chromatophores in the integument and (d) if we find transcriptional signatures that provide us with more mechanistic insights into genetic and cellular correlates as well as the genetic pathways that might be involved in triggering the morphological color change from yellow to dark morph.
Our analysis reveals substantial variation between yellow and dark morph (Fig. 1) ranging from transcriptomic differences, to differences in chromatophore morphology, number and three-dimensional organization as well as—unexpectedly—changes in neural innervation of the integument. These dramatic changes must be occurring during the morphological color change of dominant males that transition over the course of two weeks from the yellow to the dark morph (visual inspection on > 10 individuals). Below we therefore (i) comprehensively discuss the differences between the yellow and the dark morph, (ii) speculate what changes cause the transition and/or what might be merely indirect consequences of the morphological color change.
Our study demonstrates that yellow and dark morph differ greatly on different levels of biological organization ranging from gene expression differences to cellular and ultracellular characteristics of chromatophores as well as cellular composition in the three layers of the integument and on scales. The first observation is that the color of phenotypically similar regions is formed through similar mechanisms (Figs. 2 and 3): The dark stripes of the yellow morph (MLS and DLS) and the dark ventral area and interstripe (vVEN, dVEN and INT) of the dark morph are characterized by a high density of dispersed melanophores (Fig. 2g–i). In contrast, the yellow belly region of the yellow morph (vVEN) has a very low density of melanophores, but high density of large, dispersed xanthophores. A further peculiarity are thick layers of horizontally oriented iridiosomes within the stratum compactum, the deepest layer of the integument. These likely simply serve as a mirror and directly reflect light and thereby intensify the yellow coloration of the overlaying xanthophores. Such mirror-like function of iridosomes have been previously described, i.e. in the japanese Koi fish (Cyprinus carpio), where the reflectance is however generated through a fully random organization of iridosomes64. Lastly, there are several regions with iridescent white (i.e. INT in yellow morph) to blue (MLS in dark morph) coloration. The major differences to the other regions is (beside the lower density of xanthophores and melanophores) that iridosomes are not oriented horizontally but in an acute angle of about 25° to 50° (Fig. 3r). Such regularly oriented, tilted iridosomes have been previously described in the neon tetra Paracheirodon innesi, that is characterized by a single iridescent blue horizontal stripe. The periodically arranged iridosomes constitute a colour-producing microstructure that might cause the blue coloration through multilayer optical interference45,46.
At the transcriptomic level we find several pigmentation genes that are differentially expressed. Two of the genes that show higher expression in the yellow morph are gch2 and plin6 (Fig. 4; Table 2)., which have been associated with carotenoid and pteridine synthesis—the yellow to orange pigments of xanthophores The Plin6 protein targets the surface of carotenoid droplets and mediates carotenoid droplet dispersion in xanthophores49. The gene gch2 encoding a pivotal protein in the pteridine biosynthetic pathway and is required for xanthophore pigmentation in zebrafish48. Also hsd3b1 and ttc39b have been linked to lipid and carotenoid synthesis as well as carotenoid-based coloration differences in cichlids47,53,54,55. Compared to the dark morph, xanthophores are numerous in the yellow morph and densely filled with xanthosomes as supported by light microscopy (Fig. 2) and TEM images (Fig. 3), therefore these transcriptional differences are compatible with the morphological differences. We also find two pigmentation genes that have been associated with melanophore/melanocyte differentiation: foxq1 and mitfa50,51,52,56. As melanophore numbers are lower in the yellow morph, especially in vVEN (Figs. 2 and 3), it is possible that the high expression of these genes indicates that large reservoirs of undifferentiated melanoblasts exist, that only differentiate when the color change is initiated. What remains unclear is, how the melanoblasts are mechanistically kept in an undifferentiated state. Interestingly, several of the other differentially expressed genes have been previously linked to stem cell maintenance including retinol dehydrogenases65, ensconsin/map7a66, scavenger receptor class B member 167 and seipin/bscl268.
The most striking observation regarding those genes being overexpressed in the dark morph is that a substantial part of these genes (80%) are related to neuronal processes, particularly to synaptogenesis (Fig. 4; Table 1). This observation is further substantiated by immunohistochemistry for the axonal marker acetylated tubulin that demonstrate that there is a substantially higher number of neuronal fibers in scales of the dark morph than in the yellow morph (1.3 to 2-fold change; Fig. 5) across the whole trunk.
To understand the—at first surprising—link between coloration and nervous innervation, there are two facts to consider: Firstly, adult pigment cells derive from postembryonic progenitors that are located in proximity of the dorsal root ganglia. From there these progenitors migrate along peripheral nerves through the myosepta to the skin. Therefore, there is a strong developmental link between differentiation of adult pigment cells and nerve fibers69,70,71. Secondly, the connection between pigment cell remains even after differentiation. Especially melanophores, but also xanthophores and iridophores are controlled by the nervous system that can trigger the aggregation of melanosomes or xanthosomes, as well as reorientation of iridosomes11,72,73. These fast intracellular changes trigger rapid physiological color change72. But neural innervation has been also shown to induce more long-lasting changes (i.e. morphological color change) by triggering remodeling of the cytoskeleton74, apoptosis of chromatophores74,75, as well as proliferation of chromatophores and changes in pigment synthesis76. Very recently, increased neural signaling has been even linked to the depletion of melanocyte stem cell pools leading to hair greying in humans77, which supports an evolutionary conserved link between the nervous system and pigment cells.
The morphological color change of M. auratus is particularly remarkable because several, oppositely directed changes occur in different parts of the skin (Fig. 2). While in the stripes melanophores are reduced and xanthophores are increased in number, we observe the opposite trend in the interstripe and ventral skin. Additionally, we find increased numbers of iridophores, mainly in the dorsal areas, as well as reorganization of iridosomes, in particular in the MLS. The molecular processes that orchestrate such a complex rearrangement are therefore likely complex itself. Based on our results, we assume that two different processes are occurring: First, we observe changes at the level of pigment cell progenitors, either within the skin or at the postembryonic progenitor niche in vicinity of the dorsal root ganglia that lead to the generation of new chromatophores. As cell–cell interactions are an important factor for color pattern formation in fish, it is possible that these changes do not occur everywhere at the same time, but are propagated across the skin78. Recently it has been shown that long-distance signaling relayed by macrophages trigger tissue color pattern remodeling during postembryonic development79, so similar processes could certainly play a role in this instance of morphological color change. Some of the genes overexpressed in the yellow morph indicate that these fish might already have a reservoir of melanoblasts that could—triggered by an endocrine, paracrine or neuronal signal—differentiate and thereby realize the morphological color change from the yellow to the dark morph. Second, the overexpression of neural genes and increased number of neural innervations could be an indication for two processes: The nerve fibers could lead to a stronger neural control of chromatophores (i.e. physiological color change). This interpretation is supported by the fact that the melanic patterns do not fade or enhance very much in the yellow morph, while the dark morph shows substantial physiological color change with the dark melanic parts changing from light grey to black and vice versa. Second, it is also possible that the neural fibers have an instructive role and initiate the color change by promoting migration, differentiation of chromatophore progenitors, cell death as well as changes in pigment production or translocation, as shown in other organisms74,75,76,77.
One central question is how the color change of the different regions (i.e. DLS, INT, MLS, dVEN and vVEN) is orchestrated. As discussed above, changes happen in almost opposite directions, with DLS and MLS becoming lighter (i.e. grey or blue) and INT, dVEN and vVEN becoming darker (i.e. dark grey or black). However, we do not see variation in neural innervation across the dorsoventral axis that could explain these oppositely directed changes and therefore the phenotype remains difficult to explain solely by the innervation. It is possible that this is achieved by a combination of the neuronal trigger and the number of chromatophore precursors that are present in the different regions. Therefore, although our morphological and transcriptomic analyses provide important and novel insights into the molecular and cellular correlates of the color change in M. auratus, our analyses cannot indicate whether the nervous system and/or particular genes act directly or indirectly on the distribution, shape and properties of chromatophores and if they are causally involved. An even more detailed analysis will be needed that dissects the molecular and cellular changes in the different skin regions—also at different time points of the transition process—to completely understand this remarkable color change phenotype.
In conclusion, our work provides an integrative and comprehensive description of the transcriptomic, ultrastructural, cellular and neural changes that define the morphological color change of M. auratus. It provides therefore the first insights into the unexpectedly massive changes occurring at different levels of biological organization ranging from gene expression changes, to reorientation of reflecting platelets (iridosomes) in iridophores, to changes in chromatophore density and morphology, and changes in neuronal innervation. The color change M. auratus was described as “perhaps most remarkable case” of “color change reported for any cichlid” almost 50 years ago26—the complexity of the changes that shape and accompany this transition and its evolution are in no way less remarkable.
Methods
Animals
Animals used in this study were raised and kept in the animal research facility of the University of Konstanz. All animal samples used in this study were collected in accordance with relevant guidelines and regulations and sampling was approved by the authorities (Regierungspräsidium Freiburg, Anzeige T-16/13).
Light microscope image acquisition for fish scales
To quantify the change in chromatophore quantities and properties during color transition of M. auratus, we measured the coverage, density and dispersed diameter of chromatophores in scales from five homologous pigmented regions in dorsal–ventral axis. In M. auratus, chromatophores could be found in multiple layers through the integumentary system and overlaid/underlaid each other. The chromatophores only cover the posterior part of the scales and are arranged in a single layer, which makes the robust chromatophore quantification possible. Five individuals of each color morph were examined for scale chromatophore quantification. Fifteen scales from three dorsal–ventral rows (5 scales each row) were removed from each fish. Scales from most dorsal row covering DLS and INT (Fig. 2a,d); scales from middle row covering MLS and dVEN (Fig. 2b,e); the scales from most ventral row (vVEN) (Fig. 2c,f). Once being removed from the fish, scales were rinsed by Phosphate Buffered Saline (PBS, pH 7.4) and adhered to HistoBond + adhesive microscope slides (Paul Marienfeld). Photographs of M. auratus scales were captured on Leica DM6B upright microscope with a Leica DMC 2900 camera. We captured images in three different modes: brightfield for melanophore coverage and dispersed diameter, polarized light for iridophore coverage7, fluorescent with GFP filter for xanthophore coverage and dispersed diameter80,81. To count the number of melanophores and xanthophores, we treated the scales immediately after taking the above photos with 10 mg/ml adrenaline (SIGMA-ALDRICH) for 5 min at room temperature on microscope slides to aggregate the melanosomes which permit robust cell number quantification (Supplementary Fig. S1). After adrenaline treatment, microscope-slide-adhered scales were rinsed by PBS and photographed using brightfield mode. Leica Application Suite X software was used to capture the photos using the same setting.
Image and data analysis of scale chromatophores
All light microscope photos were analyzed with Fiji82. In order to obtain reliable chromatophore coverage quantification, we manually adjusted the color threshold of the brightfield photos for melanophores, polarized light photos for iridophore and fluorescent photos for xanthophore from non-treated scales (Supplementary Fig. S1). Following this setting, we executed the “Analyze Particles” function to obtain all three chromatophore coverage. For chromatophore size measurement, we randomly selected 10 melanophores from brightfield photos and 10 randomly xanthophores from fluorescent photos for each pigmented region (DLS, INT, MLS, dVEN, vVEN) and measured the dispersed diameter of pigment covered parts of the chromatophores. The number of melanophores and xanthophores was counted from the same scales after the epinephrine treatment. Although we could identify iridophores and quantified the iridophore coverage in all pigmented regions before and after epinephrine treatment (Supplementary Fig. S1), the boundaries of each iridophore could not be identified. Therefore, we were unable to quantify the iridophore number and measure the iridophore size.
Transmission electron microscope image acquisition for fish integument
Melanic and non-melanic skin with scales attached were dissected in buffer pre-fixative (2% formaldehyde, freshly depolymerized from paraformaldehyde (Merch), 2.5% glutardialdehyde (Agar Scientific) in 0.1 M HEPES (AppliChem) with 5 mM MaCl2, 5 mM CaCl2, 0.125 mM MgSO4 and 0.3 M sucrose for osmolarity, pH 7.5) and fixed in fresh-made, pre-cooled fixative (same as buffer pre-fixative) at 0 °C for 2 h. After osmification in 2% OsO4 (SERVA) (buffered with HEPES, pH 7.5) at 0 °C for 2 h, samples were pre-dehydrated in 30% and 50% Ethanol and en-bloc stained the samples by 2% uranyl acetate (Merck) in 70% Ethanol. We dehydrated the samples in graded series of acetone solutions and embedded in SPURR (TedPella) (ERL 4,206, DER 736, NSA, DMAE). 50 nm ultra-thin sections were cut by a Leica Ultracut microtome with an ultra 45° knife (Diatome, Switzerland). Sections were contrasted with uranyl acetate and lead citrate before examined with a Zeiss TEM 912. Images were processed with Fiji and stitched with TrackEM283. Iridosome orientation was estimated by measuring the angle between iridophore and skin surface. Iridosome orientation, thickness and distance between iridosomes, as well as melanosome density were analyzed with Fiji82. Plots in Fig. 3q–t and Supplementary Figs S2. S3 and S4 were generated using ggplot84 in the R-environment85 and assembled and annotated in Adobe Illustrator v.24.0.3 (Adobe Systems Software). Illustration of the chromatophore arrangement in the integument (Supplementary Fig S5) was generated using by Adobe Illustrator v.24.0.3 (Adobe Systems Software).
RNA extraction and purification
RNA extraction and purification were performed as previously described86. Briefly, dissected skin samples of eight individuals (four for each morph) were kept in RNAlater (Invitrogen) at − 20 °C. RNAlater was removed prior to homogenization. Skin samples and the appropriate amount of TRIzol (Invitrogen) (1 ml TRIzol per 100 mg sample) were homogenized in 2 ml Lysing Matrix A tube (MP Biomedicals) using FastPrep-24 Classic Instrument (MP Biomedicals). Subsequent purification and DNase treatment were performed with RNeasy Mini Kit (Qiagen) and RNase-Free DNase Set (Qiagen). Following extraction and purification, RNA was quantified using the Qubit RNA HS Assay Kit (Invitrogen) with a Qubit Fluorometer (Life Technologies). The RNA integrity number (RIN) was checked using an RNA 6000 Pico Kit (Agilent) on a 2100 Bioanalyzer System (Agilent).
RNA library preparation and sequencing
RNA-seq libraries were prepared using the TruSeq Stranded mRNA Library Prep Kit (Illumina) according to the manufacturer’s protocol. Briefly, 1 μg RNA was put into mRNA selection by poly-T oligo attached magnetic beads followed by fragmentation (94 °C for 6 min). The cleaved mRNA fragments were reverse transcribed into first-strand cDNA using GoScript Reverse Transcriptase (Promega) and random hexamer primers (Illumina). We used Illumina-supplied consumables to synthesize second-strand cDNA following by adenylating 3′ ends. Barcoded adapters from TruSeq RNA CD Index Plate (Illumina) were ligated to the ends of the double-strand cDNA. The final libraries were amplified using 15 PCR cycles and quantified and quality-assessed using an Agilent DNA 12000 Kit on a 2100 Bioanalyzer (Agilent). Indexed DNA libraries were normalized. Libraries were sequenced on a HiSeq X Ten platform (BGI Genomics, Beijing).
Mapping and data processing
For adapter trimming trimmomatic 0.38 was used87. The processed raw reads (mean: 41.7 million reads per sample) were aligned to the Maylandia zebra genome (M_zebra_UMD2a, INSDC Assembly GCA_000238955.5, Apr 2018) using the STAR RNA-seq aligner (version 2.6.1d)88. Maylandia zebra and M. auratus are very closely related, why this genome was used as a reference. Expression counts were calculated with RSEM89 using the ENSEMBL annotation. Quality control was done using MultiQC 90. 89% of the reads were uniquely mapped. The short-read data has been archived in NCBI SRA database under the Bioproject accession number PRJNA635556 (SRR11862158–SRR11862165).
Data analysis
The data was analysed in R using the DESEQ2 1.22.1 pipeline91 in R92. The following packages were additionally used: BiocParallel 1.16.293, tximport 1.10.194, stringR 1.4.095, edgeR 3.24.196, vsn 3.50.097, ggplot2 3.1.198, RColorBrewer 1.1–299, vidger 1.2.0100, pheatmap 1.0.12101, cowplot 0.9.4102 and dplyr 0.8.0.1103. GO term analyses was performed by extracting GO terms for the M. zebra ENSEMBL genome annotation using biomaRt 2.38.0104. Next, we used goseq 1.34.1105 to correct for transcript length correction and selection-unbiased testing for category enrichment amongst differentially expressed genes. P-values were adjusted with Benjamini & Hochberg correction.
Axon staining and density quantification on scales
As the RNA-seq result showing that the expression of neural genes is upregulated in the dark morph, we performed immunofluorescent staining for axon to further substantiate the expression data. Fish scales (15 scales/fish) of a total of 29 fish (5 individuals per morph except for the individuals of the ‘yellow morph’ of the amelanistic individuals for which we only had 4 individuals) were removed from the homologous regions for chromatophores measurement (5 scales from each dorsal–ventral row, 3 rows from each fish). Immunostaining was performed based on previously described with modification106. Scales were fixed in Dent’s fixative (20% dimethyl sulfoxide (DMSO), 80% Methanol) at 4 °C overnight then kept in 100% Methanol at − 20 °C. We rehydrate the samples by graded series of Methanol/PBS. After several washes in PBS and PBS 0.1% Triton X-100, we permeabilizated the samples with permeabilization buffer (1% normal goat serum (Sigma), 0.4% Triton X-100 in PBS). Scales were then blocked in 10% normal goat serum with 0.4% Triton 100-X in PBS for minimally 2 h and incubated at 4 °C overnight with mouse monoclonal anti-acetylated-tubulin antibody (1:250; clone 6-11B-1; Sigma-Aldrich # T6793) in PBS supplemented with 10% normal goat serum and 0.1% Triton X-100. Although this primary antibody was raised against acetylated tubulin of sea urchin, it has been used for detection of acetylated tubulin from several tissues of many organisms, including fish scales61,107. The next day scales were washed by PBS supplemented with 0.1% Tween-20 (PBST) several times and the staining was achieved by incubation with secondary antibody (Goat-anti-Mouse IgG (H + L) SuperClonal Secondary Antibody, Alexa Fluor 488 conjugate, Invitrogen # A28175, 1:400) at 4 °C overnight. The next day scales were washed by PBST and counterstain with 4′,6-Diamidino-2-phenylindole dihydrochloride (DAPI) (1/5000; Sigma) in PBS. We mounted the scales on HistoBond + adhesive microscope slides in Mowiol mounting medium (Sigma). Photographs were taken with a Leica DM6B microscope with a Leica DFC3000G black and white camera.
Axon density was quantified whit Fiji82. We quantified the axon density from both anterior and posterior parts of the scales (Fig. 5, Supplementary Figs. S6, S7). For axon density in anterior of scales, the two distinct endpoints of measurement line segment were located in the most dorsal and the most ventral edge of epidermis which indicated by DAPI staining. For axon density in the posterior of scales, the two distinct endpoints of the measurement line segment were located in the most dorsal and the most ventral centiis. Following this setting we executed the “Plot Profile” function and “Find Peaks” function of BAR plugin to obtain the axon density. All statistical tests were performed in R85 and plotted using ggplot84 in the R-environment85.
References
Cheney, K. L., Grutter, A. S. & Marshall, N. J. Facultative mimicry: cues for colour change and colour accuracy in a coral reef fish. Proc. Biol. Sci. 275, 117–122. https://doi.org/10.1098/rspb.2007.0966 (2008).
Mills, M. G. & Patterson, L. B. Not just black and white: pigment pattern development and evolution in vertebrates. Semin. Cell Dev. Biol. 20, 72–81. https://doi.org/10.1016/j.semcdb.2008.11.012 (2009).
Orteu, A. & Jiggins, C. D. The genomics of coloration provides insights into adaptive evolution. Nat. Rev. Genet. https://doi.org/10.1038/s41576-020-0234-z (2020).
Parichy, D. M. Animal pigment pattern: an integrative model system for studying the development, evolution, and regeneration of form. Semin. Cell Dev. Biol. 20, 63–64. https://doi.org/10.1016/j.semcdb.2008.12.010 (2009).
Singh, A. P. & Nusslein-Volhard, C. Zebrafish stripes as a model for vertebrate colour pattern formation. Curr. Biol. 25, R81–R92. https://doi.org/10.1016/j.cub.2014.11.013 (2015).
Patterson, L. B. & Parichy, D. M. Zebrafish pigment pattern formation: insights into the development and evolution of adult form. Annu. Rev. Genet. 53, 505–530. https://doi.org/10.1146/annurev-genet-112618-043741 (2019).
Liang, Y., Gerwin, J., Meyer, A. & Kratochwil, C. F. Developmental and cellular basis of vertical bar color patterns in the East African Cichlid Fish Haplochromis latifasciatus. Front. Cell Dev. Biol. https://doi.org/10.3389/fcell.2020.00062 (2020).
Fujii, R. The regulation of motile activity in fish chromatophores. Pigment Cell Res. 13, 300–319. https://doi.org/10.1034/j.1600-0749.2000.130502.x (2000).
Fujii, R. Cytophysiology of fish chromatophores. Int. Rev. Cytol. 143, 191–255 (1993).
Skold, H. N., Aspengren, S., Cheney, K. L. & Wallin, M. Fish chromatophores-from molecular motors to animal behavior. Int. Rev. Cell Mol. Biol. 321, 171–219. https://doi.org/10.1016/bs.ircmb.2015.09.005 (2016).
Fujii, R. In International Review of Cytology International Review of Cytology Vol. 143 (eds Jeon, K. W. et al.) 191–255 (Academic Press, Cambridge, 1993).
Irion, U. & Nüsslein-Volhard, C. The identification of genes involved in the evolution of color patterns in fish. Curr. Opin. Genet. Dev. 57, 31–38 (2019).
Parichy, D. M. Evolution of danio pigment pattern development. Heredity (Edinb) 97, 200–210. https://doi.org/10.1038/sj.hdy.6800867 (2006).
Nusslein-Volhard, C. & Singh, A. P. How fish color their skin: A paradigm for development and evolution of adult patterns: multipotency, plasticity, and cell competition regulate proliferation and spreading of pigment cells in Zebrafish coloration. BioEssays https://doi.org/10.1002/bies.201600231 (2017).
Kratochwil, C. F. et al. Agouti-related peptide 2 facilitates convergent evolution of stripe patterns across cichlid fish radiations. Science 362, 457–460. https://doi.org/10.1126/science.aao6809 (2018).
Santos, M. E. et al. The evolution of cichlid fish egg-spots is linked with a cis-regulatory change. Nat. Commun. 5, 5149. https://doi.org/10.1038/ncomms6149 (2014).
Santos, M. E. et al. Comparative transcriptomics of anal fin pigmentation patterns in cichlid fishes. BMC Genom. 17, 712. https://doi.org/10.1186/s12864-016-3046-y (2016).
Kratochwil, C. F., Urban, S. & Meyer, A. Genome of the Malawi golden cichlid fish (Melanochromis auratus) reveals exon loss of oca2 in an amelanistic morph. Pigment Cell Melanoma Res. 32, 719–723. https://doi.org/10.1111/pcmr.12799 (2019).
Hendrick, L. A. et al. Bar, stripe and spot development in sand-dwelling cichlids from Lake Malawi. Evodevo 10, 18. https://doi.org/10.1186/s13227-019-0132-7 (2019).
Maan, M. E. & Sefc, K. M. Colour variation in cichlid fish: developmental mechanisms, selective pressures and evolutionary consequences. Semin. Cell Dev. Biol. 24, 516–528. https://doi.org/10.1016/j.semcdb.2013.05.003 (2013).
Barlow, G. W. In Investigations of the Ichthyofauna of Nicaraguan Lakes (ed. Thorson, T. B.) 333–358 (Lincoln, University of Nebraska-Lincoln, 1976).
Henning, F., Jones, J. C., Franchini, P. & Meyer, A. Transcriptomics of morphological color change in polychromatic Midas cichlids. Bmc Genomics 14, 171 (2013).
Hofmann, H. A. & Fernald, R. D. What cichlids tell us about the social regulation of brain and behavior. J. Aquacult. Aquat. Sci 9, 17–31 (2001).
Korzan, W. J. & Fernald, R. D. Territorial male color predicts agonistic behavior of conspecifics in a color polymorphic species. Behav. Ecol. 18, 318–323. https://doi.org/10.1093/beheco/arl093 (2006).
Dijkstra, P. D. et al. The melanocortin system regulates body pigmentation and social behaviour in a colour polymorphic cichlid fish. Proc. Biol. Sci. https://doi.org/10.1098/rspb.2016.2838 (2017).
Fryer, G. & Iles, T. D. Cichlid fishes of the great lakes of Africa. (1972).
Fryer, G. The trophic interrelationships and ecology of some littoral communities of Lake Nyasa with especial reference to the fishes, and a discussion of the evolution of a group of rock-frequenting cichlidae. Proc. Zool. Soc. Lond. 132, 153–281. https://doi.org/10.1111/j.1469-7998.1959.tb05521.x (1959).
Ribbink, A. J., Marsh, B. A., Marsh, A. C., Ribbink, A. C. & Sharp, B. J. A preliminary survey of the cichlid fishes of rocky habitats in Lake Malawi. S. Afr. J. Zool. 18, 149–310. https://doi.org/10.1080/02541858.1983.11447831 (1983).
Bowers, N. J. A revision of the genus Melanochromis (Teleostei: Cichlidae) from Lake Malawi, Africa, using morphological and molecular techniques. (1994).
Markert, J. A., Arnegard, M. E., Danley, P. D. & Kocher, T. D. Biogeography and population genetics of the Lake Malawi cichlid Melanochromis auratus: habitat transience, philopatry and speciation. Mol. Ecol. 8, 1013–1026. https://doi.org/10.1046/j.1365-294x.1999.00658.x (1999).
Schneider, R. F., Rometsch, S. J., Torres-Dowdall, J. & Meyer, A. Habitat light sets the boundaries for the rapid evolution of cichlid fish vision, while sexual selection can tune it within those limits. Mol. Ecol. 29, 1476–1493. https://doi.org/10.1111/mec.15416 (2020).
Boulenger, G. A. Description of a new fish from Lake Nyassa. Ann. Mag. Nat. Hist. 6, 155 (1897).
Jones, M. R. et al. Adaptive introgression underlies polymorphic seasonal camouflage in snowshoe hares. Science 360, 1355–1358 (2018).
Duarte, R. C., Flores, A. A. V. & Stevens, M. Camouflage through colour change: mechanisms, adaptive value and ecological significance. Philos. Trans. R. Soc. B 372, 20160342. https://doi.org/10.1098/rstb.2016.0342 (2017).
Stevens, M. Color change, phenotypic plasticity, and camouflage. Front. Ecol. Evol. https://doi.org/10.3389/fevo.2016.00051 (2016).
Schneider, R. F. & Meyer, A. How plasticity, genetic assimilation and cryptic genetic variation may contribute to adaptive radiations. Mol. Ecol. 26, 330–350. https://doi.org/10.1111/mec.13880 (2017).
Kopp, A. Dmrt genes in the development and evolution of sexual dimorphism. Trends Genet. 28, 175–184. https://doi.org/10.1016/j.tig.2012.02.002 (2012).
Baerends, G. P. & Baerends-van Roon, J. An introduction to the study of the ethology of the cichlid fishes. Behav. Suppl. 1, 1–243 (1950).
Chavin, W. Fundamental aspects of morphological melanin color changes in vertebrate skin. Am. Zool. 9, 505–520. https://doi.org/10.1093/icb/9.2.505 (1969).
Hirata, M., Nakamura, K., Kanemaru, T., Shibata, Y. & Kondo, S. Pigment cell organization in the hypodermis of zebrafish. Dev Dyn 227, 497–503. https://doi.org/10.1002/dvdy.10334 (2003).
Hawkes, J. W. The structure of fish skin. II. The chromatophore unit. Cell Tissue Res 149, 159–172. https://doi.org/10.1007/bf00222271 (1974).
Hirata, M., Nakamura, K. & Kondo, S. Pigment cell distributions in different tissues of the zebrafish, with special reference to the striped pigment pattern. Dev. Dyn. 234, 293–300. https://doi.org/10.1002/dvdy.20513 (2005).
Takeuchi, I. K. Electron microscopy of two types of reflecting chromatophores (iridophores and leucophores) in the guppy Lebistes reticulatus Peters. Cell Tissue Res. 173, 17–27. https://doi.org/10.1007/bf00219263 (1976).
Gundersen, R. E. & Rivera, E. R. An ultrastructural study of the development of the dermal iridophores and structural pigmentation in Poecilia reticulata (peters). J. Morphol. 172, 349–359. https://doi.org/10.1002/jmor.1051720309 (1982).
Hiroshi, N., Noriko, O. & Ryozo, F. Light-reflecting properties of the iridophores of the neon tetra, Paracheirodon innesi. Comp. Biochem. Physiol. A 95, 337–341. https://doi.org/10.1016/0300-9629(90)90229-L (1990).
Yoshioka, S. et al. Mechanism of variable structural colour in the neon tetra: quantitative evaluation of the Venetian blind model. J. R. Soc Interface 8, 56–66. https://doi.org/10.1098/rsif.2010.0253 (2011).
Lin, J. C. et al. Two Zebrafish hsd3b genes are distinct in function, expression, and evolution. Endocrinology 156, 2854–2862. https://doi.org/10.1210/en.2014-1584 (2015).
Lister, J. A. Larval but not adult xanthophore pigmentation in zebrafish requires GTP cyclohydrolase 2 (gch2) function. Pigment Cell Melanoma Res. 32, 724–727. https://doi.org/10.1111/pcmr.12783 (2019).
Granneman, J. G. et al. Lipid droplet biology and evolution illuminated by the characterization of a novel perilipin in teleost fish. Elife https://doi.org/10.7554/eLife.21771 (2017).
Lister, J. A., Close, J. & Raible, D. W. Duplicate mitf genes in zebrafish: complementary expression and conservation of melanogenic potential. Dev. Biol. 237, 333–344. https://doi.org/10.1006/dbio.2001.0379 (2001).
Elworthy, S., Lister, J. A., Carney, T. J., Raible, D. W. & Kelsh, R. N. Transcriptional regulation of mitfa accounts for the sox10 requirement in zebrafish melanophore development. Development 130, 2809–2818. https://doi.org/10.1242/dev.00461 (2003).
Johnson, S. L., Nguyen, A. N. & Lister, J. A. mitfa is required at multiple stages of melanocyte differentiation but not to establish the melanocyte stem cell. Dev. Biol. 350, 405–413. https://doi.org/10.1016/j.ydbio.2010.12.004 (2011).
Salis, P. et al. Developmental and comparative transcriptomic identification of iridophore contribution to white barring in clownfish. Pigment Cell Melanoma Res. 32, 391–402. https://doi.org/10.1111/pcmr.12766 (2019).
Ahi, E. P. et al. Comparative transcriptomics reveals candidate carotenoid color genes in an East African cichlid fish. BMC Genom. 21, 54. https://doi.org/10.1186/s12864-020-6473-8 (2020).
Hooper, D. M., Griffith, S. C. & Price, T. D. Sex chromosome inversions enforce reproductive isolation across an avian hybrid zone. Mol. Ecol. 28, 1246–1262. https://doi.org/10.1111/mec.14874 (2019).
Bagati, A. et al. FOXQ1 controls the induced differentiation of melanocytic cells. Cell Death Differ. 25, 1040–1049. https://doi.org/10.1038/s41418-018-0066-y (2018).
Batista, A. F. R., Martinez, J. C. & Hengst, U. Intra-axonal synthesis of SNAP25 is required for the formation of presynaptic terminals. Cell Rep. 20, 3085–3098. https://doi.org/10.1016/j.celrep.2017.08.097 (2017).
Alfonso, J., Fernández, M. E., Cooper, B., Flugge, G. & Frasch, A. C. The stress-regulated protein M6a is a key modulator for neurite outgrowth and filopodium/spine formation. Proc. Natl. Acad. Sci. USA. 102, 17196. https://doi.org/10.1073/pnas.0504262102 (2005).
Monteleone, M. C., Billi, S. C., Brocco, M. A. & Frasch, A. C. Neural glycoprotein M6a is released in extracellular vesicles and modulated by chronic stressors in blood. Sci. Rep. 7, 9788. https://doi.org/10.1038/s41598-017-09713-0 (2017).
Li, C., Pan, W., Braunewell, K. H. & Ames, J. B. Structural analysis of Mg2+ and Ca2+ binding, myristoylation, and dimerization of the neuronal calcium sensor and visinin-like protein 1 (VILIP-1). J. Biol. Chem. 286, 6354–6366. https://doi.org/10.1074/jbc.M110.173724 (2011).
Rasmussen, J. P., Vo, N. T. & Sagasti, A. Fish scales dictate the pattern of adult skin innervation and vascularization. Dev Cell 46, 344–359. https://doi.org/10.1016/j.devcel.2018.06.019 (2018).
Hulsey, C. D., Zheng, J., Faircloth, B. C., Meyer, A. & Alfaro, M. E. Phylogenomic analysis of Lake Malawi cichlid fishes: further evidence that the three-stage model of diversification does not fit. Mol. Phylogenet. Evol. 114, 40–48. https://doi.org/10.1016/j.ympev.2017.05.027 (2017).
Malinsky, M. et al. Whole-genome sequences of Malawi cichlids reveal multiple radiations interconnected by gene flow. Nat. Ecol. Evol. 2, 1940–1955. https://doi.org/10.1038/s41559-018-0717-x (2018).
Gur, D., Palmer, B. A., Weiner, S. & Addadi, L. Light manipulation by guanine crystals in organisms: biogenic scatterers, mirrors, multilayer reflectors and photonic crystals. Adv. Func. Mater. 27, 1603514. https://doi.org/10.1002/adfm.201603514 (2017).
Arregi, I. et al. Retinol dehydrogenase-10 regulates pancreas organogenesis and endocrine cell differentiation via paracrine retinoic acid signaling. Endocrinology 157, 4615–4631. https://doi.org/10.1210/en.2016-1745 (2016).
Gallaud, E. et al. Ensconsin/Map7 promotes microtubule growth and centrosome separation in Drosophila neural stem cells. J. Cell Biol. 204, 1111–1121. https://doi.org/10.1083/jcb.201311094 (2014).
Gao, M. et al. Regulation of high-density lipoprotein on hematopoietic stem/progenitor cells in atherosclerosis requires scavenger receptor type BI expression. Arterioscler. Thromb. Vasc. Biol. 34, 1900–1909. https://doi.org/10.1161/ATVBAHA.114.304006 (2014).
Mori, E. et al. Impaired adipogenic capacity in induced pluripotent stem cells from lipodystrophic patients with BSCL2 mutations. Metabolism 65, 543–556. https://doi.org/10.1016/j.metabol.2015.12.015 (2016).
Dooley, C. M., Mongera, A., Walderich, B. & Nüsslein-Volhard, C. On the embryonic origin of adult melanophores: the role of ErbB and Kit signalling in establishing melanophore stem cells in zebrafish. Development 140, 1003–1013. https://doi.org/10.1242/dev.087007 (2013).
Parichy, D. M. & Spiewak, J. E. Origins of adult pigmentation: diversity in pigment stem cell lineages and implications for pattern evolution. Pigment Cell Melanoma Res. 28, 31–50. https://doi.org/10.1111/pcmr.12332 (2015).
Singh, A. P. et al. Pigment cell progenitors in zebrafish remain multipotent through metamorphosis. Dev. Cell 38, 316–330. https://doi.org/10.1016/j.devcel.2016.06.020 (2016).
Burton, D. & Burton, M. Essential Fish Biology: Diversity, Structure, and Function (Oxford University Press, Oxford, 2017).
Yoshioka, S. et al. Mechanism of variable structural colour in the neon tetra: quantitative evaluation of the Venetian blind model. J. R. Soc. Interface 8, 56–66. https://doi.org/10.1098/rsif.2010.0253 (2011).
Sugimoto, M. Morphological color changes in fish: regulation of pigment cell density and morphology. Microsc. Res. Technol. 58, 496–503. https://doi.org/10.1002/jemt.10168 (2002).
Sugimoto, M., Uchida, N. & Hatayama, M. Apoptosis in skin pigment cells of the medaka, Oryzias latipes (Teleostei), during long-term chromatic adaptation: the role of sympathetic innervation. Cell Tissue Res. 301, 205–216. https://doi.org/10.1007/s004410000226 (2000).
Hara, M. et al. Innervation of melanocytes in human skin. J. Exp. Med. 184, 1385–1395. https://doi.org/10.1084/jem.184.4.1385 (1996).
Zhang, B. et al. Hyperactivation of sympathetic nerves drives depletion of melanocyte stem cells. Nature 577, 676–681. https://doi.org/10.1038/s41586-020-1935-3 (2020).
Inoue, S., Kondo, S., Parichy, D. M. & Watanabe, M. Tetraspanin 3c requirement for pigment cell interactions and boundary formation in zebrafish adult pigment stripes. Pigment Cell Melanoma Res. 27, 190–200. https://doi.org/10.1111/pcmr.12192 (2014).
Eom, D. S. & Parichy, D. M. A macrophage relay for long-distance signaling during postembryonic tissue remodeling. Science 355, 1317–1320. https://doi.org/10.1126/science.aal2745 (2017).
Le Guyader, S. & Jesuthasan, S. Analysis of xanthophore and pterinosome biogenesis in zebrafish using methylene blue and pteridine autofluorescence. Pigments Cell Res. 15, 27–31 (2002).
Kelsh, R. N. et al. Zebrafish pigmentation mutations and the processes of neural crest development. Development 123, 369–389 (1996).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682. https://doi.org/10.1038/nmeth.2019 (2012).
Cardona, A. et al. TrakEM2 software for neural circuit reconstruction. PLoS ONE 7, e38011. https://doi.org/10.1371/journal.pone.0038011 (2012).
Wickham, H. ggplot2: elegant graphics for data analysis (Springer, Berlin, 2016).
R Development Core Team, R. (R foundation for statistical computing Vienna, Austria, 2011).
Kratochwil, C. F., Liang, Y., Urban, S., Torres-Dowdall, J. & Meyer, A. Evolutionary dynamics of structural variation at a key locus for color pattern diversification in cichlid fishes. Genome Biol. Evol. 11, 3452–3465. https://doi.org/10.1093/gbe/evz261 (2019).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 12, 323 (2011).
Ewels, P., Magnusson, M., Lundin, S. & Käller, M. MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinformatics 32, 3047–3048 (2016).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
R Development Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing. (2019).
Morgan, M., Obenchain, V., Lang, M., Thompson, R. & Turaga, N. R Package ‘BiocParallel’ v.1.16.2. Bioconductor (2019).
Soneson, C., Love, M. I. & Robinson, M. D. Differential analyses for RNA-seq: transcript-level estimates improve gene-level inferences. F1000Research 4 (2015).
Wickham, H. R Package ‘stringR’ v.1.4.0. Cran R (2019).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Huber, W., Von Heydebreck, A., Sültmann, H., Poustka, A. & Vingron, M. Variance stabilization applied to microarray data calibration and to the quantification of differential expression. Bioinformatics 18, S96–S104 (2002).
Wickham, H. et al. R Package ‘ggplot2' v.3.1.1. Cran R (2019).
Neuwirth, E. R Package ‘RColorBrewer’ v.1.1–2. Cran R (2014).
McDermaid, A., Monier, B., Zhao, J. & Ma, Q. ViDGER: An R package for integrative interpretation of differential gene expression results of RNA-seq data. bioRxiv, 268896 (2018).
Kolde, R. R Package ‘pheatmap’ v.1.0.12. Cran R (2019).
Wilke, C. O. R Package ‘cowplot’ v.0.9.4. Cran R (2019).
Wickham, H., François, R., Henry, L. & Müller, K. R Package ‘dplyr’ v.0.8.0.1. Cran R (2019).
Durinck, S., Spellman, P. T., Birney, E. & Huber, W. Mapping identifiers for the integration of genomic datasets with the R/Bioconductor package biomaRt. Nat. Protoc. 4, 1184 (2009).
Young, M. D., Wakefield, M. J., Smyth, G. K. & Oshlack, A. Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol. 11, R14 (2010).
Woltering, J. M., Holzem, M., Schneider, R. F., Nanos, V. & Meyer, A. The skeletal ontogeny of Astatotilapia burtoni—a direct-developing model system for the evolution and development of the teleost body plan. BMC Dev. Biol. 18, 8. https://doi.org/10.1186/s12861-018-0166-4 (2018).
Chitnis, A. B. & Kuwada, J. Y. Axonogenesis in the brain of zebrafish embryos. J Neurosci 10, 1892–1905 (1990).
Fontenas, L. et al. Neuronal Ndrg4 is essential for nodes of ranvier organization in zebrafish. PLoS Genet. 12, e1006459. https://doi.org/10.1371/journal.pgen.1006459 (2016).
Woods, I. G., Lyons, D. A., Voas, M. G., Pogoda, H. M. & Talbot, W. S. nsf is essential for organization of myelinated axons in zebrafish. Curr. Biol. 16, 636–648. https://doi.org/10.1016/j.cub.2006.02.067 (2006).
Bademosi, A. T. et al. In vivo single-molecule imaging of syntaxin1A reveals polyphosphoinositide- and activity-dependent trapping in presynaptic nanoclusters. Nat. Commun. 8, 13660. https://doi.org/10.1038/ncomms13660 (2017).
Lodygin, D. et al. beta-Synuclein-reactive T cells induce autoimmune CNS grey matter degeneration. Nature 566, 503–508. https://doi.org/10.1038/s41586-019-0964-2 (2019).
Yaguchi, H. et al. Sez6l2 regulates phosphorylation of ADD and neuritogenesis. Biochem. Biophys. Res. Commun. 494, 234–241. https://doi.org/10.1016/j.bbrc.2017.10.047 (2017).
Chander, P., Kennedy, M. J., Winckler, B. & Weick, J. P. Neuron-specific gene 2 (NSG2) encodes an AMPA receptor interacting protein that modulates excitatory neurotransmission. eNeuro https://doi.org/10.1523/ENEURO.0292-18.2018 (2019).
Barford, K., Yap, C. C., Dwyer, N. D. & Winckler, B. The related neuronal endosomal proteins NEEP21 (Nsg1) and P19 (Nsg2) have divergent expression profiles in vivo. J. Comp. Neurol. 525, 1861–1878. https://doi.org/10.1002/cne.24168 (2017).
Nagao, M., Ogata, T., Sawada, Y. & Gotoh, Y. Zbtb20 promotes astrocytogenesis during neocortical development. Nat. Commun. 7, 11102. https://doi.org/10.1038/ncomms11102 (2016).
Young, J. K., Heinbockel, T. & Gondre-Lewis, M. C. Astrocyte fatty acid binding protein-7 is a marker for neurogenic niches in the rat hippocampus. Hippocampus 23, 1476–1483. https://doi.org/10.1002/hipo.22200 (2013).
Schuske, K. & Jorgensen, E. M. Neuroscience. Vesicular glutamate transporter–shooting blanks. Science 304, 1750–1752. https://doi.org/10.1126/science.1100475 (2004).
Fremeau, R. T. Jr. et al. Vesicular glutamate transporters 1 and 2 target to functionally distinct synaptic release sites. Science 304, 1815–1819. https://doi.org/10.1126/science.1097468 (2004).
Tao, B. et al. Secretogranin-II plays a critical role in zebrafish neurovascular modeling. J. Mol. Cell. Biol. 10, 388–401. https://doi.org/10.1093/jmcb/mjy027 (2018).
Hashimoto, R. et al. Pituitary adenylate cyclase-activating polypeptide is associated with schizophrenia. Mol. Psychiatry 12, 1026–1032. https://doi.org/10.1038/sj.mp.4001982 (2007).
Davis, K. N. et al. GAD2 Alternative transcripts in the human prefrontal cortex, and in schizophrenia and affective disorders. PLoS ONE 11, e0148558. https://doi.org/10.1371/journal.pone.0148558 (2016).
Labouesse, M. A., Dong, E., Grayson, D. R., Guidotti, A. & Meyer, U. Maternal immune activation induces GAD1 and GAD2 promoter remodeling in the offspring prefrontal cortex. Epigenetics 10, 1143–1155. https://doi.org/10.1080/15592294.2015.1114202 (2015).
Zhang, Y. et al. Inhibition of glutamate decarboxylase (GAD) by ethyl ketopentenoate (EKP) induces treatment-resistant epileptic seizures in zebrafish. Sci. Rep. 7, 7195. https://doi.org/10.1038/s41598-017-06294-w (2017).
Liem, R. K. H. Molecular biology of neuronal intermediate filaments. Curr. Opin. Cell Biol. 5, 12–16. https://doi.org/10.1016/S0955-0674(05)80003-1 (1993).
Bai, Q., Wei, X. & Burton, E. A. Expression of a 12-kb promoter element derived from the zebrafish enolase-2 gene in the zebrafish visual system. Neurosci. Lett. 449, 252–257. https://doi.org/10.1016/j.neulet.2008.10.101 (2009).
Bai, Q., Garver, J. A., Hukriede, N. A. & Burton, E. A. Generation of a transgenic zebrafish model of Tauopathy using a novel promoter element derived from the zebrafish eno2 gene. Nucleic Acids Res. 35, 6501–6516. https://doi.org/10.1093/nar/gkm608 (2007).
Chen, N. H., Reith, M. E. & Quick, M. W. Synaptic uptake and beyond: the sodium- and chloride-dependent neurotransmitter transporter family SLC6. Pflugers Arch. 447, 519–531. https://doi.org/10.1007/s00424-003-1064-5 (2004).
Sullivan, C. S. et al. Perineuronal net protein neurocan inhibits NCAM/EphA3 repellent signaling in GABAergic interneurons. Sci. Rep. 8, 6143. https://doi.org/10.1038/s41598-018-24272-8 (2018).
Doganli, C., Beck, H. C., Ribera, A. B., Oxvig, C. & Lykke-Hartmann, K. alpha3Na+/K+-ATPase deficiency causes brain ventricle dilation and abrupt embryonic motility in zebrafish. J. Biol. Chem. 288, 8862–8874. https://doi.org/10.1074/jbc.M112.421529 (2013).
Takei, Y., Kikkawa, Y. S., Atapour, N., Hensch, T. K. & Hirokawa, N. Defects in synaptic plasticity, reduced NMDA-receptor transport, and instability of postsynaptic density proteins in mice lacking microtubule-associated protein 1A. J. Neurosci. 35, 15539–15554. https://doi.org/10.1523/JNEUROSCI.2671-15.2015 (2015).
Liu, Y., Lee, J. W. & Ackerman, S. L. Mutations in the microtubule-associated protein 1A (Map1a) gene cause Purkinje cell degeneration. J. Neurosci. 35, 4587–4598. https://doi.org/10.1523/JNEUROSCI.2757-14.2015 (2015).
Acknowledgements
This work was supported by a stipend from the China Scholarship Council (CSC, to Y.L.), the Baden-Württemberg Foundation (to C.F.K.) grants by the Deutsche Forschungsgemeinschaft (DFG, to A.M. and KR 4670/2–1, KR 4670/4-1 to C.F.K.) and the University of Konstanz (AM). The authors thank Jan Gerwin and the staff of the animal facility and the Electron Microscopy Centre of the University of Konstanz for their valuable help.
Author information
Authors and Affiliations
Contributions
C.F.K. conceived the study, C.F.K. and A.M. supervised the study, C.F.K. analyzed the RNA-seq data, Y.L. conducted all other experiments and analyses. Y.L. and C.F.K. wrote the manuscript. A.M. edited the manuscript. C.F.K. and A.M. provided funding.
Corresponding authors
Ethics declarations
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liang, Y., Meyer, A. & Kratochwil, C.F. Neural innervation as a potential trigger of morphological color change and sexual dimorphism in cichlid fish. Sci Rep 10, 12329 (2020). https://doi.org/10.1038/s41598-020-69239-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-69239-w
This article is cited by
-
East African cichlid fishes
EvoDevo (2023)
-
Thyroid hormone signaling in the evolution of carotenoid coloration in Neotropical cichlids with reversed sexual dichromatism
Environmental Biology of Fishes (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.